Quorum Quenching Agents: Resources for Antivirulence Therapy
Abstract
:1. Introduction
2. The Distribution of QS Systems in Microorganisms
| QS Signaling Type | Structure | Representative Microorganisms | Associated Phenomena | Reference |
|---|---|---|---|---|
| Intraspecies Communication Signals | ||||
| “Traditional” AHL | C4-C18, 3OC4-3OC18 and 3OHC4-3OHC18 | Various Gram-negative bacteria; only one Gram-positive bacteria: Exiguobacterium sp. MPO | Virulence, biofilm, swarming and bioluminescence | [2,20] |
| “Noncanonical” AHL | p-Coumaroyl-HSL | Rhodopseudomonas palustris CGA009 | Global gene expression | [28] |
| DSF family | Cis-unsaturated fatty acid | Xanthomonas spp., and Burkholderia cenocepacia | Virulence, biofilm and antibiotic tolerance | [24] |
| CAI-1 family | α-Hydroxyketones | Vibrio spp. and Legionella pneumophila | Virulence and biofilm | [23,31] |
| AIP family | Linear or cyclized oligopeptide | Many Gram-positive bacteria; only one Gram-negative bacterium: Thermotoga maritima | Virulence, biofilm, sporulation and exopolysaccharide production | [21,32] |
| PQS or IQS | Quinolone or thiazole compounds | Pseudomonas aeruginosa | Virulence and biofilm formation | [33,34] |
| Pyrones | α-Pyrones | Photorhabdus luminescens | Virulence | [35] |
| Interspecies and Interkingdom Communication Signals | ||||
| AI-2 | S-THMF-borate R-THMF | Many Gram-negative and Gram-positives bacteria | Virulence and biofilm formation | [15] |
| AI-3 | Unknown | Enterohemorrhagic Escherichia coli (EHEC) | Virulence | [36] |
| Indole | 2,3-Benzopyrrole | Many Gram-negative and Gram-positives bacteria | Virulence and biofilm formation | [19] |
| QS Signals in Fungi | ||||
| Farnesol or Tyrosol | Sesquiterpene or phenylethanoid | Candida albicans | Inhibition or stimulation filamentation and biofilm formation | [25,26] |
| Peptide | NH2-NFGAPGGAYPW-COOH | Cryptococcus neoformans | Colony formation in agar media | [22] |
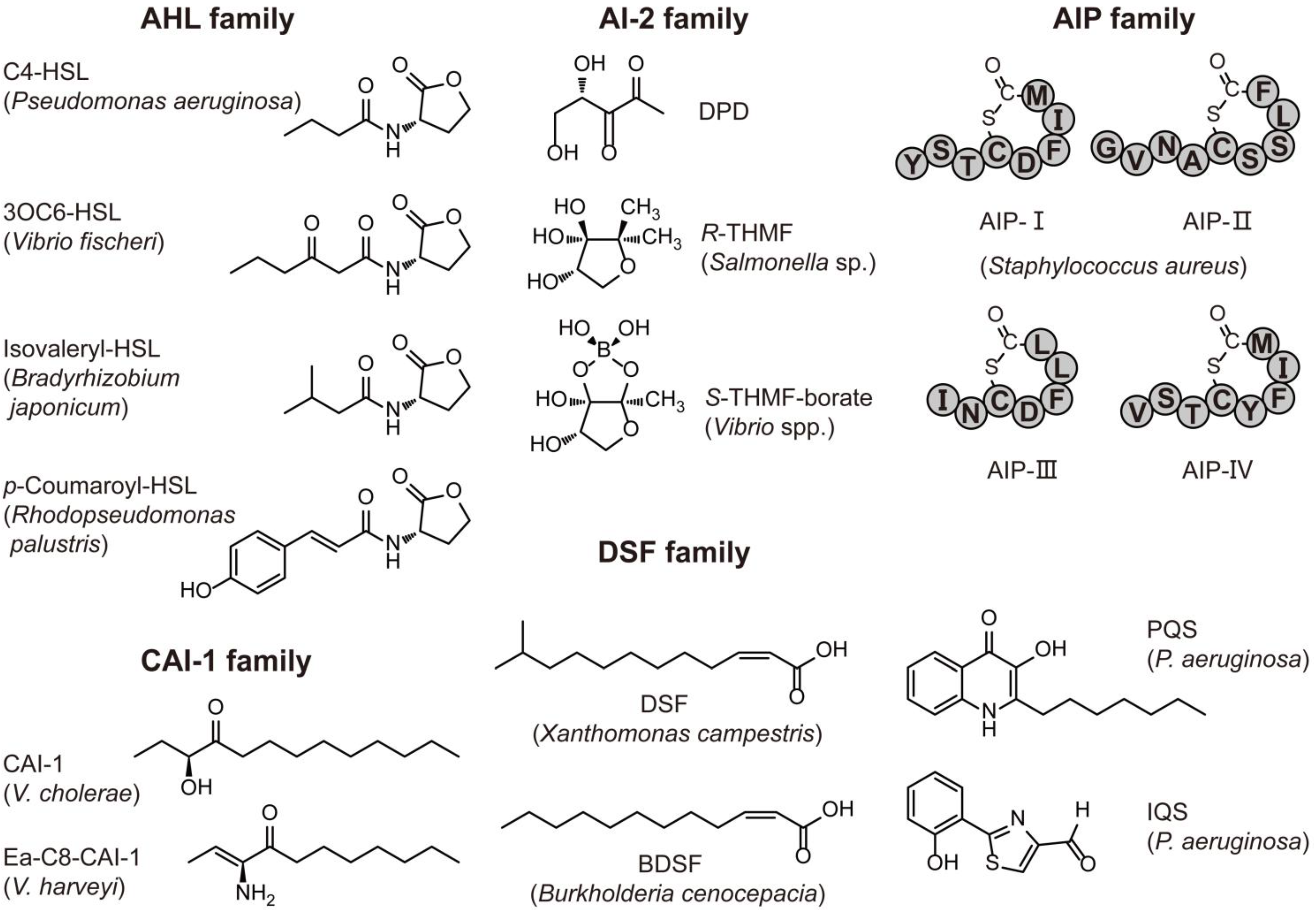
2.1. Characteristics of AHL Molecules
2.2. The Diversity and Uniformity of QS Systems
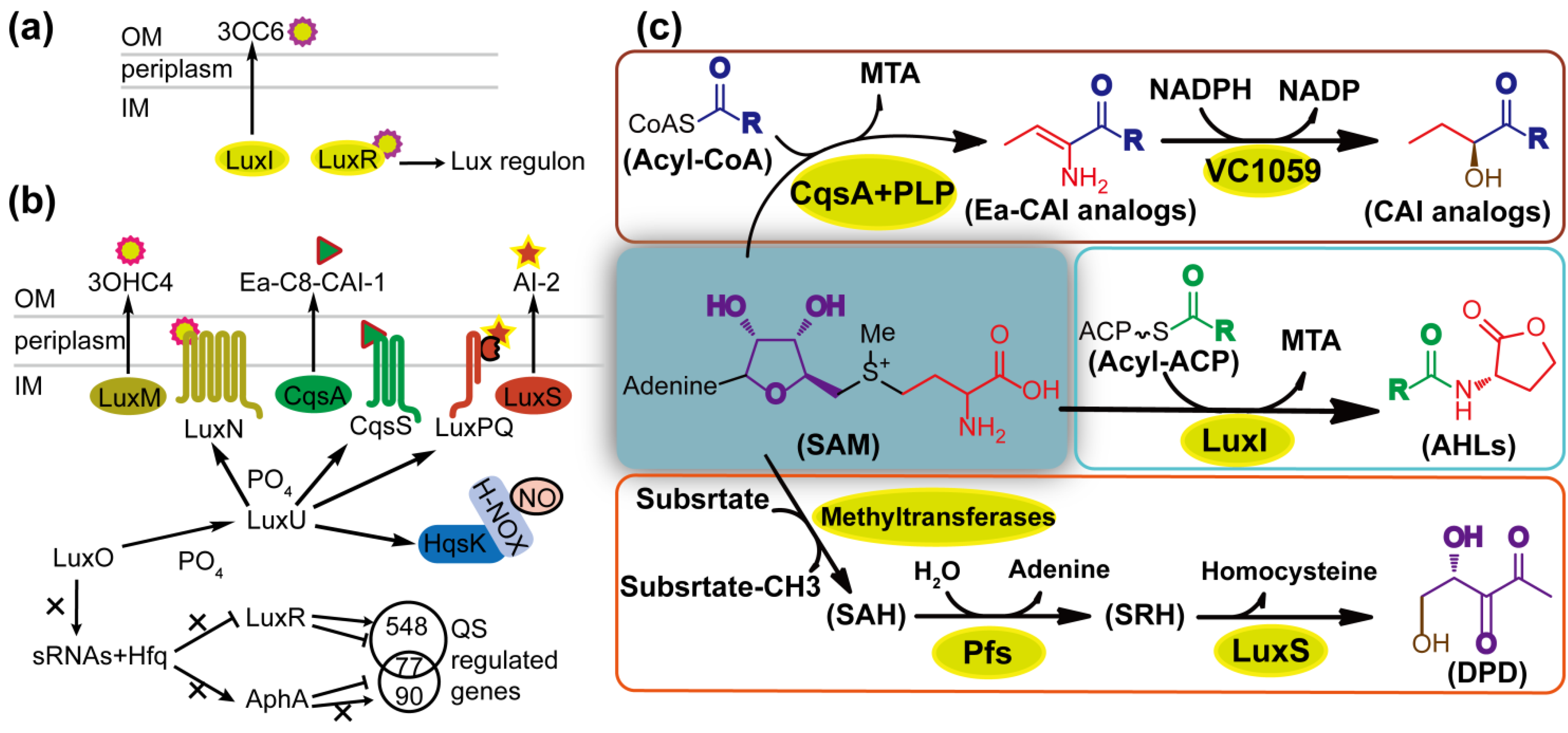
3. Natural Quorum Quenching Resources
3.1. Small Molecular QQ Agents
3.1.1. Marine-Derived Inhibitors against AHL-Dependent QS
| Category | Species | Inhibitor | Target | Reference |
|---|---|---|---|---|
| Marine-Derived Inhibitors against AHL-Dependent QS | ||||
| Actinobacteria | Streptomyces sp. | Piericidin A1 | CviR | [58] |
| Bacteria | Halobacillus salinus | N-(2-Phenylethyl)-isobutyramide and 3-Methyl-N-(2-phenylethyl)-butyramide | LuxR, CviR and Vibrio harveyi | [59,60] |
| Bacteria | Bacillus cereus D28 | Cyclo-l-proline-l-tyrosine | Chromobacterium violaceum | [60] |
| Bacteria | Marinobacter sp. SK-3 | Diketopiperazines (dkps) | CviR and LuxR | [61] |
| Bryozoan | Flustra foliacea | Brominated alkaloids compounds | CepR and LuxR | [62] |
| Coral | Pseudoplexaura flagellosa and Eunicea knighti | Cembranoids | LuxR and V. harveyi | [63,64] |
| Cyanobacteria | Blennothrix cantharidosmum | Tumonoic acid F | V. harveyi | [55] |
| Cyanobacteria | Lyngbya majuscula | 8-Epi-malyngamide C and lyngbic acid | LasR | [65] |
| Cyanobacteria | L. majuscula | Lyngbyoic acid | LasR | [66] |
| Cyanobacteria | L. majuscula | Malyngolide | CviR and LasR | [56] |
| Cyanobacteria | Leptolyngbya crossbyana | Honaucins A–C | LuxR | [57] |
| Cyanobacteria | Lyngbya sp. | Pitinoic acid A | LasR | [67] |
| Cyanobacteria | Lyngbya sp. | Pepitdes (microcolins A and B) | LuxR | [68] |
| Fungi | Penicillium atramentosum | Crude extracts | LasR | [69] |
| Red algae | Ahnfeltiopsis flabelliformes | Floridoside, betonicine and isethionic acid | TraR | [70] |
| Sponge | Luffareilla variabilis | Manoalide, manoalide monoacetate, and secomanoalide | LuxR and LasR | [71] |
| Sponge | Hymeniacidon aldis | Alkaloid (hymenialdisin) | LuxR and LasR | [68] |
| Terrestrial-Derived Inhibitors against AHL-Dependent QS | ||||
| Bacteria | Staphylococcus delphini | Yayurea A and B | LuxN | [72] |
| Bacteria | Stenotrophomonas maltophilia BJ01 | Cis-9-octadecenoic acid | CviR | [73] |
| Bacteria | Pseudomonas spp. | Protoanemonin | LasR | [74] |
| Fungi | Aspergillus spp. | Kojic acid | LuxR | [68] |
| Fungi | Penicillium spp. | Patulin and Penilillic acid | LasR and RhlR | [75] |
| Insect productions | Bee | Honey and propolis | C. violaceum, LasR and RhlR | [76,77] |
| Insect: fire ant | Solenopis invicta | Solenopsin A | rhl circuit | [78] |
| Plant | Baccharis cassinaefolia | Benzopyran | CviR, LuxR and LasR | [68] |
| Plant | Syzygium aromaticum | Eugenol | CviR, LasR and PQS | [79] |
| Plant: alfalfa | Medicago sativa | l-Canavanine | CviR and ExpR | [80] |
| Plant: Combretaceae | Combretum albiflorum | Catachin and naringenin | CviR and RhlR | [81,82] |
| Plant: Compositae | Centratherum punctatum | Sesquiterpene lactones | Pseudomonas aeruginosa | [83] |
| Plant: garlic | Allium sativum | Ajoene | LuxR family | [84] |
| Plant: horseradish | Armoracia rusticana | Iberin | LasR and RhlR | [85] |
| Plant: Myristicaceae | Myristica cinnamomea | Malabaricone C | CviR | [86] |
| Plant: tree | Drimys winteri | A drimane sesquiterpene | C. violaceum | [87] |
| Plant: turmeric | Curcuma longa | Curcumin | CviR | [88] |
| Plants | Various plants | p-Coumaric acid | PpuR, CviR and TraR | [89] |
| Plants and bacteria | Conocarpus erectus | Ellagitannins and urolithins | P. aeruginosa and Yersinia enterocolitica | [90,91] |
| Roundworm | Caenorhabditis elegans | Exudates | LuxR | [92] |
| Soil-freshwater alga | Chlamydomonas reinhardtii | Lumichrome | Sinorhizobium meliloti | [93,94] |
| TCMs * | Rhubarb | Emodin | P. aeruginosa and S. maltophilia | [95] |
| TCMs | Scutellaria baicalensis | Flavonoid (baicalein) | TraR and RhlR | [96] |
| Inhibitors against PQS System | ||||
| Fungi | Candida albicans | Farnesol (sesquiterpene) | PqsA | [97] |
| Inhibitors against AI-2 and AI-3 System | ||||
| Marine alga | Delisea pulchra | Furanone and its derivatives | LuxR and LuxS | [98,99] |
| Plant | Many plants | Cinnamaldehyde and its derivatives | LuxR and AI-2 | [100,101,102] |
| Plant | Broccoli | Quercetin | AI-2 and AI-3 | [103] |
| Plant | Grapefruit | Limonoids (obacunone) | EHEC | [104] |
| Inhibitors against AIP System | ||||
| Bacteria | Lactobacillus reuteri | Cyclic dipeptides: cyclo(l-Phe-l-Pro) and cyclo(l-Tyr-l-Pro) | agr system | [105] |
| Marine bacteria | Photobacterium | Cyclodepsipeptides (solonomide a, b) | agr system | [106] |
| Plant: witch hazel | Hamamelis virginiana | 2,5-di-O-galloyl-d hamamelose | RNAIII | [107] |
3.1.2. Terrestrial-Derived Inhibitors against AHL-Dependent QS
3.1.3. Natural Inhibitors against Other QS Systems
3.1.4. Evaluation of Natural QS Inhibitors
3.2. Macromolecular QQ Agents
3.2.1. AHL Lactonases
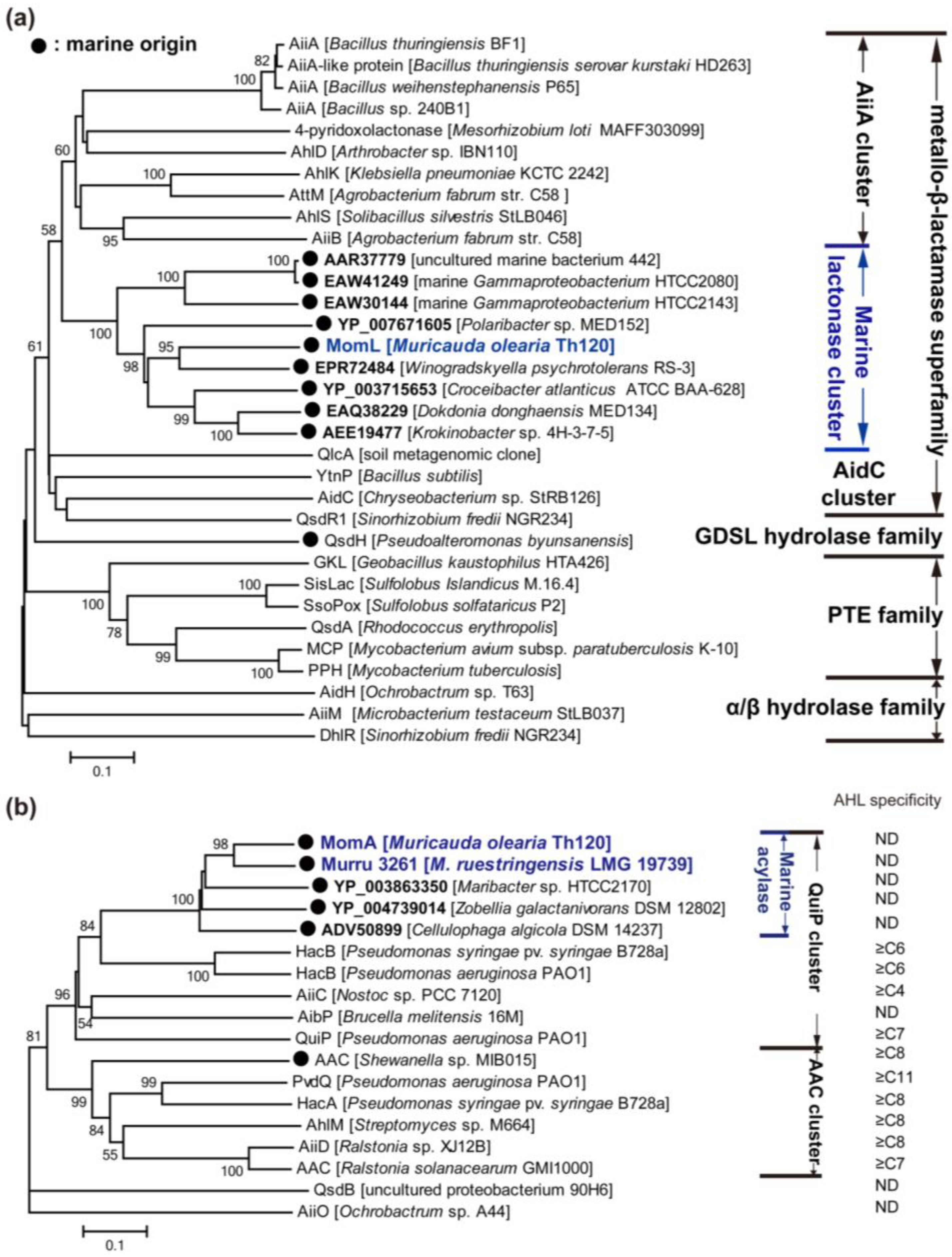
3.2.2. AHL Acylases
3.2.3. AHL Oxidoreductases
4. Microorganisms May Produce QQ Agents to Gain Benefits in a Competitive Environment
5. QQ in the Marine Environment: A Tremendous Resource to Be Developed
| Strain | AHL-Degrading Ability * | Activity ** | Origin | Reference |
|---|---|---|---|---|
| Actinobacteria | ||||
| Rhodococcus erythropolis strains | C4, C6, C10 and 3OC12 | Lactonase | Fucus vesiculosus and sediment | [188] |
| Alphaproteobacteria | ||||
| Hyphomonas sp. USC2 | C4, C6, C10 and 3OC12 | Lactonase | Fucus vesiculosus | [188] |
| Marivita sp. Th30 | C6, C12 and C14 | ND | Flounder | [129] |
| Novosphingobium sp. Th20 | C6-C14 and 3OC6-3OC14 | ND | Flounder | [129] |
| Paracoccus sp. PP2-663 | C4-C12 | ND | Manila clam | [189] |
| Phaeobacter sp. USC177 | C4, C6, C10 and 3OC12 | ND | Fucus vesiculosus | [188] |
| Rhodobacter sp. Th15 | C8-C14 and 3OC14 | ND | Flounder | [129] |
| Roseovarius aestuarii USC61 | C4, C6, C10 and 3OC12 | Lactonase | Water tank | [188] |
| Sphingopyxis flavimaris T51 | C6-C14 and 3OC10-3OC14 | ND | Flounder | [129] |
| Sphingopyxis litoris th8 | C6-C14 and 3OC6-3OC14 | ND | Flounder | [129] |
| Stappia sp. USC176 | C4, C6, C10 and 3OC12 | Lactonase | Fucus vesiculosus | [188] |
| Stappia sp. USC5 | C4, C6, C10 and 3OC12 | Lactonase | Fucus vesiculosus | [188] |
| Firmicutes | ||||
| Bacillus circulans USC24 | C4, C6, C10 and 3OC12 | Lactonase | Sediment | [188] |
| Bacillus sp. KT7 | C6-C14, 3OC8-3OC12 and 3OHC8-3OHC12 | ND | Intertidal rocks colonized by Ulva | [190] |
| Bacillus spp. | C6 | ND | Shrimp and bass | [191] |
| Oceanobacillus spp. | C4, C6, C10 and 3OC12 | Lactonase | Fucus vesiculosus | [188] |
| Flavobacteria | ||||
| Flaviramulus ichthyoenteri Th78 | C6-C14 and 3OC6-3OC14 | Lactonase | Flounder | [129] |
| Maribacter sp. 139 | C4, C6, C10 and 3OC12 | Lactonase | Ocean water | [184] |
| Muricauda olearia Th120 | C6-C14 and 3OC6-3OC14 | Latonase and acylase | Flounder | [129] |
| Olleya marilimosa 138E | C4, C6, C10 and 3OC12 | Lactonase | Ocean water | [184] |
| O. marilimosa t168 | C6-C14 and 3OC6-3OC14 | Lactonase | Marine | [129] |
| Tenacibaculum discolor 20J | C4, C6, C10 and 3OC12 | Lactonase | Sediment | [188] |
| T. discolor t84 | C6-C14 and 3OC6-3OC14 | ND | Gill of flounder | [129] |
| T. maritimum 2154t | C10 | Acylase | Fish farm disease | [192] |
| T. soleae strains | C6-C14 and 3OC6-3OC14 | Lactonase | Gill of flounder | [129] |
| Gammaproteobacteria | ||||
| Alteromonas marina PP2-67 | C4-C12 | ND | Pod razor clam | [189] |
| Alteromonas sp. USC168 | C4, C6, C10 and 3OC12 | ND | Fucus vesiculosus | [188] |
| A. stellipolaris pp2-644 | C4-C12 | ND | Carpet-shell clam | [189] |
| Colwellia aestuarii T171 | C8-C14 and 3OC10-3OC14 | ND | Gill of flounder | [129] |
| Glaciecola sp. B20 | C10-C14, 3OC10-, 3OHC10, 3OC12 and 3OHC12 | ND | Intertidal rocks | [190] |
| Halomonas taeanensis USC33 | C4, C6, C10 and 3OC12 | Lactonase | Sediment | [188] |
| Marinobacterium sp. B2 | 3OC10, C12, 3OC12, 3OHC12 and C14 | ND | Intertidal rocks | [190] |
| Pseudoalteromonas byunsanensis 1A01261 | C4-C14 and 3OC4-3OC12 | Lactonase | Marine | [193] |
| P. rydzensis Th125 | C10-C14 and 3OC10-3OC14 | ND | Flounder | [129] |
| Salinimonas sp. T194 | C8-C14 and 3OC10-3OC14 | ND | Gill of flounder | [129] |
| Salinicola salarius 131 | C4, C6, C10 and 3OC12 | Lactonase | Ocean water | [184] |
| Shewanella sp. B21 | C8-C14, 3OC8-3OC12 and 3OHC8-3OHC12 | ND | Intertidal rocks | [190] |
| Thalassomonas sp. PP2-459 | C4-C12 | ND | Carpet-shell clam | [189] |
| Thalassomonas sp. T202 | C8-C14 and 3OC10-3OC14 | ND | Gill of flounder | [129] |
6. Further Issues of Concern for the Application of QQ Agents
7. Concluding Remarks
Abbreviations
| AHL | N-Acyl-homoserine lactonse |
| AI-2 | Autoinducter-2 |
| AI-3 | Autoinducter-3 |
| AIP | Autoinducing peptides |
| DSF | Diffusible signal factor |
| BDSF | Burkholderia cenocepacia diffusible signal factor |
| CAI-1 | Cholerae autoinducer-1 |
| Ea-C8-CAI-1 | (Z)-3-Aminoundec-2-en-4-one |
| PQS | Pseudomonas quinolone signal |
| IQS | Integrating QS signal |
| R-THMF | (2R,4S)-2-Methyl-2,3,3,4-tetrahydroxytetrahydrofuran |
| S-THMF-borate | (2S,4S)-2-Methyl-2,3,3,4-tetrahydroxytetrahydrofuran-borate |
| DPD | 4,5-Dihydroxy-2,3-pentanedione |
Acknowledgments
Conflicts of Interest
References
- Clatworthy, A.E.; Pierson, E.; Hung, D.T. Targeting virulence: A new paradigm for antimicrobial therapy. Nat. Chem. Biol. 2007, 3, 541–548. [Google Scholar] [CrossRef]
- Ng, W.L.; Bassler, B.L. Bacterial quorum-sensing network architectures. Annu. Rev. Genet. 2009, 43, 197–222. [Google Scholar] [CrossRef]
- Waters, C.M.; Bassler, B.L. Quorum sensing: Cell-to-cell communication in bacteria. Annu. Rev. Cell. Dev. Biol. 2005, 21, 319–346. [Google Scholar] [CrossRef]
- Fuqua, W.C.; Winans, S.C.; Greenberg, E.P. Quorum sensing in bacteria: The LuxR-LuxI family of cell density-responsive transcriptional regulators. J. Bacteriol. 1994, 176, 269–275. [Google Scholar]
- Redfield, R.J. Is quorum sensing a side effect of diffusion sensing? Trends Microbiol. 2002, 10, 365–370. [Google Scholar] [CrossRef]
- Hense, B.A.; Kuttler, C.; Muller, J.; Rothballer, M.; Hartmann, A.; Kreft, J.U. Does efficiency sensing unify diffusion and quorum sensing? Nat. Rev. Microbiol. 2007, 5, 230–239. [Google Scholar] [CrossRef]
- Boyer, M.; Wisniewski-Dye, F. Cell-cell signalling in bacteria: Not simply a matter of quorum. FEMS Microbiol. Ecol. 2009, 70, 1–19. [Google Scholar] [CrossRef]
- West, S.A.; Winzer, K.; Gardner, A.; Diggle, S.P. Quorum sensing and the confusion about diffusion. Trends Microbiol. 2012, 20, 586–594. [Google Scholar] [CrossRef]
- Decho, A.W.; Norman, R.S.; Visscher, P.T. Quorum sensing in natural environments: Emerging views from microbial mats. Trends Microbiol. 2010, 18, 73–80. [Google Scholar] [CrossRef]
- Boedicker, J.Q.; Vincent, M.E.; Ismagilov, R.F. Microfluidic confinement of single cells of bacteria in small volumes initiates high-density behavior of quorum sensing and growth and reveals its variability. Angew. Chem. Int. Ed. 2009, 48, 5908–5911. [Google Scholar] [CrossRef]
- Carnes, E.C.; Lopez, D.M.; Donegan, N.P.; Cheung, A.; Gresham, H.; Timmins, G.S.; Brinker, C.J. Confinement-induced quorum sensing of individual Staphylococcus aureus bacteria. Nat. Chem. Biol. 2010, 6, 41–45. [Google Scholar] [CrossRef]
- Schuster, M.; Sexton, D.J.; Diggle, S.P.; Greenberg, E.P. Acyl-homoserine lactone quorum sensing: From evolution to application. Annu. Rev. Microbiol. 2013, 67, 43–63. [Google Scholar] [CrossRef]
- Platt, T.G.; Fuqua, C. What’s in a name? The semantics of quorum sensing. Trends Microbiol. 2010, 18, 383–387. [Google Scholar] [CrossRef]
- Surette, M.G.; Miller, M.B.; Bassler, B.L. Quorum sensing in Escherichia coli, Salmonella typhimurium, and Vibrio harveyi: A new family of genes responsible for autoinducer production. Proc. Natl. Acad. Sci. USA 1999, 96, 1639–1644. [Google Scholar] [CrossRef]
- Pereira, C.S.; Thompson, J.A.; Xavier, K.B. AI-2-mediated signalling in bacteria. FEMS Microbiol. Rev. 2013, 37, 156–181. [Google Scholar]
- Rezzonico, F.; Duffy, B. Lack of genomic evidence of AI-2 receptors suggests a non-quorum sensing role for luxS in most bacteria. BMC Microbiol. 2008, 8, 154. [Google Scholar] [CrossRef]
- Pereira, C.S.; de Regt, A.K.; Brito, P.H.; Miller, S.T.; Xavier, K.B. Identification of functional LsrB-like autoinducer-2 receptors. J. Bacteriol. 2009, 191, 6975–6987. [Google Scholar] [CrossRef]
- Winzer, K.; Hardie, K.R.; Burgess, N.; Doherty, N.; Kirke, D.; Holden, M.T.; Linforth, R.; Cornell, K.A.; Taylor, A.J.; Hill, P.J.; et al. LuxS: Its role in central metabolism and the in vitro synthesis of 4-hydroxy-5-methyl-3(2H)-furanone. Microbiology 2002, 148, 909–922. [Google Scholar]
- Lee, J.H.; Lee, J. Indole as an intercellular signal in microbial communities. FEMS Microbiol. Rev. 2010, 34, 426–444. [Google Scholar]
- Biswa, P.; Doble, M. Production of acylated homoserine lactone by Gram-positive bacteria isolated from marine water. FEMS Microbiol. Lett. 2013, 343, 34–41. [Google Scholar] [CrossRef]
- Johnson, M.R.; Montero, C.I.; Conners, S.B.; Shockley, K.R.; Bridger, S.L.; Kelly, R.M. Population density-dependent regulation of exopolysaccharide formation in the hyperthermophilic bacterium Thermotoga maritima. Mol. Microbiol. 2005, 55, 664–674. [Google Scholar]
- Lee, H.; Chang, Y.C.; Nardone, G.; Kwon-Chung, K.J. TUP1 disruption in Cryptococcus neoformans uncovers a peptide-mediated density-dependent growth phenomenon that mimics quorum sensing. Mol. Microbiol. 2007, 64, 591–601. [Google Scholar] [CrossRef]
- Kelly, R.C.; Bolitho, M.E.; Higgins, D.A.; Lu, W.; Ng, W.-L.; Jeffrey, P.D.; Rabinowitz, J.D.; Semmelhack, M.F.; Hughson, F.M.; Bassler, B.L. The Vibrio cholerae quorum-sensing autoinducer CAI-1: Analysis of the biosynthetic enzyme CqsA. Nat. Chem. Biol. 2009, 5, 891–895. [Google Scholar] [CrossRef]
- Deng, Y.; Wu, J.; Tao, F.; Zhang, L.H. Listening to a new language: DSF-based quorum sensing in Gram-negative bacteria. Chem. Rev. 2011, 111, 160–173. [Google Scholar] [CrossRef]
- Chen, H.; Fujita, M.; Feng, Q.; Clardy, J.; Fink, G.R. Tyrosol is a quorum-sensing molecule in Candida albicans. Proc. Natl. Acad. Sci. USA 2004, 101, 5048–5052. [Google Scholar] [CrossRef]
- Hornby, J.M.; Jensen, E.C.; Lisec, A.D.; Tasto, J.J.; Jahnke, B.; Shoemaker, R.; Dussault, P.; Nickerson, K.W. Quorum sensing in the dimorphic fungus Candida albicans is mediated by farnesol. Appl. Environ. Microbiol. 2001, 67, 2982–2992. [Google Scholar] [CrossRef]
- Zhang, G.; Zhang, F.; Ding, G.; Li, J.; Guo, X.; Zhu, J.; Zhou, L.; Cai, S.; Liu, X.; Luo, Y.; et al. Acyl homoserine lactone-based quorum sensing in a methanogenic archaeon. ISME J. 2012, 6, 1336–1344. [Google Scholar] [CrossRef]
- Schaefer, A.L.; Greenberg, E.P.; Oliver, C.M.; Oda, Y.; Huang, J.J.; Bittan-Banin, G.; Peres, C.M.; Schmidt, S.; Juhaszova, K.; Sufrin, J.R.; et al. A new class of homoserine lactone quorum-sensing signals. Nature 2008, 454, 595–599. [Google Scholar] [CrossRef]
- Ahlgren, N.A.; Harwood, C.S.; Schaefer, A.L.; Giraud, E.; Greenberg, E.P. Aryl-homoserine lactone quorum sensing in stem-nodulating photosynthetic bradyrhizobia. Proc. Natl. Acad. Sci. USA 2011, 108, 7183–7188. [Google Scholar] [CrossRef]
- Lindemann, A.; Pessi, G.; Schaefer, A.L.; Mattmann, M.E.; Christensen, Q.H.; Kessler, A.; Hennecke, H.; Blackwell, H.E.; Greenberg, E.P.; Harwood, C.S. Isovaleryl-homoserine lactone, an unusual branched-chain quorum-sensing signal from the soybean symbiont Bradyrhizobium japonicum. Proc. Natl. Acad. Sci. USA 2011, 108, 16765–16770. [Google Scholar] [CrossRef]
- Gooding, J.R.; May, A.L.; Hilliard, K.R.; Campagna, S.R. Establishing a quantitative definition of quorum sensing provides insight into the information content of the autoinducer signals in Vibrio harveyi and Escherichia coli. Biochemistry 2010, 49, 5621–5623. [Google Scholar] [CrossRef]
- Saenz, H.L.; Augsburger, V.; Vuong, C.; Jack, R.W.; Gotz, F.; Otto, M. Inducible expression and cellular location of AgrB, a protein involved in the maturation of the staphylococcal quorum-sensing pheromone. Arch. Microbiol. 2000, 174, 452–455. [Google Scholar] [CrossRef]
- Lee, J.; Wu, J.; Deng, Y.; Wang, J.; Wang, C.; Wang, J.; Chang, C.; Dong, Y.; Williams, P.; Zhang, L.H. A cell-cell communication signal integrates quorum sensing and stress response. Nat. Chem. Biol. 2013, 9, 339–343. [Google Scholar] [CrossRef]
- Pesci, E.C.; Milbank, J.B.; Pearson, J.P.; McKnight, S.; Kende, A.S.; Greenberg, E.P.; Iglewski, B.H. Quinolone signaling in the cell-to-cell communication system of Pseudomonas aeruginosa. Proc. Natl. Acad. Sci. USA 1999, 96, 11229–11234. [Google Scholar] [CrossRef]
- Brachmann, A.O.; Brameyer, S.; Kresovic, D.; Hitkova, I.; Kopp, Y.; Manske, C.; Schubert, K.; Bode, H.B.; Heermann, R. Pyrones as bacterial signaling molecules. Nat. Chem. Biol. 2013, 9, 573–578. [Google Scholar] [CrossRef]
- Walters, M.; Sperandio, V. Autoinducer 3 and epinephrine signaling in the kinetics of locus of enterocyte effacement gene expression in enterohemorrhagic Escherichia coli. Infect. Immun. 2006, 74, 5445–5455. [Google Scholar] [CrossRef]
- Milton, D.L. Quorum sensing in vibrios: Complexity for diversification. Int. J. Med. Microbiol. 2006, 296, 61–71. [Google Scholar] [CrossRef]
- Henares, B.M.; Higgins, K.E.; Boon, E.M. Discovery of a nitric oxide responsive quorum sensing circuit in Vibrio harveyi. ACS Chem. Biol. 2012, 7, 1331–1336. [Google Scholar] [CrossRef]
- Lasarre, B.; Federle, M.J. Exploiting quorum sensing to confuse bacterial pathogens. Microbiol. Mol.Biol. Rev. 2013, 77, 73–111. [Google Scholar] [CrossRef]
- Yates, E.A.; Philipp, B.; Buckley, C.; Atkinson, S.; Chhabra, S.R.; Sockett, R.E.; Goldner, M.; Dessaux, Y.; Camara, M.; Smith, H.; et al. N-Acylhomoserine lactones undergo lactonolysis in a pH-, temperature-, and acyl chain length-dependent manner during growth of Yersinia pseudotuberculosis and Pseudomonas aeruginosa. Infect. Immun. 2002, 70, 5635–5646. [Google Scholar] [CrossRef]
- Taga, M.E.; Miller, S.T.; Bassler, B.L. Lsr-mediated transport and processing of AI-2 in Salmonella typhimurium. Mol. Microbiol. 2003, 50, 1411–1427. [Google Scholar] [CrossRef]
- Ng, W.L.; Perez, L.J.; Wei, Y.; Kraml, C.; Semmelhack, M.F.; Bassler, B.L. Signal production and detection specificity in Vibrio CqsA/CqsS quorum-sensing systems. Mol. Microbiol. 2011, 79, 1407–1417. [Google Scholar] [CrossRef]
- Tu, K.C.; Bassler, B.L. Multiple small RNAs act additively to integrate sensory information and control quorum sensing in Vibrio harveyi. Genes Dev. 2007, 21, 221–233. [Google Scholar] [CrossRef]
- Rutherford, S.T.; van Kessel, J.C.; Shao, Y.; Bassler, B.L. AphA and LuxR/HapR reciprocally control quorum sensing in vibrios. Genes Dev. 2011, 25, 397–408. [Google Scholar] [CrossRef]
- Van Kessel, J.C.; Rutherford, S.T.; Shao, Y.; Utria, A.F.; Bassler, B.L. Individual and combined roles of the master regulators AphA and LuxR in control of the Vibrio harveyi quorum-sensing regulon. J. Bacteriol. 2013, 195, 436–443. [Google Scholar] [CrossRef]
- Schauder, S.; Shokat, K.; Surette, M.G.; Bassler, B.L. The LuxS family of bacterial autoinducers: Biosynthesis of a novel quorum-sensing signal molecule. Mol. Microbiol. 2001, 41, 463–476. [Google Scholar] [CrossRef]
- Minogue, T.D.; Wehland-von Trebra, M.; Bernhard, F.; von Bodman, S.B. The autoregulatory role of EsaR, a quorum-sensing regulator in Pantoea stewartii ssp. stewartii: Evidence for a repressor function. Mol. Microbiol. 2002, 44, 1625–1635. [Google Scholar] [CrossRef]
- Dong, Y.H.; Wang, L.H.; Xu, J.L.; Zhang, H.B.; Zhang, X.F.; Zhang, L.H. Quenching quorum-sensing-dependent bacterial infection by an N-acyl homoserine lactonase. Nature 2001, 411, 813–817. [Google Scholar] [CrossRef]
- Eberhard, A.; Widrig, C.A.; McBath, P.; Schineller, J.B. Analogs of the autoinducer of bioluminescence in Vibrio fischeri. Arch. Microbiol. 1986, 146, 35–40. [Google Scholar] [CrossRef]
- Fuqua, C. The QscR quorum-sensing regulon of Pseudomonas aeruginosa: An orphan claims its identity. J. Bacteriol. 2006, 188, 3169–3171. [Google Scholar] [CrossRef]
- Kovacikova, G.; Skorupski, K. Regulation of virulence gene expression in Vibrio cholerae by quorum sensing: HapR functions at the aphA promoter. Mol. Microbiol. 2002, 46, 1135–1147. [Google Scholar] [CrossRef]
- Mattmann, M.E.; Shipway, P.M.; Heth, N.J.; Blackwell, H.E. Potent and selective synthetic modulators of a quorum sensing repressor in Pseudomonas aeruginosa identified from second-generation libraries of N-acylated L-homoserine lactones. ChemBioChem 2011, 12, 942–949. [Google Scholar] [CrossRef]
- Ng, W.L.; Perez, L.; Cong, J.; Semmelhack, M.F.; Bassler, B.L. Broad spectrum pro-quorum-sensing molecules as inhibitors of virulence in vibrios. PLoS Pathog. 2012, 8, e1002767. [Google Scholar] [CrossRef]
- Perez, L.J.; Karagounis, T.K.; Hurley, A.; Bassler, B.L.; Semmelhack, M.F. Highly potent, chemically stable quorum sensing agonists for Vibrio cholerae. Chem. Sci. 2014, 5, 151–155. [Google Scholar] [CrossRef]
- Clark, B.R.; Engene, N.; Teasdale, M.E.; Rowley, D.C.; Matainaho, T.; Valeriote, F.A.; Gerwick, W.H. Natural products chemistry and taxonomy of the marine cyanobacterium Blennothrix cantharidosmum. J. Nat. Prod. 2008, 71, 1530–1537. [Google Scholar] [CrossRef]
- Dobretsov, S.; Teplitski, M.; Alagely, A.; Gunasekera, S.P.; Paul, V.J. Malyngolide from the cyanobacterium Lyngbya majuscula interferes with quorum sensing circuitry. Environ. Microbiol. Rep. 2010, 2, 739–744. [Google Scholar] [CrossRef]
- Choi, H.; Mascuch, S.J.; Villa, F.A.; Byrum, T.; Teasdale, M.E.; Smith, J.E.; Preskitt, L.B.; Rowley, D.C.; Gerwick, L.; Gerwick, W.H. Honaucins A–C, potent inhibitors of inflammation and bacterial quorum sensing: Synthetic derivatives and structure-activity relationships. Chem. Biol. 2012, 19, 589–598. [Google Scholar] [CrossRef]
- Ooka, K.; Fukumoto, A.; Yamanaka, T.; Shimada, K.; Ishihara, R.; Anzai, Y.; Kato, F. Piericidins, novel quorum-sensing inhibitors against Chromobacterium violaceum CV026, from Streptomyces sp. TOHO-Y209 and TOHO-O348. Open J. Med. Chem. 2013, 3, 93–99. [Google Scholar]
- Teasdale, M.E.; Liu, J.Y.; Wallace, J.; Akhlaghi, F.; Rowley, D.C. Secondary metabolites produced by the marine bacterium Halobacillus salinus that inhibit quorum sensing-controlled phenotypes in Gram-negative bacteria. Appl. Environ. Microbiol. 2009, 75, 567–572. [Google Scholar]
- Teasdale, M.E.; Donovan, K.A.; Forschner-Dancause, S.R.; Rowley, D.C. Gram-positive marine bacteria as a potential resource for the discovery of quorum sensing inhibitors. Mar. Biotechnol. 2011, 13, 722–732. [Google Scholar] [CrossRef]
- Abed, R.M.; Dobretsov, S.; Al-Fori, M.; Gunasekera, S.P.; Sudesh, K.; Paul, V.J. Quorum-sensing inhibitory compounds from extremophilic microorganisms isolated from a hypersaline cyanobacterial mat. J. Ind. Microbiol. Biotechnol. 2013, 40, 759–772. [Google Scholar] [CrossRef]
- Peters, L.; König, G.M.; Wright, A.D.; Pukall, R.; Stackebrandt, E.; Eberl, L.; Riedel, K. Secondary metabolites of Flustra foliacea and their influence on bacteria. Appl. Environ. Microbiol. 2003, 69, 3469–3475. [Google Scholar] [CrossRef]
- Tello, E.; Castellanos, L.; Arevalo-Ferro, C.; Rodríguez, J.; Jiménez, C.; Duque, C. Absolute stereochemistry of antifouling cembranoid epimers at C-8 from the Caribbean octocoral Pseudoplexaura flagellosa. Revised structures of plexaurolones. Tetrahedron 2011, 67, 9112–9121. [Google Scholar] [CrossRef]
- Tello, E.; Castellanos, L.; Arevalo-Ferro, C.; Duque, C. Disruption in quorum-sensing systems and bacterial biofilm inhibition by cembranoid diterpenes isolated from the octocoral Eunicea knighti. J. Nat. Prod. 2012, 75, 1637–1642. [Google Scholar] [CrossRef]
- Kwan, J.C.; Teplitski, M.; Gunasekera, S.P.; Paul, V.J.; Luesch, H. Isolation and biological evaluation of 8-epi-malyngamide C from the Floridian marine cyanobacterium Lyngbya majuscula. J. Nat. Prod. 2010, 73, 463–466. [Google Scholar] [CrossRef]
- Kwan, J.C.; Meickle, T.; Ladwa, D.; Teplitski, M.; Paul, V.; Luesch, H. Lyngbyoic acid, a “tagged” fatty acid from a marine cyanobacterium, disrupts quorum sensing in Pseudomonas aeruginos. Mol. Biosyst. 2011, 7, 1205–1216. [Google Scholar] [CrossRef]
- Montaser, R.; Paul, V.J.; Luesch, H. Modular strategies for structure and function employed by marine cyanobacteria: Characterization and synthesis of pitinoic acids. Org. Lett. 2013, 15, 4050–4053. [Google Scholar] [CrossRef]
- Dobretsov, S.; Teplitski, M.; Bayer, M.; Gunasekera, S.; Proksch, P.; Paul, V.J. Inhibition of marine biofouling by bacterial quorum sensing inhibitors. Biofouling 2011, 27, 893–905. [Google Scholar] [CrossRef]
- Wang, L.; Zou, S.; Yin, S.; Liu, H.; Yu, W.; Gong, Q. Construction of an effective screening system for detection of Pseudomonas aeruginosa quorum sensing inhibitors and its application in bioautographic thin-layer chromatography. Biotechnol. Lett. 2011, 33, 1381–1387. [Google Scholar] [CrossRef]
- Liu, H.B.; Koh, K.P.; Kim, J.S.; Seo, Y.; Park, S. The effects of betonicine, floridoside, and isethionic acid from the red alga Ahnfeltiopsis flabelliformis on quorum-sensing activity. Biotechnol. Bioprocess Eng. 2008, 13, 458–463. [Google Scholar] [CrossRef]
- Skindersoe, M.E.; Ettinger-Epstein, P.; Rasmussen, T.B.; Bjarnsholt, T.; de Nys, R.; Givskov, M. Quorum sensing antagonism from marine organisms. Mar. Biotechnol. (N. Y.) 2008, 10, 56–63. [Google Scholar] [CrossRef]
- Chu, Y.Y.; Nega, M.; Wolfle, M.; Plener, L.; Grond, S.; Jung, K.; Gotz, F. A new class of quorum quenching molecules from Staphylococcus species affects communication and growth of Gram-negative bacteria. PLoS Pathog. 2013, 9, e1003654. [Google Scholar] [CrossRef]
- Singh, V.K.; Kavita, K.; Prabhakaran, R.; Jha, B. Cis-9-octadecenoic acid from the rhizospheric bacterium Stenotrophomonas maltophilia BJ01 shows quorum quenching and anti-biofilm activities. Biofouling 2013, 29, 855–867. [Google Scholar] [CrossRef]
- Bobadilla Fazzini, R.A.; Skindersoe, M.E.; Bielecki, P.; Puchalka, J.; Givskov, M.; Martins sos Santos, V.A. Protoanemonin: A natural quorum sensing inhibitor that selectively activates iron starvation response. Environ. Microbiol. 2013, 15, 111–120. [Google Scholar] [CrossRef]
- HjelmgaardRasmussen, T.B.; Skindersoe, M.E.; Bjarnsholt, T.; Phipps, R.K.; Christensen, K.B.; Jensen, P.O.; Andersen, J.B.; Koch, B.; Larsen, T.O.; Hentzer, M.; et al. Identity and effects of quorum-sensing inhibitors produced by Penicillium species. Microbiology 2005, 151, 1325–1340. [Google Scholar] [CrossRef]
- Truchado, P.; Lopez-Galvez, F.; Gil, M.I.; Tomas-Barberan, F.A.; Allende, A. Quorum sensing inhibitory and antimicrobial activities of honeys and the relationship with individual phenolics. Food Chem. 2009, 115, 1337–1344. [Google Scholar] [CrossRef]
- Bulman, Z.; Le, P.; Hudson, A.O.; Savka, M.A. A novel property of propolis (bee glue): Anti-pathogenic activity by inhibition of N-acyl-homoserine lactone mediated signaling in bacteria. J. Ethnopharmacol. 2011, 138, 788–797. [Google Scholar] [CrossRef]
- Park, J.; Kaufmann, G.F.; Bowen, J.P.; Arbiser, J.L.; Janda, K.D. Solenopsin A, a venom alkaloid from the fire ant Solenopsis invicta, inhibits quorum-sensing signaling in Pseudomonas aeruginosa. J. Infect. Dis. 2008, 198, 1198–1201. [Google Scholar] [CrossRef]
- Zhou, L.; Zheng, H.; Tang, Y.; Yu, W.; Gong, Q. Eugenol inhibits quorum sensing at sub-inhibitory concentrations. Biotechnol. Lett. 2013, 35, 631–637. [Google Scholar] [CrossRef]
- Keshavan, N.D.; Chowdhary, P.K.; Haines, D.C.; Gonzalez, J.E. l-canavanine made by Medicago sativa interferes with quorum sensing in Sinorhizobium meliloti. J. Bacteriol. 2005, 187, 8427–8436. [Google Scholar] [CrossRef]
- Vandeputte, O.M.; Kiendrebeogo, M.; Rajaonson, S.; Diallo, B.; Mol, A.; El Jaziri, M.; Baucher, M. Identification of catechin as one of the flavonoids from Combretum albiflorum bark extract that reduces the production of quorum-sensing-controlled virulence factors in Pseudomonas aeruginosa PAO1. Appl. Environ. Microbiol. 2010, 76, 243–253. [Google Scholar] [CrossRef]
- Vandeputte, O.M.; Kiendrebeogo, M.; Rasamiravaka, T.; Stevigny, C.; Duez, P.; Rajaonson, S.; Diallo, B.; Mol, A.; Baucher, M.; El Jaziri, M. The flavanone naringenin reduces the production of quorum sensing-controlled virulence factors in Pseudomonas aeruginosa PAO1. Microbiology 2011, 157, 2120–2132. [Google Scholar] [CrossRef]
- Amaya, S.; Pereira, J.A.; Borkosky, S.A.; Valdez, J.C.; Bardon, A.; Arena, M.E. Inhibition of quorum sensing in Pseudomonas aeruginosa by sesquiterpene lactones. Phytomedicine 2012, 19, 1173–1177. [Google Scholar] [CrossRef]
- Jakobsen, T.H.; van Gennip, M.; Phipps, R.K.; Shanmugham, M.S.; Christensen, L.D.; Alhede, M.; Skindersoe, M.E.; Rasmussen, T.B.; Friedrich, K.; Uthe, F.; et al. Ajoene, a sulfur-rich molecule from garlic, inhibits genes controlled by quorum sensing. Antimicrob. Agents Chemother. 2012, 56, 2314–2325. [Google Scholar] [CrossRef] [Green Version]
- Jakobsen, T.H.; Bragason, S.K.; Phipps, R.K.; Christensen, L.D.; van Gennip, M.; Alhede, M.; Skindersoe, M.; Larsen, T.O.; Hoiby, N.; Bjarnsholt, T.; et al. Food as a source for quorum sensing inhibitors: Iberin from horseradish revealed as a quorum sensing inhibitor of Pseudomonas aeruginosa. Appl. Environ. Microbiol. 2012, 78, 2410–2421. [Google Scholar] [CrossRef]
- Chong, Y.M.; Yin, W.F.; Ho, C.Y.; Mustafa, M.R.; Hadi, A.H.A.; Awang, K.; Narrima, P.; Koh, C.L.; Appleton, D.R.; Chan, K.G. Malabaricone C from Myristica cinnamomea exhibits anti-quorum sensing activity. J. Nat. Prod. 2011, 74, 2261–2264. [Google Scholar]
- Paza, C.; Carcamo, G.; Silva, M.; Becerra, J.; Urrutia, H.; Sossa, K. Drimendiol, a drimane sesquiterpene with quorum sensing inhibition activity. Nat. Prod. Commun. 2013, 8, 147–148. [Google Scholar]
- Packiavathy, I.A.; Priya, S.; Pandian, S.K.; Ravi, A.V. Inhibition of biofilm development of uropathogens by curcumin-an anti-quorum sensing agent from Curcuma longa. Food Chem. 2014, 148, 453–460. [Google Scholar] [CrossRef]
- Bodini, S.F.; Manfredini, S.; Epp, M.; Valentini, S.; Santori, F. Quorum sensing inhibition activity of garlic extract and p-coumaric acid. Lett. Appl. Microbiol. 2009, 49, 551–555. [Google Scholar] [CrossRef]
- Giménez-Bastida, J.A.; Truchado, P.; Larrosa, M.; Espín, J.C.; Tomás-Barberán, F.A.; Allende, A.; García-Conesa, M.T. Urolithins, ellagitannin metabolites produced by colon microbiota, inhibit quorum sensing in Yersinia enterocolitica: Phenotypic response and associated molecular changes. Food Chem. 2012, 132, 1465–1474. [Google Scholar]
- Adonizio, A.; Dawlaty, J.; Ausubel, F.M.; Clardy, J.; Mathee, K. Ellagitannins from Conocarpus erectus exhibit anti-quorum sensing activity against Pseudomonas aeruginosa. Planta Med. 2008, 74, 1035–1035. [Google Scholar]
- Kaplan, F.; Badri, D.V.; Zachariah, C.; Ajredini, R.; Sandoval, F.J.; Roje, S.; Levine, L.H.; Zhang, F.L.; Robinette, S.L.; Alborn, H.T.; et al. Bacterial attraction and quorum sensing inhibition in Caenorhabditis elegans exudates. J. Chem. Ecol. 2009, 35, 878–892. [Google Scholar] [CrossRef]
- Teplitski, M.; Chen, H.; Rajamani, S.; Gao, M.; Merighi, M.; Sayre, R.T.; Robinson, J.B.; Rolfe, B.G.; Bauer, W.D. Chlamydomonas reinhardtii secretes compounds that mimic bacterial signals and interfere with quorum sensing regulation in bacteria. Plant Physiol. 2004, 134, 137–146. [Google Scholar] [CrossRef]
- Rajamani, S.; Bauer, W.D.; Robinson, J.B.; Farrow, J.M.; Pesci, E.C.; Teplitski, M.; Gao, M.S.; Sayre, R.T.; Phillips, D.A. The vitamin riboflavin and its derivative lumichrome activate the LasR bacterial quorum-sensing receptor. Mol. Plant-Microbe Interact. 2008, 21, 1184–1192. [Google Scholar] [CrossRef]
- Ding, X.; Yin, B.; Qian, L.; Zeng, Z.R.; Yang, Z.L.; Li, H.X.; Lu, Y.J.; Zhou, S.N. Screening for novel quorum-sensing inhibitors to interfere with the formation of Pseudomonas aeruginosa biofilm. J. Med. Microbiol. 2011, 60, 1827–1834. [Google Scholar] [CrossRef]
- Zeng, Z.; Qian, L.; Cao, L.; Tan, H.; Huang, Y.; Xue, X.; Shen, Y.; Zhou, S. Virtual screening for novel quorum sensing inhibitors to eradicate biofilm formation of Pseudomonas aeruginosa. Appl. Microbiol. Biotechnol. 2008, 79, 119–126. [Google Scholar] [CrossRef]
- Cugini, C.; Calfee, M.W.; Farrow, J.M., 3rd.; Morales, D.K.; Pesci, E.C.; Hogan, D.A. Farnesol, a common sesquiterpene, inhibits PQS production in Pseudomonas aeruginosa. Mol. Microbiol. 2007, 65, 896–906. [Google Scholar] [CrossRef]
- De Nys, R.; Wright, A.D.; König, G.M.; Sticher, O. New halogenated furanones from the marine alga Delisea pulchra (cf. fimbriata). Tetrahedron 1993, 49, 11213–11220. [Google Scholar] [CrossRef]
- Hentzer, M.; Riedel, K.; Rasmussen, T.B.; Heydorn, A.; Andersen, J.B.; Parsek, M.R.; Rice, S.A.; Eberl, L.; Molin, S.; Hoiby, N.; et al. Inhibition of quorum sensing in Pseudomonas aeruginosa biofilm bacteria by a halogenated furanone compound. Microbiology 2002, 148, 87–102. [Google Scholar]
- Brackman, G.; Celen, S.; Hillaert, U.; van Calenbergh, S.; Cos, P.; Maes, L.; Nelis, H.J.; Coenye, T. Structure-activity relationship of cinnamaldehyde analogs as inhibitors of AI-2 based quorum sensing and their effect on virulence of Vibrio spp. PLoS One 2011, 6, e16084. [Google Scholar] [CrossRef] [Green Version]
- Niu, C.; Afre, S.; Gilbert, E.S. Subinhibitory concentrations of cinnamaldehyde interfere with quorum sensing. Lett. Appl. Microbiol. 2006, 43, 489–494. [Google Scholar] [CrossRef]
- Brackman, G.; Defoirdt, T.; Miyamoto, C.; Bossier, P.; van Calenbergh, S.; Nelis, H.; Coenye, T. Cinnamaldehyde and cinnamaldehyde derivatives reduce virulence in Vibrio spp. by decreasing the DNA-binding activity of the quorum sensing response regulator LuxR. BMC Microbiol. 2008, 8, 149. [Google Scholar] [CrossRef]
- Lee, K.M.; Lim, J.; Nam, S.; Yoon, M.Y.; Kwon, Y.K.; Jung, B.Y.; Park, Y.; Park, S.; Yoon, S.S. Inhibitory effects of broccoli extract on Escherichia coli O157:H7 quorum sensing and in vivo virulence. FEMS Microbiol. Lett. 2011, 321, 67–74. [Google Scholar] [CrossRef]
- Vikram, A.; Jesudhasan, P.R.; Jayaprakasha, G.K.; Pillai, B.S.; Patil, B.S. Grapefruit bioactive limonoids modulate E. coli O157:H7 TTSS and biofilm. Int. J. Food Microbiol. 2010, 140, 109–116. [Google Scholar] [CrossRef]
- Li, J.; Wang, W.; Xu, S.X.; Magarvey, N.A.; McCormick, J.K. Lactobacillus reuteri-produced cyclic dipeptides quench agr-mediated expression of toxic shock syndrome toxin-1 in staphylococci. Proc. Natl. Acad. Sci. USA 2011, 108, 3360–3365. [Google Scholar]
- Mansson, M.; Nielsen, A.; Kjaerulff, L.; Gotfredsen, C.H.; Wietz, M.; Ingmer, H.; Gram, L.; Larsen, T.O. Inhibition of virulence gene expression in Staphylococcus aureus by novel depsipeptides from a marine Photobacterium. Mar. Drugs 2011, 9, 2537–2552. [Google Scholar] [CrossRef]
- Kiran, M.D.; Adikesavan, N.V.; Cirioni, O.; Giacometti, A.; Silvestri, C.; Scalise, G.; Ghiselli, R.; Saba, V.; Orlando, F.; Shoham, M.; et al. Discovery of a quorum-sensing inhibitor of drug-resistant staphylococcal infections by structure-based virtual screening. Mol. Pharmacol. 2008, 73, 1578–1586. [Google Scholar] [CrossRef]
- Duncan, M.C.; Wong, W.R.; Dupzyk, A.J.; Bray, W.M.; Linington, R.G.; Auerbuch, V. An NF-kappaB-based high-throughput screen identifies piericidins as inhibitors of the Yersinia pseudotuberculosis type III secretion system. Antimicrob. Agents Chemother. 2014, 58, 1118–1126. [Google Scholar] [CrossRef]
- De Nys, R.; Steinberg, P.; Willemsen, P.; Dworjanyn, S.; Gabelish, C.; King, R. Broad spectrum effects of secondary metabolites from the red alga Delisea pulchra in antifouling assays. Biofouling 1995, 8, 259–271. [Google Scholar] [CrossRef]
- Martinelli, D.; Grossmann, G.; Sequin, U.; Brandl, H.; Bachofen, R. Effects of natural and chemically synthesized furanones on quorum sensing in Chromobacterium violaceum. BMC Microbiol. 2004, 4, 25. [Google Scholar] [CrossRef] [Green Version]
- Manefield, M.; Rasmussen, T.B.; Henzter, M.; Andersen, J.B.; Steinberg, P.; Kjelleberg, S.; Givskov, M. Halogenated furanones inhibit quorum sensing through accelerated LuxR turnover. Microbiology 2002, 148, 1119–1127. [Google Scholar]
- Defoirdt, T.; Miyamoto, C.M.; Wood, T.K.; Meighen, E.A.; Sorgeloos, P.; Verstraete, W.; Bossier, P. The natural furanone (5Z)-4-bromo-5-(bromomethylene)-3-butyl-2(5H)-furanone disrupts quorum sensing-regulated gene expression in Vibrio harveyi by decreasing the DNA-binding activity of the transcriptional regulator protein luxR. Environ. Microbiol. 2007, 9, 2486–2495. [Google Scholar] [CrossRef]
- Zang, T.; Lee, B.W.; Cannon, L.M.; Ritter, K.A.; Dai, S.; Ren, D.; Wood, T.K.; Zhou, Z.S. A naturally occurring brominated furanone covalently modifies and inactivates LuxS. Bioorg. Med. Chem. Lett. 2009, 19, 6200–6204. [Google Scholar] [CrossRef]
- Defoirdt, T.; Crab, R.; Wood, T.K.; Sorgeloos, P.; Verstraete, W.; Bossier, P. Quorum sensing-disrupting brominated furanones protect the gnotobiotic brine shrimp Artemia franciscana from pathogenic Vibrio harveyi, Vibrio campbellii, and Vibrio parahaemolyticus isolates. Appl. Environ. Microbiol. 2006, 72, 6419–6423. [Google Scholar] [CrossRef] [Green Version]
- Rasch, M.; Buch, C.; Austin, B.; Slierendrecht, W.J.; Ekmann, K.S.; Larsen, J.L.; Johansen, C.; Riedel, K.; Eberl, L.; Givskov, M.; et al. An inhibitor of bacterial quorum sensing reduces mortalities caused by vibriosis in rainbow trout (Oncorhynchus mykiss, Walbaum). Syst. Appl. Microbiol. 2004, 27, 350–359. [Google Scholar] [CrossRef]
- Tinh, N.T.N.; Linh, N.D.; Wood, T.K.; Dierckens, K.; Sorgeloos, P.; Bossier, P. Interference with the quorum sensing systems in a Vibrio harveyi strain alters the growth rate of gnotobiotically cultured rotifer Brachionus plicatilis. J. Appl. Microbiol. 2007, 103, 194–203. [Google Scholar] [CrossRef]
- Wu, H.; Song, Z.; Hentzer, M.; Andersen, J.B.; Molin, S.; Givskov, M.; Hoiby, N. Synthetic furanones inhibit quorum-sensing and enhance bacterial clearance in Pseudomonas aeruginosa lung infection in mice. J. Antimicrob. Chemother. 2004, 53, 1054–1061. [Google Scholar] [CrossRef]
- Natrah, F.M.I.; Kenmegne, M.M.; Wiyoto, W.; Sorgeloos, P.; Bossier, P.; Defoirdt, T. Effects of micro-algae commonly used in aquaculture on acyl-homoserine lactone quorum sensing. Aquaculture 2011, 317, 53–57. [Google Scholar] [CrossRef]
- Havsteen, B.H. The biochemistry and medical significance of the flavonoids. Pharmacol. Ther. 2002, 96, 67–202. [Google Scholar] [CrossRef]
- Vikram, A.; Jayaprakasha, G.K.; Jesudhasan, P.R.; Pillai, S.D.; Patil, B.S. Suppression of bacterial cell-cell signalling, biofilm formation and type III secretion system by citrus flavonoids. J. Appl. Microbiol. 2010, 109, 515–527. [Google Scholar]
- Pande, G.S.J.; Scheie, A.A.; Benneche, T.; Wille, M.; Sorgeloos, P.; Bossier, P.; Defoirdt, T. Quorum sensing-disrupting compounds protect larvae of the giant freshwater prawn Macrobrachium rosenbergii from Vibrio harveyi infection. Aquaculture 2013, 406–407, 121–124. [Google Scholar] [CrossRef]
- Defoirdt, T.; Brackman, G.; Coenye, T. Quorum sensing inhibitors: How strong is the evidence? Trends Microbiol. 2013, 21, 619–624. [Google Scholar] [CrossRef]
- Ni, N.; Choudhary, G.; Li, M.; Wang, B. Pyrogallol and its analogs can antagonize bacterial quorum sensing in Vibrio harveyi. Bioorg. Med. Chem. Lett. 2008, 18, 1567–1572. [Google Scholar] [CrossRef]
- Defoirdt, T.; Pande, G.S.; Baruah, K.; Bossier, P. The apparent quorum-sensing inhibitory activity of pyrogallol is a side effect of peroxide production. Antimicrob. Agents Chemother. 2013, 57, 2870–2873. [Google Scholar] [CrossRef]
- Newman, K.L.; Chatterjee, S.; Ho, K.A.; Lindow, S.E. Virulence of plant pathogenic bacteria attenuated by degradation of fatty acid cell-to-cell signaling factors. Mol. Plant Microbe Interact. 2008, 21, 326–334. [Google Scholar] [CrossRef]
- Pustelny, C.; Albers, A.; Buldt-Karentzopoulos, K.; Parschat, K.; Chhabra, S.R.; Camara, M.; Williams, P.; Fetzner, S. Dioxygenase-mediated quenching of quinolone-dependent quorum sensing in Pseudomonas aeruginosa. Chem. Biol. 2009, 16, 1259–1267. [Google Scholar] [CrossRef]
- Roy, V.; Fernandes, R.; Tsao, C.Y.; Bentley, W.E. Cross species quorum quenching using a native AI-2 processing enzyme. ACS Chem. Biol. 2010, 5, 223–232. [Google Scholar] [CrossRef]
- Draganov, D.I.; Teiber, J.F.; Speelman, A.; Osawa, Y.; Sunahara, R.; La Du, B.N. Human paraoxonases (PON1, PON2, and PON3) are lactonases with overlapping and distinct substrate specificities. J. Lipid Res. 2005, 46, 1239–1247. [Google Scholar] [CrossRef]
- Tang, K.; Zhang, Y.; Yu, M.; Shi, X.; Coenye, T.; Bossier, P.; Zhang, X.H. Evaluation of a new high-throughput method for identifying quorum quenching bacteria. Sci. Rep. 2013, 3, 2935. [Google Scholar] [Green Version]
- Zhang, X.H.; Ocean University of China, Qingdao, China. MomL, a novel marine-derived N-acyl homeserine lactonase in Muricauda olearia. 2014; Unpublished work. [Google Scholar]
- Dong, Y.H.; Xu, J.L.; Li, X.Z.; Zhang, L.H. AiiA, an enzyme that inactivates the acylhomoserine lactone quorum-sensing signal and attenuates the virulence of Erwinia carotovora. Proc. Natl. Acad. Sci. USA 2000, 97, 3526–3531. [Google Scholar] [CrossRef]
- Thomas, P.W.; Stone, E.M.; Costello, A.L.; Tierney, D.L.; Fast, W. The quorum-quenching lactonase from Bacillus thuringiensis is a metalloprotein. Biochemistry 2005, 44, 7559–7569. [Google Scholar] [CrossRef]
- Kim, M.H.; Choi, W.C.; Kang, H.O.; Lee, J.S.; Kang, B.S.; Kim, K.J.; Derewenda, Z.S.; Oh, T.K.; Lee, C.H.; Lee, J.K. The molecular structure and catalytic mechanism of a quorum-quenching N-acyl-L-homoserine lactone hydrolase. Proc. Natl. Acad. Sci. USA 2005, 102, 17606–17611. [Google Scholar] [CrossRef]
- Liu, D.; Momb, J.; Thomas, P.W.; Moulin, A.; Petsko, G.A.; Fast, W.; Ringe, D. Mechanism of the quorum-quenching lactonase (AiiA) from Bacillus thuringiensis. 1. product-bound structures. Biochemistry 2008, 47, 7706–7714. [Google Scholar] [CrossRef]
- Momb, J.; Wang, C.; Liu, D.; Thomas, P.W.; Petsko, G.A.; Guo, H.; Ringe, D.; Fast, W. Mechanism of the quorum-quenching lactonase (AiiA) from Bacillus thuringiensis. 2. substrate modeling and active site mutations. Biochemistry 2008, 47, 7715–7725. [Google Scholar] [CrossRef]
- Liu, C.F.; Liu, D.; Momb, J.; Thomas, P.W.; Lajoie, A.; Petsko, G.A.; Fast, W.; Ringe, D. A phenylalanine clamp controls substrate specificity in the quorum-quenching metallo-gamma-lactonase from Bacillus thuringiensis. Biochemistry 2013, 52, 1603–1610. [Google Scholar] [CrossRef]
- Momb, J.; Thomas, P.W.; Breece, R.M.; Tierney, D.L.; Fast, W. The quorum-quenching metallo-gamma-lactonase from Bacillus thuringiensis exhibits a leaving group thio effect. Biochemistry 2006, 45, 13385–13393. [Google Scholar] [CrossRef]
- Carlier, A.; Uroz, S.; Smadja, B.; Fray, R.; Latour, X.; Dessaux, Y.; Faure, D. The Ti plasmid of Agrobacterium tumefaciens harbors an attM-paralogous gene, aiiB, also encoding N-acyl homoserine lactonase activity. Appl. Environ. Microbiol. 2003, 69, 4989–4993. [Google Scholar] [CrossRef]
- Liu, D.; Thomas, P.W.; Momb, J.; Hoang, Q.Q.; Petsko, G.A.; Ringe, D.; Fast, W. Structure and specificity of a quorum-quenching lactonase (AiiB) from Agrobacterium tumefaciens. Biochemistry 2007, 46, 11789–11799. [Google Scholar] [CrossRef]
- Piper, K.R.; Beck von Bodman, S.; Farrand, S.K. Conjugation factor of Agrobacterium tumefaciens regulates Ti plasmid transfer by autoinduction. Nature 1993, 362, 448–450. [Google Scholar] [CrossRef]
- Haudecoeur, E.; Planamente, S.; Cirou, A.; Tannieres, M.; Shelp, B.J.; Morera, S.; Faure, D. Proline antagonizes GABA-induced quenching of quorum-sensing in Agrobacterium tumefaciens. Proc. Natl. Acad. Sci. USA 2009, 106, 14587–14592. [Google Scholar] [CrossRef]
- Wang, W.Z.; Morohoshi, T.; Someya, N.; Ikeda, T. AidC, a novel N-acylhomoserine lactonase from the potato root-associated cytophaga-flavobacteria-bacteroides (CFB) group bacterium Chryseobacterium sp. strain StRB126. Appl. Environ. Microbiol. 2012, 78, 7985–7992. [Google Scholar] [CrossRef]
- Krysciak, D.; Schmeisser, C.; Preuss, S.; Riethausen, J.; Quitschau, M.; Grond, S.; Streit, W.R. Involvement of multiple loci in quorum quenching of autoinducer I molecules in the nitrogen-fixing symbiont Rhizobium (Sinorhizobium) sp. strain StRB126. Appl. Environ. Microbiol. 2011, 77, 5089–5099. [Google Scholar]
- Schneider, J.; Yepes, A.; Garcia-Betancur, J.C.; Westedt, I.; Mielich, B.; Lopez, D. Streptomycin-induced expression in Bacillus subtilis of YtnP, a lactonase-homologous protein that inhibits development and streptomycin production in Streptomyces griseus. Appl. Environ. Microbiol. 2012, 78, 599–603. [Google Scholar] [CrossRef]
- Afriat, L.; Roodveldt, C.; Manco, G.; Tawfik, D.S. The latent promiscuity of newly identified microbial lactonases is linked to a recently diverged phosphotriesterase. Biochemistry 2006, 45, 13677–13686. [Google Scholar] [CrossRef]
- Elias, M.; Dupuy, J.; Merone, L.; Mandrich, L.; Porzio, E.; Moniot, S.; Rochu, D.; Lecomte, C.; Rossi, M.; Masson, P.; et al. Structural basis for natural lactonase and promiscuous phosphotriesterase activities. J. Mol. Biol. 2008, 379, 1017–1028. [Google Scholar]
- Ng, F.S.; Wright, D.M.; Seah, S.Y. Characterization of a phosphotriesterase-like lactonase from Sulfolobus solfataricus and its immobilization for disruption of quorum sensing. Appl. Environ. Microbiol. 2011, 77, 1181–1186. [Google Scholar] [CrossRef]
- Xue, B.; Chow, J.Y.; Baldansuren, A.; Yap, L.L.; Gan, Y.H.; Dikanov, S.A.; Robinson, R.C.; Yew, W.S. Structural evidence of a productive active site architecture for an evolved quorum-quenching GKL lactonase. Biochemistry 2013, 52, 2359–2370. [Google Scholar] [CrossRef]
- Hiblot, J.; Gotthard, G.; Chabriere, E.; Elias, M. Structural and enzymatic characterization of the lactonase SisLac from Sulfolobus islandicus. PLoS One 2012, 7, e47028. [Google Scholar]
- Mei, G.Y.; Yan, X.X.; Turak, A.; Luo, Z.Q.; Zhang, L.Q. AidH, an alpha/beta-hydrolase fold family member from an Ochrobactrum sp. strain, is a novel N-acylhomoserine lactonase. Appl. Environ. Microbiol. 2010, 76, 4933–4942. [Google Scholar] [CrossRef]
- Wang, W.Z.; Morohoshi, T.; Ikenoya, M.; Someya, N.; Ikeda, T. AiiM, a novel class of N-acylhomoserine lactonase from the leaf-associated bacterium Microbacterium testaceum. Appl. Environ. Microbiol. 2010, 76, 2524–2530. [Google Scholar] [CrossRef]
- Lin, Y.H.; Xu, J.L.; Hu, J.; Wang, L.H.; Ong, S.L.; Leadbetter, J.R.; Zhang, L.H. Acyl-homoserine lactone acylase from Ralstonia strain XJ12B represents a novel and potent class of quorum-quenching enzymes. Mol. Microbiol. 2003, 47, 849–860. [Google Scholar] [CrossRef]
- Leadbetter, J.R.; Greenberg, E.P. Metabolism of acyl-homoserine lactone quorum-sensing signals by Variovorax paradoxus. J. Bacteriol. 2000, 182, 6921–6926. [Google Scholar]
- Park, S.Y.; Kang, H.O.; Jang, H.S.; Lee, J.K.; Koo, B.T.; Yum, D.Y. Identification of extracellular N-acylhomoserine lactone acylase from a Streptomyces sp. and its application to quorum quenching. Appl. Environ. Microbiol. 2005, 71, 2632–2641. [Google Scholar] [CrossRef]
- Han, J.I.; Choi, H.K.; Lee, S.W.; Orwin, P.M.; Kim, J.; Laroe, S.L.; Kim, T.G.; O’Neil, J.; Leadbetter, J.R.; Lee, S.Y.; et al. Complete genome sequence of the metabolically versatile plant growth-promoting endophyte Variovorax paradoxus S110. J. Bacteriol. 2011, 193, 1183–1190. [Google Scholar] [CrossRef]
- Sio, C.F.; Otten, L.G.; Cool, R.H.; Diggle, S.P.; Braun, P.G.; Bos, R.; Daykin, M.; Cámara, M.; Williams, P.; Quax, W.J. Quorum quenching by an N-acyl-homoserine lactone acylase from Pseudomonas aeruginosa PAO1. Infect. Immun. 2006, 74, 1673–1682. [Google Scholar] [CrossRef]
- Huang, J.J.; Petersen, A.; Whiteley, M.; Leadbetter, J.R. Identification of QuiP, the product of gene PA1032, as the second acyl-homoserine lactone acylase of Pseudomonas aeruginosa PAO1. Appl. Environ. Microbiol. 2006, 72, 1190–1197. [Google Scholar] [CrossRef]
- Wahjudi, M.; Papaioannou, E.; Hendrawati, O.; van Assen, A.H.G.; van Merkerk, R.; Cool, R.H.; Poelarends, G.J.; Quax, W.J. PA0305 of Pseudomonas aeruginosa is a quorum quenching acylhomoserine lactone acylase belonging to the Ntn hydrolase superfamily. Microbiology 2011, 157, 2042–2055. [Google Scholar] [CrossRef]
- Nadal Jimenez, P.; Koch, G.; Papaioannou, E.; Wahjudi, M.; Krzeslak, J.; Coenye, T.; Cool, R.H.; Quax, W.J. Role of PvdQ in Pseudomonas aeruginosa virulence under iron-limiting conditions. Microbiology 2010, 156, 49–59. [Google Scholar] [CrossRef]
- Clevenger, K.D.; Wu, R.; Er, J.A.; Liu, D.; Fast, W. Rational design of a transition state analogue with picomolar affinity for Pseudomonas aeruginosa PvdQ, a siderophore biosynthetic enzyme. ACS Chem. Biol. 2013, 8, 2192–2200. [Google Scholar] [CrossRef]
- Czajkowski, R.; Krzyżanowska, D.; Karczewska, J.; Atkinson, S.; Przysowa, J.; Lojkowska, E.; Williams, P.; Jafra, S. Inactivation of AHLs by Ochrobactrum sp. A44 depends on the activity of a novel class of AHL acylase. Environ. Microbiol. Rep. 2011, 3, 59–68. [Google Scholar] [CrossRef]
- Tannieres, M.; Beury-Cirou, A.; Vigouroux, A.; Mondy, S.; Pellissier, F.; Dessaux, Y.; Faure, D. A metagenomic study highlights phylogenetic proximity of quorum-quenching and xenobiotic-degrading amidases of the AS-family. PLoS One 2013, 8, e65473. [Google Scholar]
- Uroz, S.; Chhabra, S.R.; Camara, M.; Williams, P.; Oger, P.; Dessaux, Y. N-Acylhomoserine lactone quorum-sensing molecules are modified and degraded by Rhodococcus erythropolis W2 by both amidolytic and novel oxidoreductase activities. Microbiology 2005, 151, 3313–3322. [Google Scholar] [CrossRef]
- Chowdhary, P.K.; Keshavan, N.; Nguyen, H.Q.; Peterson, J.A.; Gonzalez, J.E.; Haines, D.C. Bacillus megaterium CYP102A1 oxidation of acyl homoserine lactones and acyl homoserines. Biochemistry 2007, 46, 14429–14437. [Google Scholar] [CrossRef]
- Bijtenhoorn, P.; Mayerhofer, H.; Muller-Dieckmann, J.; Utpatel, C.; Schipper, C.; Hornung, C.; Szesny, M.; Grond, S.; Thurmer, A.; Brzuszkiewicz, E.; et al. A novel metagenomic short-chain dehydrogenase/reductase attenuates Pseudomonas aeruginosa biofilm formation and virulence on Caenorhabditis elegans. PLoS One 2011, 6, e26278. [Google Scholar] [CrossRef]
- Chan, K.G.; Atkinson, S.; Mathee, K.; Sam, C.K.; Chhabra, S.R.; Camara, M.; Koh, C.L.; Williams, P. Characterization of N-acylhomoserine lactone-degrading bacteria associated with the Zingiber officinale (ginger) rhizosphere: Co-existence of quorum quenching and quorum sensing in Acinetobacter and Burkholderia. BMC Microbiol. 2011, 11, 51. [Google Scholar] [CrossRef]
- Hong, K.-W.; Koh, C.-L.; Sam, C.-K.; Yin, W.-F.; Chan, K.-G. Complete genome sequence of Burkholderia sp. strain GG4, a Betaproteobacterium that reduces 3-oxo-N-acylhomoserine lactones and produces different N-acylhomoserine lactones. J. Bacteriol. 2012, 194, 6317. [Google Scholar] [CrossRef]
- Sandy, M.; Carter-Franklin, J.N.; Martin, J.D.; Butler, A. Vanadium bromoperoxidase from Delisea pulchra: Enzyme-catalyzed formation of bromofuranone and attendant disruption of quorum sensing. Chem. Commun. 2011, 47, 12086–12088. [Google Scholar] [CrossRef]
- Borchardt, S.A.; Allain, E.J.; Michels, J.J.; Stearns, G.W.; Kelly, R.F.; McCoy, W.F. Reaction of acylated homoserine lactone bacterial signaling molecules with oxidized halogen antimicrobials. Appl. Environ. Microbiol. 2001, 67, 3174–3179. [Google Scholar] [CrossRef]
- Syrpas, M.; Ruysbergh, E.; Blommaert, L.; Vanelslander, B.; Sabbe, K.; Vyverman, W.; de Kimpe, N.; Mangelinckx, S. Haloperoxidase mediated quorum quenching by Nitzschia cf pellucida: Study of the metabolization of N-acyl homoserine lactones by a benthic diatom. Mar. Drugs 2014, 12, 352–367. [Google Scholar] [CrossRef] [Green Version]
- Hibbing, M.E.; Fuqua, C.; Parsek, M.R.; Peterson, S.B. Bacterial competition: Surviving and thriving in the microbial jungle. Nat. Rev. Microbiol. 2010, 8, 15–25. [Google Scholar] [CrossRef]
- D’Angelo-Picard, C.; Faure, D.; Penot, I.; Dessaux, Y. Diversity of N-acyl homoserine lactone-producing and -degrading bacteria in soil and tobacco rhizosphere. Environ. Microbiol. 2005, 7, 1796–1808. [Google Scholar] [CrossRef]
- Chemistry, A.; July, R.; Sepetember, A.; Shewanella, I. N-acyl homoserine lactone-producing or -degrading bacteria Isolated from the intestinal microbial flora of Ayu fish (Plecoglossus altivelis). ASM News 2005, 20, 264–268. [Google Scholar]
- Biswas, L.; Biswas, R.; Schlag, M.; Bertram, R.; Götz, F. Small-colony variant selection as a survival strategy for Staphylococcus aureus in the presence of Pseudomonas aeruginosa. Appl. Environ. Microbiol. 2009, 75, 6910–6912. [Google Scholar] [CrossRef]
- Kerr, J.; Taylor, G.; Rutman, A.; Høiby, N.; Cole, P.; Wilson, R. Pseudomonas aeruginosa pyocyanin and 1-hydroxyphenazine inhibit fungal growth. J. Clin. Pathol. 1999, 52, 385–387. [Google Scholar] [CrossRef]
- McClean, K.H.; Winson, M.K.; Fish, L.; Taylor, A.; Chhabra, S.R.; Camara, M.; Daykin, M.; Lamb, J.H.; Swift, S.; Bycroft, B.W.; et al. Quorum sensing and Chromobacterium violaceum: Exploitation of violacein production and inhibition for the detection of N-acylhomoserine lactones. Microbiology 1997, 143, 3703–3711. [Google Scholar] [CrossRef]
- Morohoshi, T.; Kato, M.; Fukamachi, K.; Kato, N.; Ikeda, T. N-acylhomoserine lactone regulates violacein production in Chromobacterium violaceum type strain ATCC 12472. FEMS Microbiol. Lett. 2008, 279, 124–130. [Google Scholar] [CrossRef]
- Lyon, G.J.; Wright, J.S.; Muir, T.W.; Novick, R.P. Key determinants of receptor activation in the agr autoinducing peptides of Staphylococcus aureus. Biochemistry 2002, 41, 10095–10104. [Google Scholar] [CrossRef]
- Dufour, P.; Jarraud, S.; Vandenesch, F.; Greenland, T.; Novick, R.P.; Bes, M.; Etienne, J.; Lina, G. High genetic variability of the agr locus in Staphylococcus species. J. Bacteriol. 2002, 184, 1180–1186. [Google Scholar] [CrossRef]
- Jarraud, S.; Lyon, G.J.; Figueiredo, A.M.; Lina, G.; Vandenesch, F.; Etienne, J.; Muir, T.W.; Novick, R.P. Exfoliatin-producing strains define a fourth agr specificity group in Staphylococcus aureus. J. Bacteriol. 2000, 182, 6517–6522. [Google Scholar]
- Ji, G.; Beavis, R.; Novick, R.P. Bacterial interference caused by autoinducing peptide variants. Science 1997, 276, 2027–2030. [Google Scholar] [CrossRef]
- Hong, K.W.; Koh, C.L.; Sam, C.K.; Yin, W.F.; Chan, K.G. Quorum quenching revisited-from signal decays to signalling confusion. Sensors (Basel) 2012, 12, 4661–4696. [Google Scholar] [CrossRef]
- Czajkowski, R.; Jafra, S. Quenching of acyl-homoserine lactone-dependent quorum sensing by enzymatic disruption of signal molecules. Acta Biochim. Pol. 2009, 56, 1–16. [Google Scholar]
- Romero, M.; Martin-Cuadrado, A.B.; Otero, A. Determination of whether quorum quenching is a common activity in marine bacteria by analysis of cultivable bacteria and metagenomic sequences. Appl. Environ. Microbiol. 2012, 78, 6345–6348. [Google Scholar] [CrossRef]
- Khan, S.R.; Farrand, S.K. The BlcC (AttM) lactonase of Agrobacterium tumefaciens does not quench the quorum-sensing system that regulates Ti plasmid conjugative transfer. J. Bacteriol. 2009, 191, 1320–1329. [Google Scholar] [CrossRef]
- Flagan, S.; Ching, W.K.; Leadbetter, J.R. Arthrobacter strain VAI-A utilizes acyl-homoserine lactone inactivation products and stimulates quorum signal biodegradation by Variovorax paradoxus. Appl. Environ. Microbiol. 2003, 69, 909–916. [Google Scholar] [CrossRef]
- Park, S.J.; Park, S.Y.; Ryu, C.M.; Park, S.H.; Lee, J.K. The role of AiiA, a quorum-quenching enzyme from Bacillus thuringiensis, on the rhizosphere competence. J. Microbiol. Biotechnol. 2008, 18, 1518–1521. [Google Scholar]
- Romero, M.; Martin-Cuadrado, A.B.; Roca-Rivada, A.; Cabello, A.M.; Otero, A. Quorum quenching in cultivable bacteria from dense marine coastal microbial communities. FEMS Microbiol. Ecol. 2011, 75, 205–217. [Google Scholar] [CrossRef]
- Torres, M.; Romero, M.; Prado, S.; Dubert, J.; Tahrioui, A.; Otero, A.; Llamas, I. N-acylhomoserine lactone-degrading bacteria isolated from hatchery bivalve larval cultures. Microbiol. Res. 2013, 168, 547–554. [Google Scholar] [CrossRef]
- Tait, K.; Williamson, H.; Atkinson, S.; Williams, P.; Camara, M.; Joint, I. Turnover of quorum sensing signal molecules modulates cross-kingdom signalling. Environ. Microbiol. 2009, 11, 1792–1802. [Google Scholar] [CrossRef]
- Defoirdt, T.; Thanh, L.D.; Van Delsen, B.; De Schryver, P.; Sorgeloos, P.; Boon, N.; Bossier, P. N-acylhomoserine lactone-degrading Bacillus strains isolated from aquaculture animals. Aquaculture 2011, 311, 258–260. [Google Scholar] [CrossRef]
- Romero, M.; Avendaño-Herrera, R.; Magariños, B.; Cámara, M.; Otero, A. Acylhomoserine lactone production and degradation by the fish pathogen Tenacibaculum maritimum, a member of the Cytophaga–Flavobacterium–Bacteroides (CFB) group. FEMS Microbiol. Lett. 2010, 304, 131–139. [Google Scholar] [CrossRef]
- Huang, W.; Lin, Y.; Yi, S.; Liu, P.; Shen, J.; Shao, Z.; Liu, Z. QsdH, a novel AHL lactonase in the RND-type inner membrane of marine Pseudoalteromonas byunsanensis strain 1A01261. PLoS One 2012, 7, e46587. [Google Scholar]
- Swem, L.R.; Swem, D.L.; O’Loughlin, C.T.; Gatmaitan, R.; Zhao, B.X.; Ulrich, S.M.; Bassler, B.L. A quorum-sensing antagonist targets both membrane-bound and cytoplasmic receptors and controls bacterial pathogenicity. Mol. Cell 2009, 35, 143–153. [Google Scholar] [CrossRef]
- O’Loughlin, C.T.; Miller, L.C.; Siryaporn, A.; Drescher, K.; Semmelhack, M.F.; Bassler, B.L. A quorum-sensing inhibitor blocks Pseudomonas aeruginosa virulence and biofilm formation. Proc. Natl. Acad. Sci. USA 2013, 110, 17981–17986. [Google Scholar]
- Whitehead, N.A.; Welch, M.; Salmond, G.P. Silencing the majority. Nat. Biotechnol. 2001, 19, 735–736. [Google Scholar] [CrossRef]
- Defoirdt, T.; Boon, N.; Bossier, P. Can bacteria evolve resistance to quorum sensing disruption? PLoS Pathog. 2010, 6, e1000989. [Google Scholar] [CrossRef] [Green Version]
- Maeda, T.; Garcia-Contreras, R.; Pu, M.; Sheng, L.; Garcia, L.R.; Tomas, M.; Wood, T.K. Quorum quenching quandary: Resistance to antivirulence compounds. ISME J. 2012, 6, 493–501. [Google Scholar] [CrossRef]
- Garcia-Contreras, R.; Maeda, T.; Wood, T.K. Resistance to quorum-quenching compounds. Appl. Environ. Microbiol. 2013, 79, 6840–6846. [Google Scholar] [CrossRef]
- Fernandes, R.; Roy, V.; Wu, H.-C.; Bentley, W.E. Engineered biological nanofactories trigger quorum sensing response in targeted bacteria. Nat. Nano 2010, 5, 213–217. [Google Scholar] [CrossRef]
- Komnatnyy, V.V.; Chiang, W.C.; Tolker-Nielsen, T.; Givskov, M.; Nielsen, T.E. Bacteria-triggered release of antimicrobial agents. Angew. Chem. Int. Ed. Engl. 2014, 53, 439–441. [Google Scholar] [CrossRef]
- Park, K.; Kwon, I.C.; Park, K. Oral protein delivery: Current status and future prospect. React. Funct. Polym. 2011, 71, 280–287. [Google Scholar] [CrossRef]
© 2014 by the authors; licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution license (http://creativecommons.org/licenses/by/3.0/).
Share and Cite
Tang, K.; Zhang, X.-H. Quorum Quenching Agents: Resources for Antivirulence Therapy. Mar. Drugs 2014, 12, 3245-3282. https://doi.org/10.3390/md12063245
Tang K, Zhang X-H. Quorum Quenching Agents: Resources for Antivirulence Therapy. Marine Drugs. 2014; 12(6):3245-3282. https://doi.org/10.3390/md12063245
Chicago/Turabian StyleTang, Kaihao, and Xiao-Hua Zhang. 2014. "Quorum Quenching Agents: Resources for Antivirulence Therapy" Marine Drugs 12, no. 6: 3245-3282. https://doi.org/10.3390/md12063245




