Harnessing the Potential of Halogenated Natural Product Biosynthesis by Mangrove-Derived Actinomycetes
Abstract
:1. Introduction
2. Results and Discussion
2.1. Potential of Biosynthesis of Halogenated Metabolites in Mangrove-Derived Actinomycetes
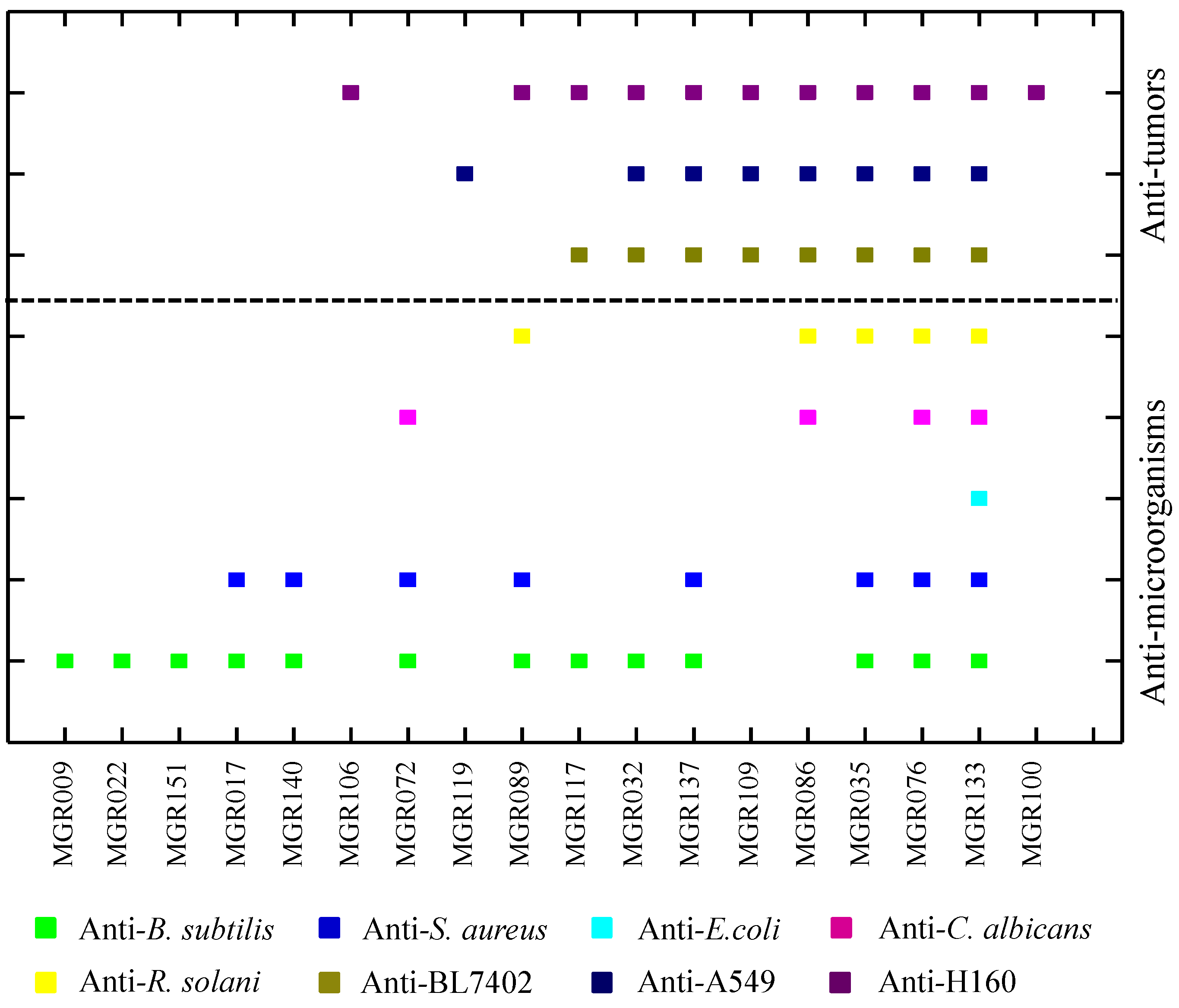
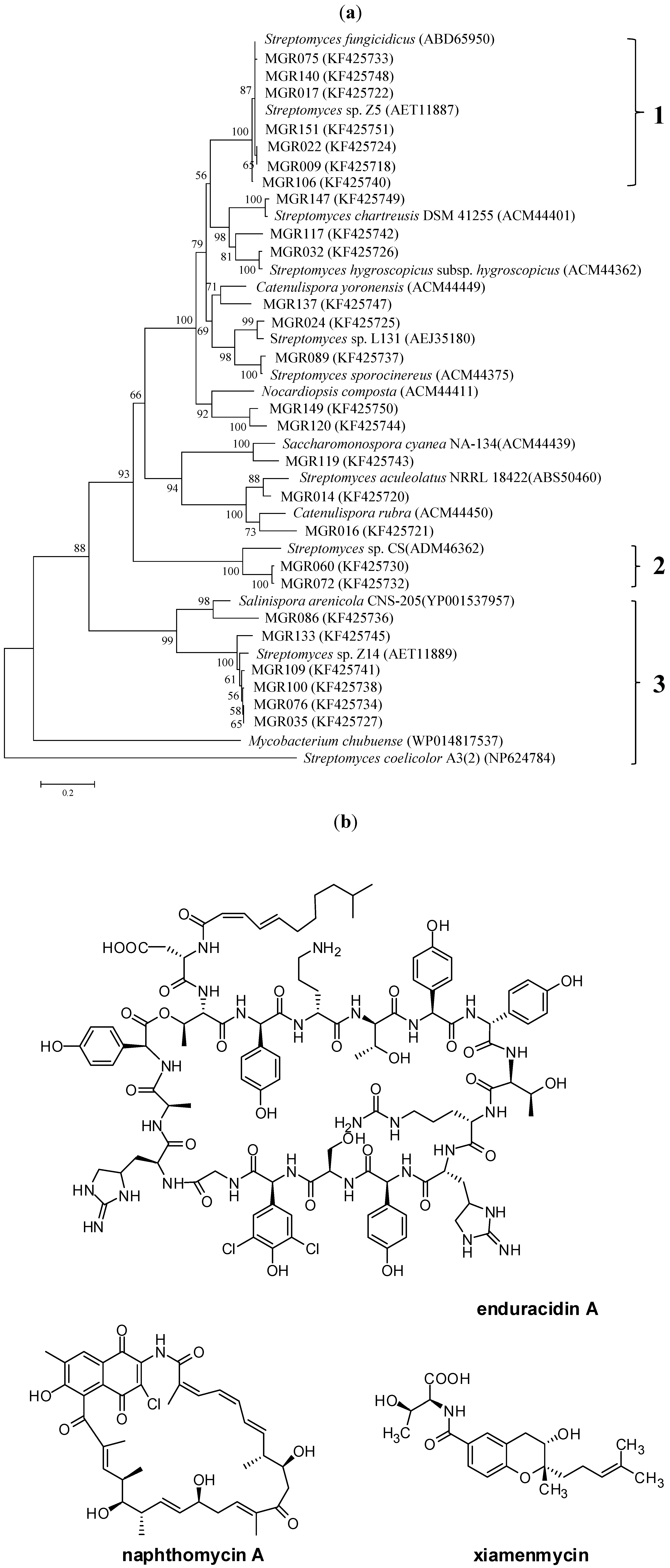
2.2. A New Enduracidin Producer S. atrovirens MGR140
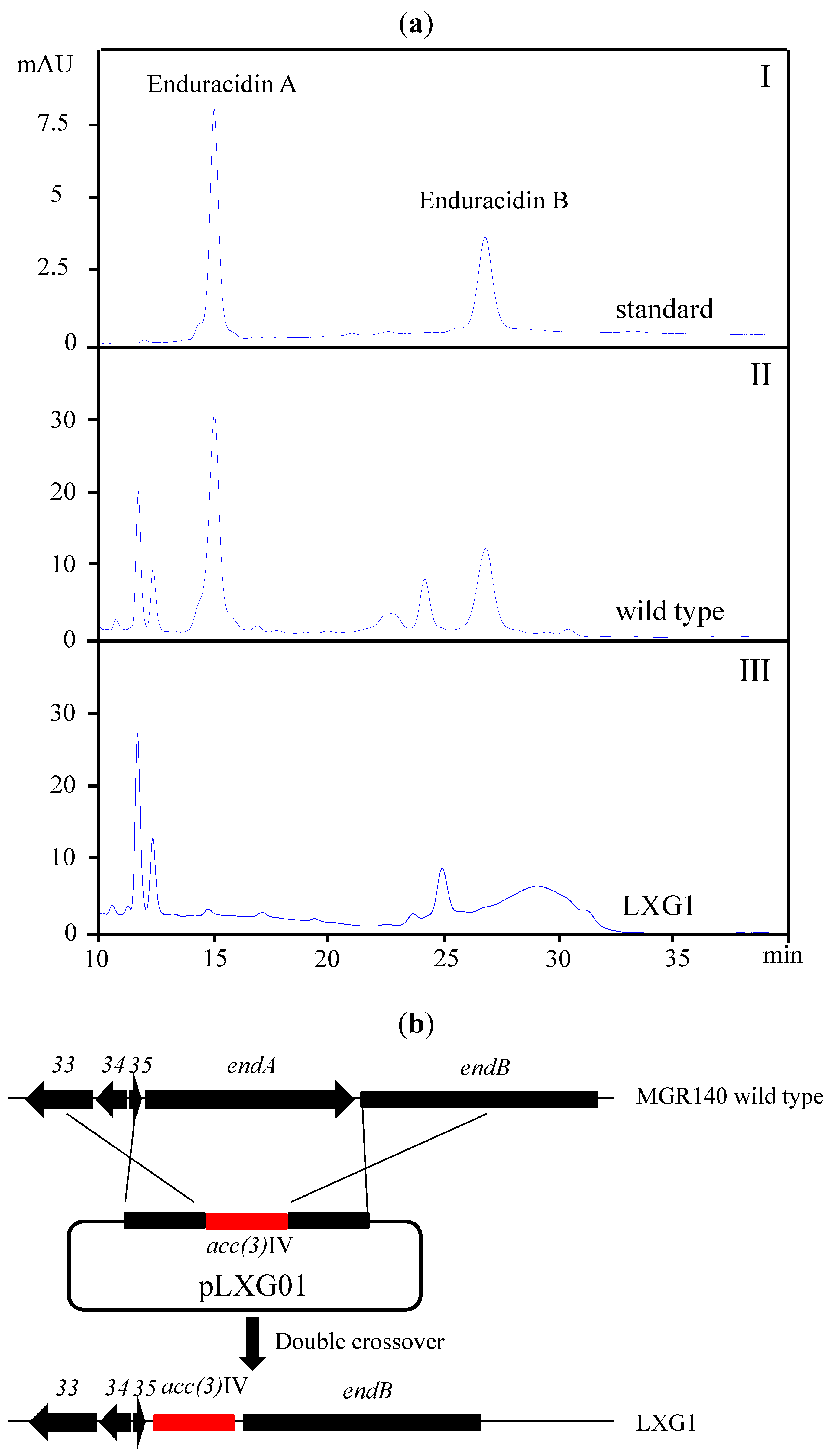
2.3. Identification of a Putative Halogenated Ansamycin Gene-Cluster from S. albogriseolus MGR072
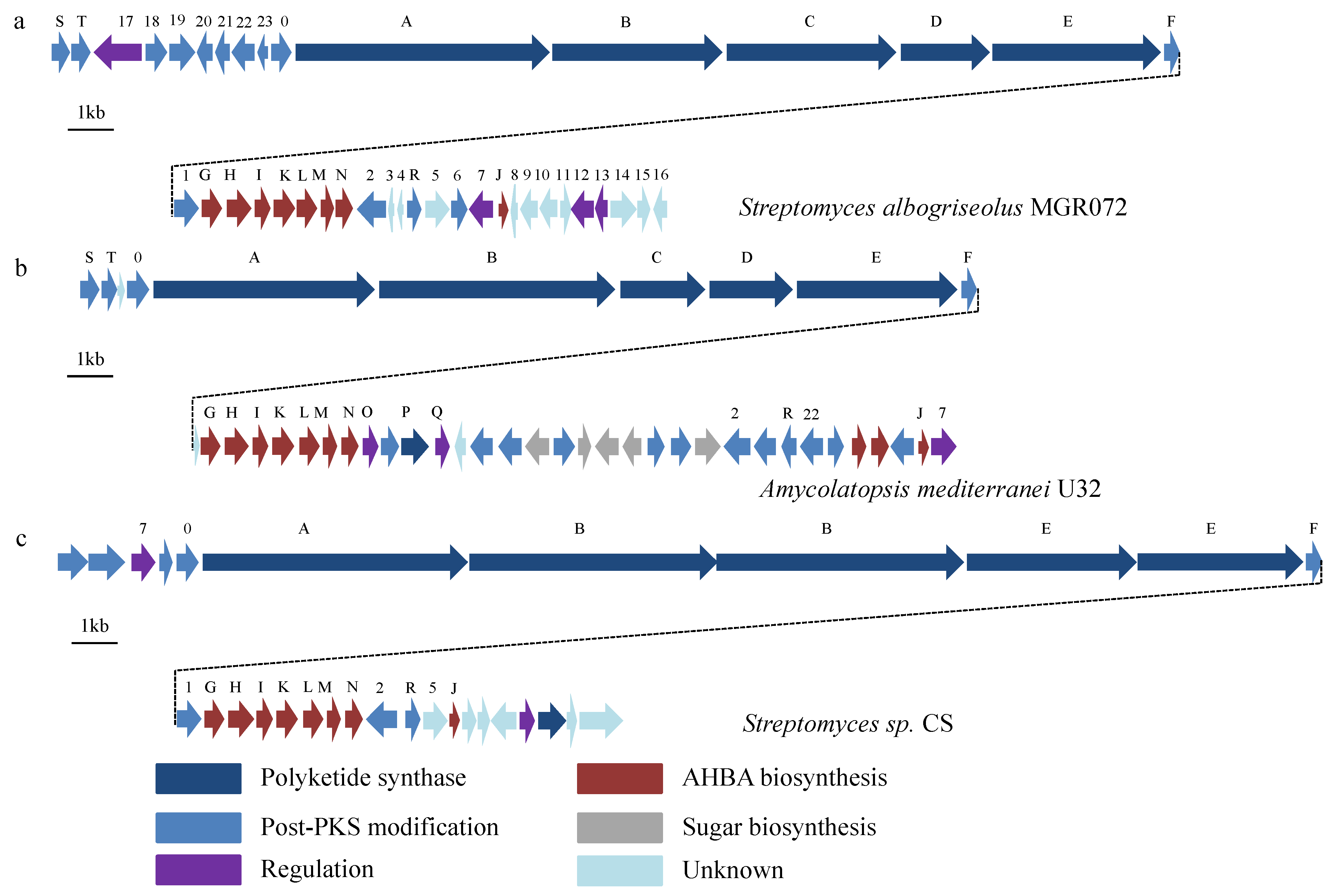
3. Discussion
4. Experimental Section
4.1. Strain and Culture Conditions
4.2. Detection of PKS I, PKS II, and NRPS Genes by PCR Amplifications
4.3. Fosmid Library Construction and Screening
4.4. Gene Inactivation
4.5. DNA Sequencing and Bioinformatic Analysis
4.6. Phylogenetic Analysis
5. Conclusions
Acknowledgments
Conflicts of Interest
References and Notes
- Costanza, R.; d’Arge, R.; De Groot, R.; Farber, S.; Grasso, M.; Hannon, B.; Limburg, K.; Naeem, S.; O’neill, R.V.; Paruelo, J. The value of the world’s ecosystem services and natural capital. Nature 1997, 387, 253–260. [Google Scholar] [CrossRef]
- Kathiresan, K.; Bingham, B.L. Biology of mangroves and mangrove ecosystems. Adv. Mar. Biol. 2001, 40, 81–251. [Google Scholar] [CrossRef]
- Remya, M.; Vijayakumar, R. Isolation and characterization of marine antagonistic actinomycetes from west coast of India. Facta Universitatis Ser. Med. Biol. 2008, 15, 13–19. [Google Scholar]
- Hong, K.; Gao, A.-H.; Xie, Q.-Y.; Gao, H.G.; Zhuang, L.; Lin, H.-P.; Yu, H.-P.; Li, J.; Yao, X.-S.; Goodfellow, M. Actinomycetes for marine drug discovery isolated from mangrove soils and plants in China. Mar. Drugs 2009, 7, 24–44. [Google Scholar] [CrossRef]
- Dias, A.C.; Andreote, F.D.; Dini-Andreote, F.; Lacava, P.T.; Sá, A.L.; Melo, I.S.; Azevedo, J.L.; Araújo, W.L. Diversity and biotechnological potential of culturable bacteria from Brazilian mangrove sediment. World J. Microbiol. Biotechnol. 2009, 25, 1305–1311. [Google Scholar] [CrossRef]
- Huang, H.; Lv, J.; Hu, Y.; Fang, Z.; Zhang, K.; Bao, S. Micromonospora rifamycinica sp. nov., a novel actinomycete from mangrove sediment. Int. J. Syst. Evol. Microbiol. 2008, 58, 17–20. [Google Scholar] [CrossRef]
- Hornung, A.; Bertazzo, M.; Dziarnowski, A.; Schneider, K.; Welzel, K.; Wohlert, S.E.; Holzenkämpfer, M.; Nicholson, G.J.; Bechthold, A.; Süssmuth, R.D. A Genomic Screening Approach to the Structure-Guided Identification of Drug Candidates from Natural Sources. Chembiochem 2007, 8, 757–766. [Google Scholar] [CrossRef]
- Wang, G.; Zhang, H.; Sun, G.; Wu, L.; Zhang, J.; Wang, Y. A new method for rapid identification of ansamycin compounds by inactivating KLM gene clusters in potential ansamycin-producing actinomyces. J. Appl. Microbiol. 2012, 112, 353–362. [Google Scholar] [CrossRef]
- Zerikly, M.; Challis, G.L. Strategies for the discovery of new natural products by genome mining. Chembiochem 2009, 10, 625–633. [Google Scholar] [CrossRef]
- Walsh, C.T.; Fischbach, M.A. Natural products version 2.0: Connecting genes to molecules. J. Am. Chem. Soc. 2010, 132, 2469–2493. [Google Scholar] [CrossRef]
- Du, L.; Lou, L. PKS and NRPS release mechanisms. Nat. Prod. Rep. 2010, 27, 255–278. [Google Scholar] [CrossRef]
- Olano, C.; Méndez, C.; Salas, J.A. Post-PKS tailoring steps in natural product-producing actinomycetes from the perspective of combinatorial biosynthesis. Nat. Prod. Rep. 2010, 27, 571–616. [Google Scholar] [CrossRef]
- Ouyang, Y.; Wu, H.; Xie, L.; Wang, G.; Dai, S.; Chen, M.; Yang, K.; Li, X. A method to type the potential angucycline producers in actinomycetes isolated from marine sponges. Antonie van Leeuwenhoek 2011, 99, 807–815. [Google Scholar] [CrossRef]
- Liu, T.; Cane, D.E.; Deng, Z. The enzymology of polyether biosynthesis. Methods Enzymol. 2009, 459, 187–214. [Google Scholar] [CrossRef]
- Jiang, C.; Wang, H.; Kang, Q.; Liu, J.; Bai, L. Cloning and Characterization of the Polyether Salinomycin Biosynthesis Gene Cluster of Streptomyces albus XM211. Appl. Environ. Microbiol. 2012, 78, 994–1003. [Google Scholar] [CrossRef]
- Neumann, C.S.; Fujimori, D.G.; Walsh, C.T. Halogenation strategies in natural product biosynthesis. Chem. Biol. 2008, 15, 99–109. [Google Scholar] [CrossRef]
- Gao, P.; Huang, Y. Detection, distribution, and organohalogen compound discovery implications of the reduced flavin adenine dinucleotide-dependent halogenase gene in major filamentous actinomycete taxonomic groups. Appl. Environ. Microbiol. 2009, 75, 4813–4820. [Google Scholar] [CrossRef]
- Shen, B. Polyketide biosynthesis beyond the type I, II and III polyketide synthase paradigms. Curr. Opin. Chem. Biol. 2003, 7, 285–295. [Google Scholar] [CrossRef]
- Schwarzer, D.; Finking, R.; Marahiel, M.A. Nonribosomal peptides: From genes to products. Nat. Prod. Rep. 2003, 20, 275–287. [Google Scholar] [CrossRef]
- González, I.; Ayuso-Sacido, A.; Anderson, A.; Genilloud, O. Actinomycetes isolated from lichens: Evaluation of their diversity and detection of biosynthetic gene sequences. FEMS Microbiol. Ecol. 2005, 54, 401–415. [Google Scholar] [CrossRef]
- Li, X.-L.; Xu, M.-J.; Zhao, Y.-L.; Xu, J. A Novel Benzo [f][1,7] Naphthyridine Produced by Streptomyces Albogriseolus from Mangrove Sediments. Molecules 2010, 15, 9298–9307. [Google Scholar] [CrossRef]
- Xu, J.; Wang, Y.; Xie, S.-J.; Xu, J.; Xiao, J.; Ruan, J.-S. Streptomyces xiamenensis sp. nov., isolated from mangrove sediment. Int. J. Syst. Evol. Microbiol. 2009, 59, 472–476. [Google Scholar] [CrossRef]
- Xu, M.-J.; Liu, X.-J.; Zhao, Y.-L.; Liu, D.; Xu, Z.-H.; Lang, X.-M.; Ao, P.; Lin, W.-H.; Yang, S.-L.; Zhang, Z.-G. Identification and characterization of an anti-fibrotic benzopyran compound isolated from mangrove-derived Streptomyces xiamenensis. Mar. Drugs 2012, 10, 639–654. [Google Scholar] [CrossRef]
- Liu, X.-J.; Xu, M.-J.; Fan, S.-T.; Wu, Z.; Li, J.; Yang, X.-M.; Wang, Y.-H.; Xu, J.; Zhang, Z.-G. Xiamenmycin Attenuates Hypertrophic Scars by Suppressing Local Inflammation and the Effects of Mechanical Stress. J. Invest. Dermatol. 2013, 133, 1351–1360. [Google Scholar] [CrossRef]
- Zhang, X.; Sun, Y.; Bao, J.; He, F.; Xu, X.; Qi, S. Phylogenetic survey and antimicrobial activity of culturable microorganisms associated with the South China Sea black coral Antipathes dichotoma. FEMS Microbiol. Lett. 2012, 336, 122–130. [Google Scholar] [CrossRef]
- Yin, X.; Zabriskie, T.M. The enduracidin biosynthetic gene cluster from Streptomyces fungicidicus. Microbiology 2006, 152, 2969–2983. [Google Scholar] [CrossRef]
- Wu, Y.; Kang, Q.; Shen, Y.; Su, W.; Bai, L. Cloning and functional analysis of the naphthomycin biosynthetic gene cluster in Streptomyces sp. CS. Mol. Biosyst. 2011, 7, 2459–2469. [Google Scholar] [CrossRef]
- Marahiel, M.A.; Stachelhaus, T.; Mootz, H.D. Modular peptide synthetases involved in nonribosomal peptide synthesis. Chem. Rev. 1997, 97, 2651–2674. [Google Scholar] [CrossRef]
- Zhao, W.; Zhong, Y.; Yuan, H.; Wang, J.; Zheng, H.; Wang, Y.; Cen, X.; Xu, F.; Bai, J.; Han, X. Complete genome sequence of the rifamycin SV-producing Amycolatopsis mediterranei U32 revealed its genetic characteristics in phylogeny and metabolism. Cell Res. 2010, 20, 1096–1108. [Google Scholar] [CrossRef]
- Floss, H.G.; Yu, T.-W.; Arakawa, K. The biosynthesis of 3-amino-5-hydroxybenzoic acid (AHBA), the precursor of mC7N units in ansamycin and mitomycin antibiotics: A review. J. Antibiot. (Tokyo) 2010, 64, 35–44. [Google Scholar]
- Floss, H.G.; Yu, T.W. Rifamycin-mode of action, resistance, and biosynthesis. Chem. Rev. 2005, 105, 621–632. [Google Scholar] [CrossRef]
- Gribble, G.W. The diversity of naturally produced organohalogens. Chemosphere 2003, 52, 289–297. [Google Scholar] [CrossRef]
- Gribble, G.W. Naturally occurring organohalogen compounds—a comprehensive survey. Fortschr. Chem. Org. Naturst. 1996, 68, 1–423. [Google Scholar] [CrossRef]
- Auffinger, P.; Hays, F.A.; Westhof, E.; Ho, P.S. Halogen bonds in biological molecules. Proc. Natl. Acad. Sci. USA 2004, 101, 16789–16794. [Google Scholar] [CrossRef]
- Smith, D.R.; Gruschow, S.; Goss, R.J. Scope and potential of halogenases in biosynthetic applications. Curr. Opin. Chem. Biol. 2013, 17, 276–283. [Google Scholar] [CrossRef]
- Pistorius, D.; Muller, R. Discovery of the rhizopodin biosynthetic gene cluster in Stigmatella aurantiaca Sg a15 by genome mining. Chembiochem 2012, 13, 416–426. [Google Scholar] [CrossRef]
- Vaillancourt, F.H.; Vosburg, D.A.; Walsh, C.T. Dichlorination and Bromination of a Threonyl-S-Carrier Protein by the Non-heme FeII Halogenase SyrB2. Chembiochem 2006, 7, 748–752. [Google Scholar] [CrossRef]
- Bayer, K.; Scheuermayer, M.; Fieseler, L.; Hentschel, U. Genomic mining for novel FADH2-dependent halogenases in marine sponge-associated microbial consortia. Mar. Biotechnol. 2013, 15, 63–72. [Google Scholar] [CrossRef]
- Murphy, C.D. Recent developments in enzymatic chlorination. Nat. Prod. Rep. 2006, 23, 147–152. [Google Scholar] [CrossRef]
- Ayuso-Sacido, A.; Genilloud, O. New PCR primers for the screening of NRPS and PKS-I systems in actinomycetes: Detection and distribution of these biosynthetic gene sequences in major taxonomic groups. Microb. Ecol. 2005, 49, 10–24. [Google Scholar] [CrossRef]
- Schirmer, A.; Gadkari, R.; Reeves, C.D.; Ibrahim, F.; DeLong, E.F.; Hutchinson, C.R. Metagenomic analysis reveals diverse polyketide synthase gene clusters in microorganisms associated with the marine sponge Discodermia dissoluta. Appl. Environ. Microbiol. 2005, 71, 4840–4849. [Google Scholar] [CrossRef]
- Metsä-Ketelä, M.; Salo, V.; Halo, L.; Hautala, A.; Hakala, J.; Mäntsälä, P.; Ylihonko, K. An efficient approach for screening minimal PKS genes from Streptomyces. FEMS Microbiol. Lett. 1999, 180, 1–6. [Google Scholar]
- Kieser, T.; Bibb, M.J.; Buttner, M.J.; Chater, K.F.; Hopwood, D.A. Practical Streptomyces Genetics; John Innes Foundation Norwich: Norwich, UK, 2000; p. 412. [Google Scholar]
- Ishikawa, J.; Hotta, K. FramePlot: A new implementation of the frame analysis for predicting protein-coding regions in bacterial DNA with a high G + C content. FEMS Microbiol. Lett. 1999, 174, 251–253. [Google Scholar] [CrossRef]
- Yadav, G.; Gokhale, R.S.; Mohanty, D. SEARCHPKS: A program for detection and analysis of polyketide synthase domains. Nucleic Acids Res. 2003, 31, 3654–3658. [Google Scholar] [CrossRef]
Supplementary Files
© 2013 by the authors; licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution license (http://creativecommons.org/licenses/by/3.0/).
Share and Cite
Li, X.-G.; Tang, X.-M.; Xiao, J.; Ma, G.-H.; Xu, L.; Xie, S.-J.; Xu, M.-J.; Xiao, X.; Xu, J. Harnessing the Potential of Halogenated Natural Product Biosynthesis by Mangrove-Derived Actinomycetes. Mar. Drugs 2013, 11, 3875-3890. https://doi.org/10.3390/md11103875
Li X-G, Tang X-M, Xiao J, Ma G-H, Xu L, Xie S-J, Xu M-J, Xiao X, Xu J. Harnessing the Potential of Halogenated Natural Product Biosynthesis by Mangrove-Derived Actinomycetes. Marine Drugs. 2013; 11(10):3875-3890. https://doi.org/10.3390/md11103875
Chicago/Turabian StyleLi, Xue-Gong, Xiao-Min Tang, Jing Xiao, Guang-Hui Ma, Li Xu, Shu-Jie Xie, Min-Juan Xu, Xiang Xiao, and Jun Xu. 2013. "Harnessing the Potential of Halogenated Natural Product Biosynthesis by Mangrove-Derived Actinomycetes" Marine Drugs 11, no. 10: 3875-3890. https://doi.org/10.3390/md11103875



