Anti-Microbial, Anti-Biofilm Activities and Cell Selectivity of the NRC-16 Peptide Derived from Witch Flounder, Glyptocephalus cynoglossus
Abstract
:1. Introduction
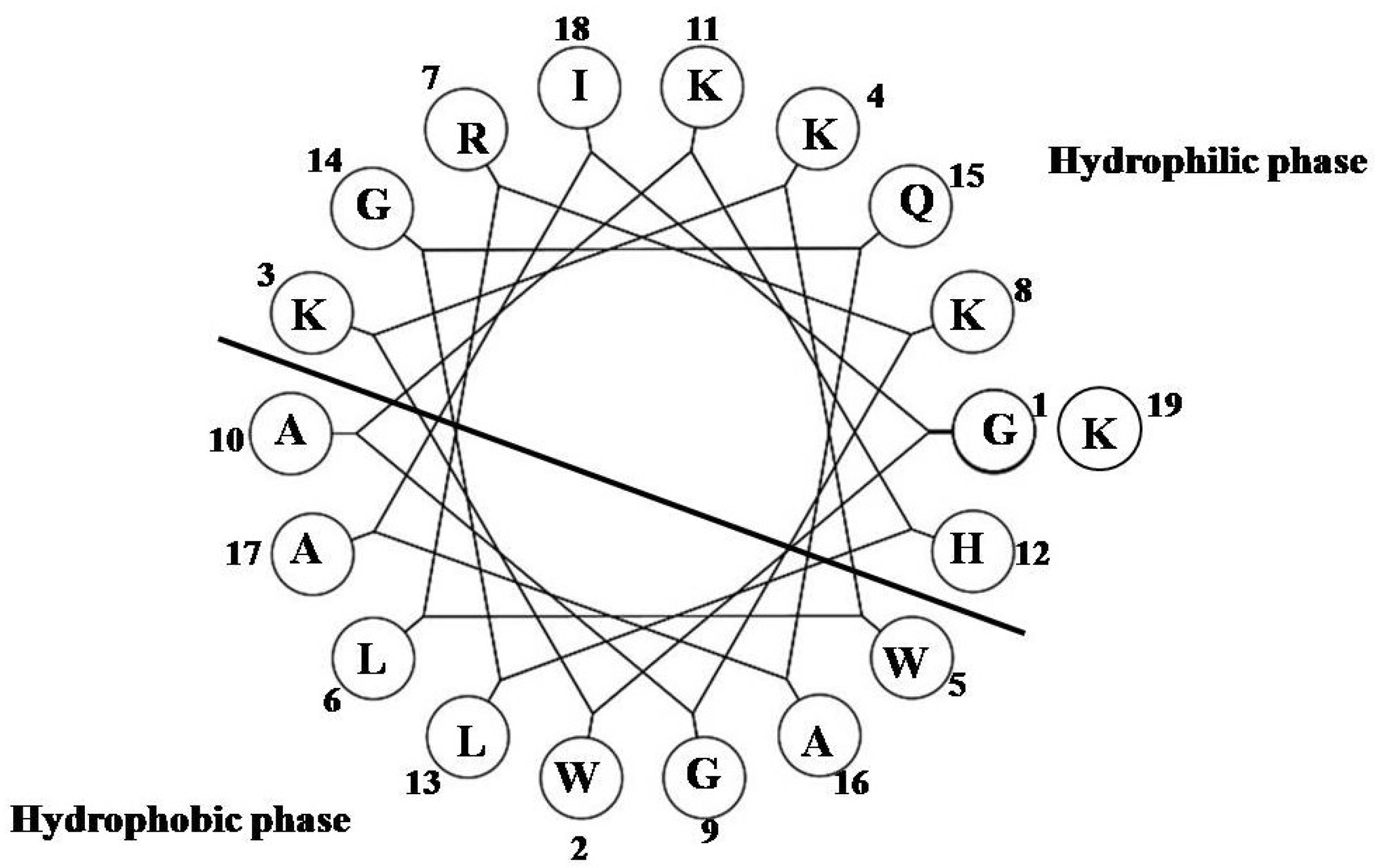
2. Results and Discussion
2.1. Lytic Effects of NRC-16
| MIC (µM) | ||
|---|---|---|
| Microorganism | NRC-16 | Melittin |
| Gram (−) bacteria | ||
| E. coli | 2(4) | 2(4) |
| S. typhimurium | 1(2) | 2(2) |
| P. aeruginosa | 4(8) | 8(16) |
| Gram (+) bacteria | ||
| S. aureus | 4(8) | 2(2) |
| B. subtilis | 2(8) | 1(1) |
| Yeast | ||
| C. albicans | 8(16) | 8(16) |
| T. beigelli | 4(8) | 2(4) |
| Resistant strains a | ||
| E. coli CCARM 1229 b | 8 | 2 |
| E. coli CCARM 1238 b | 4 | 2 |
| S. typhimurium CCARM 8007 c | 4 | 8 |
| S. typhimurium CCARM 8009 c | 16 | 16 |
| S. typhimurium CCARM 8013 c | 4 | 8 |
| S. aureus CCARM 3089 d | 2 | 2 |
| S. aureus CCARM 3090 d | 8 | 8 |
| S. aureus CCARM 3108 d | 2 | 2 |
| S. aureus CCARM 3114 d | 4 | 2 |
| S. aureus CCARM 3126 d | 4 | 8 |
| C. albicans CCARM 14001 e | 8 | 4 |
| MIC (µM) | ||
|---|---|---|
| Resistant strains | NRC-16 | Melittin |
| P. aeruginosa 1034 a | 4 | 4 |
| P. aeruginosa 1162 a | 2 | 2 |
| P. aeruginosa 3399 a | 2 | 2 |
| P. aeruginosa 3547 a | 4 | 8 |
| P. aeruginosa 3592 a | 8 | 8 |
| P. aeruginosa 4007 a | 2 | 2 |
| P. aeruginosa 4076 a | 8 | 8 |
| P. aeruginosa 5018 a | 4 | 8 |
| FRPA b | 8 | 16 |
| CRPSP c | 8 | 16 |
| IRPA d | 4 | 16 |
| S. aureus 254348 e | 2 | 2 |
| S. aureus 254422 e | 1 | 1 |
| S. aureus 691054 e | 2 | 4 |
| S. aureus 949987 e | 2 | 2 |
| S. aureus 950805 e | 1 | 8 |
| S. aureus 2-660 e | 8 | 2 |
| S. aureus 3518 e | 8 | 4 |
| S. aureus 2-3566 e | 4 | 4 |
| S. aureus 2-777 e | 4 | 2 |
| S. aureus 2-3122 e | 4 | 2 |
| S. aureus 2-254 e | 4 | 2 |
2.2. Effect of AMPs on the P. aeruginosa Biofilm
| MBIC (μM) | ||||||||
|---|---|---|---|---|---|---|---|---|
| Strains | Amp | Chl | Ery | Lev | Cip | Pip | NRC-16 | Melittin |
| 1162 | >512 | >512 | >512 | >512 | 256 | 128 | 8 | 4 |
| 3547 | >512 | >512 | >512 | >512 | 512 | 256 | 8 | 16 |
| 4007 | >512 | >512 | >512 | >512 | 512 | 128 | 16 | 4 |
| 3399 | >512 | >512 | >512 | >512 | >512 | 256 | 8 | 4 |
| 1034 | >512 | >512 | >512 | >512 | >512 | 128 | 16 | 8 |
2.3. Hemolytic and Cytotoxicity Activity of Peptides

2.4. NRC-16 is Non-Selective against Eukaryotic Membranes Using Liposomes
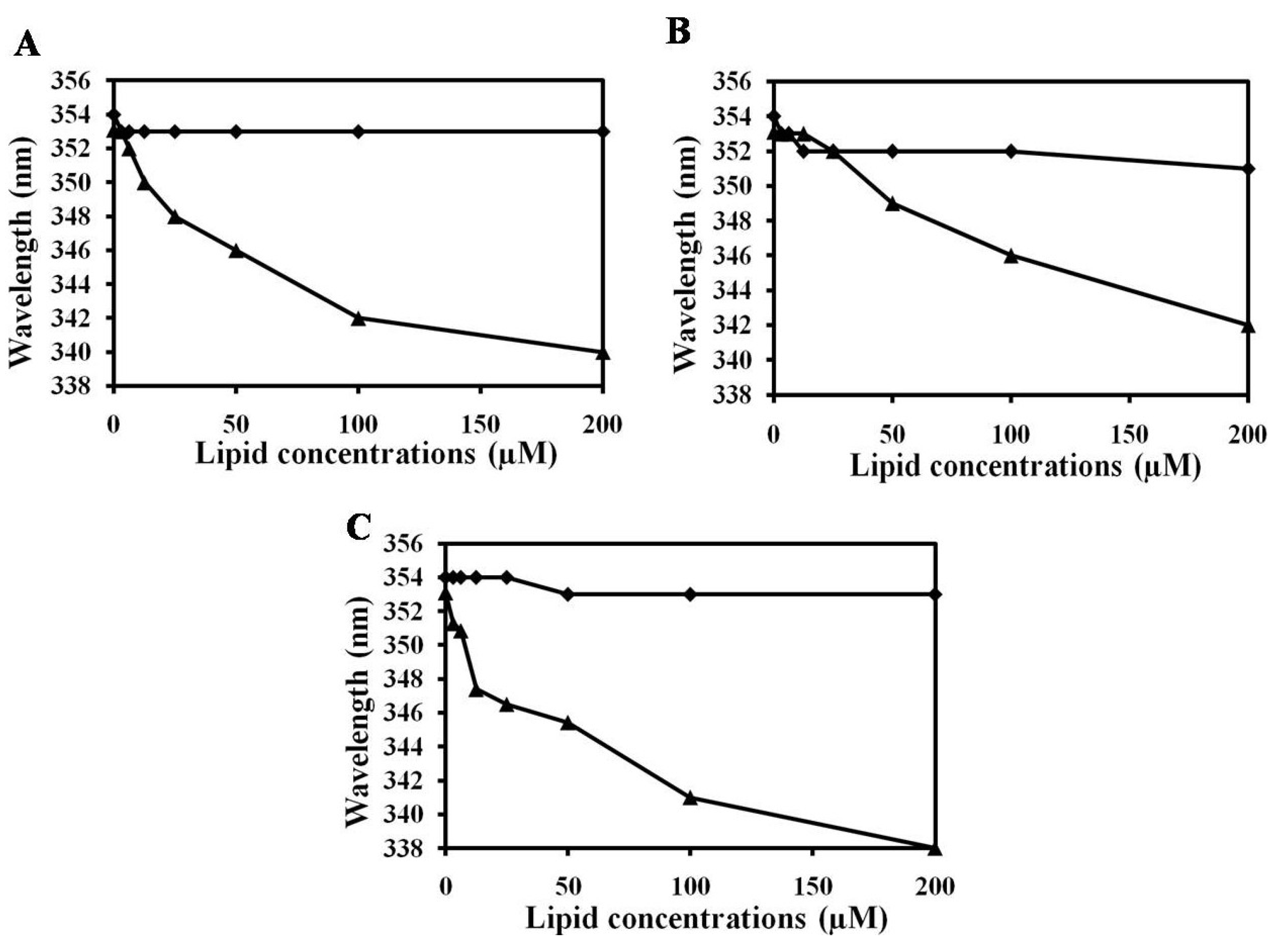
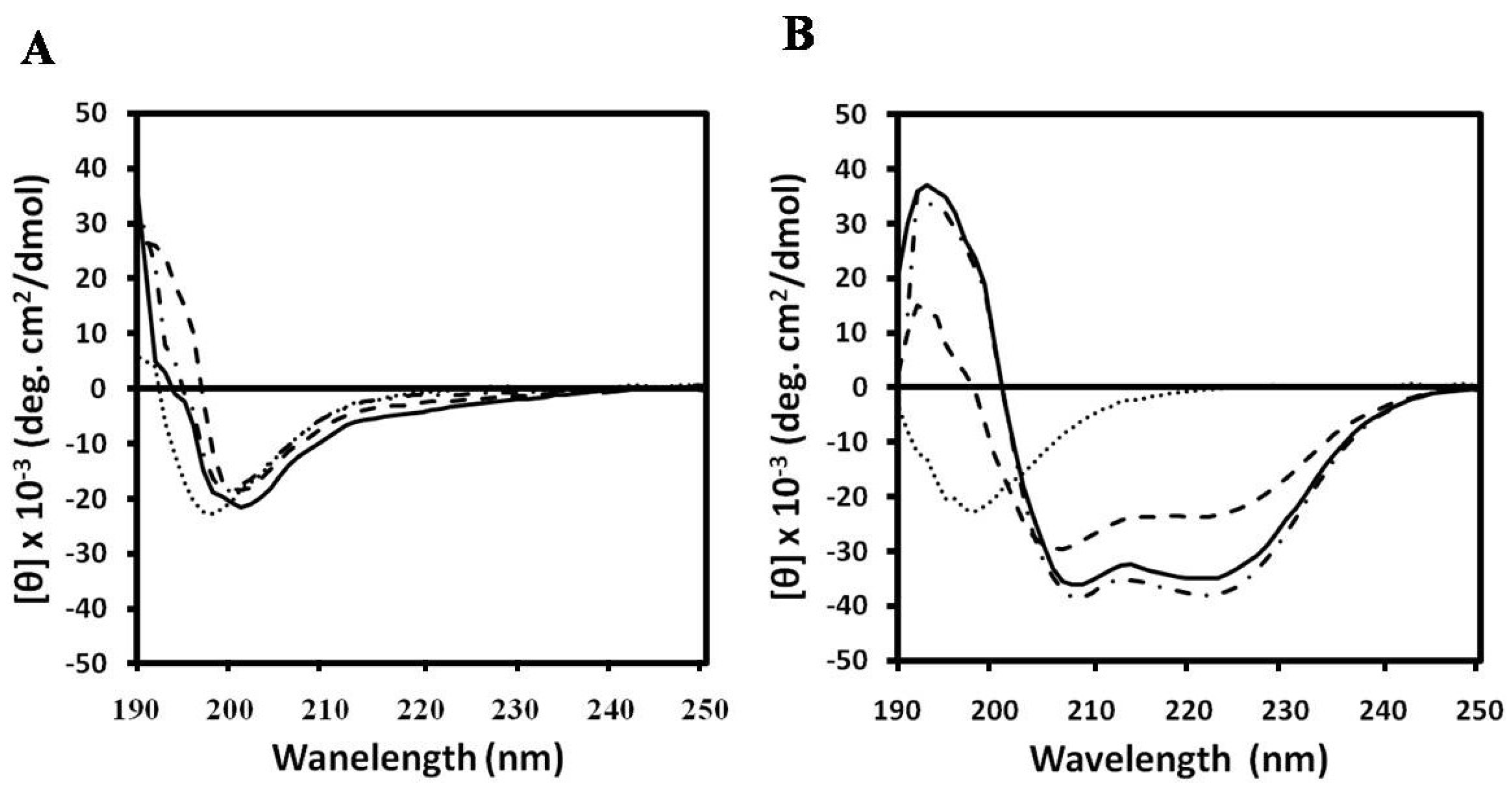
3. Experimental Section
3.1. Materials
3.2. Peptides Synthesis
3.3. Antibacterial Activity
3.4. Antifungal Activity
3.5. In Vitro Activity of the NRC-16 Peptide against Clinical Isolated P. aeruginosa and S. aureus Strains
3.6. Biofilm Forming Strains Subjected to Susceptibility Assay with NRC-16, Melittin and Some Conventional Antibiotics
3.7. Hemolysis and Cytotoxicity
3.8. Trp Fluorescence Assay
3.9. Calcein Leakage
3.10. CD Spectroscopy
4. Conclusions
Acknowledgements
References
- Park, S.C.; Park, Y.; Hahm, K.S. The role of antimicrobial peptides in preventing multidrug resistant bacterial infections and biofilm formation. Int. J. Mol. Sci. 2011, 12, 5971–5992. [Google Scholar] [CrossRef]
- Dzidic, S.; Suskovic, J.; Kos, B. Antibiotic resistance mechanisms in bacteria: Biochemical and genetic aspects. Food Technol. Biotechnol. 2008, 46, 11–21. [Google Scholar]
- Wright, G.D. Bacterial resistance to antibiotics: Enzymatic degradation and modification. Adv. Drug Deliv. Rev. 2005, 57, 1451–1470. [Google Scholar] [CrossRef]
- Lambert, P.A. Bacterial resistance to antibiotics: Modified target sites. Adv. Drug Deliv. Rev. 2005, 57, 1471–1485. [Google Scholar] [CrossRef]
- Kumar, A.; Schweizer, H.P. Bacterial resistance to antibiotics: Active efflux and reduced uptake. Adv. Drug Deliv. Rev. 2005, 57, 1486–1513. [Google Scholar] [CrossRef]
- Costerton, J.W.; Stewart, P.S.; Greenberg, E.P. Bacterial biofilms: A common cause of persistent infections. Science 1999, 284, 1318–1322. [Google Scholar] [CrossRef]
- Drenkard, E.; Ausubel, F.M. Pseudomonas biofilm formation and antibiotic resistance are linked to phenotypic variation. Nature 2002, 416, 740–743. [Google Scholar] [CrossRef]
- Ehrlich, G.D.; Veeh, R.; Wang, X.; Costerton, J.W.; Hayes, J.D.; Hu, F.Z.; Daigle, B.J.; Ehrlich, M.D.; Post, J.C. Mucosal biofilm formation on middle-ear mucosa in the chinchilla model of otitis media. JAMA 2002, 287, 1710–1715. [Google Scholar] [CrossRef]
- Singh, P.K.; Schaefer, A.L.; Parsek, M.R.; Moninger, T.O.; Welsh, M.J.; Greenberg, E.P. Quorum-sensing signals indicate that cystic fibrosis lungs are infected with bacterial biofilms. Nature 2000, 407, 762–764. [Google Scholar] [CrossRef]
- Perron, C.G.; Zasloff, M.; Bell, G. Experimental evolution of resistance to an antimicrobial peptide. Proc. Biol. Sci. 2006, 273, 251–256. [Google Scholar] [CrossRef]
- Yeaman, M.R.; Yount, N.Y. Mechanisms of antimicrobial peptide action and resistance. Pharmacol. Rev. 2003, 55, 27–55. [Google Scholar] [CrossRef]
- Giuliani, A.; Pirri, G.; Nicoletto, S.F. Antimicrobial peptides: An overview of a promising class of therapeutics. Cent. Eur. J. Biol. 2007, 2, 1–33. [Google Scholar] [CrossRef]
- Diaz, G.A. Defensins and cystein rich peptides: Two types of antimicrobial peptide in marine molluscs. Invert. Surviv. J. 2010, 7, 157–164. [Google Scholar]
- Rosa, R.D.; Barracco, M.A. Antimicrobial peptides in crustaceans. Invert. Surviv. J. 2010, 7, 262–284. [Google Scholar]
- Matsunaga, S.; Fusetani, N.; Konosu, S. Bioactive marine metabolites, IV. Isolation and the amino acid composition of discodermin A, an antimicrobial peptide, from the marine sponge Discodermia kiiensis. J. Nat. Prod. 1985, 48, 236–241. [Google Scholar] [CrossRef]
- Otero-González, A.J.; Magalhães, B.S.; Garcia-Villarino, M.; López-Abarratequi, C.; Sousa, D.A.; Dias, S.C.; Franco, O.L. Antimicrobial peptides from marine invertebrates as a new frontier for microbial infection control. FASEB J. 2010, 24, 1320–1334. [Google Scholar] [CrossRef]
- Noga, E.J.; Ullal, A.J.; Corrales, J.; Fernandes, J.M. Application of antimicrobial polypeptide host defenses to aquaculture: Exploitation of downregulation and upregulation responses. Comp. Biochem. Physiol. Part D 2011, 6, 44–54. [Google Scholar]
- Rakers, S.; Niklasson, L.; Steinhagen, D.; Kruse, C.; Schauber, J.; Sundell, K.; Paus, R. Antimicrobial peptides (AMPs) from fish epidermis: Perspective for investigative dermatology. J. Invest. Dermatol. 2013, 133, 1140–1149. [Google Scholar] [CrossRef]
- Patrzykat, A.; Gallant, J.W.; Seo, J.K.; Pytyck, J.; Douglas, S.E. Novel antimicrobial peptides derived from flatfish genes. Antimicrob. Agents Chemother. 2003, 47, 2464–2470. [Google Scholar] [CrossRef]
- Kim, J.Y.; Park, S.C.; Yoon, M.Y.; Hahm, K.S.; Park, Y. C-Terminal amidation of PMAP-23: Translocation to the inner membrane of Gram-negative bacteria. Amino Acids 2011, 40, 183–195. [Google Scholar] [CrossRef]
- Gopal, R.; Park, J.S.; Seo, C.H.; Park, Y. Applications of circular dichroism for structural analysis of gelatin and antimicrobial peptides. Int. J. Mol. Sci. 2012, 13, 3229–3244. [Google Scholar] [CrossRef]
- Findlay, B.; Zhanel, G.G.; Schweizer, F. Cationic amphiphiles, a new generation of antimicrobials inspired by the natural antimicrobial peptide scaffold. Antimicrob. Agents Chemother. 2010, 54, 4049–4058. [Google Scholar] [CrossRef]
- Cole, A.M.; Darouiche, R.O.; Legarda, D.; Connell, N.; Diamond, G. Characterisation of a fish and antimicrobial peptide: Gene expression, subcellular localization, and spectrum of activity. Antimicrob. Agents Chemother. 2000, 44, 2039–2045. [Google Scholar] [CrossRef]
- Cho, J.; Choi, H.; Lee, D.G. Influence of the N- and C-terminal regions of antimicrobial peptide plurocidin on antibacterial activity. J. Microbiol. Biotechnol. 2012, 22, 1367–1374. [Google Scholar] [CrossRef]
- Kolmer, H.L.; Taketomi, E.A.; Hazen, K.C.; Hughs, E.; Wilson, B.B.; Platts-Mills, T.A. Effect of combined and antifungal treatment in severe atopic dermatitis. J. Allergy Clin. Immunol. 1996, 98, 702–707. [Google Scholar] [CrossRef]
- National Committee for Clinical Laboratory Standards, Methods for Dilution Antimicrobial Susceptibility Tests for Bacteria that Grow Aerobically, Approved Standard M7-A6; National Committee for Clinical Laboratory Standards: Wayne, PA, USA, 2003.
- Jeong, N.; Kim, J.Y.; Park, S.C.; Lee, J.K.; Gopal, R.; Yoo, S.; Son, B.K.; Hahm, J.S.; Park, Y.; Hahm, K.S. Antibiotic and synergistic effect of Leu-Lys rich peptide against antibiotic resistant microoganisms isolated from patients with cholelithiasis. Biochem. Biophys. Res. Commun. 2010, 399, 581–586. [Google Scholar]
- Maisetta, G.; di Luca, M.; Esin, S.; Florio, W.; Brancatisano, F.L.; Bottai, D.; Campa, M.; Batoni, G. Evaluation of the inhibitory effects of human serum components on bactericidal activity of human beta defensin 3. Peptides 2008, 29, 1–6. [Google Scholar] [CrossRef]
- Goldman, M.J.; Anderson, G.M.; Stolzenberg, E.D.; Kari, U.P.; Zasloff, M.; Wilson, J.M. Human beta-defensin-1 is a salt-sensitive antibiotic in lung that is inactivated in cystic fibrosis. Cell 1997, 88, 553–560. [Google Scholar] [CrossRef]
- Lee, I.H.; Cho, Y.; Lehrer, R.I. Effects of pH and salinity on the antimicrobial properties of clavanins. Infect. Immun. 1997, 65, 2898–2903. [Google Scholar]
- Bowdish, D.M.; Davidson, D.J.; Lau, Y.E.; Lee, K.; Scott, M.G.; Hancock, R.E. Impact of LL-37 on anti-infective immunity. J. Leukoc. Biol. 2005, 77, 451–459. [Google Scholar]
- Tam, J.P.; Lu, Y.A.; Yang, J.L. Correlations of cationic charges with salt sensitivity and microbial specificity of cystine-stabilized β-strand antimicrobial peptides. J. Biol. Chem. 2002, 277, 50450–50456. [Google Scholar] [CrossRef]
- Høiby, N.; Krogh Johansen, H.; Moser, C.; Song, Z.; Ciofu, O.; Kharazmi, A. Pseudomonas aeruginosa and the in vitro and in vivo biofilm mode of growth. Microbes Infect. 2001, 3, 23–35. [Google Scholar] [CrossRef]
- Pruitt, B.A., Jr.; McManus, A.T.; Kim, S.H.; Goodwin, C.W. Burn wound infections: Current status. World J. Surg. 1998, 22, 135–145. [Google Scholar] [CrossRef]
- Tredget, E.E.; Shankowsky, H.A.; Rennie, R.; Burrell, R.E.; Logsetty, S. Pseudomonas infections in the thermally injured patient. Burns 2004, 30, 3–26. [Google Scholar] [CrossRef]
- Steven, L.P.; Philip, G.B. Biofilms and their potential role in wound healing. Wounds 2004, 16, 234–240. [Google Scholar]
- Jabalameli, F.; Mirsalehian, A.; Khoramian, B.; Aligholi, M.; Khoramrooz, S.S.; Asadollahi, P.; Taherikalani, M.; Emaneini, M. Evaluation of biofilm production and characterization of genes encoding type III secretion system among Pseudomonas aeruginosa isolated from burn patients. Burns 2012, 38, 1192–1197. [Google Scholar] [CrossRef]
- Harrison-Balestra, C.; Cazzaniga, A.L.; Davis, S.C.; Mertz, P.M. A wound-isolated Pseudomonas aeruginosa grows a biofilm in vitro within 10 hours and is visualized by light microscopy. Dermatol. Surg. 2003, 29, 631–635. [Google Scholar] [CrossRef]
- Serralta, V.W.; Harrison-Balestra, C.; Cazzaniga, A.L.; Davis, S.C.; Mertz, P.M. Lifestyles of bacteria in wounds: Presence of biofilms? Wounds 2001, 13, 29–34. [Google Scholar]
- Schaber, J.A.; Triffo, W.J.; Suh, S.J.; Oliver, J.W.; Hastert, M.C.; Griswold, J.A.; Auer, M.; Hamood, A.N.; Rumbaugh, K.P. Pseudomonas aeruginosa forms biofilms in acute infection independent of cell-to-cell signaling. Infect. Immun. 2007, 75, 3715–3721. [Google Scholar] [CrossRef]
- Sauer, K.; Camper, K.; Ehrlich, G.D.; Costerton, J.W.; Davies, D.G. Pseudomonas aeruginosa displays multiple phenotypes during development as a biofilm. J. Bacteriol. 2002, 184, 1140–1154. [Google Scholar] [CrossRef]
- Whiteley, M.; Bangera, M.G.; Bumgarner, R.E.; Parsek, M.R.; Teitzel, G.M.; Lory, S.; Greenberg, E.P. Gene expression in Pseudomonas aeruginosa biofilms. Nature 2001, 413, 860–864. [Google Scholar] [CrossRef]
- Vidaillac, C.; Benichou, L.; Duval, R.E. In vitro synergy of colistin combinations against colistin resistant Acinetobacter baumannii, Pseudomonas aeruginosa, and Klebsiella pneumoniae isolates. Antimicrob. Agents Chemother. 2012, 56, 4856–4861. [Google Scholar] [CrossRef]
- Donlan, R.M.; Costerton, J.W. Biofilms: Survival mechanisms of clinically relevant microorganisms. Clin. Microbiol. Rev. 2002, 15, 167–193. [Google Scholar] [CrossRef]
- Sandoe, J.A.; Wysome, J.; West, A.P.; Heritage, J.; Wilcox, M.H. Measurement of ampicillin, vancomycin, linezolid and gentamicin activity against Enterococcal biofilms. J. Antimicrob. Chemother. 2006, 57, 767–770. [Google Scholar] [CrossRef]
- Evans, R.C.; Holmes, C.J. Effect of vancomycin hydrochloride on Staphylococcus epidermidis biofilm associated with silicone elastomer. Antimicro. Agents Chemother. 1987, 31, 889–894. [Google Scholar] [CrossRef]
- Overhage, J.; Campisano, A.; Bains, M.; Torfs, E.C.; Rehm, B.H.; Hancock, R.E. Human host defense peptide LL-37 prevents bacterial biofilm formation. Infect. Immun. 2008, 76, 4176–4182. [Google Scholar] [CrossRef]
- Wei, G.X.; Campagna, A.N.; Bobek, L.A. Effect of MUC7 peptides on the growth of bacteria and on Streptococcus mutans biofilm. J. Antimicrob. Chemother. 2006, 57, 1100–1109. [Google Scholar] [CrossRef]
- Eckert, R.; Brady, K.M.; Greenberg, E.P.; Qi, F.; Yarbrough, D.K.; He, J.; McHardy, I.; Anderson, M.H.; Shi, W. Enhancement of antimicrobial activity against Pseudomonas aeruginosa by coadministration of G10KHc and tobramycin. Antimicrob. Agents Chemother. 2006, 50, 3833–3838. [Google Scholar] [CrossRef]
- Pamp, S.J.; Gjermansen, M.; Johansen, H.K.; Tolker-Nielsen, T. Tolerance to the antimicrobial peptide colistin in Pseudomonas aeruginosa biofilms is linked to metabolically active cells, and depends on the pmr and mexAB-oprM genes. Mol. Microbiol. 2008, 68, 223–240. [Google Scholar] [CrossRef]
- Nagant, C.; Pitts, B.; Nazmi, K.; Vandenbranden, M.; Bolscher, J.G.; Stewart, P.S.; Dehaye, J.P. Identification of peptides derived from the human antimicrobial peptide LL-37 active against biofilms formed by Pseudomonas aeruginosa using a library of truncated fragments. Antimicrob. Agents Chemother. 2012, 56, 5698–5708. [Google Scholar] [CrossRef]
- Choi, H.; Lee, D.G. Antimicrobial peptide pleurocidin synergizes with antibiotics through hydroxyl radical formation and membrane damage, and exerts antibiofilm activity. Biochim. Biophys. Acta 2012, 1820, 1831–1838. [Google Scholar] [CrossRef]
- Dartois, V.; Sanchez-Quesada, J.; Cabezas, E.; Chi, E.; Dubbelde, C.; Dunn, C.; Granja, J.; Gritzen, C.; Weinberger, D.; Ghadiri, M.R.; et al. Systemic antibacterial activity of novel synthetic cyclic peptides. Antimicrob. Agents Chemother. 2005, 49, 3302–3310. [Google Scholar] [CrossRef]
- Bowler, P.G.; Welsby, S.; Towers, V.; Booth, R.; Hogarth, A.; Rowlands, V.; Joseph, A.; Jones, S.A. Multidrug reistant organisms, wounds and topical application. Int. Wound J. 2012, 9, 387–396. [Google Scholar] [CrossRef]
- Nidadavolu, P.; Amor, W.; Tran, P.L.; Dertien, J.; Colmer-Hamood, J.A.; Hamood, A.N. Garlic ointment inhibits biofilm formation by bacterial pathogens from burn wounds. J. Med. Microbiol. 2012, 61, 662–671. [Google Scholar] [CrossRef]
- Ngo, Q.D.; Vickery, K.; Deva, A.K. The effect of topical negative pressure on wound biofilms using an in vitro wound model. Wound Repair Regen. 2012, 20, 83–90. [Google Scholar] [CrossRef]
- Ryu, S.; Choi, S.Y.; Acharya, S.; Chun, Y.J.; Gurley, C.; Park, Y.; Armstrong, C.A.; Song, P.I.; Kim, B.J. Antimicrobial and anti-inflammatory effects of cecropin A (1-8)-magainin 2(1-12) hybrid peptide analog P5 against Malassezia furfur infection in human ketatinocytes. J. Invest. Dermatol. 2011, 131, 1677–1683. [Google Scholar] [CrossRef]
- Javadpour, M.M.; Juban, M.M.; Lo, W.C.; Bishop, S.M.; Alberty, J.B.; Cowell, S.M.; Becker, C.L.; Mclaughlin, M.L. De novo antimicrobial peptides with low mammalian cell toxicity. J. Med. Chem. 1996, 39, 3107–3113. [Google Scholar] [CrossRef]
- Fernandez-Lopez, S.; Kim, H.S.; Choi, E.C.; Delgado, M.; Granja, J.R.; Khasanov, A.; Kraehenbuehl, K.; Long, G.; Weinberger, D.A.; Wilcoxen, K.M.; et al. Antibacterial agents based on the cyclic d,l-α-peptide architecture. Nature 2001, 412, 452–455. [Google Scholar] [CrossRef]
- Chan, D.I.; Prenner, E.J.; Vogel, H.J. Tryptophan- and arginine-rich antimicrobial peptides: Structures and mechanisms of action. Biochim. Biophys. Acta 2006, 1758, 1184–1202. [Google Scholar] [CrossRef]
- Schibli, D.J.; Hwang, P.M.; Vogel, H.J. The structure of the antimicrobial active center of lactoferricin B bound to sodium dodecyl sulfate micelles. FEBS Lett. 1999, 446, 213–217. [Google Scholar] [CrossRef]
- Jing, W.; Hunter, H.N.; Hagel, J.; Vogel, H.J. The structure of the antimicrobial peptide Ac-RRWWRF-NH2 bound to micelles and its interactions with phospholipid bilayers. J. Pept. Res. 2003, 61, 219–229. [Google Scholar]
- Glukhov, E.; Stark, M.; Burrows, L.L.; Deber, C.M. Basis for selectivity of cationic antimicrobial peptides for bacterial versus mammalian membranes. J. Biol. Chem. 2005, 280, 33960–33967. [Google Scholar]
- Andra, J.; Monreal, D.; Martinez de Tejada, G.; Olak, C.; Brezesinski, G.; Gomez, S.S.; Goldmann, T.; Bartels, R.; Brandenburg, K.; Moriyon, I. Rationale for the design of shortened derivatives of the NK-lysin-derived antimicrobial peptide NK-2 with improved activity against Gram-negative pathogens. J. Biol. Chem. 2007, 282, 14719–14728. [Google Scholar] [CrossRef]
- Hawrani, A.; Howe, R.A.; Walsh, T.R.; Dempsey, C.E. Origin of low mammalian cell toxicity in a class of highly active antimicrobial amphipathic helical peptides. J. Biol. Chem. 2008, 283, 18636–18645. [Google Scholar]
- Gopal, R.; Park, S.C.; Ha, K.J.; Cho, S.J.; Kim, S.W.; Song, P.I.; Nah, J.W.; Park, Y.; Hahm, K.S. Effect of Leucine and Lysine substitution on the antimicrobial activity and evaluation of the mechanism of the HPA3NT3 analog peptide. J. Pept. Sci. 2009, 15, 589–594. [Google Scholar] [CrossRef]
- Pandey, B.K.; Ahmad, A.; Asthana, N.; Azmi, S.; Srivastava, R.M.; Srivastava, S.; Verma, R.; Vishwakarma, A.L.; Ghosh, J.K. Cell-selective lysis by novel analogues of melittin against human red blood cells and Escherichia coli. Biochemistry 2010, 49, 7920–7929. [Google Scholar] [CrossRef]
- Gopal, R.; Seo, C.H.; Song, P.I.; Park, Y. Effect of repetitive lysine-tryptophan motifs on the bactericidal activity of antimicrobial peptides. Amino Acids 2013, 44, 645–660. [Google Scholar] [CrossRef]
- Gill, S.C.; von Hippel, P.H. Calculation of protein extinction coefficients from amino acid sequence data. Anal. Biochem. 1989, 182, 319–326. [Google Scholar] [CrossRef]
- Park, S.C.; Kim, J.Y.; Lee, J.K.; Hwang, I.; Cheong, H.; Nah, J.W.; Hahm, K.S.; Park, Y. Antifungal mechanism of a novel antifungal protein from pumpkin rinds against various fungal pathogens. J. Agric. Food. Chem. 2009, 57, 9299–9304. [Google Scholar]
- Rolli, E.; Ragni, E.; de Medina-Redondo, M.; Arroyo, J.; de Aldana, C.R.; Popolo, L. Expression, stability, and replacement of glucan-remodeling enzymes during developmental transition in Saccharomyces cerevisiae. Mol. Biol. Cell. 2011, 22, 1585–1598. [Google Scholar] [CrossRef]
- Gopal, R.; Na, H.; Seo, C.H.; Park, Y. Antifungal activity of (KW)n or (RW)n peptide against Fusarium solani and Fusarium oxysporum. Int. J. Mol. Sci. 2012, 13, 15042–15053. [Google Scholar] [CrossRef]
- Christensen, G.D.; Simpson, W.A.; Younger, J.J.; Baddour, F.F.; Barrett, D.M.; Melton, D.M.; Beachey, E.H. Adherence of coagulase-negative staphylococci to plastic tissue culture plates: A quantitative model for the adherence of staphylococci to medical devices. J. Clin. Microbiol. 1985, 22, 996–1006. [Google Scholar]
- Mayer, L.D.; Hope, M.J.; Cullis, P.R. Vesicles of variable sizes produced by a rapid extrusion procedure. Biochem. Biophys. Acta 1986, 858, 161–168. [Google Scholar] [CrossRef]
- Stewart, J.C.M. Colorimetric determination of phospholipids with ammonium ferrothiocyanate. Anal. Biochem. 1980, 104, 10–14. [Google Scholar]
- Gopal, R.; Lee, J.K.; Lee, J.H.; Kim, Y.G.; Oh, G.C.; Seo, C.H.; Park, Y. Effect of repetitive lysine-tryptophan motifs on the eukaryotic membrane. Int. J. Mol. Sci. 2013, 14, 2190–2202. [Google Scholar] [CrossRef]
- Chongsiriwatana, N.P.; Barron, A.E. Comparing bacterial membrane interactions of antimicrobial peptides and their mimics. Methods Mol. Biol. 2010, 618, 171–182. [Google Scholar] [CrossRef]
- Allen, T.M.; Cleland, L.G. Serum-induced leakage of liposome contents. Biochim. Biophys. Acta 1980, 10, 418–426. [Google Scholar]
© 2013 by the authors; licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution license (http://creativecommons.org/licenses/by/3.0/).
Share and Cite
Gopal, R.; Lee, J.H.; Kim, Y.G.; Kim, M.-S.; Seo, C.H.; Park, Y. Anti-Microbial, Anti-Biofilm Activities and Cell Selectivity of the NRC-16 Peptide Derived from Witch Flounder, Glyptocephalus cynoglossus. Mar. Drugs 2013, 11, 1836-1852. https://doi.org/10.3390/md11061836
Gopal R, Lee JH, Kim YG, Kim M-S, Seo CH, Park Y. Anti-Microbial, Anti-Biofilm Activities and Cell Selectivity of the NRC-16 Peptide Derived from Witch Flounder, Glyptocephalus cynoglossus. Marine Drugs. 2013; 11(6):1836-1852. https://doi.org/10.3390/md11061836
Chicago/Turabian StyleGopal, Ramamourthy, Jun Ho Lee, Young Gwon Kim, Myeong-Sun Kim, Chang Ho Seo, and Yoonkyung Park. 2013. "Anti-Microbial, Anti-Biofilm Activities and Cell Selectivity of the NRC-16 Peptide Derived from Witch Flounder, Glyptocephalus cynoglossus" Marine Drugs 11, no. 6: 1836-1852. https://doi.org/10.3390/md11061836





