Development of Pedigree Classification Using Microsatellite and Mitochondrial Markers for Giant Grouper Broodstock (Epinephelus lanceolatus) Management in Taiwan
Abstract
:1. Introduction
2. Results and Discussion
2.1. Analysis of Mitochondrial D-Loop Region
| Parameter to the Genetic Diversity | |
|---|---|
| Farms | 3 |
| Samples | 118 |
| Haplotypes | 42 |
| Length (base pair) | 776 |
| Nucleotide variations | 56 |
| Haplotype diversity(Hd) | 0.983 |
| Nucleotide diversity(π) | 0.00773 |
| Maximum distance between each two haplotypes | 0.0215 |
| Minimum distance between each two haplotypes | 0.0013 |
| Mean distance between each two haplotypes | 0.0079 |
2.2. Analysis of Giant Grouper Microsatellite Loci
| Locus * | Accession Number | Repeat Motif | Primer Sequences (5′→3′) | Allele Size | Allele Number | Expected Heterozygosity (EH) | Observed Heterozygosity (OH) |
|---|---|---|---|---|---|---|---|
| Efu02 a | EU016533 | (CA)n | F: CTGTCTCAGCTGATTTATGG | 345–371 | 10 | 0.667815 | 0.484375 |
| R: TTTACAGTCTCGTGGTTTCA | |||||||
| Efu06 a | EU016535 | (GACA)n | F: CATTGTCATTGTTGCTGTTTCTGTC | 308 | 1 | 0 | 0 |
| R: CCCTTTGGCCAATTGATGTGAT | |||||||
| Efu08 a | EU016537 | (CA)n | F: TGGAGAAGCCTGTAGATTATTGTG | 292–330 | 9 | 0.783465 | 0.796875 |
| R: AAGCAGGAGAGGAGTTGAAGGAGT | |||||||
| Efu18 a | EU016543 | (CA)n | F: ACTGGCTCCCTTCTGTTCC | 370–384 | 2 | 0.361713 | 0.4375 |
| R: ATTGCCACCATCGCTACC | |||||||
| Efu19 a | EU016544 | (CA)n | F: GGGCGGTAACCTCTCCAG | 93–115 | 7 | 0.746063 | 0.8125 |
| R: AGCAGCAACACCTTCTTCTCA | |||||||
| Efu41 a | EU016545 | (CA)n | F: CAGCACGCAGTTTAATTTACCAG | 243–249 | 2 | 0.353223 | 0.390625 |
| R: CAGGACCCGAGCTTCAGAA | |||||||
| ELMS009 b | EF607131 | (CA)n | F: TTCCACAGCAATTAGCAGCA | 260–278 | 8 | 0.77842 | 0.875 |
| R: TTTCCTCCCACAGTCCAAAG | |||||||
| ELMS015 b | EF607136 | (TG)n | F: AAGCTGAGCCGAATTTTTCA | 335–369 | 12 | 0.83157 | 0.640625 |
| R: GCTCCTCGTGTTTCCGATTA | |||||||
| Epaw3 c | EU684479 | (GT)n | F: GTCGTGTCTGTGACCATGAG | 72–76 | 2 | 0.503445 | 0 |
| R: TAAGGAGGGGGCTAAATGAT | |||||||
| Epaw6 c | EU684482 | (GT)n | F: ATGGTGTGGGAAAAGAGAGT | 146–233 | 5 | 0.619218 | 0.453125 |
| R: TTGTTTCAGGACAAGTGAGC | |||||||
| Epaw19 c | EU684495 | (GT)n | F: AGGTGGCTTGTGTGTGTATT | 243–247 | 3 | 0.215428 | 0.234375 |
| R: GCTTCCTTGACTGCTATGAC | |||||||
| Epaw34 c | EU684510 | (TG)n | F: ACAGCACCTCTACCATGAAC | 224–248 | 3 | 0.212968 | 0.078125 |
| R: CGTCCCCGTATATATCTCTG | |||||||
| CA-2 d | AF539606 | (CA)n | F: GACTTGATTCAGCAAAATAAAGATG | 150–262 | 6 | 0.350271 | 0.0625 |
| R: AGAGACGGTGCCAGTAAATGAA | |||||||
| CA-3 d | AF539605 | (CA)n | F: ATGTGACACGTTGACAGGCAAGT | 300 | 1 | 0 | 0 |
| R: GACCTTGATATTTTCATTGCTTG | |||||||
| CA-6 d | AF539608 | (CA)n | F: GTGTTGCTGGGGTTACTAATGAAG | 266–290 | 4 | 0.495325 | 0.5 |
| R: TTAGACACATTGTCACGATGGTCC | |||||||
| RH_CA_1 e | DQ223785 | (CA)n | F: CGAGATAAGCCCTGGTGAAA | 376–388 | 3 | 0.452879 | 0.46875 |
| R: AGTCCCGATGTGGTAACGAG | |||||||
| RH_GATA_2 e | DQ223791 | (GATA)n | F: CTCGACAGTGGACAAGGTCA | 132 | 1 | 0 | 0 |
| R: AAGGGCATGATGGGAAATG | |||||||
| RH_CA_2 e | DQ223785 | (CA)n | F: CTCGTTACCACATCGGGACT | 135–175 | 13 | 0.822466 | 0.890625 |
| R: AACACTGGCTGGTTTGCACT | |||||||
| RH_GATA_3 e | DQ223790 | (GATA)n | F: GGGCAATTTGGTTCTTCACA | 225–273 | 10 | 0.845349 | 0.828125 |
| R: TGTCAATGCCACAGGATACA | |||||||
| RH_CA_7 e | DQ223786 | (CA)n | F: CAGAAACATCTCCCCCAAAA | 259–335 | 12 | 0.639887 | 0.34375 |
| R: CTGGCAGAGCAATTAGAGGC | |||||||
| RH_CA_8 e | DQ223787 | (CA)n | F: AGTTGCCCAGGTTACACGAG | 219–227 | 4 | 0.554995 | 0.515625 |
| R: TTGGGTCCTGGCATTTAGAG |
| Paternity Exclusion Probabilities | |
|---|---|
| Combined non-exclusion probability (first parent) | 0.01138738 |
| Combined non-exclusion probability (second parent) | 0.00089627 |
3. Experimental Section
3.1. Fish Samples
3.2. DNA Extraction
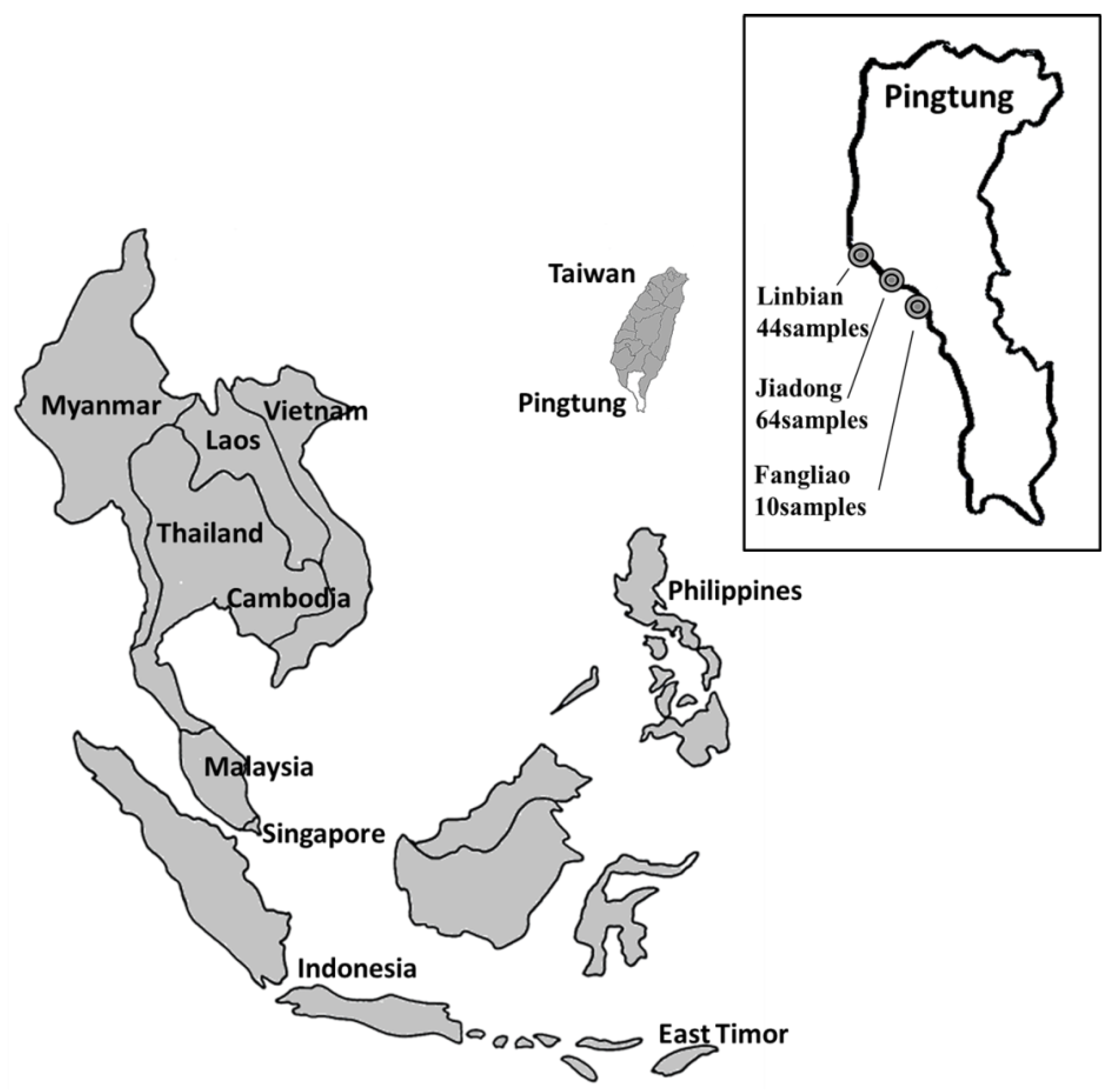
3.3. Establishment of Haplotype Database
3.4. PCR and Analysis of Microsatellite Loci
4. Conclusions
Acknowledgments
Author Contributions
Conflicts of Interest
References
- Duarte, C.M.; Marbá, N.; Holmer, M. Rapid domestication of marine species. Science 2007, 316, 382–383. [Google Scholar] [CrossRef]
- Kuo, H.C.; Wang, T.Y.; Chen, P.P.; Chen, Y.M.; Chuang, H.C.; Chen, T.Y. Real-time quantitative PCR assay for monitoring of nervous necrosis virus infection in grouper aquaculture. J. Clin. Microbiol. 2011, 49, 1090–1096. [Google Scholar] [CrossRef]
- Kuo, H.C.; Wang, T.Y.; Hsu, H.H.; Lee, S.H.; Chen, Y.M.; Tsai, T.J.; Ou, M.C.; Ku, H.T.; Lee, G.B.; Chen, T.Y. An automated microfluidic chip system for detection of piscine nodavirus and characterization of its potential carrier in grouper farms. PLoS One 2012, 7, e42203. [Google Scholar] [CrossRef]
- Kuo, H.C.; Wang, T.Y.; Hsu, H.H.; Chen, P.P.; Lee, S.H.; Chen, Y.M.; Tsai, T.J.; Wang, C.K.; Ku, H.T.; Lee, G.B.; et al. Nervous necrosis virus replicates following the embryo development and dual infection with iridovirus at juvenile stage in grouper. PLoS One 2012, 7, e36183. [Google Scholar] [CrossRef]
- Pierre, S.; Gaillard, S.; Prevot-D’Alvise, N.; Aubert, J.; Rostaing-Capaillon, O.; Leung-Tack, D.; Grillasca, J.P. Grouper aquaculture: Asian success and Mediterranean trials. Aquat. Conserv. Mar. Freshw. Ecosyst. 2008, 18, 297–308. [Google Scholar] [CrossRef]
- Gjerde, B.; Gunnes, K.; Gjedrem, T. Effect of inbreeding on survival and growth in rainbow trout. Aquaculture 1983, 34, 327–332. [Google Scholar] [CrossRef]
- Pante, M.J.R.; Gjerde, B.; McMillan, I. Effect of inbreeding on body weight at harvest in rainbow trout, Oncorhynchus mykiss. Aquaculture 2001, 192, 201–211. [Google Scholar] [CrossRef]
- Chiang, T.Y.; Lee, T.W.; Hsu, K.C.; Kuo, C.H.; Lin, D.Y.; Lin, H.D. Population structure in the endangered cyprinid fish Pararasbora moltrechti in Taiwan, based on mitochondrial and microsatellite DNAs. Zool. Sci. 2011, 28, 642–651. [Google Scholar] [CrossRef]
- Jackson, T.R.; Martin-Robichaud, D.J.; Reith, M.E. Application of DNA markers to the management of Atlantic halibut (Hippoglossus hippoglossus) broodstock. Aquaculture 2003, 220, 245–259. [Google Scholar] [CrossRef]
- Jonsson, B.; Jonsson, N. Cultured Atlantic salmon in nature: A review of their ecology and interaction with wild fish. ICES J. Mar. Sci. 2006, 63, 1162–1181. [Google Scholar] [CrossRef]
- Sunnucks, P. Efficient genetic markers for population biology. Trends Ecol. Evol. 2000, 15, 199–203. [Google Scholar] [CrossRef]
- Sekino, M.; Takagi, N.; Hara, M.; Takahashi, H. Analysis of microsatellite DNA polymorphisms in rockfish Sebastes thompsoni and application to population genetics studies. Mar. Biotechnol. 2001, 3, 45–52. [Google Scholar] [CrossRef]
- Powell, W.; Morgante, M.; Andre, C.; McNicol, J.W.; Machray, G.C.; Doyle, J.J.; Tingey, S.V.; Rafalski, J.A. Hypervariable microsatellites provide a general source of polymorphic DNA markers for the chloroplast genome. Curr. Biol. 1995, 5, 1023–1029. [Google Scholar] [CrossRef]
- Beacham, T.D.; Pollard, S.; Le, K.D. Microsatellite DNA population structure and stock identification of steelhead trout (Oncorhynchus mykiss) in the Nass and Skeena Rivers in northern British Columbia. Mar. Biotechnol. 2000, 2, 587–600. [Google Scholar] [CrossRef]
- Olsen, J.B.; Bentzen, P.; Banks, M.A.; Shaklee, J.B.; Young, S. Microsatellites reveal population identity of individual pink salmon to allow supportive breeding of a population at risk of extinction. Trans. Am. Fish. Soc. 2000, 129, 232–242. [Google Scholar] [CrossRef]
- Bierne, N.; Beuzart, I.; Vonau, V.; Bonhomme, F.; Bédier, E.A. Microsatellite-associated heterosis in hatchery-propagated stocks of the shrimp Penaeus stylirostris. Aquaculture 2000, 184, 203–219. [Google Scholar] [CrossRef]
- Sang, T.K.; Chang, H.Y.; Chen, C.T.; Hui, C.F. Population structure of the Japanese eel, Anguilla japonica. Mol. Biol. Evol. 1994, 11, 250–260. [Google Scholar]
- Zhu, Z.Y.; Lo, L.C.; Lin, G.; Xu, Y.X.; Yue, G.H. Isolation and characterization of polymorphic microsatellites from red coral grouper (Plectropomus maculatus). Mol. Ecol. Notes 2005, 5, 579–581. [Google Scholar] [CrossRef]
- An, H.S.; Kim, J.W.; Lee, J.W.; Kim, S.K.; Lee, B.I.; Kim, D.J.; Kim, Y.C. Development and characterization of microsatellite markers for an endangered species, Epinephelus bruneus, to establish a conservation program. Anim. Cell Syst. 2012, 16, 50–56. [Google Scholar] [CrossRef]
- Wang, L.; Meng, Z.; Liu, X.; Zhang, Y.; Lin, H. Genetic diversity and differentiation of the orange-spotted grouper (Epinephelus coioides) between and within cultured stocks and wild populations inferred from microsatellite DNA analysis. Int. J. Mol. Sci. 2011, 12, 4378–4394. [Google Scholar] [CrossRef]
- Rivera, M.A.; Graham, G.C.; Roderick, G.K. Isolation and characterization of nine microsatellite loci from the Hawaiian grouper Epinephelus quernus (Serranidae) for population genetic analyses. Mar. Biotechnol. 2003, 5, 126–129. [Google Scholar] [CrossRef]
- Han, J.; Lv, F.; Cai, H. Detection of species-specific long VNTRs in mitochondrial control region and their application to identifying sympatric Hong Kong grouper (Epinephelus akaara) and yellow grouper (Epinephelus awoara). Mol. Ecol. Resour. 2011, 11, 215–218. [Google Scholar] [CrossRef]
- Ramírez, M.A.; Patricia-Acevedo, J.; Planas, S.; Carlin, J.L.; Funk, S.M.; McMillan, W.O. New microsatellite resources for groupers (Serranidae). Mol. Ecol. Notes 2006, 6, 813–817. [Google Scholar] [CrossRef]
- Xia, X.; Xie, Z. DAMBE: Software package for data analysis in molecular biology and evolution. J. Heredity 2001, 92, 371–373. [Google Scholar] [CrossRef]
- Lo, L.C.; Yue, G.H. Microsatellites for broodstock management of the Tiger grouper, Epinephelus fuscoguttatus. Anim. Genet. 2008, 39, 90–91. [Google Scholar] [CrossRef]
- Zeng, H.S.; Ding, S.X.; Wang, J.; Su, Y.Q. Characterization of eight polymorphic microsatellite loci for the giant grouper (Epinephelus lanceolatus Bloch). Mol. Ecol. Resour. 2008, 8, 805–807. [Google Scholar] [CrossRef]
- Lin, L.Y.; Cheng, I.P.; Tzeng, C.S.; Huang, P.C. Maternal transmission of mitochondrial DNA in ducks. Biochem. Biophys. Res. Comm. 1990, 168, 188–193. [Google Scholar] [CrossRef]
- Mapsof.net. Available online: http://mapsof.net (accessed on 15 February 2013).
- Nei, M.; Gojobori, T. Simple methods for estimating the numbers of synonymous and nonsynonymous nucleotide substitutions. Mol. Biol. Evol. 1986, 3, 418–426. [Google Scholar]
- Nei, M.; Tajima, F. DNA polymorphism detectable by restriction endonucleases. Genetics 1981, 97, 145–163. [Google Scholar]
- Tamura, K.; Dudley, J.; Nei, M.; Kumar, S. MEGA4: Molecular Evolutionary Genetics Analysis (MEGA) software version 4.0. Mol. Biol. Evol. 2007, 24, 1596–1599. [Google Scholar] [CrossRef]
- Zhao, L.; Shao, C.; Liao, X.; Ma, H.; Zhu, X.; Chen, S. Twelve novel polymorphic microsatellite loci for the Yellow grouper (Epinephelus awoara) and cross-species amplifications. Conserv. Genet. 2009, 10, 743–745. [Google Scholar] [CrossRef]
- Dieringer, D.; Schlötterer, C. MICROSATELLITE ANALYSER (MSA): A platform independent analysis tool for large microsatellite data sets. Mol. Ecol. Notes 2003, 3, 167–169. [Google Scholar] [CrossRef]
- Kalinowski, S.T.; Taper, M.L.; Marshall, T.C. Revising how the computer program CERVUS accommodates genotyping error increases success in paternity assignment. Mol. Ecol. 2007, 16, 1099–1106. [Google Scholar] [CrossRef]
© 2014 by the authors; licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution license (http://creativecommons.org/licenses/by/3.0/).
Share and Cite
Kuo, H.-C.; Hsu, H.-H.; Chua, C.S.; Wang, T.-Y.; Chen, Y.-M.; Chen, T.-Y. Development of Pedigree Classification Using Microsatellite and Mitochondrial Markers for Giant Grouper Broodstock (Epinephelus lanceolatus) Management in Taiwan. Mar. Drugs 2014, 12, 2397-2407. https://doi.org/10.3390/md12052397
Kuo H-C, Hsu H-H, Chua CS, Wang T-Y, Chen Y-M, Chen T-Y. Development of Pedigree Classification Using Microsatellite and Mitochondrial Markers for Giant Grouper Broodstock (Epinephelus lanceolatus) Management in Taiwan. Marine Drugs. 2014; 12(5):2397-2407. https://doi.org/10.3390/md12052397
Chicago/Turabian StyleKuo, Hsiao-Che, Hao-Hsuan Hsu, Chee Shin Chua, Ting-Yu Wang, Young-Mao Chen, and Tzong-Yueh Chen. 2014. "Development of Pedigree Classification Using Microsatellite and Mitochondrial Markers for Giant Grouper Broodstock (Epinephelus lanceolatus) Management in Taiwan" Marine Drugs 12, no. 5: 2397-2407. https://doi.org/10.3390/md12052397




