Subtilomycin: A New Lantibiotic from Bacillus subtilis Strain MMA7 Isolated from the Marine Sponge Haliclona simulans
Abstract
:1. Introduction
2. Results
2.1. Antimicrobial Activity of B. subtilis Strain MMA7
| Indicators | MMA7 culture | Peptide |
|---|---|---|
| B. cereus | +++ | ++ |
| B. megaterium | ++++ | ++ |
| L. monocytogenes | ++++ | ++ |
| L. innocua | ++++ | ++ |
| C. sporogenes | +++++ | +++ |
| C. perfringens | ++++ | - * |
| C. difficile | ++ | - * |
| E. faecium | ++ | - * |
| S. aureus | ++ | + |
| MRSA | ++ | + |
| hVISA | + | + |
| VRE | + | - * |
| L. lactis HP | ++++ | + |
| A. hydrophila | ++++ | + * |
| V. anguillarum | + | + * |
| Alteromonas sp. | +++ | + * |
| P. aeruginosa | - | + * |
| C. albicans | ++++ | - * |
| C. dubliniensis | ++ | - * |
| C. lusitaniae | ++ | - * |
| C. parapsilosis | + | - * |

Kinetics of Antimicrobial Activity
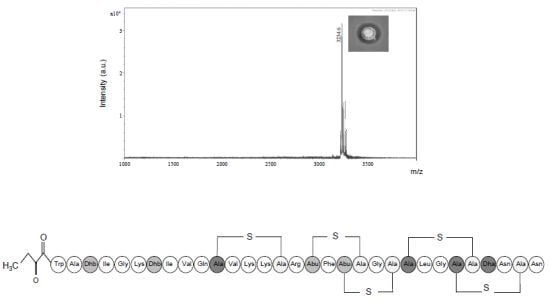
2.2. Purification and Characterisation of the Antimicrobial Compound from 12 h Cultures
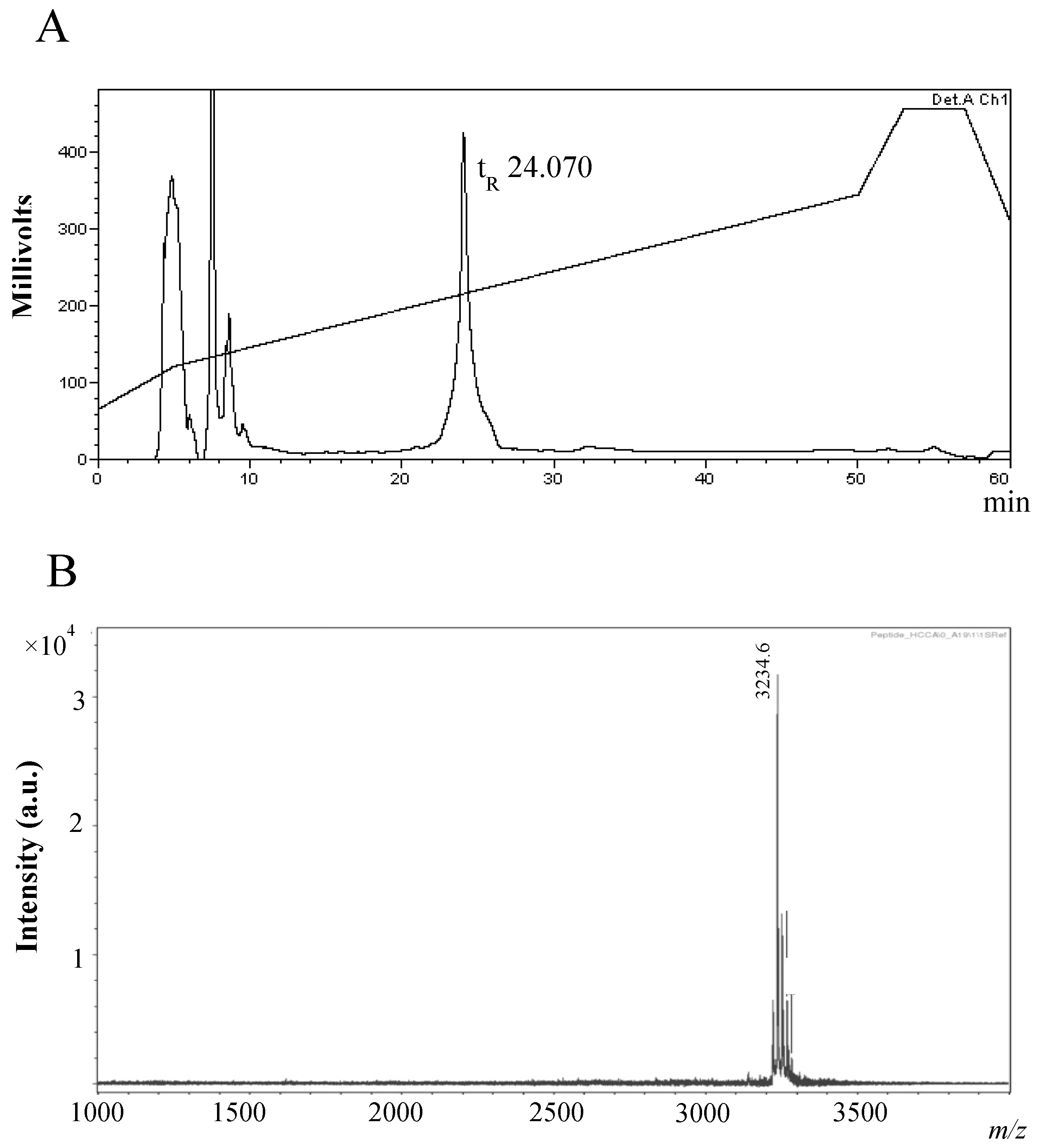
Physicochemical Characterisation of the Antimicrobial Peptide

2.3. Genetic and Structural Characterisation of the Antimicrobial Peptide
2.3.1. Primary Amino Acid Sequence of the Antimicrobial Peptide
2.3.2. Genetic Organisation and Location of the Subtilomycin Biosynthetic Gene Cluster

2.3.3. Further Structural Characterisation of Subtilomycin
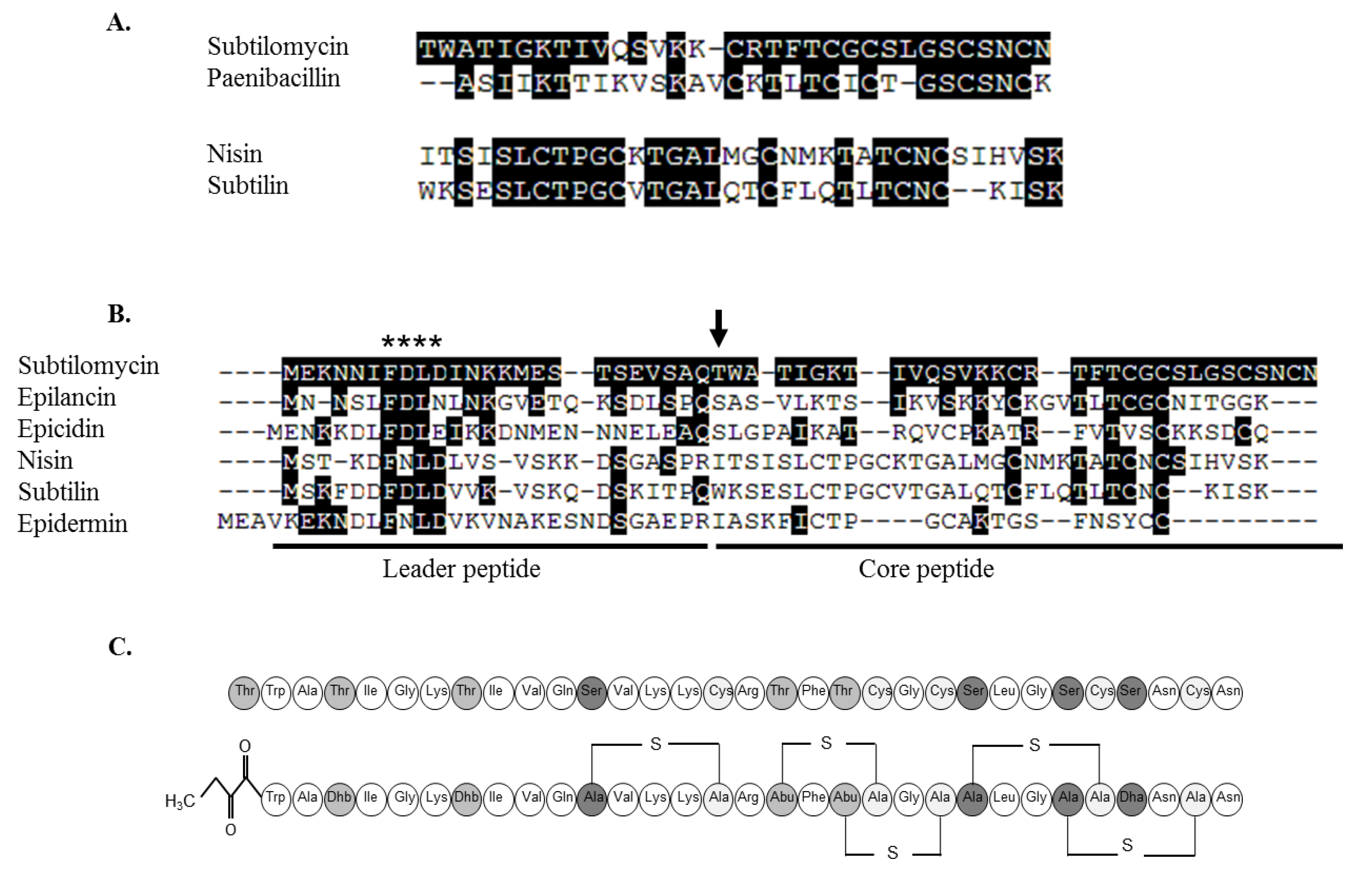
2.4. Distribution of the Subtilomycin Biosynthetic Gene Cluster
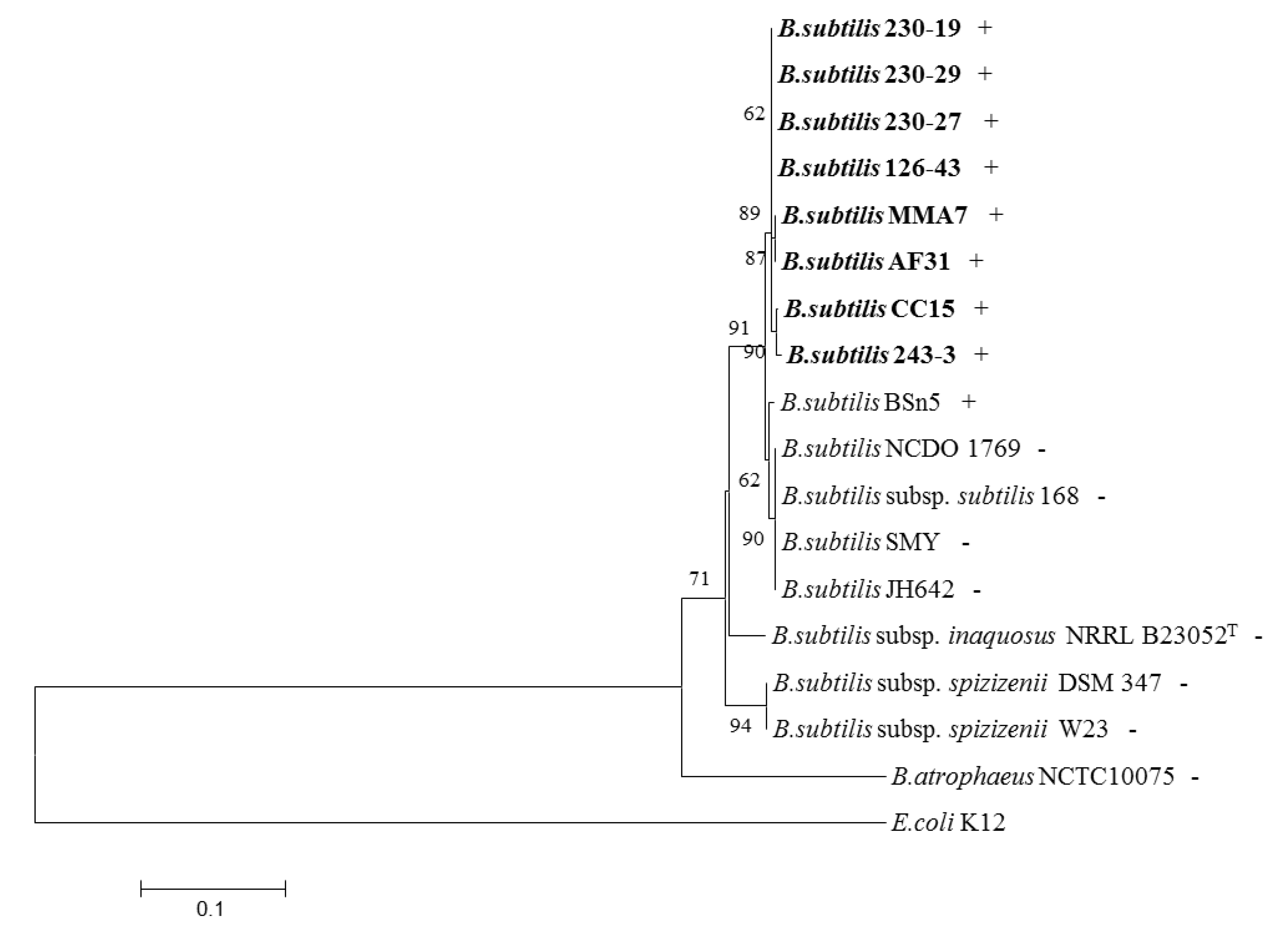
3. Discussion
4. Experimental Section
4.1. Bacterial Strains, Media and Growth Conditions
4.2. Antimicrobial Activity Screening Assays
4.3. DNA Extraction, PCR Amplification, DNA Sequencing and Phylogenetic Analysis
4.4. Construction of Strain MMA7 ∆Sbo-albF::Cat Mutant
4.5. Kinetics of Antimicrobial Production
4.6. Purification and Characterisation of the Antimicrobial Compound
4.7. Heat, pH and Proteolytic Treatment of the Antimicrobial Peptide
4.8. Genome Sequencing of B. subtilis MMA7 and Sequence Analysis of the Subtilomycin Biosynthetic Gene Cluster
5. Conclusions
Acknowledgments
Conflict of Interest
References
- Pelaez, F. The historical delivery of antibiotics from microbial natural products-can history repeat? Biochem. Pharmacol. 2006, 71, 981–990. [Google Scholar] [CrossRef]
- Taylor, M.W.; Radax, R.; Steger, D.; Wagner, M. Sponge-associated microorganisms: evolution, ecology, and biotechnological potential. Microbiol. Mol. Biol. Rev. 2007, 71, 295–347. [Google Scholar] [CrossRef]
- Kennedy, J.; Baker, P.; Piper, C.; Cotter, P.D.; Walsh, M.; Mooij, M.J.; Bourke, M.B.; Rea, M.C.; O’Connor, P.M.; Ross, R.P. Isolation and analysis of bacteria with antimicrobial activities from the marine sponge Haliclona simulans collected from Irish waters. Mar. Biotechnol. (NY) 2009, 11, 384–396. [Google Scholar] [CrossRef]
- Thomas, T.R.; Kavlekar, D.P.; LokaBharathi, P.A. Marine drugs from sponge-microbe association—A review. Mar. Drugs 2010, 8, 1417–1468. [Google Scholar] [CrossRef]
- Muscholl-Silberhorn, A.; Thiel, V.; Imhoff, J.F. Abundance and bioactivity of cultured sponge-associated bacteria from the Mediterranean sea. Microb. Ecol. 2008, 55, 94–106. [Google Scholar] [CrossRef]
- Zhu, P.; Li, Q.; Wang, G. Unique microbial signatures of the alien Hawaiian marine sponge Suberites zeteki. Microb. Ecol. 2008, 55, 406–414. [Google Scholar] [CrossRef]
- Lee, O.O.; Wong, Y.H.; Qian, P.Y. Inter- and intraspecific variations of bacterial communities associated with marine sponges from san juan island, washington. Appl. Environ. Microbiol. 2009, 75, 3513–3521. [Google Scholar] [CrossRef]
- Menezes, C.B.; Bonugli-Santos, R.C.; Miqueletto, P.B.; Passarini, M.R.; Silva, C.H.; Justo, M.R.; Leal, R.R.; Fantinatti-Garboggini, F.; Oliveira, V.M.; Berlinck, R.G.; Sette, L.D. Microbial diversity associated with algae, ascidians and sponges from the north coast of Sao Paulo state, Brazil. Microbiol. Res. 2009, 165, 466–482. [Google Scholar]
- Phelan, R.W.; O’Halloran, J.A.; Kennedy, J.; Morrissey, J.P.; Dobson, A.D.; O’Gara, F.; Barbosa, T.M. Diversity and bioactive potential of endospore-forming bacteria cultured from the marine sponge Haliclona simulans. J. Appl. Microbiol. 2012, 112, 65–78. [Google Scholar] [CrossRef]
- Stein, T. Bacillus subtilis antibiotics: Structures, syntheses and specific functions. Mol. Microbiol. 2005, 56, 845–857. [Google Scholar] [CrossRef]
- Westers, L.; Westers, H.; Quax, W.J. Bacillus subtilis as cell factory for pharmaceutical proteins: A biotechnological approach to optimize the host organism. Biochim. Biophys. Acta 2004, 1694, 299–310. [Google Scholar] [CrossRef]
- Alcaraz, L.D.; Moreno-Hagelsieb, G.; Eguiarte, L.E.; Souza, V.; Herrera-Estrella, L.; Olmedo-Alvarez, G. Understanding the evolutionary relationships and major traits of Bacillus through comparative genomics. BMC Genomics 2010, 11, 332. [Google Scholar]
- Barbosa, T.M.; Serra, C.R.; Henriques, A.O. Gut Sporeformers. In Bacterial Spore Formers: Probiotics and Emerging Applications; Ricca, E., Henriques, A.O., Cutting, S.M., Eds.; Horizon Bioscience: Norfolk, UK, 2004; pp. 183–191. [Google Scholar]
- Barbosa, T.M.; Serra, C.R.; La Ragione, R.M.; Woodward, M.J.; Henriques, A.O. Screening for Bacillus isolates in the broiler gastrointestinal tract. Appl. Environ. Microbiol. 2005, 71, 968–978. [Google Scholar]
- Tam, N.K.; Uyen, N.Q.; Hong, H.A.; Le Duc, H.; Hoa, T.T.; Serra, C.R.; Henriques, A.O.; Cutting, S.M. The intestinal life cycle of Bacillus subtilis and close relatives. J. Bacteriol. 2006, 188, 2692–2700. [Google Scholar] [CrossRef]
- Hong, H.A.; Khaneja, R.; Tam, N.M.; Cazzato, A.; Tan, S.; Urdaci, M.; Brisson, A.; Gasbarrini, A.; Barnes, I.; Cutting, S.M. Bacillus subtilis isolated from the human gastrointestinal tract. Res. Microbiol. 2009, 160, 134–143. [Google Scholar] [CrossRef]
- Hong, H.A.; To, E.; Fakhry, S.; Baccigalupi, L.; Ricca, E.; Cutting, S.M. Defining the natural habitat of Bacillus spore-formers. Res. Microbiol. 2009, 160, 375–379. [Google Scholar] [CrossRef]
- Knerr, P.J.; van der Donk, W.A. Discovery, biosynthesis, and engineering of lantipeptides. Annu. Rev. Biochem. 2012, 81, 479–505. [Google Scholar] [CrossRef]
- Willey, J.M.; van der Donk, W.A. Lantibiotics: Peptides of diverse structure and function. Annu. Rev. Microbiol. 2007, 61, 477–501. [Google Scholar] [CrossRef]
- Cotter, P.D.; Hill, C.; Ross, R.P. Bacterial lantibiotics: Strategies to improve therapeutic potential. Curr. Protein Pept. Sci. 2005, 6, 61–75. [Google Scholar] [CrossRef]
- Brotz, H.; Sahl, H.G. New insights into the mechanism of action of lantibiotics—Diverse biological effects by binding to the same molecular target. J. Antimicrob. Chemother. 2000, 46, 1–6. [Google Scholar] [CrossRef]
- Suda, S.; Westerbeek, A.; O’Connor, P.M.; Ross, R.P.; Hill, C.; Cotter, P.D. Effect of bioengineering lacticin 3147 lanthionine bridges on specific activity and resistance to heat and proteases. Chem. Biol. 2010, 17, 1151–1160. [Google Scholar] [CrossRef]
- Kim, T.W.; Kim, Y.H.; Kim, S.E.; Lee, J.H.; Park, C.S.; Kim, H.Y. Identification and distribution of Bacillus species in doenjang by whole-cell protein patterns and 16S rRNA gene sequence analysis. J. Microbiol. Biotechnol. 2010, 20, 1210–1214. [Google Scholar] [CrossRef]
- Kim, T.W.; Lee, J.H.; Park, M.H.; Kim, H.Y. Analysis of bacterial and fungal communities in Japanese- and Chinese-fermented soybean pastes using nested PCR-DGGE. Curr. Microbiol. 2010, 60, 315–320. [Google Scholar]
- Hong, H.A.; Le Duc, H.; Cutting, S.M. The use of bacterial spore formers as probiotics. FEMS Microbiol. Rev. 2005, 29, 813–835. [Google Scholar] [CrossRef]
- Lalloo, R.; Maharajh, D.; Gorgens, J.; Gardiner, N.; Gorgens, J.F. High-density spore production of a B. cereus aquaculture biological agent by nutrient supplementation. Appl. Microbiol. Biotechnol. 2009, 83, 59–66. [Google Scholar] [CrossRef]
- Anand, T.P.; Bhat, A.W.; Shouche, Y.S.; Roy, U.; Siddharth, J.; Sarma, S.P. Antimicrobial activity of marine bacteria associated with sponges from the waters off the coast of South East India. Microbiol. Res. 2006, 161, 252–262. [Google Scholar] [CrossRef]
- Hentschel, U.; Schmid, M.; Wagner, M.; Fieseler, L.; Gernert, C.; Hacker, J. Isolation and phylogenetic analysis of bacteria with antimicrobial activities from the Mediterranean sponges Aplysina aerophoba and Aplysina cavernicola. FEMS Microbiol. Ecol. 2001, 35, 305–312. [Google Scholar]
- Pabel, C.T.; Vater, J.; Wilde, C.; Franke, P.; Hofemeister, J.; Adler, B.; Bringmann, G.; Hacker, J.; Hentschel, U. Antimicrobial activities and matrix-assisted laser desorption/ionization mass spectrometry of Bacillus isolates from the marine sponge Aplysina aerophoba. Mar. Biotechnol. (NY) 2003, 5, 424–434. [Google Scholar] [CrossRef]
- Gonzalez-Pastor, J.E.; Hobbs, E.C.; Losick, R. Cannibalism by sporulating bacteria. Science 2003, 301, 510–513. [Google Scholar] [CrossRef]
- Sahl, H.G.; Jack, R.W.; Bierbaum, G. Biosynthesis and biological activities of lantibiotics with unique post-translational modifications. Eur. J. Biochem. 1995, 230, 827–853. [Google Scholar] [CrossRef]
- He, Z.; Yuan, C.; Zhang, L.; Yousef, A.E. N-Terminal acetylation in paenibacillin, a novel lantibiotic. FEBS Lett. 2008, 582, 2787–2792. [Google Scholar] [CrossRef]
- He, Z.; Kisla, D.; Zhang, L.; Yuan, C.; Green-Church, K.B.; Yousef, A.E. Isolation and identification of a Paenibacillus polymyxa strain that coproduces a novel lantibiotic and polymyxin. Appl. Environ. Microbiol. 2007, 73, 168–178. [Google Scholar] [CrossRef]
- Deng, Y.; Zhu, Y.; Wang, P.; Zhu, L.; Zheng, J.; Li, R.; Ruan, L.; Peng, D.; Sun, M. Complete genome sequence of Bacillus subtilis BSn5, an endophytic bacterium of Amorphophallus konjac with antimicrobial activity for the plant pathogen Erwinia carotovora subsp. carotovora. J. Bacteriol. 2011, 193, 2070–2071. [Google Scholar] [CrossRef]
- Foulston, L.C.; Bibb, M.J. Microbisporicin gene cluster reveals unusual features of lantibiotic biosynthesis in actinomycetes. Proc. Natl. Acad. Sci. USA 2010, 107, 13461–13466. [Google Scholar] [CrossRef]
- Xie, L.; van der Donk, W.A. Post-translational modifications during lantibiotic biosynthesis. Curr. Opin. Chem. Biol. 2004, 8, 498–507. [Google Scholar] [CrossRef]
- Kleerebezem, M. Quorum sensing control of lantibiotic production; nisin and subtilin autoregulate their own biosynthesis. Peptides 2004, 25, 1405–1414. [Google Scholar] [CrossRef]
- Field, D.; Hill, C.; Cotter, P.D.; Ross, R.P. The dawning of a 'Golden era' in lantibiotic bioengineering. Mol. Microbiol. 2010, 78, 1077–1087. [Google Scholar] [CrossRef]
- Borisova, S.A.; Circello, B.T.; Zhang, J.K.; van der Donk, W.A.; Metcalf, W.W. Biosynthesis of rhizocticins, antifungal phosphonate oligopeptides produced by Bacillus subtilis ATCC6633. Chem. Biol. 2010, 17, 28–37. [Google Scholar] [CrossRef]
- Pitcher, D.G.; Saunders, N.A.; Owen, R.J. Rapid extraction of bacterial genomic DNA with guanidium thiocyanate. Lett. Appl. Microbiol. 1989, 8, 151–156. [Google Scholar] [CrossRef]
- Larkin, M.A.; Blackshields, G.; Brown, N.P.; Chenna, R.; McGettigan, P.A.; McWilliam, H.; Valentin, F.; Wallace, I.M.; Wilm, A.; Lopez, R. Clustal W and Clustal X version 2.0. Bioinformatics 2007, 23, 2947–2948. [Google Scholar] [CrossRef]
- Tamura, K.; Dudley, J.; Nei, M.; Kumar, S. MEGA4: Molecular evolutionary genetics analysis (MEGA) software version 4.0. Mol. Biol. Evol. 2007, 24, 1596–1599. [Google Scholar] [CrossRef]
- Sambrook, J.; Fritsch, E.F.; Maniatis, T. Molecular Cloning: A Laboratory Manual; Cold Spring Harbor Laboratory Press: Woodbury, NY, USA, 1989. [Google Scholar]
- Cutting, S.M.; Vander Horn, P.B. Genetic analysis. In Molecular Biology Methods for Bacillus; Harwood, C.R., Cutting, S.M., Eds.; John Wiley and Sons: Chichester, UK, 1990; pp. 53–54. [Google Scholar]
- Zilhao, R.; Serrano, M.; Isticato, R.; Ricca, E.; Moran, C.P.; Henriques, A.O. Interactions among CotB, CotG, and CotH during assembly of the Bacillus subtilis spore coat. J. Bacteriol. 2004, 186, 1110–1119. [Google Scholar] [CrossRef]
- Altschul, S.F.; Gish, W.; Miller, W.; Myers, E.W.; Lipman, D.J. Basic local alignment search tool. J. Mol. Biol. 1990, 215, 403–410. [Google Scholar]
- Tomancak, P.; Beaton, A.; Weiszmann, R.; Kwan, E.; Shu, S.; Lewis, S.E.; Richards, S.; Ashburner, M.; Hartenstein, V.; Celniker, S.E.; Rubin, G.M. Systematic determination of patterns of gene expression during Drosophila embryogenesis. Genome Biol. 2002, 3, 1–14. [Google Scholar]
- Tomancak, P.; Berman, B.P.; Beaton, A.; Weiszmann, R.; Kwan, E.; Hartenstein, V.; Celniker, S.E.; Rubin, G.M. Global analysis of patterns of gene expression during Drosophila embryogenesis. Genome Biol. 2007, 8. [Google Scholar] [CrossRef]
Supplementary Files
© 2013 by the authors; licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution license (http://creativecommons.org/licenses/by/3.0/).
Share and Cite
Phelan, R.W.; Barret, M.; Cotter, P.D.; O'Connor, P.M.; Chen, R.; Morrissey, J.P.; Dobson, A.D.W.; O'Gara, F.; Barbosa, T.M. Subtilomycin: A New Lantibiotic from Bacillus subtilis Strain MMA7 Isolated from the Marine Sponge Haliclona simulans. Mar. Drugs 2013, 11, 1878-1898. https://doi.org/10.3390/md11061878
Phelan RW, Barret M, Cotter PD, O'Connor PM, Chen R, Morrissey JP, Dobson ADW, O'Gara F, Barbosa TM. Subtilomycin: A New Lantibiotic from Bacillus subtilis Strain MMA7 Isolated from the Marine Sponge Haliclona simulans. Marine Drugs. 2013; 11(6):1878-1898. https://doi.org/10.3390/md11061878
Chicago/Turabian StylePhelan, Robert W., Matthieu Barret, Paul D. Cotter, Paula M. O'Connor, Rui Chen, John P. Morrissey, Alan D. W. Dobson, Fergal O'Gara, and Teresa M. Barbosa. 2013. "Subtilomycin: A New Lantibiotic from Bacillus subtilis Strain MMA7 Isolated from the Marine Sponge Haliclona simulans" Marine Drugs 11, no. 6: 1878-1898. https://doi.org/10.3390/md11061878