Pathways of Lipid Metabolism in Marine Algae, Co-Expression Network, Bottlenecks and Candidate Genes for Enhanced Production of EPA and DHA in Species of Chromista
Abstract
:1. Introduction
2. Long Chain Polyunsaturated Fatty Acids (LC-PUFAs) for Human Health
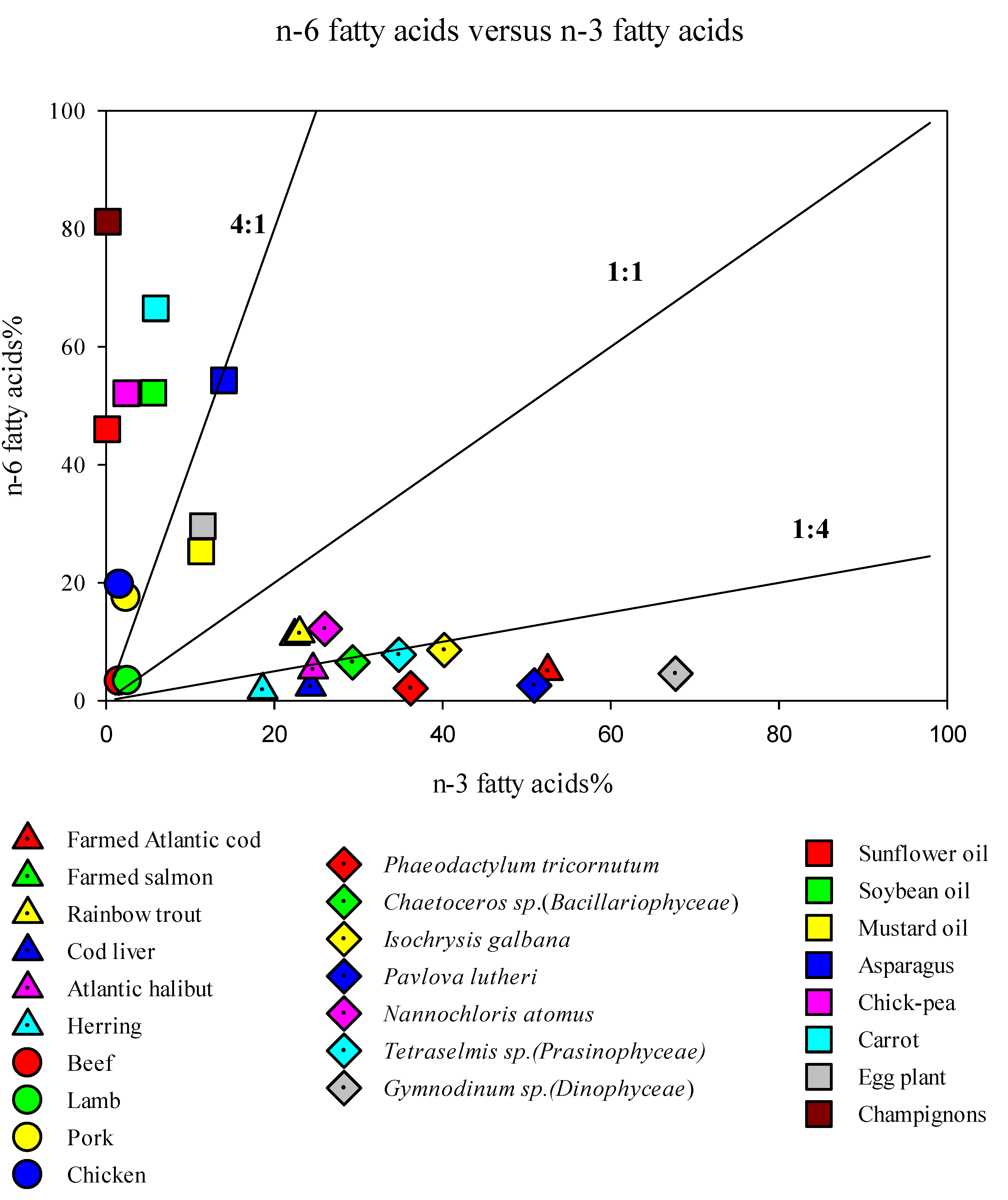
3. LC-PUFA Sources and the Need for Alternatives
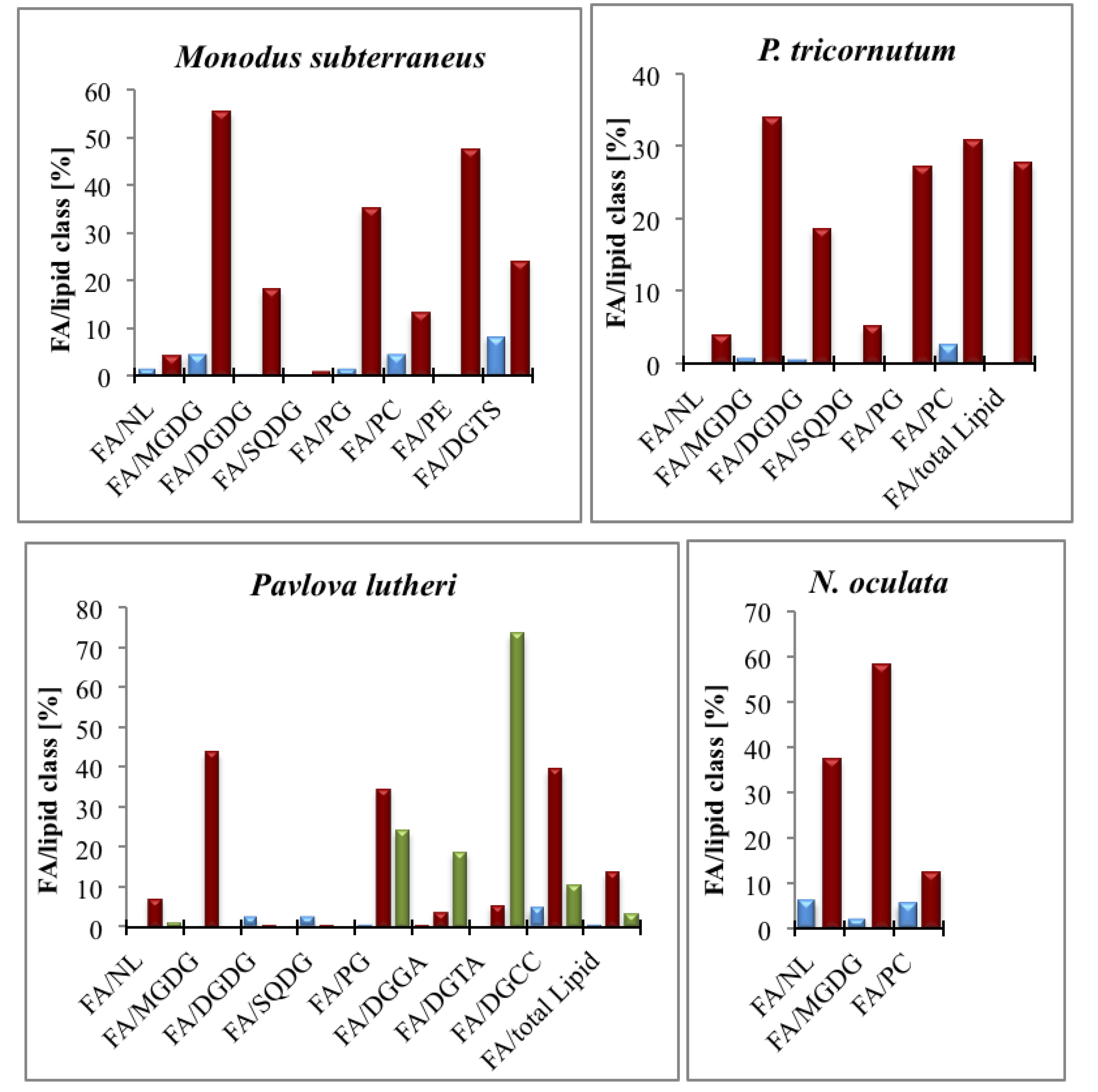
4. Chromista and Their LC-PUFA Content and Function
5. Eicosapentaenoic Acid (EPA, 20:5n-3) Synthesis in Phaedoactylum tricornutum
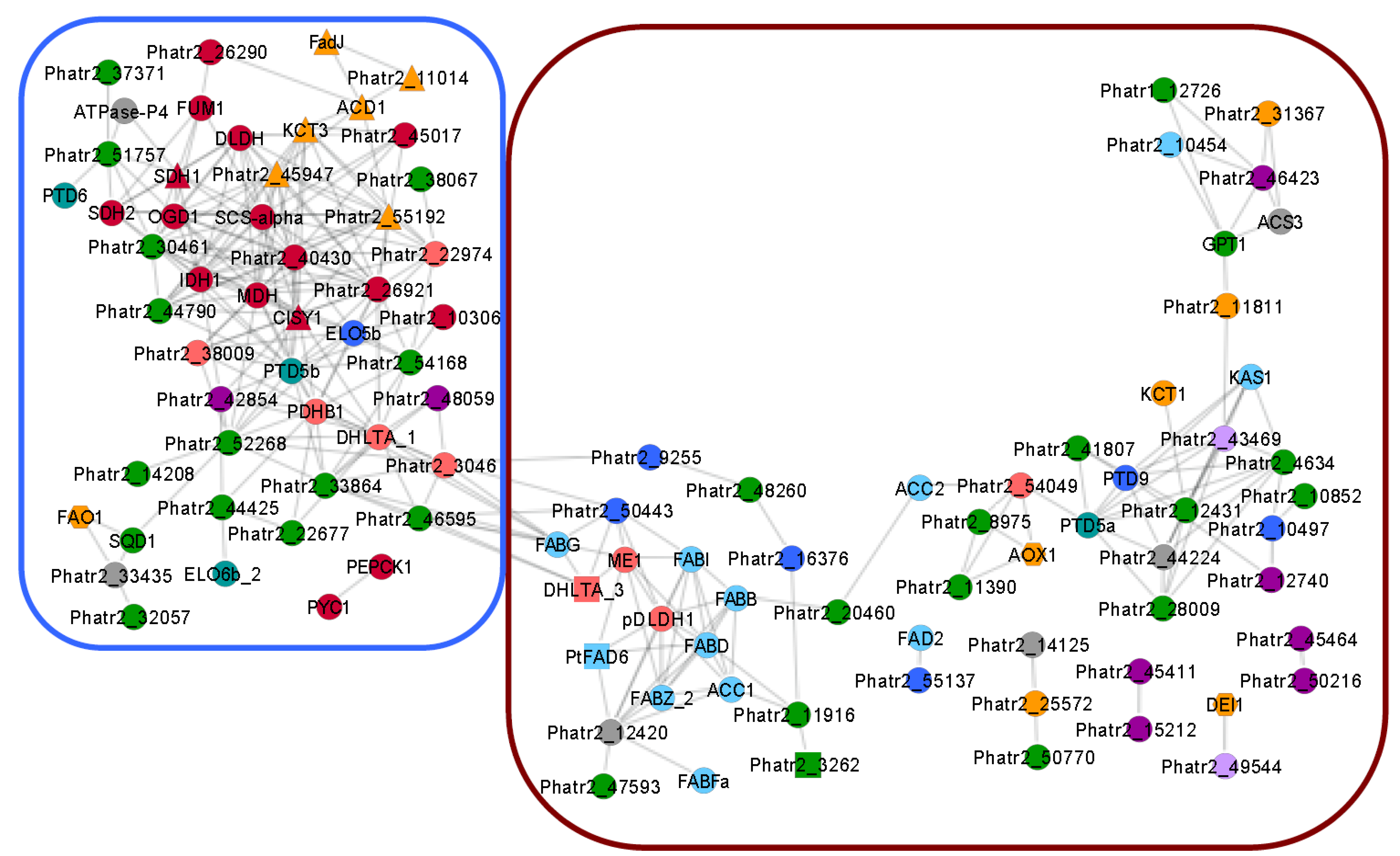
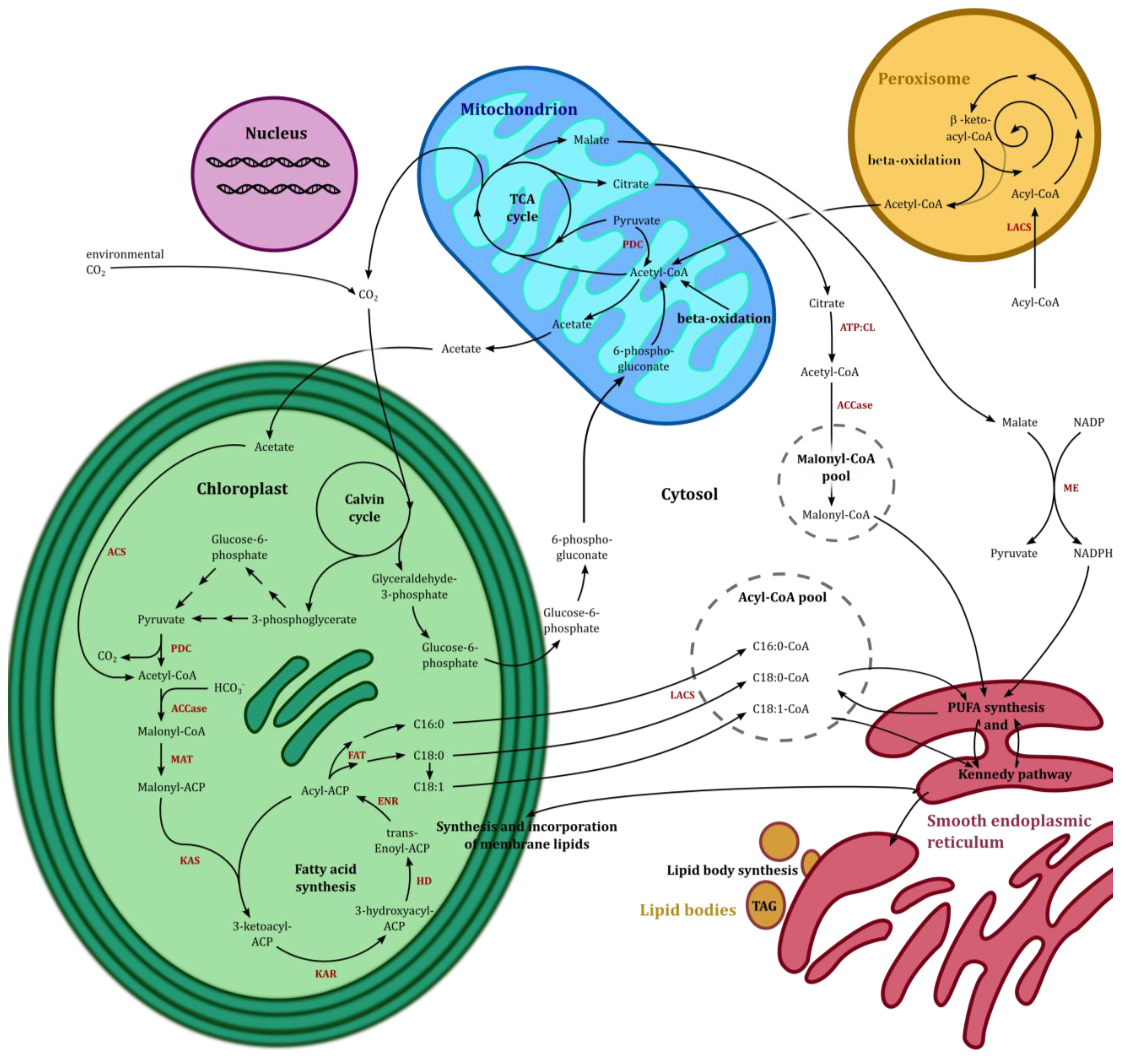
5.1. The TCA Cycle and β-Oxidation
5.2. De Novo Fatty Acid Synthesis and Hexadecatrienoic Acid Synthesis

5.3. Acyl-CoA Activator and EPA Synthesis
| Gene Number Enzyme Name Annotation | Present in Cluster (Figure 3) | |||
|---|---|---|---|---|
| Elongases predicted to be membrane bound | ||||
| Phatr2_16376 | - | Long chain fatty acid elongase | √ | |
| Phatr2_9255 | - | Polyunsaturated fatty acid elongase | √ | |
| Phatr2_49867 | - | Long chain fatty acid elongase | ||
| Phatr2_34485 | Elo5b | Long chain fatty acid elongases, membrane bound | √ | |
| Phatr2_25360 | - | Probably a short-chain dehydrogenase/reductase | ||
| Desaturases predicted to be microsomal or membrane bound | ||||
| Phatr2_55137 | - | Fatty acid desaturase | √ | |
| Phatr2_50443 | - | Fatty acid desaturase, cytochrome b5 motif | √ | |
| Phatr2_46275 | - | Fatty acid desaturase, putative | ||
| Phatr2_44622 | - | Fatty acid desaturase | ||
| Phatr2_22510 | - | Fatty acid desaturase, cytochrome b5 motif | ||
5.4. EPA Incorporation into Galactosylglycerides and Triacylglycerides
5.5. Genetic Drivers for n-3 LC-PUFA Synthesis
6. EPA Synthesis in Other Chromista
7. DHA Synthesis in Other Chromista
8. Conclusions
Fatty Acid Nomenclature and Abbreviations
| ADA | adrenic acid (22:4n-6) |
| ALA | α-linolenic acid (18:3n-3) |
| ARA | arachidonic acid (20:4n-6) |
| DGLA | dihomo-γ-linolenic acid (20:3n-6) |
| DHA | docosahexaenoic acid (20:6n-3) |
| DPA | docosapentaenoic acid (22:5n-3) |
| EDA | eicosadienoic acid (20:2n-6) |
| EPA | eicosapentaenoic acid (20:5n-3) |
| ETA | eicosatetraenoic acid (20:4n-3) |
| ETrA | eicosatrienoic acid (20:3n-3) |
| FA(s) | fatty acid(s) |
| GLA | γ-linolenic acid (18:3n-6) |
| HAD | hexadecadienoic acid (16:2n-4) |
| HTA | hexadecatrienoic acid (16:3n-4) |
| LA | linoleic acid (18:2n-6) |
| LC-PUFA(s) | long chain polyunsaturated fatty acid(s), >C20 |
| MA | myristic acid (14:0) |
| OA | oleic acid (18:1n-9) |
| PA | Palmitic acid (16:0) |
| POA | Palmitoleic acid (16:1n-7) |
| PONA | palmitolenic acid (16:2n-7) |
| PUFA(s) | polyunsaturated fatty acid(s), >C18 |
| SA | stearic acid (18:0) |
| STA | stearidonic acid (18:4n-3) |
| THA | tetracosahexaenoic acid (24:6n-3) |
Acknowledgments
Conflicts of Interest
References
- Kelly, P.B.; Reiser, R.; Hood, D.W. The origin of the marine polyunsaturated fatty acids. Composition of some marine plankton. J. Am. Oil Chem. Soc. 1959, 36, 104–106. [Google Scholar] [CrossRef]
- Yaguchi, T.; Tanaka, S.; Yokochi, T.; Nakahara, T.; Higashihara, T. Production of high yields of docosahexaenoic acid by Schizochytrium sp. strain SR21. J. Am. Oil Chem. Soc. 1997, 74, 1431–1434. [Google Scholar] [CrossRef]
- Yongmanitchai, W.; Ward, O.P. Growth of and omega-3 fatty acid production by Phaeodactylum tricornutum under different culture conditions. Appl. Environ. Microbiol. 1990, 57, 419–425. [Google Scholar]
- Sukenik, A. Ecophysiological considerations in the optimization of eicosapentaenoic acid production by Nannochloropsis sp. (Eustigmatophyceae). Bioresour. Technol. 1991, 35, 263–269. [Google Scholar] [CrossRef]
- Wan, C.; Bai, F.-W.; Zhao, X.-Q. Effects of nitrogen concentration and media replacement on cell growth and lipid production of oleaginous marine microalga Nannochloropsis oceanica DUT01. Biochem. Eng. J. 2013, 78, 32–38. [Google Scholar] [CrossRef]
- Cavalier-Smith, T. Kingdoms protozoa and chromista and the eozoan root of the eukaryotic tree. Biol. Lett. 2010, 6, 342–345. [Google Scholar] [CrossRef]
- Cavalier-Smith, T. Principles of protein and lipid targeting in secondary symbiogenesis: euglenoid, dinoflagellate, and sporozoan plastid origins and the eukaryote family tree. J. Eukaryot. Microbiol. 1999, 46, 347–366. [Google Scholar] [CrossRef]
- Chang, G.; Luo, Z.; Gu, S.; Wu, Q.; Chang, M.; Wang, X. Fatty acid shifts and metabolic activity changes of Schizochytrium sp. S31 cultured on glycerol. Bioresour. Technol. 2013, 142, 255–260. [Google Scholar] [CrossRef]
- Ward, O.P.; Singh, A. Omega-3/6 fatty acids: Alternative sources of production. Process Biochem. 2005, 40, 3627–3652. [Google Scholar] [CrossRef]
- Hu, Q.; Sommerfeld, M.R.; Jarvis, E.; Ghirardi, M.; Posewitz, M.; Seibert, M.; Darzins, A. Microalgal triacylglycerols as feedstocks for biofuel production: Perspectives and advances. Plant J. 2008, 54, 521–639. [Google Scholar]
- Adarme-Vega, T.C.; Lim, D.K.; Timmins, M.; Vernen, F.; Li, Y.; Schenk, P.M. Microalgal biofactories: A promising approach towards sustainable omega-3 fatty acid production. Microb. Cell Fact. 2012, 11, 96. [Google Scholar] [CrossRef]
- Barclay, W.R. Method of aquaculture comprising feeding microflora having a small cell aggregate size. US Patent 5,688,500, 18 November 1997. [Google Scholar]
- Martins, D.A.; Custodio, L.; Barreira, L.; Pereira, H.; Ben-Hamadou, R.; Varela, J.; Abu-Salah, K.M. Alternative sources of n-3 long-chain polyunsaturated fatty acids in marine microalgae. Mar. Drugs 2013, 11, 2259–2281. [Google Scholar] [CrossRef]
- Hu, H.; Gao, K. Optimization of growth and fatty acid composition of a unicellular marine picoplankton, Nannochloropsis sp., with enriched carbon sources. Biotechnol. Lett. 2003, 25, 421–425. [Google Scholar] [CrossRef]
- Lu, C.; Rao, K.; Hall, D.; Vonshak, A. Production of eicosapentaenoic acid (EPA) in Monodus subterraneus grown in a helical tubular photobioreactor as affected by cell density and light intensity. J. Appl. Phycol. 2001, 13, 517–522. [Google Scholar] [CrossRef]
- Wen, Z.-Y.; Chen, F. Production potential of eicosapentaenoic acid by the diatom Nitzschia laevis. Biotechnol. Lett. 2000, 22, 727–733. [Google Scholar] [CrossRef]
- Yang, Z.-K.; Niu, Y.-F.; Ma, Y.-H.; Xue, J.; Zhang, M.-H.; Yang, W.-D.; Liu, J.-S.; Lu, S.-H.; Guan, Y.; Li, H.-Y. Molecular and cellular mechanisms of neutral lipid accumulation in diatom following nitrogen deprivation. Biotechnol. Biofuels 2013, 6, 67. [Google Scholar] [CrossRef]
- Fidalgo, J.P.; Cid, A.; Torres, E.; Sukenik, A.; Herrero, C. Effects of nitrogen source and growth phase on proximate biochemical composition, lipid classes and fatty acid profile of the marine microalga Isochrysis galbana. Aquaculture 1998, 166, 105–116. [Google Scholar] [CrossRef]
- Taoka, Y.; Nagano, N.; Okita, Y.; Izumida, H.; Sugimoto, S.; Hayashi, M. Influences of culture temperature on the growth, lipid content and fatty acid composition of Aurantiochytrium sp. strain mh0186. Mar. Biotechnol. 2009, 11, 368–374. [Google Scholar] [CrossRef]
- Yokochi, T.; Honda, D.; Higashihara, T.; Nakahara, T. Optimization of docosahexaenoic acid production by Schizochytrium limacinum SR21. Appl. Microbiol. Biotechnol. 1998, 49, 72–76. [Google Scholar] [CrossRef]
- Chepurnov, V.A.; Mann, D.G.; von Dassow, P.; Vanormelingen, P.; Gillard, J.; Inze, D.; Sabbe, K.; Vyverman, W. In search of new tractable diatoms for experimental biology. BioEssays 2008, 30, 692–702. [Google Scholar] [CrossRef]
- Radakovits, R.; Jinkerson, R.E.; Darzins, A.; Posewitz, M.C. Genetic engineering of algae for enhanced biofuel production. Eukaryot. Cell 2010, 9, 486–501. [Google Scholar] [CrossRef]
- Graham, I.A.; Larson, T.; Napier, J.A. Rational metabolic engineering of transgenic plants for biosynthesis of omega-3 polyunsaturates. Curr. Opin. Biotechnol. 2007, 18, 142–147. [Google Scholar] [CrossRef]
- Drexler, H.; Spiekermann, P.; Meyer, A.; Domergue, F.; Zank, T.; Sperling, P.; Abbadi, A.; Heinz, E. Metabolic engineering of fatty acids for breeding of new oilseed crops: Strategies, problems and first results. J. Plant Physiol. 2003, 160, 779–802. [Google Scholar]
- Hoffmann, M.; Wagner, M.; Abbadi, A.; Fulda, M.; Feussner, I. Metabolic engineering of omega 3-very long chain polyunsaturated fatty acid production by an exclusively acyl-CoA-dependent pathway. J. Biol. Chem. 2008, 283, 22352–22362. [Google Scholar]
- Sharma, K.K.; Schuhmann, H.; Schenk, P.M. High lipid induction in microalgae for biodiesel production. Energies 2012, 5, 1532–1553. [Google Scholar]
- Harwood, J.L.; Guschina, I.A. The versatility of algae and their lipid metabolism. Biochimie 2009, 91, 679–684. [Google Scholar] [CrossRef]
- Guschina, I.A.; Harwood, J.L. Lipids and lipid metabolism in eukaryotic algae. Prog. Lipid Res. 2006, 45, 160–186. [Google Scholar] [CrossRef]
- Khozin-Goldberg, I.; Cohen, Z. Unraveling algal lipid metabolism: Recent advances in gene identification. Biochemie 2011, 93, 91–100. [Google Scholar] [CrossRef]
- Liu, B.S.; Benning, C. Lipid metabolism in microalgae distinguishes itself. Curr. Opin. Biotechnol. 2013, 24, 300–309. [Google Scholar] [CrossRef]
- Das, U.N. Essential fatty acids—A review. Curr. Pharm. Biotechnol. 2006, 7, 467–482. [Google Scholar]
- Tocher, D.R. Fatty acid requirements in ontogeny of marine and freshwater fish. Aquac. Res. 2010, 41, 717–732. [Google Scholar] [CrossRef]
- Plourde, M.; Cunnane, S.C. Extremely limited synthesis of long chain polyunsaturates in adults: Implications for their dietary essentiality and use as supplements. Appl. Physiol. Nutr. Metab. 2007, 32, 619–634. [Google Scholar] [CrossRef]
- Khozin-Goldberg, I.; Iskandarov, U.; Cohen, Z. LC-PUFA from photosynthetic microalgae: Occurrence, biosynthesis, and prospects in biotechnology. Appl. Microbiol. Biotechnol. 2011, 91, 905–915. [Google Scholar] [CrossRef]
- Bell, J.G.; McEvoy, L.A.; Estevez, A.; Shields, R.J.; Sargent, J.R. Optimising lipid nutrition in first-feeding flatfish larvae. Aquaculture 2003, 227, 211–220. [Google Scholar] [CrossRef]
- Harel, M.; Koven, W.; Lein, I.; Bar, Y.; Behrens, P.; Stubblefield, J.; Zohar, Y.; Place, A.R. Advanced DHA, EPA and ArA enrichment materials for marine aquaculture using single cell heterotrophs. Aquaculture 2002, 213, 347–362. [Google Scholar] [CrossRef]
- Li, Q.; Ai, Q.; Mai, K.; Xu, W.; Zheng, Y. A comparative study: Invitro effects of EPA and DHA on immune functions of head-kidney macrophages isolated from large yellow croaker (Larmichthys crocea). Fish Shellfish Immunol. 2013, 35, 933–940. [Google Scholar] [CrossRef]
- Kuratko, C.N.; Barrett, E.C.; Nelson, E.B.; Salem, N., Jr. The relationship of docosahexaenoic acid (DHA) with learning and behavior in healthy children: A review. Nutrients 2013, 5, 2777–2810. [Google Scholar] [CrossRef]
- Kremmyda, L.S.; Tvrzicka, E.; Stankova, B.; Zak, A. Fatty acids as biocompounds: Their role in human metabolism, health and disease-a review. Part 2: Fatty acid physiological roles and applications in human health and disease. Biomed. Pap. Med. Fac. Univ. Palacky Olomouc. Czech. Repub. 2011, 155, 195–218. [Google Scholar] [CrossRef]
- Simopoulos, A.P. The importance of the ratio of omega-6/omega-3 essential fatty acids. Biomed. Pharmacother. 2002, 56, 365–379. [Google Scholar] [CrossRef]
- Sinn, N.; Milte, C.M.; Street, S.J.; Buckley, J.D.; Coates, A.M.; Petkov, J.; Howe, P.R.C. Effects of n-3 fatty acids, EPA v. DHA, on depressive symptoms, quality of life, memory and executive function in older adults with mild cognitive impairment: A 6-month randomised controlled trial. Br. J. Nutr. 2012, 107, 1682–1693. [Google Scholar] [CrossRef]
- Das, U.N. Essential fatty acids in health and disease. J. Assoc. Physicians India 1999, 47, 906–911. [Google Scholar]
- Ormarsson, O.T.; Geirsson, T.; Bjornsson, E.S.; Jonsson, T.; Moller, P.H.; Loftsson, T.; Stefansson, E. Clinical trial: marine lipid suppositories as laxatives. Mar. Drugs 2012, 10, 2047–2054. [Google Scholar] [CrossRef] [Green Version]
- Riediger, N.D.; Othman, R.A.; Suh, M.; Moghadasian, M.H. A systemic review of the roles of n-3 fatty acids in health and disease. J. Am. Diet. Assoc. 2009, 109, 668–679. [Google Scholar] [CrossRef]
- Hibbeln, J.R.; Nieminen, L.R.G.; Blasbalg, T.L.; Riggs, J.A.; Lands, W.E.M. Healthy intakes of n-3 and n-6 fatty acids: estimations considering worldwide diversity. Am. J. Clin. Nutr. 2006, 83, 1483S–1493S. [Google Scholar]
- Kris-Etherton, P.M.; Grieger, J.A.; Etherton, T.D. Dietary reference intakes for DHA and EPA. Prostag. Leukotr. Ess. 2009, 81, 99–104. [Google Scholar] [CrossRef]
- Kris-Etherton, P.M.; Innis, S. Position of the American dietetic association and dietitians of Canada: Dietary fatty acids. J. Am. Diet. Assoc. 2007, 107, 1599–1611. [Google Scholar]
- Norwegian Scientific Committee for Food Safety (VKM). Evaluation of Negative and Positive Health Effects of N-3 Fatty Acids as Constituents of Food Supplements and Fortified Foods; VKM: Oslo, Norway, 2011. [Google Scholar]
- Sargent, J.; McEvoy, L.; Estevez, A.; Bell, G.; Bell, M.; Henderson, J.; Tocher, D. Lipid nutrition of marine fish during early development: Current status and future directions. Aquaculture 1999, 179, 217–229. [Google Scholar] [CrossRef]
- Kang, J.X.; Liu, A. The role of the tissue omega-6/omega-3 fatty acid ratio in regulating tumor angiogenesis. Cancer Metastasis Rev. 2013, 32, 201–210. [Google Scholar] [CrossRef]
- Seafood data. Available online: http://www.nifes.no/index.php?page_id=164&lang_id=2 (accessed on 28August 2013).
- Reitan, K.I.; Rainuzzo, J.R.; Olsen, Y. Effect of nutrient limitation on fatty acid and lipid content of marine microalgae. J. Phycol. 1994, 30, 972–979. [Google Scholar]
- Enser, M.; Hallett, K.; Hewitt, B.; Fursey, G.A.J.; Wood, J.D. Fatty acid content and composition of English beef, lamb and pork at retail. Meat Sci. 1996, 42, 443–456. [Google Scholar] [CrossRef]
- Carnevale de Almeida, J.; Perassolo, M.S.; Camargo, J.L.; Bragagno, N.; Gross, J.L. Fatty acid composition and cholesterol content of beef and chicken meat in Southern Brazil. Rev. Bras. Cienc. Farm. 2006, 42, 109–117. [Google Scholar]
- Chowdhury, K.; Banu, L.A.; Khan, S.; Latif, A. Studies on the Fatty Acid Composition of Edible Oil. Bangladesh J. Sci. Ind. Res. 2007, 42, 311–316. [Google Scholar]
- Vidrih, R.; Filip, S.; Hribar, J. Content of Higher Fatty Acids in Green Vegetables. Czech J. Food Sci. 2009, 27, S125–S129. [Google Scholar]
- FAO Fisheries and Aquaculture Department. The State of World Fisheries and Aquaculture; Food and Agriculture Organization of the United Nations: Rome, Italy, 2012. [Google Scholar]
- Olsen, Y. Resources for fish feed in future mariculture. Aquacult. Environ. Interact. 2011, 1, 187–200. [Google Scholar] [CrossRef]
- Nasopoulou, C.; Zabetakis, I. Benefits of fish oil replacement by plant originated oils in compounded fish feeds. A review. LWT-Food Sci. Technol. 2012, 47, 217–224. [Google Scholar] [CrossRef]
- Torstensen, B.E.; Bell, J.G.; Rosenlund, G.; Henderson, R.J.; Graff, I.E.; Tocher, D.R.; Lie, O.; Sargent, J.R. Tailoring of Atlantic salmon (Salmo salar L.) flesh lipid composition and sensory quality by replacing fish oil with a vegetable oil blend. J. Agric. Food Chem. 2005, 53, 10166–10178. [Google Scholar] [CrossRef]
- Benedito-Palos, L.; Navarro, J.C.; Sitja-Bobadilla, A.; Bell, J.G.; Kaushik, S.; Perez-Sanchez, J. High levels of vegetable oils in plant protein-rich diets fed to gilthead sea bream (Sparus aurata L.): Growth performance, muscle fatty acid profiles and histological alterations of target tissues. Br. J. Nutr. 2008, 100, 992–1003. [Google Scholar] [CrossRef]
- Tur, J.A.; Bibiloni, M.M.; Sureda, A.; Pons, A. Dietary sources of omega 3 fatty acids: Public health risks and benefits. Br. J. Nutr. 2012, 107, S23–S52. [Google Scholar] [CrossRef]
- Dembitsky, V.M. Betaine ether-linked glycerolipids: Chemistry and biology. Prog. Lipid Res. 1996, 35, 1–51. [Google Scholar] [CrossRef]
- Loll, B.; Kern, J.; Saenger, W.; Zouni, A.; Biesiadka, J. Lipids in photosystem II: Interactions with protein and cofactors. Biochim. Biophys. Acta 2007, 1767, 509–519. [Google Scholar] [CrossRef]
- Guschina, I.A.; Harwood, J.L. Algal lipids and their metabolism. In Developments in Applied Phycology 5, Algae for Biofuels and Energy; Moheimani, N.R., Borowitzka, M.A., Eds.; Springer Science+Business Media Dordrecht: Dordrecht, The Netherlands, 2013; pp. 17–36. [Google Scholar]
- Lepetit, B.; Goss, R.; Jakob, T.; Wilhelm, C. Molecular dynamics of the diatom thylakoid membrane under different light conditions. Photosyn. Res. 2012, 111, 245–257. [Google Scholar] [CrossRef]
- Khozin-Goldberg, I.; Didi-Cohen, S.; Shayakhmetova, I.; Cohen, Z. Biosynthesis of eicosapentaenoic acid (EPA) in the freshwater eustigmatophyte Monodus subterraneus (Eustigmatophyceae). J. Phycol. 2002, 38, 745–756. [Google Scholar] [CrossRef]
- Hodgson, P.A.; Henderson, R.J.; Sargent, J.R.; Leftley, J.W. Patterns of variation in the lipid class and fatty acid composition of Nannochloropsis oculata (Eustigmatophyceae) during batch culture. J. Appl. Phycol. 1991, 3, 169–181. [Google Scholar] [CrossRef]
- Eichenberger, W.; Gribi, C. Lipids of Pavlova lutheri: Cellular site and metabolic role of DGCC. Phytochemistry 1997, 45, 1561–1567. [Google Scholar] [CrossRef]
- Arao, T.; Kawaguchi, A.; Yamada, M. Positional distribution of fatty-acids in lipids of the marine diatom Phaeodactylum tricornutum. Phytochemistry 1987, 26, 2573–2576. [Google Scholar] [CrossRef]
- Dodson, V.J.; Dahmen, J.L.; Mouget, J.-L.; Leblond, J.D. Mono- and digalactosyldiacylglycerol composition of the marennine-producing diatom, Haslea ostrearia: Comparison to a selection of pennate and centric diatoms. Phycol. Res. 2013, 61, 199–207. [Google Scholar] [CrossRef]
- Roughan, P.G.; Slack, C.R. Cellular organization of glycerolipid metabolism. Plant Physiol. 1982, 33, 97–132. [Google Scholar] [CrossRef]
- Khozin, I.; Adlerstein, D.; Bigongo, C.; Heimer, Y.M.; Cohen, Z. Elucidation of the biosynthesis of eicosapentaenoic acid in the microalga Porphyridium cruentum (II. studies with radiolabeled precursors). Plant Physiol. 1997, 114, 223–230. [Google Scholar]
- Khozin-Goldberg, I.; Yu, H.Z.; Adlerstein, D.; Didi-Cohen, S.; Heimer, Y.M.; Cohen, Z. Triacylglycerols of the red microalga Porphyridium cruentum can contribute to the biosynthesis of eukaryotic galactolipids. Lipids 2000, 35, 881–889. [Google Scholar] [CrossRef]
- Sato, N.; Aoki, M.; Maru, Y.; Sonoike, K.; Minoda, A.; Tsuzuki, M. Involvement of sulfoquinovosyl diacylglycerol in the structural integrity and heat-tolerance of photosystem II. Planta 2003, 217, 245–251. [Google Scholar]
- Jiang, H.M.; Gao, K.S. Effects of lowering temperature during culture on the production of polyunsaturated fatty acids in the marine diatom Phaeodactylum tricornutum (Bacillariophyceae). J. Phycol. 2004, 40, 651–654. [Google Scholar] [CrossRef]
- Tonon, T.; Harvey, D.; Larson, T.R.; Graham, I.A. Long chain polyunsaturated fatty acid production and partitioning to triacylglycerols in four microalgae. Phytochemistry 2002, 61, 15–24. [Google Scholar]
- Khozin-Goldberg, I.; Cohen, Z. The effect of phosphate starvation on the lipid and fatty acid composition of the fresh water eustigmatophyte Monodus subterraneus. Phytochemistry 2006, 67, 696–701. [Google Scholar] [CrossRef]
- Pal, D.; Khozin-Goldberg, I.; Cohen, Z.; Boussiba, S. The effect of light, salinity, and nitrogen availability on lipid production by Nannochloropsis sp. Appl. Microbiol. Biotechnol. 2011, 90, 1429–1441. [Google Scholar] [CrossRef]
- Palmucci, M.; Ratti, S.; Giordano, M. Ecological and evolutionary implications of carbon allocation in marine phytoplankton as a function of nitrogen availability: A fourier transform infrared spectroscopy approach. J. Phyciol. 2011, 47, 313–323. [Google Scholar] [CrossRef]
- Markou, G.; Angelidaki, I.; Georgakakis, D. Microalgal carbohydrates: An overview of the factors influencing carbohydrates production, and of main bioconversion technologies for production of biofuels. Appl. Microbiol. Biotechnol. 2012, 96, 631–645. [Google Scholar] [CrossRef]
- Obata, T.; Fernie, A.R.; Nunes-Nesi, A. The Central Carbon and Energy Metabolism of Marine Diatoms. Metabolites 2013, 3, 325–346. [Google Scholar] [CrossRef]
- Beattie, A.; Hirst, E.L.; Percival, E. Studies on the metabolism of the chrysophyceae. Comparative structural investigations on leucosin (chrysolaminarin) separated from diatoms and laminarin from the brown algae. Biochem. J. 1961, 79, 531–537. [Google Scholar]
- Wilhelm, C.; Jakob, T. From photons to biomass and biofuels: evaluation of different strategies for the improvement of algal biotechnology based on comparative energy balances. Appl. Microbiol. Biotechnol. 2011, 92, 909–919. [Google Scholar] [CrossRef]
- Wang, K.S.; Chai, T.-J. Reduction in omega-3 fatty acids by UV-B irradiation in microalgae. J. Appl. Phycol. 1994, 6, 415–421. [Google Scholar] [CrossRef]
- Bowler, C.; Allen, A.E.; Badger, J.H.; Grimwood, J.; Jabbari, K.; Kuo, A.; Maheswari, U.; Martens, C.; Maumus, F.; Otillar, R.P.; et al. The Phaeodactylum genome reveals the evolutionary history of diatom genomes. Nature 2008, 456, 239–244. [Google Scholar] [CrossRef]
- Mus, F.; Toussaint, J.P.; Cooksey, K.E.; Fields, M.W.; Gerlach, R.; Peyton, B.M.; Carlson, R.P. Physiological and molecular analysis of carbon source supplementation and pH stress-induced lipid accumulation in the marine diatom Phaeodactylum tricornutum. Appl. Microbiol. Biotechnol. 2013, 97, 3625–3642. [Google Scholar] [CrossRef]
- De Martino, A.; Bartual, A.; Willis, A.; Meichenin, A.; Villazan, B.; Maheswari, U.; Bowler, C. Physiological and molecular evidence that environmental changes elicit morphological interconversion in the model diatom Phaeodactylum tricornutum. Protist 2011, 162, 462–481. [Google Scholar] [CrossRef]
- Valenzuela, J.; Mazurie, A.; Carlson, R.P.; Gerlach, R.; Cooksey, K.E.; Peyton, B.M.; Fields, M.W. Potential role of multiple carbon fixation pathways during lipid accumulation in Phaeodactylum tricornutum. Biotechnol. Biofuels 2012, 5, 40. [Google Scholar] [CrossRef]
- Gardner, R.D.; Cooksey, K.E.; Mus, F.; Macur, R.; Moll, K.; Eustance, E.; Carlson, R.P.; Gerlach, R.; Fields, M.W.; Peyton, B.M. Use of sodium bicarbonate to stimulate triacylglycerol accumulation in the chlorophyte Scenedesmus sp. and the diatom Phaeodactylum tricornutum. J. Appl. Phycol. 2012, 24, 1311–1320. [Google Scholar] [CrossRef]
- Fabris, M.; Matthijs, M.; Rombauts, S.; Vyverman, W.; Goossens, A.; Baart, G.J. The metabolic blueprint of Phaeodactylum tricornutum reveals a eukaryotic Entner-Doudoroff glycolytic pathway. Plant J. 2012, 70, 1004–1014. [Google Scholar] [CrossRef]
- Kanehisa, M.; Goto, S.; Sato, Y.; Furumichi, M.; Tanabe, M. KEGG for integration and interpretation of large-scale molecular data sets. Nucl. Acids Res. 2012, 40, D109–D114. [Google Scholar] [CrossRef]
- Brembu, T.; Jorstad, M.; Winge, P.; Valle, K.C.; Bones, A.M. Genome-Wide profiling of responses to cadmium in the diatom Phaeodactylum tricornutum. Environ. Sci. Technol. 2011, 45, 7640–7647. [Google Scholar]
- Nymark, M.; Valle, K.C.; Brembu, T.; Hancke, K.; Winge, P.; Andresen, K.; Johnsen, G.; Bones, A.M. An integrated analysis of molecular acclimation to high light in the marine diatom Phaeodactylum tricornutum. PLoS One 2009, 4, e7743. [Google Scholar]
- Nymark, M.; Valle, K.C.; Hancke, K.; Winge, P.; Andresen, K.; Johnsen, G.; Bones, A.M.; Brembu, T. Molecular and photosynthetic responses to prolonged darkness and subsequent acclimation to re-illumination in the diatom Phaeodactylum tricornutum. PLoS One 2013, 8, e58722. [Google Scholar]
- Sapriel, G.; Quinet, M.; Heijde, M.; Jourdren, L.; Tanty, V.; Luo, G.Z.; Le Crom, S.; Lopez, P.J. Genome-Wide transcriptome analyses of silicon metabolism in Phaeodactylum tricornutum reveal the multilevel regulation of silicic acid transporters. PLoS One 2009, 4, e7458. [Google Scholar]
- Chauton, M.S.; Winge, P.; Brembu, T.; Vadstein, O.; Bones, A.M. Gene regulation of carbon fixation, storage, and utilization in the diatom Phaeodactylum tricornutum acclimated to light/dark cycles. Plant Physiol. 2013, 161, 1034–1048. [Google Scholar] [CrossRef]
- Shannon, P.; Markiel, A.; Ozier, O.; Baliga, N.S.; Wang, J.T.; Ramage, D.; Amin, N.; Schwikowski, B.; Ideker, T. Cytoscape a software envirnoment for inetgrated models of biomolecular interaction networks. Genome Res. 2003, 13, 2498–2504. [Google Scholar] [CrossRef]
- Bellou, S.; Aggelis, G. Biochemical activities in Chlorella sp. and Nannochloropsis salina during lipid and sugar synthesis in a lab-scale open pond simulating reactor. J. Biotechnol. 2012, 164, 318–329. [Google Scholar] [CrossRef]
- Kroth, P.G.; Chiovitti, A.; Gruber, A.; Martin-Jezequel, V.; Mock, T.; Parker, M.S.; Stanley, M.S.; Kaplan, A.; Caron, L.; Weber, T.; et al. A model for carbohydrate metabolism in the diatom Phaeodactylum tricornutum deduced from comparative whole genome analysis. PLoS One 2008, 3, e1426. [Google Scholar] [CrossRef]
- Zhang, Y.; Adams, I.P.; Ratledge, C. Malic enzyme: The controlling activity for lipid production? Overexpression of malic enzyme in Mucor circinelloides leads to a 2.5-fold increase in lipid accumulation. Microbiology 2007, 153, 2013–2025. [Google Scholar] [CrossRef]
- Kitano, M.; Matsukawa, R.; Karube, I. Enhanced eicosapentaenoic acid production by Navicula saprophila. J. Appl. Phycol. 1998, 10, 101–105. [Google Scholar] [CrossRef]
- Schneider, J.C.; Roessler, P. Radiolabeling Studies of Lipids and Fatty-Acids in Nannochloropsis (Eustigmatophyceae), an Oleaginous Marine Alga. J. Phycol. 1994, 30, 594–598. [Google Scholar]
- Hofmann, M.; Eichenberger, W. Radiolabelling studies on the lipid metabolism in the marine brown alga Dictyopteris membranacea. Plant Cell Physiol. 1998, 39, 508–515. [Google Scholar] [CrossRef]
- Domergue, F.; Spiekermann, P.; Lerchl, J.; Beckmann, C.; Kilian, O.; Kroth, P.G.; Boland, W.; Zähringer, U.; Heinz, E. New insight into Phaeodactylum tricornutum fatty acid metabolism. Cloning and functional characterization of plastidial and microsomal Δ12-fatty acid sesaturases. Plant Physiol. 2003, 131, 1648–1660. [Google Scholar] [CrossRef]
- Huerlimann, R.; Heimann, K. Comprehensive guide to acetyl-carboxylases in algae. Crit. Rev. Biotechnol. 2013, 33, 49–65. [Google Scholar] [CrossRef]
- Nikolau, B.J.; Ohlrogge, J.B.; Wurtele, E.S. Plant biotin-containing carboxylases. Arch. Biochem. Biophys. 2003, 414, 211–222. [Google Scholar] [CrossRef]
- Dunahay, T.G.; Jarvis, E.E.; Dais, S.S.; Roessler, P.G. Manipulation of microalgal lipid production using genetic engineering. Appl. Biochemi. Biotechnol. 1996, 57, 223–231. [Google Scholar]
- Radakovits, R.; Eduafo, P.M.; Posewitz, M.C. Genetic engineering of fatty acid chain length in Phaeodactylum tricornutum. Metab. Eng. 2011, 13, 89–95. [Google Scholar] [CrossRef]
- Ryall, K.; Harper, J.T.; Keeling, P.J. Plastid-derived Type II fatty acid biosynthetic enzymes in chromists. Gene 2003, 313, 139–148. [Google Scholar] [CrossRef]
- Harwood, J.L.; Guschina, I.A. Regulation of lipid synthesis in oil crops. FEBS Lett. 2013, 587, 2079–2081. [Google Scholar] [CrossRef]
- Subrahmanyam, S.; Cronan, J.E. Overproduction of a functional fatty acid biosynthetic enzyme blocks fatty acid synthesis in Escherichia coli. J. Bacteriol. 1998, 180, 4596–4602. [Google Scholar]
- Domergue, F.; Abbadi, A.; Ott, C.; Zank, T.K.; Zahringer, U.; Heinz, E. Acyl carriers used as substrates by the desaturases and elongases involved in very long-chain polyunsaturated fatty acids biosynthesis reconstituted in yeast. J. Biol. Chem. 2003, 278, 35115–35126. [Google Scholar]
- Sperling, P.; Heinz, E. Desaturases fused to their electron donor. Eur. J. Lipid Sci. Technol. 2001, 103, 158–180. [Google Scholar] [CrossRef]
- Groot, P.H.; Scholte, H.R.; Hulsmann, W.C. Fatty acid activation: specificity, localization, and function. Adv. Lipid Res. 1976, 14, 75–126. [Google Scholar]
- Fulda, M.; Shockey, J.; Werber, M.; Wolter, F.P.; Heinz, E. Two long-chain acyl-CoA synthetases from Arabidopsis thaliana involved in peroxisomal fatty acid beta-oxidation. Plant J. 2002, 32, 93–103. [Google Scholar] [CrossRef]
- Sørensen, B.M.; Furukawa-Stoffer, T.L.; Marshall, K.S.; Page, E.K.; Mir, Z.; Forster, R.J.; Weselake, R.J. Storage lipid accumulation and acyltransferase action in developing flaxseed. Lipids 2005, 40, 1043–1049. [Google Scholar] [CrossRef]
- Snyder, C.S.; Yurchenko, O.P.; Siloto, R.M.P.; Chen, X.; Liu, Q.; Mietkiewska, E.; Weselake, R.J. Acyltransferase action in the modification of seed oil biosynthesis. New Biotechnol. 2009, 26, 11–14. [Google Scholar]
- Kennedy, E.P. Biosynthesis of complex lipids. Fed. Proc. 1961, 20, 934–940. [Google Scholar]
- Wendel, A.A.; Lewin, T.M.; Coleman, R.A. Glycerol-3-phosphate acyltransferases: Rate limiting enzymes of triacylglycerol biosynthesis. Biochim. Biophys. Acta Mol. Cell. Biol. Lipids 2009, 1791, 501–506. [Google Scholar] [CrossRef]
- Domergue, F.; Lerchl, J.; Zahringer, U.; Heinz, E. Cloning and functional characterization of Phaeodactylum tricornutum front-end desaturases involved in eicosapentaenoic acid biosynthesis. Eur. J. Biochem. 2002, 269, 4105–4113. [Google Scholar] [CrossRef]
- Arao, T.; Yamada, M. Biosynthesis of Polyunsaturated Fatty-Acids in the Marine Diatom, Phaeodactylum tricornutum. Phytochemistry 1993, 35, 1177–1181. [Google Scholar] [CrossRef]
- Yongmanitchai, W.; Ward, O.P. Positional distribution of fatty acids, and molecular species of polar lipids, in the diatom Phaeodactylum tricornutum. J. Gen Microbiol. 1993, 139, 465–472. [Google Scholar] [CrossRef]
- Joyard, J.; Ferro, M.; Masselon, C.; Seigneurin-Berny, D.; Salvi, D.; Garin, J.; Rolland, N. Chloroplast proteomics highlights the subcellular compartmentation of lipid metabolism. Prog. Lipid Res. 2010, 49, 128–158. [Google Scholar] [CrossRef]
- Kobayashi, K.; Awai, K.; Nakamura, M.; Nagatani, A.; Masuda, T.; Ohta, H. Type-B monogalactosyldiacylglycerol synthases are involved in phosphate starvation-induced lipid remodeling, and are crucial for low-phosphate adaptation. Plant J. 2009, 57, 322–331. [Google Scholar] [CrossRef]
- De Riso, V.; Raniello, R.; Maumus, F.; Rogato, A.; Bowler, C.; Falciatore, A. Gene silencing in the marine diatom Phaeodactylum tricornutum. Nucleic Acids Res. 2009, 37, e96. [Google Scholar] [CrossRef]
- Siaut, M.; Heijde, M.; Mangogna, M.; Montsant, A.; Coesel, S.; Allen, A.; Manfredonia, A.; Falciatore, A.; Bowler, C. Molecular toolbox for studying diatom biology in Phaeodactylum tricornutum. Gene 2007, 406, 23–35. [Google Scholar] [CrossRef]
- Xu, J.Y.; Zheng, Z.F.; Zou, J.T. A membrane-bound glycerol-3-phosphate acyltransferase from Thalassiosira pseudonana regulates acyl composition of glycerolipids. Botany 2009, 87, 544–551. [Google Scholar] [CrossRef]
- Zhang, L.; Ma, X.L.; Yang, G.P.; Zhu, B.H.; Han, J.C.; Yu, W.G.; Pan, K.H. Isolation and characterization of a long-chain acyl-coenzyme A synthetase encoding gene from the marine microalga Nannochloropsis oculata. J. Appl. Phycol. 2012, 24, 873–880. [Google Scholar] [CrossRef]
- Tonon, T.; Qing, R.; Harvey, D.; Li, Y.; Larson, T.R.; Graham, I.A. Identification of a long-chain polyunsaturated fatty acid acyl-coenzyme A synthetase from the diatom Thalassiosira pseudonana. Plant. Physiol. 2005, 138, 402–408. [Google Scholar] [CrossRef]
- Fulda, M.; Schnurr, J.; Abbadi, A.; Heinz, E.; Browse, J. Peroxisomal Acyl-CoA synthetase activity is essential for seedling development in Arabidopsis thaliana. Plant Cell 2004, 16, 394–405. [Google Scholar] [CrossRef]
- Hettema, E.H.; van Roermund, C.W.; Distel, B.; van den Berg, M.; Vilela, C.; Rodrigues-Pousada, C.; Wanders, R.J.; Tabak, H.F. The ABC transporter proteins Pat1 and Pat2 are required for import of long-chain fatty acids into peroxisomes of Saccharomyces cerevisiae. EMBO J. 1996, 15, 3813–3822. [Google Scholar]
- Gong, Y.; Zhang, J.; Guo, X.; Wan, X.; Liang, Z.; Hu, C.J.; Jiang, M. Identification and characterization of PtDGAT2B, an acyltransferase of the DGAT2 acyl-coenzyme A: diacylglycerol acyltransferase family in the diatom Phaeodactylum tricornutum. FEBS Lett. 2013, 587, 481–487. [Google Scholar] [CrossRef]
- Gong, Y.; Guo, X.; Wan, X.; Liang, Z.; Jiang, M. Characterization of a novel thioesterase (PtTE) from Phaeodactylum tricornutum. J. Basic Microbiol. 2011, 51, 666–672. [Google Scholar] [CrossRef]
- Muto, M.; Kubota, C.; Tanaka, M.; Satoh, A.; Matsumoto, M.; Yoshino, T.; Tanaka, T. Identification and functional analysis of delta-9 desaturase, a key enzyme in PUFA synthesis, isolated from the oleaginous diatom Fistulifera. PLoS One 2013, 8, e73507. [Google Scholar]
- Petrie, J.R.; Shrestha, P.; Mansour, M.P.; Nichols, P.D.; Liu, Q.; Singh, S.P. Metabolic engineering of omega-3 long-chain polyunsaturated fatty acids in plants using an acyl-CoA Δ6-desaturase with omega 3-preference from the marine microalga Micromonas pusilla. Metab. Eng. 2010, 12, 233–240. [Google Scholar] [CrossRef]
- Senger, T.; Marty, L.; Stymne, S.; Lindberg Yilmaz, J.; Napier, J.A.; Sayanova, O.; Haslam, R.; Noemi, R.L. Acyltransferases and uses therof in fatty acid production. US Patent 20,120,060,242 A1, 29 December 2011. [Google Scholar]
- Bates, P.D.; Ohlrogge, J.B.; Pollard, M. Incorporation of newly synthesized fatty acids into cytosolic glycerolipids in pea leaves occurs via acyl editing. J. Biol. Chem. 2007, 282, 31206–31216. [Google Scholar] [CrossRef]
- Hu, Z.; Ren, Z.; Lu, C. The phosphatidylcholine diacylglycerol cholinephosphotransferase is required for efficient hydroxy fatty acid accumulation in transgenic Arabidopsis. Plant Physiol. 2012, 158, 1944–1954. [Google Scholar] [CrossRef]
- Pan, K.; Qin, J.; Li, S.; Dai, W.; Zhu, B.; Jin, Y.; Yu, W.; Yang, G.; Li, D. Nuclear monoploidy and asexual propagation of Nannochloropsis oceanica (Eustigmatophyceae) as revealed by its genome sequence. J. Phyciol. 2011, 47, 1425–1432. [Google Scholar] [CrossRef]
- Radakovits, R.; Jinkerson, R.E.; Fuerstenberg, S.I.; Tae, H.; Settlage, R.E.; Boore, J.L.; Posewitz, M.C. Draft genome sequence and genetic transformation of the oleaginous alga Nannochloropis gaditana. Nat. Commun. 2012, 3, 686. [Google Scholar]
- Kilian, O.; Vick, B. Homologous recombination in an algal nuclear genome. US Patent 20,110,091,977 A1, 21 April 2011. [Google Scholar]
- Jinkerson, R.E.; Radakovits, R.; Posewitz, M.C. Genomic insights from the oleaginous model alga Nannochloropsis gaditana. Bioengineered 2013, 4, 37–43. [Google Scholar] [CrossRef]
- Vieler, A.; Wu, G.; Tsai, C.H.; Bullard, B.; Cornish, A.J.; Harvey, C.; Reca, I.B.; Thornburg, C.; Achawanantakun, R.; Buehl, C.J.; et al. Genome, functional gene annotation, and nuclear transformation of the heterokont oleaginous alga Nannochloropsis oceanica CCMP1779. PLoS Genet. 2012, 8, e1003064. [Google Scholar] [CrossRef]
- Bondioli, P.; Bella, L.D.; Rivolta, G.; Zittelli, G.C.; Bassi, N.; Rodolfi, L.; Casini, D.; Prussi, M.; Chiaramonti, D.; Tredici, M.R. Oil production by the marine microalgae Nannochloropsis sp. F&M-M24 and Tetraselmis suecica F&M-M33. Bioresour. Technol. 2012, 114, 567–572. [Google Scholar] [CrossRef]
- Kilian, O.; Vick, B. Algal desaturases. Patent US8440805B2, 14 May 2013. [Google Scholar]
- Kilian, O.; Vick, B. Algal elongases. Patent US2012/0277418A1, 1 November 2012. [Google Scholar]
- Schneider, J.C.; Roessler, P.G. A novel acyltransferase activity in an oleaginous alga. Plant Lipid Meta. 1995, 105–107. [Google Scholar] [CrossRef]
- Kato, M.; Sakai, M.; Adachi, K.; Ikemoto, H.; Sano, H. Distribution of betaine lipids in marine algae. Phytochemistry 1996, 42, 1341–1345. [Google Scholar] [CrossRef]
- Sprecher, H. Metabolism of highly unsaturated n-3 and n-6 fatty acids. Biochim. Biophys. Acta Mol. Cell. Biol. Lipids 2000, 1486, 219–231. [Google Scholar] [CrossRef]
- Metz, J.G.; Roessler, P.; Facciotti, D.; Levering, C.; Dittrich, F.; Lassner, M.; Valentine, R.; Lardizabal, K.; Domergue, F.; Yamada, A.; et al. Production of polyunsaturated fatty acids by polyketide synthases in both prokaryotes and eukaryotes. Science 2001, 293, 290–293. [Google Scholar] [CrossRef]
- Metz, J.G.; Kuner, J.; Rosenzweig, B.; Lippmeier, J.C.; Roessler, P.; Zirkle, R. Biochemical characterization of polyunsaturated fatty acid synthesis in Schizochytrium: release of the products as free fatty acids. Plant Physiol. Biochem. 2009, 47, 472–478. [Google Scholar] [CrossRef]
- Hauvermale, A.; Kuner, J.; Rosenzweig, B.; Guerra, D.; Diltz, S.; Metz, J.G. Fatty acid production in Schizochytrium sp.: Involvement of a polyunsaturated fatty acid synthase and a type I fatty acid synthase. Lipids 2006, 41, 739–747. [Google Scholar] [CrossRef]
- Chang, G.; Gao, N.; Tian, G.; Wu, Q.; Chang, M.; Wang, X. Improvement of docosahexaenoic acid production on glycerol by Schizochytrium sp. S31 with constantly high oxygen transfer coefficient. Bioresour. Technol. 2013, 142, 400–406. [Google Scholar] [CrossRef]
- Matsuda, T.; Sakaguchi, K.; Hamaguchi, R.; Kobayashi, T.; Abe, E.; Hama, Y.; Hayashi, M.; Honda, D.; Okita, Y.; Sugimoto, S.; et al. Analysis of Δ12-fatty acid desaturase function revealed that two distinct pathways are active for the synthesis of PUFAs in T. aureum ATCC 34304. J. Lipid Res. 2012, 53, 2806–2806. [Google Scholar] [CrossRef]
- Ohara, J.; Sakaguchi, K.; Okita, Y.; Okino, N.; Ito, M. Two fatty acid elongases possessing C18-Δ6/C18-Δ9/C20-Δ5 or C16-Δ9 elongase activity in Thraustochytrium sp. ATCC 26185. Marine Biotechnol. 2013, 15, 476–486. [Google Scholar] [CrossRef]
- Qiu, X.; Hong, H.P.; MacKenzie, S.L. Identification of a Δ4 fatty acid desaturase from Thraustochytrium sp. involved in the biosynthesis of docosahexanoic acid by heterologous expression in Saccharomyces cerevisiae and Brassica juncea. J. Biol. Chem. 2001, 276, 31561–31566. [Google Scholar] [CrossRef]
- Nagano, N.; Sakaguchi, K.; Taoka, Y.; Okita, Y.; Honda, D.; Ito, M.; Hayashi, M. Detection of genes Involved in fatty acid elongation and desaturation in thraustochytrid marine eukaryotes. J. Oleo Sci. 2011, 60, 475–481. [Google Scholar] [CrossRef]
- Shi, T.L.; Yu, A.Q.; Li, M.; Ou, X.Y.; Xing, L.J.; Li, M.C. Identification of a novel C22-Δ4-producing docosahexaenoic acid (DHA) specific polyunsaturated fatty acid desaturase gene from Isochrysis galbana and its expression in Saccharomyces cerevisiae. Biotechnol. Lett. 2012, 34, 2265–2274. [Google Scholar] [CrossRef]
- Guihéneuf, F.; Ulmann, L.; Mimouni, V.; Tremblin, G. Use of radiolabeled substrates to determine the desaturase and elongase activities involved in eicosapentaenoic acid and docosahexaenoic acid biosynthesis in the marine microalga Pavlova lutheri. Phytochemistry 2013, 90, 43–49. [Google Scholar]
- Pereira, S.L.; Leonard, A.E.; Huang, Y.S.; Chuang, L.T.; Mukerji, P. Identification of two novel microalgal enzymes involved in the conversion of the omega3-fatty acid, eicosapentaenoic acid, into docosahexaenoic acid. Biochem. J. 2004, 384, 357–366. [Google Scholar] [CrossRef]
- Tonon, T.; Harvey, D.; Larson, T.R.; Graham, I.A. Identification of a very long chain polyunsaturated fatty acid Δ4-desaturase from the microalga Pavlova lutheri. FEBS Lett. 2003, 553, 440–444. [Google Scholar] [CrossRef]
- Daboussia, F.; Collinb, S.; Leduca, S.; Cavareca, L.; Marechala, A.; Falciatorec, A.; Laeufferb, F.; Fourageb, L.; Duchateaua, P. Diatoms genome engineering using targeted nucleases approaches. In The Molecular Life of Diatoms; EMBO Workshop: Paris, France, 2013. [Google Scholar]
- Doan, T.T.Y.; Obbard, J.P. Enhanced intracellular lipid in Nannochloropsis sp. via random mutagenesis and flow cytometric cell sorting. Algal Res. 2012, 1, 17–21. [Google Scholar] [CrossRef]
- Chaturvedi, R.; Uppalapati, S.R.; Alamsjah, M.A.; Fujita, Y. Isolation of quizalofop-resistant mutants of Nannochloropsis oculata (Eustigmatophyceae) with high eicosapentaenoic acid following N-methyl-N-nitrosourea-induced random mutagenesis. J. Appl. Phycol. 2004, 16, 135–144. [Google Scholar] [CrossRef]
- Meireles, L.A.; Guedes, A.C.; Malcata, F.X. Increase of the yields of eicosapentaenoic and docosahexaenoic acids by the microalga Pavlova lutheri following random mutagenesis. Biotechnol. Bioeng. 2003, 81, 50–55. [Google Scholar] [CrossRef]
Appendix: Definitions and Abbreviations
© 2013 by the authors; licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution license (http://creativecommons.org/licenses/by/3.0/).
Share and Cite
Mühlroth, A.; Li, K.; Røkke, G.; Winge, P.; Olsen, Y.; Hohmann-Marriott, M.F.; Vadstein, O.; Bones, A.M. Pathways of Lipid Metabolism in Marine Algae, Co-Expression Network, Bottlenecks and Candidate Genes for Enhanced Production of EPA and DHA in Species of Chromista. Mar. Drugs 2013, 11, 4662-4697. https://doi.org/10.3390/md11114662
Mühlroth A, Li K, Røkke G, Winge P, Olsen Y, Hohmann-Marriott MF, Vadstein O, Bones AM. Pathways of Lipid Metabolism in Marine Algae, Co-Expression Network, Bottlenecks and Candidate Genes for Enhanced Production of EPA and DHA in Species of Chromista. Marine Drugs. 2013; 11(11):4662-4697. https://doi.org/10.3390/md11114662
Chicago/Turabian StyleMühlroth, Alice, Keshuai Li, Gunvor Røkke, Per Winge, Yngvar Olsen, Martin F. Hohmann-Marriott, Olav Vadstein, and Atle M. Bones. 2013. "Pathways of Lipid Metabolism in Marine Algae, Co-Expression Network, Bottlenecks and Candidate Genes for Enhanced Production of EPA and DHA in Species of Chromista" Marine Drugs 11, no. 11: 4662-4697. https://doi.org/10.3390/md11114662