Construction and Characterization of an Escherichia coli Mutant Producing Kdo2-Lipid A
Abstract
:1. Introduction

2. Results and Discussion
2.1. Construction of E. coli Mutant WJW00 that Produces Kdo2-Lipid A by Deletion of the rfaD Gene
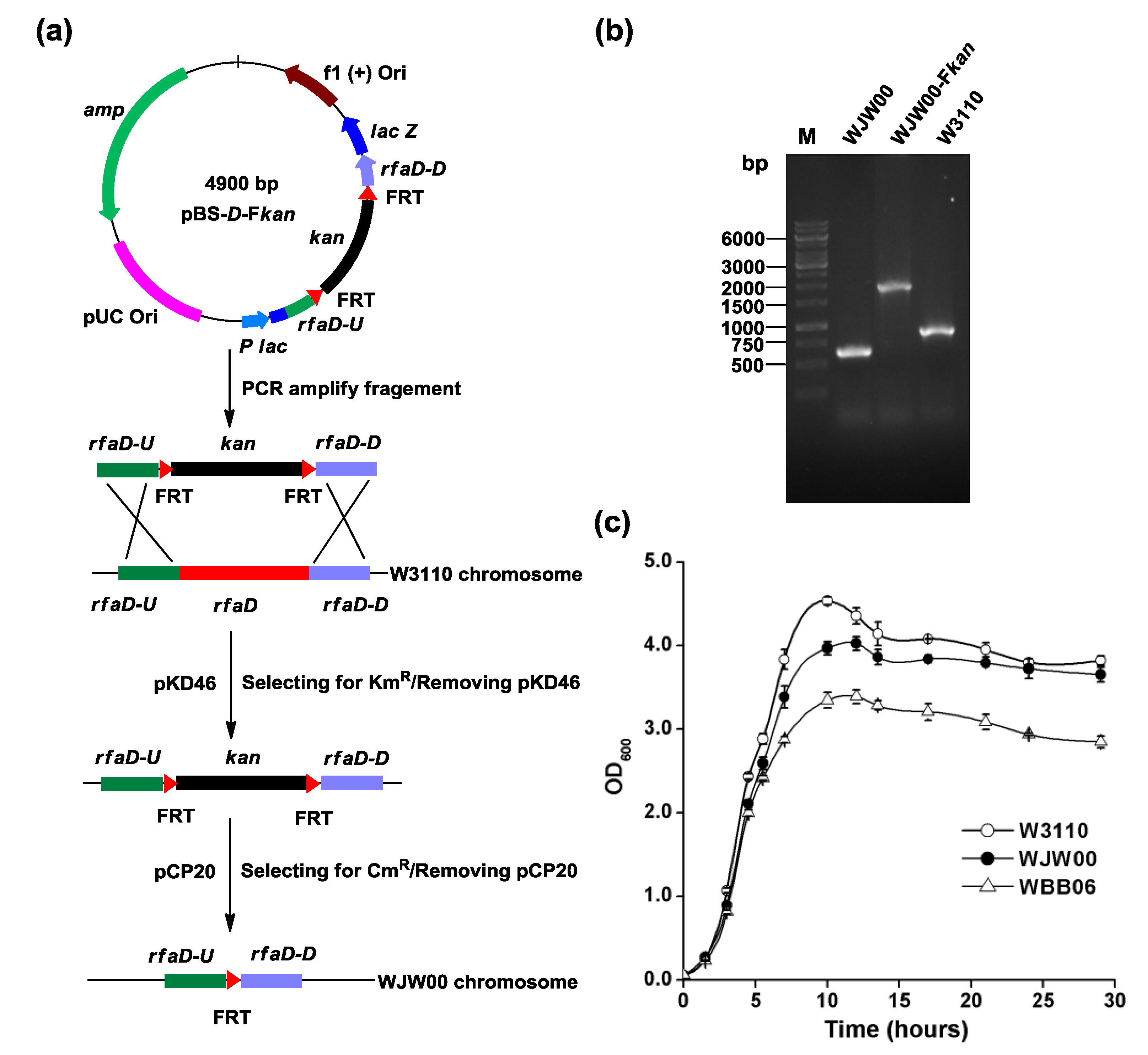
2.2. SDS-PAGE and TLC Analysis of Kdo2-Lipid A Produced by WJW00

2.3. ESI/MS Analysis of Kdo2-Lipid A Produced by WJW00
2.4. The Cell Surface Properties of WJW00

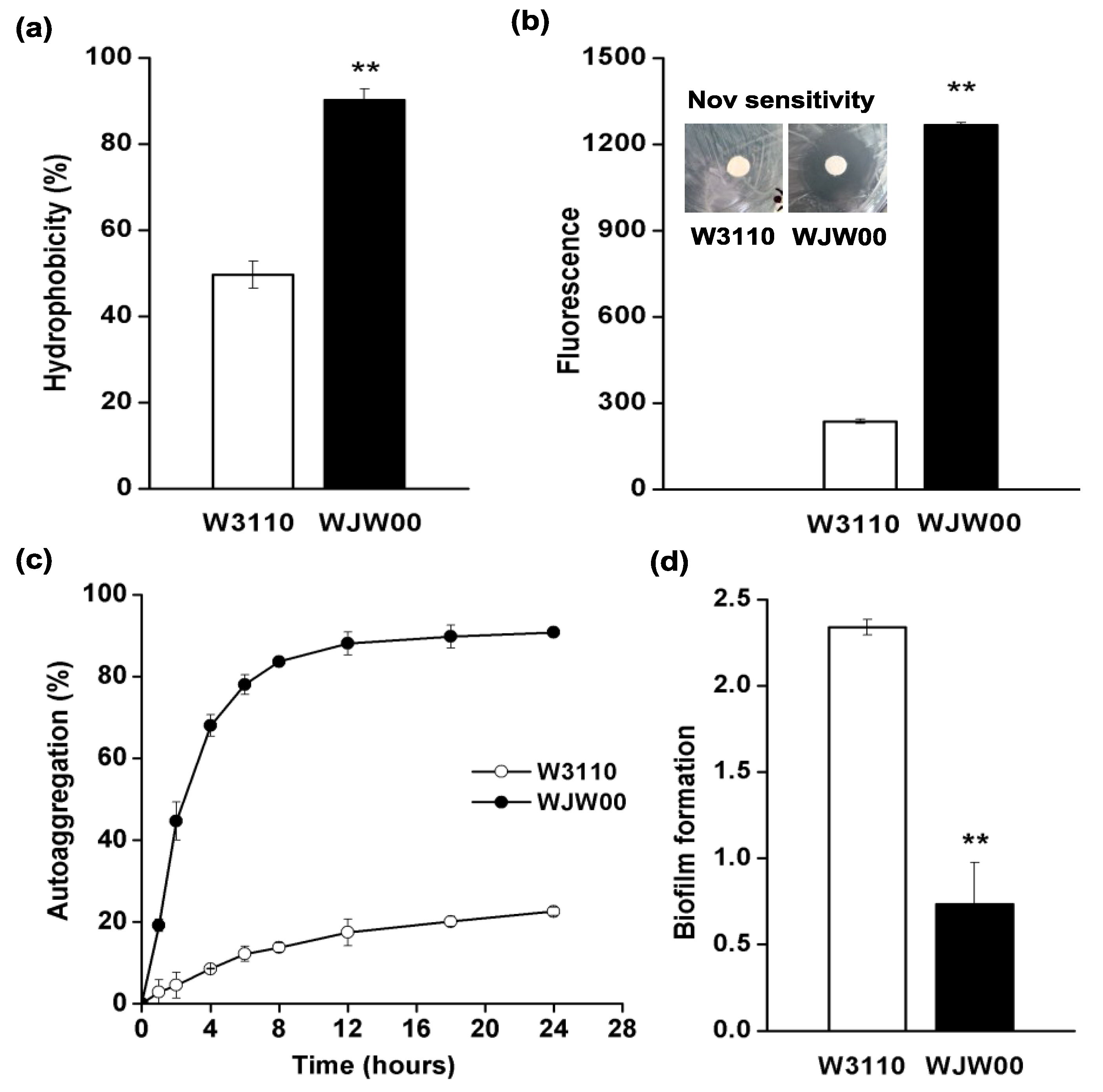
3. Experimental Section
3.1. DNA Preparation and PCR Techniques
| Primers | Sequence (5′–3′) | Restriction Site |
|---|---|---|
| rfaD-U-F | CCGCTCGAGTCCGTTACACCTTCAGCA | XhoI |
| rfaD-U-R | CGGAATTCGTGGCTGATGTGAATCTGTGGT | EcoRI |
| rfaD-D-F | CCCAAGCTTTACTTGCCGTCCCACTCG | HindIII |
| rfaD-D-R | AAAACTGCAGGCTTTATCGGCAGCAACA | PstI |
| kan-FRT-F | CGGAATTCGTGTAGGCTGGAGCTGCTTCG | EcoRI |
| kan-FRT-R | CCCAAGCTTGCCATTAATTCACTGATCAG | HindIII |
3.2. Construction of WJW00 and Growth Analysis
| Strains or Plasmids | Description | Source |
|---|---|---|
| Strains | ||
| W3110 | Wild-type E. coli, F−, λ− | Laboratory strain |
| W3110/pKD46 | W3110 transformed by pKD46 | Laboratory strain |
| WBB06 | W3110 mutant with a mutation of waaC and waaF genes (rfaC-rfaF::tet6) | [22] |
| WJW00 | W3110 ΔrfaD | This Study |
| Plasmids | ||
| pBluscript II SK+ | Cloning vector, ColE1, lacZ, AmpR | Stratagene |
| pBS-D-Fkan | Plasmid for deleting rfaD in E. coli | This Study |
| pKD46 | ParaBγβ exo, Repts, AmpR | [23] |
| pKD13 | oriR6K, FRT KanR FRT, AmpR | [23] |
| pCP20 | FLP+, λ cI857, λpRRepts, CamR, AmpR | [24] |
3.3. Extraction and SDS-PAGE Analysis of LPS
3.4. Isolation and TLC Analysis of Lipids
3.5. ESI/MS Analysis of Kdo2-Lipid A
3.6. Surface Hydrophobicity Assay
3.7. Outer Membrane Permeability and Novobiocin Sensitivity Assay
3.8. Autoaggregation Assay
3.9. Biofilm Assay
4. Conclusions
Acknowledgments
Author Contributions
Conflicts of Interest
References
- Raetz, C.R.; Whitfield, C. Lipopolysaccharide endotoxins. Annu. Rev. Biochem. 2002, 71, 635–700. [Google Scholar] [CrossRef]
- Wang, X.; Quinn, P.J. Lipopolysaccharide: Biosynthetic pathway and structure modification. Progr. Lipid Res. 2010, 49, 97–107. [Google Scholar] [CrossRef]
- Wang, X.; Quinn, P.J. Endotoxins: Lipopolysaccharides of Gram-Negative Bacteria. In Endotoxins: Structure, Function and Recognition; Springer: Heidelberg, Germany, 2010; Volume 53, pp. 3–26. [Google Scholar]
- Opiyo, S.O.; Pardy, R.L.; Moriyama, H.; Moriyama, E.N. Evolution of the Kdo2-lipid A biosynthesis in bacteria. BMC Evol. Biol. 2010, 10, 362. [Google Scholar] [CrossRef]
- Beutler, B. Inferences, questions and possibilities in Toll-like receptor signalling. Nature 2004, 430, 257–263. [Google Scholar] [CrossRef]
- Beutler, B.; Rietschel, E.T. Innate immune sensing and its roots: the story of endotoxin. Nat. Rev. Immunol. 2003, 3, 169–176. [Google Scholar] [CrossRef]
- Miller, S.I.; Ernst, R.K.; Bader, M.W. LPS, TLR4 and infectious disease diversity. Nat. Rev. Microbiol. 2005, 3, 36–46. [Google Scholar] [CrossRef]
- Akira, S.; Uematsu, S.; Takeuchi, O. Pathogen recognition and innate immunity. Cell 2006, 124, 783–801. [Google Scholar] [CrossRef]
- Passarelli, M.K.; Ewing, A.G.; Winograd, N. C(60)-SIMS studies of glycerophospholipid in a LIPID MAPS model system: KDO(2)-Lipid A stimulated RAW 264.7 cells. Surf. Interface Anal. 2013, 45, 298–301. [Google Scholar] [CrossRef]
- Srivastava, H.C.; Breuninger, E.; Creech, H.J.; Adams, G.A. Preparation and properties of polysaccharide-lipid complexes from Serratia marcescens. Can. J. Biochem. Physiol. 1962, 40, 905–918. [Google Scholar] [CrossRef]
- Raetz, C.R.; Garrett, T.A.; Reynolds, C.M.; Shaw, W.A.; Moore, J.D.; Smith, D.C., Jr.; Ribeiro, A.A.; Murphy, R.C.; Ulevitch, R.J.; Fearns, C.; et al. Kdo2-Lipid A of Escherichia coli, a defined endotoxin that activates macrophages via TLR-4. J. Lipid Res. 2006, 47, 1097–1111. [Google Scholar] [CrossRef]
- Murphy, R.C.; Raetz, C.R.; Reynolds, C.M.; Barkley, R.M. Mass spectrometry advances in lipidomica: collision-induced decomposition of Kdo2-lipid A. Prostaglandins Other Lipid Mediat. 2005, 77, 131–140. [Google Scholar] [CrossRef]
- Sims, K.; Haynes, C.A.; Kelly, S.; Allegood, J.C.; Wang, E.; Momin, A.; Leipelt, M.; Reichart, D.; Glass, C.K.; Sullards, M.C.; et al. Kdo2-lipid A, a TLR4-specific agonist, induces de novo sphingolipid biosynthesis in RAW264.7 macrophages, which is essential for induction of autophagy. J. Biol. Chem. 2010, 285, 38568–38579. [Google Scholar] [CrossRef]
- Kim, E.Y.; Shin, H.Y.; Kim, J.Y.; Kim, D.G.; Choi, Y.M.; Kwon, H.K.; Rhee, D.K.; Kim, Y.S.; Choi, S. ATF3 plays a key role in Kdo2-lipid A-induced TLR4-dependent gene expression via NF-kappaB activation. PLoS One 2010, 5, e14181. [Google Scholar]
- Reynolds, C.M.; Raetz, C.R. Replacement of lipopolysaccharide with free lipid A molecules in Escherichia coli mutants lacking all core sugars. Biochemistry 2009, 48, 9627–9640. [Google Scholar] [CrossRef]
- Kadrmas, J.L.; Raetz, C.R. Enzymatic synthesis of lipopolysaccharide in Escherichia coli. Purification and properties of heptosyltransferase I. J. Biol. Chem. 1998, 273, 2799–2807. [Google Scholar] [CrossRef]
- Gronow, S.; Brabetz, W.; Brade, H. Comparative functional characterization in vitro of heptosyltransferase I (WaaC) and II (WaaF) from Escherichia coli. Eur. J. Biochem. 2000, 267, 6602–6611. [Google Scholar] [CrossRef]
- Grizot, S.; Salem, M.; Vongsouthi, V.; Durand, L.; Moreau, F.; Dohi, H.; Vincent, S.; Escaich, S.; Ducruix, A. Structure of the Escherichia coli heptosyltransferase WaaC: Binary complexes with ADP and ADP-2-deoxy-2-fluoro heptose. J. Mol. Biol. 2006, 363, 383–394. [Google Scholar] [CrossRef]
- Coleman, W.G., Jr. The rfaD gene codes for ADP-l-glycero-d-mannoheptose-6-epimerase. An enzyme required for lipopolysaccharide core biosynthesis. J. Biol. Chem. 1983, 258, 1985–1990. [Google Scholar]
- Deacon, A.M.; Ni, Y.S.; Coleman, W.G., Jr.; Ealick, S.E. The crystal structure of ADP-l-glycero-d-mannoheptose 6-epimerase: Catalysis with a twist. Structure 2000, 8, 453–462. [Google Scholar] [CrossRef]
- Chang, P.C.; Wang, C.J.; You, C.K.; Kao, M.C. Effects of a HP0859 (rfaD) knockout mutation on lipopolysaccharide structure of Helicobacter pylori 26695 and the bacterial adhesion on AGS cells. Biochem. Biophys. Res. Commun. 2011, 405, 497–502. [Google Scholar] [CrossRef]
- Brabetz, W.; Muller-Loennies, S.; Holst, O.; Brade, H. Deletion of the heptosyltransferase genes rfaC and rfaF in Escherichia coli K-12 results in an Re-type lipopolysaccharide with a high degree of 2-aminoethanol phosphate substitution. Eur. J. Biochem. 1997, 247, 716–724. [Google Scholar]
- Datsenko, K.A.; Wanner, B.L. One-step inactivation of chromosomal genes in Escherichia coli K-12 using PCR products. Proc. Natl. Acad. Sci. USA 2000, 97, 6640–6645. [Google Scholar] [CrossRef]
- Cherepanov, P.P.; Wackernagel, W. Gene disruption in Escherichia coli: TcR and KmR cassettes with the option of Flp-catalyzed excision of the antibiotic-resistance determinant. Gene 1995, 158, 9–14. [Google Scholar]
- Ding, L.; Seto, B.L.; Ahmed, S.A.; Coleman, W.G., Jr. Purification and properties of the Escherichia coli K-12 NAD-dependent nucleotide diphosphosugar epimerase, ADP-l-glycero-d-mannoheptose 6-epimerase. J. Biol. Chem. 1994, 269, 24384–24390. [Google Scholar]
- Kanipes, M.I.; Papp-Szabo, E.; Guerry, P.; Monteiro, M.A. Mutation of waaC, encoding heptosyltransferase I in Campylobacter jejuni 81–176, affects the structure of both lipooligosaccharide and capsular carbohydrate. J. Bacteriol. 2006, 188, 3273–3279. [Google Scholar] [CrossRef]
- Parker, C.T.; Kloser, A.W.; Schnaitman, C.A.; Stein, M.A.; Gottesman, S.; Gibson, B.W. Role of the rfaG and rfaP genes in determining the lipopolysaccharide core structure and cell-surface properties of Escherichia-coli K-12. J. Bacteriol. 1992, 174, 2525–2538. [Google Scholar]
- Fujishima, H.; Nishimura, A.; Wachi, M.; Takagi, H.; Hirasawa, T.; Teraoka, H.; Nishimori, K.; Kawabata, T.; Nishikawa, K.; Nagai, K. kdsA mutations affect FtsZ-ring formation in Escherichia coli K-12. Microbiology 2002, 148, 103–112. [Google Scholar]
- Kido, N.; Ohta, M.; Kato, N. Detection of lipopolysaccharides by ethidium bromide staining after sodium dodecyl sulfate-polyacrylamide gel electrophoresis. J. Bacteriol. 1990, 172, 1145–1147. [Google Scholar]
- Tsai, C.M.; Frasch, C.E. A sensitive silver stain for detecting lipopolysaccharides in polyacrylamide gels. Anal. Biochem. 1982, 119, 115–119. [Google Scholar] [CrossRef]
- Bligh, E.G.; Dyer, W.J. A rapid method of total lipid extraction and purification. Can. J. Biochem. Physiol. 1959, 37, 911–917. [Google Scholar] [CrossRef]
- Zhao, J.; Raetz, C.R. A two-component Kdo hydrolase in the inner membrane of Francisella novicida. Mol. Microbiol. 2010, 78, 820–836. [Google Scholar] [CrossRef]
- Wang, X.; Ribeiro, A.A.; Guan, Z.; Raetz, C.R. Identification of undecaprenyl phosphate-beta-d-galactosamine in Francisella novicida and its function in lipid A modification. Biochemistry 2009, 48, 1162–1172. [Google Scholar] [CrossRef]
- Delcour, A.H. Outer membrane permeability and antibiotic resistance. Biochim. Biophys. Acta 2009, 1794, 808–816. [Google Scholar] [CrossRef]
- Nikaido, H. Molecular basis of bacterial outer membrane permeability revisited. Microbiol. Mol. Biol. Rev. 2003, 67, 593–656. [Google Scholar] [CrossRef]
- Vaara, M. Agents that increase the permeability of the outer-membrane. Microbiol. Rev. 1992, 56, 395–411. [Google Scholar]
- Bandara, H.M.; Lam, O.L.; Watt, R.M.; Jin, L.J.; Samaranayake, L.P. Bacterial lipopolysaccharides variably modulate in vitro biofilm formation of Candida species. J. Med. Microbiol. 2010, 59, 1225–1234. [Google Scholar] [CrossRef]
- Lau, P.C.; Lindhout, T.; Beveridge, T.J.; Dutcher, J.R.; Lam, J.S. Differential lipopolysaccharide core capping leads to quantitative and correlated modifications of mechanical and structural properties in Pseudomonas aeruginosa biofilms. J. Bacteriol. 2009, 191, 6618–6631. [Google Scholar] [CrossRef]
- Beloin, C.; Roux, A.; Ghigo, J.M. Escherichia coli biofilms. Curr. Top. Microbiol. Immunol. 2008, 322, 249–289. [Google Scholar]
- Wood, T.K.; Gonzalez Barrios, A.F.; Herzberg, M.; Lee, J. Motility influences biofilm architecture in Escherichia coli. Appl. Microbiol. Biotechnol. 2006, 72, 361–367. [Google Scholar] [CrossRef]
- Schembri, M.A.; Kjaergaard, K.; Klemm, P. Global gene expression in Escherichia coli biofilms. Mol. Microbiol. 2003, 48, 253–267. [Google Scholar] [CrossRef]
- Wang, L.; Hu, X.; Tao, G.; Wang, X. Outer membrane defect and stronger biofilm formation caused by inactivation of a gene encoding for heptosyltransferase I in Cronobacter sakazakii ATCC BAA-894. J. Appl. Microbiol. 2012, 112, 985–997. [Google Scholar] [CrossRef]
- Han, Y.; Li, Y.; Chen, J.; Tan, Y.; Guan, F.; Wang, X. Construction of monophosphoryl lipid A producing Escherichia coli mutants and comparison of immuno-stimulatory activities of their lipopolysaccharides. Mar. Drugs 2013, 11, 363–376. [Google Scholar] [CrossRef]
- Vaara, M. Antimicrobial susceptibility of Salmonella typhimurium carrying the outer membrane permeability mutation SS-B. Antimicrob. Agents Chemother. 1990, 34, 853–857. [Google Scholar] [CrossRef]
- Wang, X.; Ribeiro, A.A.; Guan, Z.; Abraham, S.N.; Raetz, C.R. Attenuated virulence of a Francisella mutant lacking the lipid A 4′-phosphatase. Proc. Natl. Acad. Sci. USA 2007, 104, 4136–4141. [Google Scholar] [CrossRef]
- Rahman, M.; Kim, W.S.; Kumura, H.; Shimazaki, K. In vitro effects of bovine lactoferrin on autoaggregation ability and surface hydrophobicity of bifidobacteria. Anaerobe 2008, 14, 73–77. [Google Scholar] [CrossRef]
- Raetz, C.R.; Reynolds, C.M.; Trent, M.S.; Bishop, R.E. Lipid A modification systems in gram-negative bacteria. Annu. Rev. Biochem. 2007, 76, 295–329. [Google Scholar] [CrossRef]
- Needham, B.D.; Carroll, S.M.; Giles, D.K.; Georgiou, G.; Whiteley, M.; Trent, M.S. Modulating the innate immune response by combinatorial engineering of endotoxin. Proc. Natl. Acad. Sci. USA 2013, 110, 1464–1469. [Google Scholar] [CrossRef]
- Vorachek-Warren, M.K.; Ramirez, S.; Cotter, R.J.; Raetz, C.R. A triple mutant of Escherichia coli lacking secondary acyl chains on lipid A. J. Biol. Chem. 2002, 277, 14194–14205. [Google Scholar] [CrossRef]
- Klein, G.; Lindner, B.; Brabetz, W.; Brade, H.; Raina, S. Escherichia coli K-12 suppressor-free mutants lacking early glycosyltransferases and late acyltransferases: Minimal lipopolysaccharide structure and induction of envelope stress response. J. Biol. Chem. 2009, 284, 15369–15389. [Google Scholar] [CrossRef]
- Wang, X.; Karbarz, M.J.; McGrath, S.C.; Cotter, R.J.; Raetz, C.R. MsbA transporter-dependent lipid A 1-dephosphorylation on the periplasmic surface of the inner membrane: topography of Francisella novicida LpxE expressed in Escherichia coli. J. Biol. Chem. 2004, 279, 49470–49478. [Google Scholar]
© 2014 by the authors; licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution license (http://creativecommons.org/licenses/by/3.0/).
Share and Cite
Wang, J.; Ma, W.; Wang, Z.; Li, Y.; Wang, X. Construction and Characterization of an Escherichia coli Mutant Producing Kdo2-Lipid A. Mar. Drugs 2014, 12, 1495-1511. https://doi.org/10.3390/md12031495
Wang J, Ma W, Wang Z, Li Y, Wang X. Construction and Characterization of an Escherichia coli Mutant Producing Kdo2-Lipid A. Marine Drugs. 2014; 12(3):1495-1511. https://doi.org/10.3390/md12031495
Chicago/Turabian StyleWang, Jianli, Wenjian Ma, Zhou Wang, Ye Li, and Xiaoyuan Wang. 2014. "Construction and Characterization of an Escherichia coli Mutant Producing Kdo2-Lipid A" Marine Drugs 12, no. 3: 1495-1511. https://doi.org/10.3390/md12031495





