Epigenetic Control and Cancer: The Potential of Histone Demethylases as Therapeutic Targets
Abstract
:1. Introduction
 ), phosphorylation (
), phosphorylation (  ), ubiquitination (
), ubiquitination (  ), acetylation (
), acetylation (  ) or methylation (
) or methylation (  ) are shown as one-letter code, together with the different enzymes able to add (in blue) each functional group. The histone demethylases are shown in red, besides the lysine residues susceptible to their specific enzymatic activity.
) are shown as one-letter code, together with the different enzymes able to add (in blue) each functional group. The histone demethylases are shown in red, besides the lysine residues susceptible to their specific enzymatic activity.
 ), phosphorylation (
), phosphorylation (  ), ubiquitination (
), ubiquitination (  ), acetylation (
), acetylation (  ) or methylation (
) or methylation (  ) are shown as one-letter code, together with the different enzymes able to add (in blue) each functional group. The histone demethylases are shown in red, besides the lysine residues susceptible to their specific enzymatic activity.
) are shown as one-letter code, together with the different enzymes able to add (in blue) each functional group. The histone demethylases are shown in red, besides the lysine residues susceptible to their specific enzymatic activity.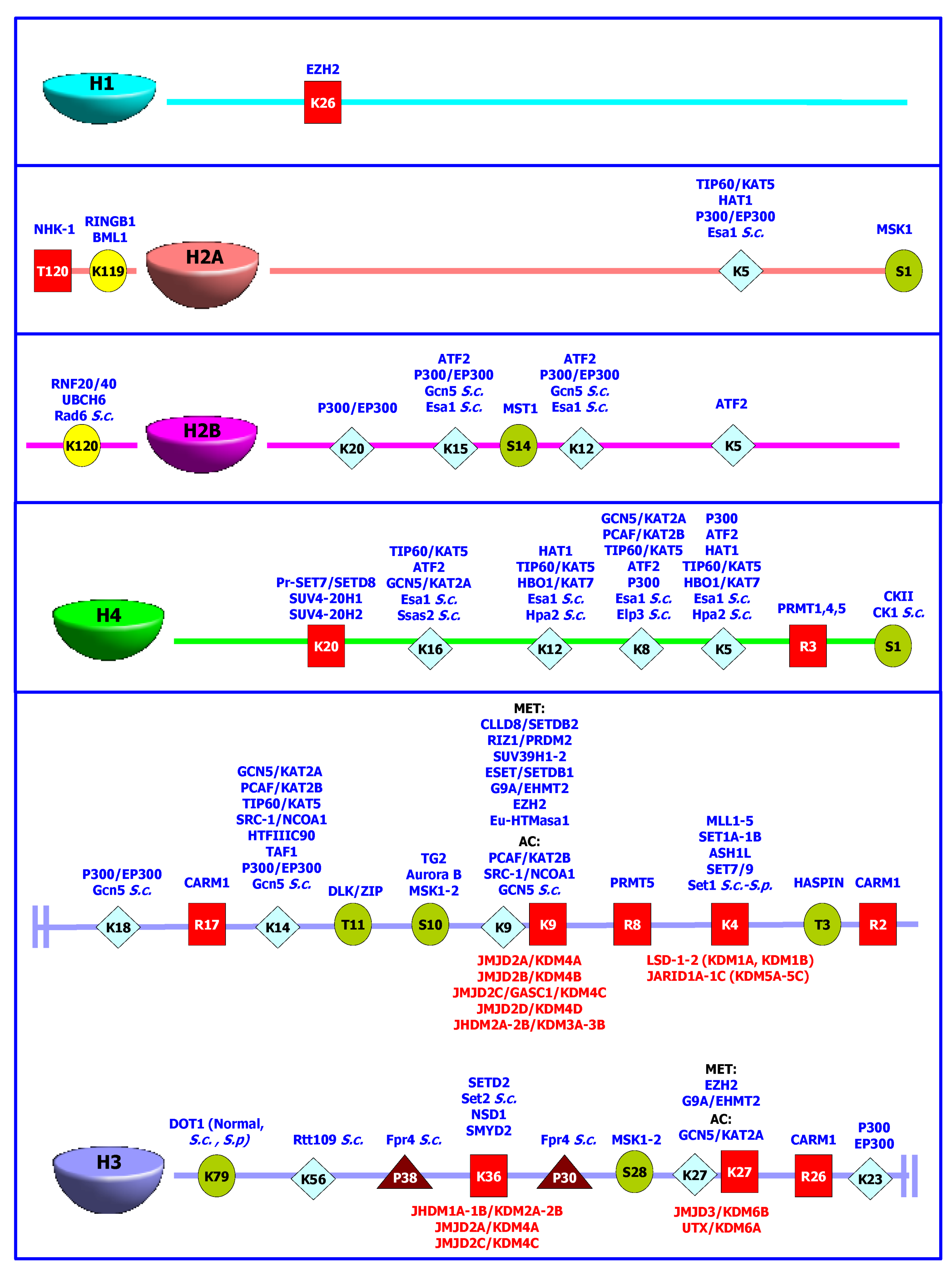
2. Connection between Histone Methylation and Disease
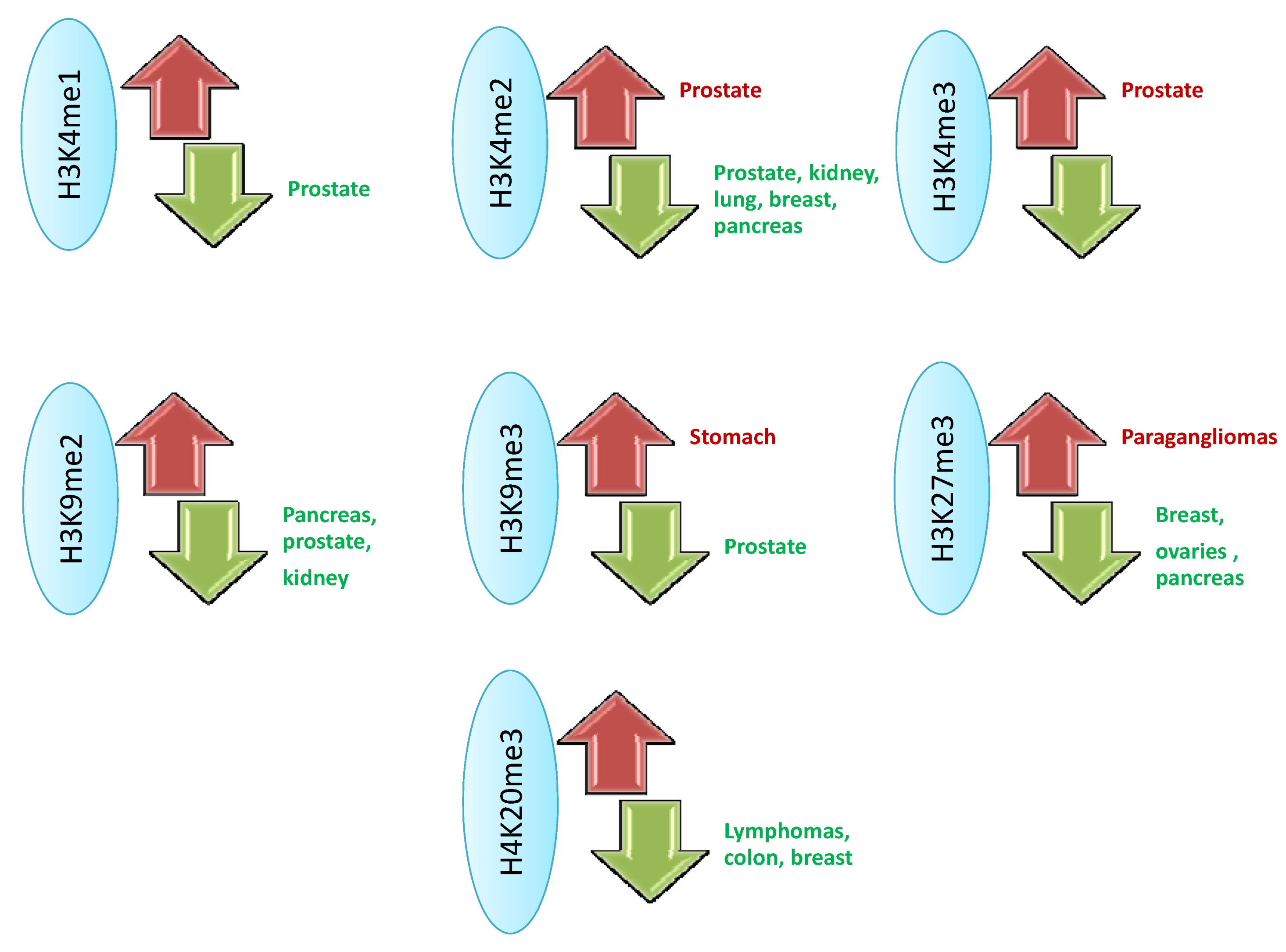
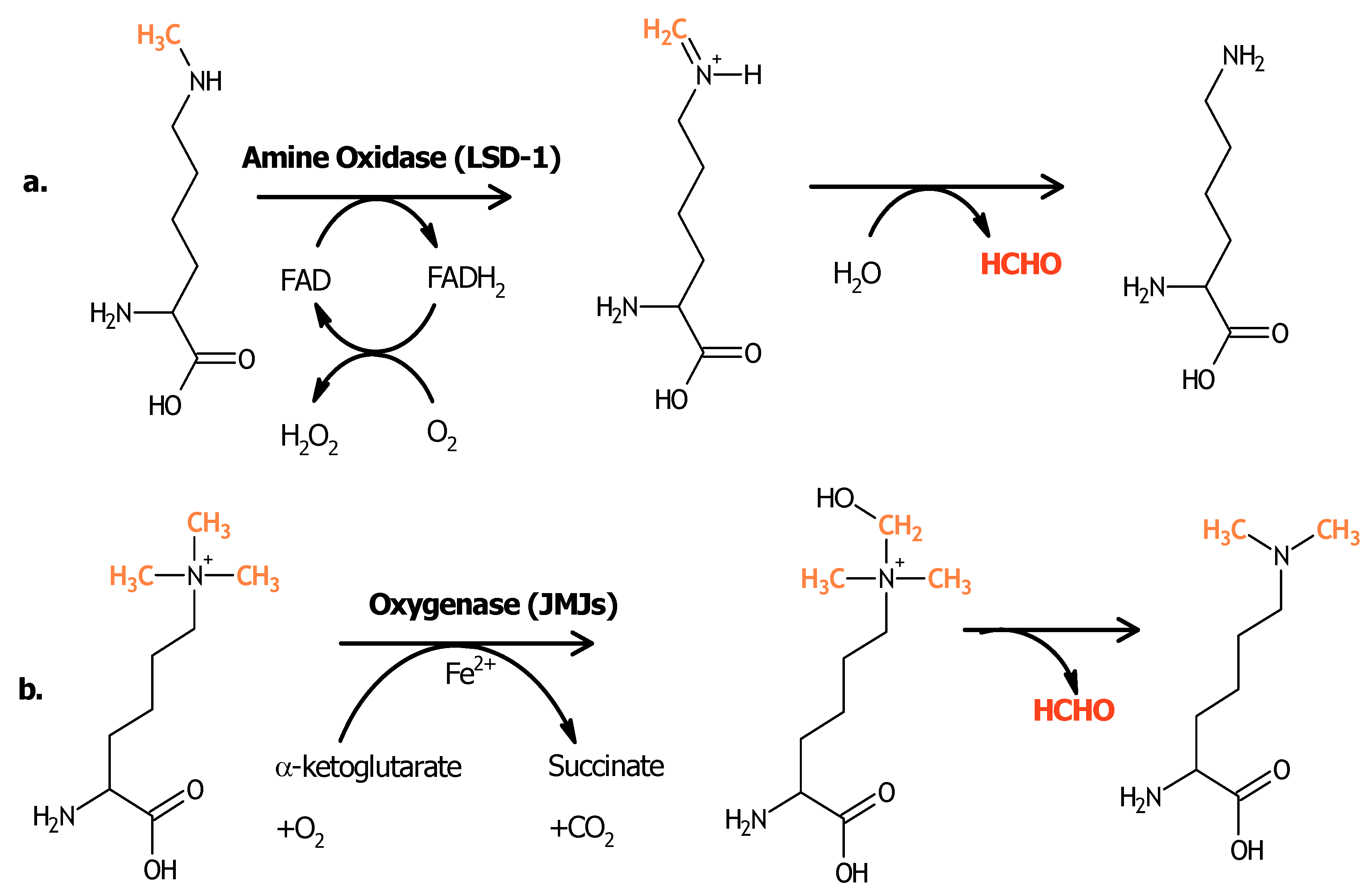
3. Histone Demethylases and Cancer
| Histone demethylase | Genebank Acc. Number | Alteration | Associated cancer | References |
|---|---|---|---|---|
| LSD-1/KDM1A | NM_015013 | Over-expression | ER-negative breast, Prostate, Leukemia, Sarcomas, Lung, neuroblastoma | [31,41,42,43,44,45,46,47,48,49] |
| Low expression | Hepatocellular, gastrointestinal, Hepatobiliary and metastatic breast carcinomas | [50,51] | ||
| JARID1A/KDM5A | NM_001042603 | Over-expression | Gastric, Leukemia | [52,53] |
| JARID1B/KDM5B | NM_006618 | Over-expression | Testicular, Ovaries, Breast, Prostate, leukemia | [54,55,56,57,58,59,60,61] |
| JARID1C/KDM5C | NM_004187.3 NM_001146702.1 | Over-expression | Renal Carcinoma, Cervical | [62,63,64] |
| JHDM1B/FBXL10/KDM2B | NM_032590 | Over-expression/Mutation | Lymphomas, Acute myeloid leukemia | [65,66,67] |
| Low expression | Gliobastoma multiforme, brain | [68] | ||
| JMJD2C/KDM4C | NM_015061; NM_001146694; NM_001146695; NM_001146696 | Over-expression | MALT and Hodgkin Lymphomas, myeloid leukemia, Breast, gliobastoma multiforme, prostate, desmoplasic meduloblastoma, sarcomatoid and esophageal carcinoma. | [69,70,71,72,73,74,75,76,77,78,79,80,81,82] |
| JMJD2A/KDM4A | NM_014663 | Over-expression | Kaposi Sarcoma, Prostate, Breast, Colon, Bladder. | [18,47,83,84,85,86,87] |
| JMJD3/KDM6B | NM_001080424 | Over-expression | Colon, Lymphomas | [88,89,90] |
| UTX/KDM6A | NM_021140 | Mutation/Low expression | Advanced forms of cancer, Multiple mieloma, Leukemia, renal and bladder carcinomas. | [91,92,93,94,95,96] |
3.1. LSD-1 and LSD-2
3.2. JARID1 Family
3.3. JHDM1B
3.4. JMJD2C
3.5. JMJD2A
3.6. JMJD3
3.7. UTX
4. Therapeutic Approaches

5. Conclusions
Acknowledgments
Conflict of Interest
References
- World Health Organization, WHO World Cancer Report 2008; WHO: Geneva, Switzerland, 2008.
- Kornberg, R.D. Chromatin structure: A repeating unit of histones and DNA. Science 1974, 184, 868–871. [Google Scholar]
- Li, G.; Reinberg, D. Chromatin higher-order structures and gene regulation. Curr. Opin. Genet. Dev. 2011, 21, 175–186. [Google Scholar] [CrossRef]
- Luger, K.; Mader, A.W.; Richmond, R.K.; Sargent, D.F.; Richmond, T.J. Crystal structure of the nucleosome core particle at 2.8 A resolution. Nature 1997, 389, 251–260. [Google Scholar]
- Xu, M.; Zhu, B. Nucleosome assembly and epigenetic inheritance. Protein Cell 2010, 1, 820–829. [Google Scholar] [CrossRef]
- Chen, P.; Li, G. Dynamics of the higher-order structure of chromatin. Protein Cell 2010, 1, 967–971. [Google Scholar] [CrossRef]
- Bassett, A.; Cooper, S.; Wu, C.; Travers, A. The folding and unfolding of eukaryotic chromatin. Curr. Opin. Genet. Dev. 2009, 19, 159–165. [Google Scholar]
- Berger, S.L. The complex language of chromatin regulation during transcription. Nature 2007, 447, 407–412. [Google Scholar] [CrossRef]
- Latham, J.A.; Dent, S.Y. Cross-regulation of histone modifications. Nat. Struct. Mol. Biol. 2007, 14, 1017–1024. [Google Scholar] [CrossRef]
- Arrowsmith, C.H.; Bountra, C.; Fish, P.V.; Lee, K.; Schapira, M. Epigenetic protein families: A new frontier for drug discovery. Nat. Rev. Drug Discov. 2012, 11, 384–400. [Google Scholar] [CrossRef]
- Bayarsaihan, D. Epigenetic mechanisms in inflammation. J. Dent. Res. 2011, 90, 9–17. [Google Scholar] [CrossRef]
- Hamm, C.A.; Costa, F.F. The impact of epigenomics on future drug design and new therapies. Drug Discov. Today 2011, 16, 626–635. [Google Scholar] [CrossRef]
- Tsai, H.C.; Baylin, S.B. Cancer epigenetics: Linking basic biology to clinical medicine. Cell Res. 2011, 21, 502–517. [Google Scholar] [CrossRef]
- Zahir, F.R.; Brown, C.J. Epigenetic impacts on neurodevelopment: Pathophysiological mechanisms and genetic modes of action. Pediatr. Res. 2011, 69, 92R–100R. [Google Scholar] [CrossRef]
- Zhang, X.; Ho, S.M. Epigenetics meets endocrinology. J. Mol. Endocrinol. 2011, 46, R11–R32. [Google Scholar] [CrossRef]
- Castellano, S.; Kuck, D.; Viviano, M.; Yoo, J.; Lopez-Vallejo, F.; Conti, P.; Tamborini, L.; Pinto, A.; Medina-Franco, J.L.; Sbardella, G. Synthesis and biochemical evaluation of delta(2)-isoxazoline derivatives as DNA methyltransferase 1 inhibitors. J. Med. Chem. 2011, 54, 7663–7677. [Google Scholar] [CrossRef]
- Yi, T.Z.; Li, J.; Han, X.; Guo, J.; Qu, Q.; Guo, L.; Sun, H.D.; Tan, W.H. DNMT Inhibitors and HDAC Inhibitors Regulate E-Cadherin and Bcl-2 Expression in Endometrial Carcinoma in vitro and in vivo. Chemotherapy 2012, 58, 19–29. [Google Scholar] [CrossRef]
- Chen, M.; Shabashvili, D.; Nawab, A.; Yang, S.X.; Dyer, L.M.; Brown, K.D.; Hollingshead, M.; Hunter, K.W.; Kaye, F.J.; Hochwald, S.N.; et al. DNA methyltransferase inhibitor, zebularine, delays tumor growth and induces apoptosis in a genetically engineered mouse model of breast cancer. Mol. Cancer Ther. 2012, 11, 370–382. [Google Scholar] [CrossRef]
- Ahmad, M.; Hamid, A.; Hussain, A.; Majeed, R.; Qurishi, Y.; Bhat, J.A.; Najar, R.A.; Qazi, A.K.; Zargar, M.A.; Singh, S.K.; et al. Understanding histone deacetylases in the cancer development and treatment: An epigenetic perspective of cancer chemotherapy. DNA Cell Biol. 2012. [Google Scholar]
- Mehdi, O.; Francoise, S.; Sofia, C.L.; Urs, G.; Kevin, Z.; Bernard, S.; Igor, S.; Anabela, C.D.; Dominique, L.; Eric, M.; Ali, O. HDAC gene expression in pancreatic tumor cell lines following treatment with the HDAC inhibitors panobinostat (LBH589) and trichostatine (TSA). Pancreatology 2012, 12, 146–155. [Google Scholar]
- Munster, P.N.; Thurn, K.T.; Thomas, S.; Raha, P.; Lacevic, M.; Miller, A.; Melisko, M.; Ismail-Khan, R.; Rugo, H.; Moasser, M.; et al. A phase II study of the histone deacetylase inhibitor vorinostat combined with tamoxifen for the treatment of patients with hormone therapy-resistant breast cancer. Br. J. Cancer 2011, 104, 1828–1835. [Google Scholar] [CrossRef]
- Oike, T.; Ogiwara, H.; Torikai, K.; Nakano, T.; Yokota, J.; Kohno, T. Garcinol, a histone acetyltransferase inhibitor, radiosensitizes cancer cells by inhibiting non-homologous end joining. Int. J. Radiat. Oncol. Biol. Phys. 2012, in press.. [Google Scholar]
- Thurn, K.T.; Thomas, S.; Moore, A.; Munster, P.N. Rational therapeutic combinations with histone deacetylase inhibitors for the treatment of cancer. Future Oncol. 2011, 7, 263–283. [Google Scholar] [CrossRef]
- Uehara, N.; Yoshizawa, K.; Tsubura, A. Vorinostat enhances protein stability of p27 and p21 through negative regulation of Skp2 and Cks1 in human breast cancer cells. Oncol. Rep. 2012, 28, 105–110. [Google Scholar]
- Hamada, S.; Suzuki, T.; Mino, K.; Koseki, K.; Oehme, F.; Flamme, I.; Ozasa, H.; Itoh, Y.; Ogasawara, D.; Komaarashi, H.; et al. Design, synthesis, enzyme-inhibitory activity, and effect on human cancer cells of a novel series of jumonji domain-containing protein 2 histone demethylase inhibitors. J. Med. Chem. 2010, 53, 5629–5638. [Google Scholar] [CrossRef]
- Rao, M.; Chinnasamy, N.; Hong, J.A.; Zhang, Y.; Zhang, M.; Xi, S.; Liu, F.; Marquez, V.E.; Morgan, R.A.; Schrump, D.S. Inhibition of histone lysine methylation enhances cancer-testis antigen expression in lung cancer cells: Implications for adoptive immunotherapy of cancer. Cancer Res. 2011, 71, 4192–4204. [Google Scholar] [CrossRef]
- Toyokawa, G.; Cho, H.S.; Iwai, Y.; Yoshimatsu, M.; Takawa, M.; Hayami, S.; Maejima, K.; Shimizu, N.; Tanaka, H.; Tsunoda, T.; et al. The histone demethylase JMJD2B plays an essential role in human carcinogenesis through positive regulation of cyclin-dependent kinase 6. Cancer Prev. Res. (Phila.) 2011, 4, 2051–2061. [Google Scholar] [CrossRef]
- Toyokawa, G.; Cho, H.S.; Masuda, K.; Yamane, Y.; Yoshimatsu, M.; Hayami, S.; Takawa, M.; Iwai, Y.; Daigo, Y.; Tsuchiya, E.; et al. Histone lysine methyltransferase Wolf-Hirschhorn syndrome candidate 1 is involved in human carcinogenesis through regulation of the Wnt pathway. Neoplasia 2011, 13, 887–898. [Google Scholar]
- Wang, J.; Lu, F.; Ren, Q.; Sun, H.; Xu, Z.; Lan, R.; Liu, Y.; Ward, D.; Quan, J.; Ye, T.; et al. Novel histone demethylase LSD1 inhibitors selectively target cancer cells with pluripotent stem cell properties. Cancer Res. 2011, 71, 7238–7249. [Google Scholar] [CrossRef]
- Chen, X.; Hu, Y.; Zhou, D.X. Epigenetic gene regulation by plant Jumonji group of histone demethylase. Biochim. Biophys. Acta 2011, 1809, 421–426. [Google Scholar] [CrossRef]
- Li, Y.; Deng, C.; Hu, X.; Patel, B.; Fu, X.; Qiu, Y.; Brand, M.; Zhao, K.; Huang, S. Dynamic interaction between TAL1 oncoprotein and LSD1 regulates TAL1 function in hematopoiesis and leukemogenesis. Oncogene 2012. [Google Scholar]
- Rodova, M.; Lu, Q.; Li, Y.; Woodbury, B.G.; Crist, J.D.; Gardner, B.M.; Yost, J.G.; Zhong, X.B.; Anderson, H.C.; Wang, J. Nfat1 regulates adult articular chondrocyte function through its age-dependent expression mediated by epigenetic histone methylation. J. Bone Miner. Res. 2011, 26, 1974–1986. [Google Scholar] [CrossRef]
- Ura, H.; Murakami, K.; Akagi, T.; Kinoshita, K.; Yamaguchi, S.; Masui, S.; Niwa, H.; Koide, H.; Yokota, T. Eed/Sox2 regulatory loop controls ES cell self-renewal through histone methylation and acetylation. EMBO J. 2011, 30, 2190–2204. [Google Scholar]
- Vicent, G.P.; Nacht, A.S.; Font-Mateu, J.; Castellano, G.; Gaveglia, L.; Ballare, C.; Beato, M. Four enzymes cooperate to displace histone H1 during the first minute of hormonal gene activation. Genes Dev. 2011, 25, 845–862. [Google Scholar]
- Bannister, A.J.; Schneider, R.; Kouzarides, T. Histone methylation: Dynamic or static? Cell 2002, 109, 801–806. [Google Scholar] [CrossRef]
- Shi, Y.; Lan, F.; Matson, C.; Mulligan, P.; Whetstine, J.R.; Cole, P.A.; Casero, R.A. Histone demethylation mediated by the nuclear amine oxidase homolog LSD1. Cell 2004, 119, 941–953. [Google Scholar] [CrossRef]
- Barski, A.; Cuddapah, S.; Cui, K.; Roh, T.Y.; Schones, D.E.; Wang, Z.; Wei, G.; Chepelev, I.; Zhao, K. High-resolution profiling of histone methylations in the human genome. Cell 2007, 129, 823–837. [Google Scholar] [CrossRef]
- Rotili, D.; Mai, A. Targeting histone demethylases: A new avenue for the fight against cancer. Genes Cancer 2011, 2, 663–679. [Google Scholar]
- Varier, R.A.; Timmers, H.T. Histone lysine methylation and demethylation pathways in cancer. Biochim. Biophys. Acta 2011, 1815, 75–89. [Google Scholar]
- Hou, H.; Yu, H. Structural insights into histone lysine demethylation. Curr. Opin. Struct. Biol. 2010, 20, 739–748. [Google Scholar]
- Kahl, P.; Gullotti, L.; Heukamp, L.C.; Wolf, S.; Friedrichs, N.; Vorreuther, R.; Solleder, G.; Bastian, P.J.; Ellinger, J.; Metzger, E.; et al. Androgen receptor coactivators lysine-specific histone demethylase 1 and four and a half LIM domain protein 2 predict risk of prostate cancer recurrence. Cancer Res. 2006, 66, 11341–11347. [Google Scholar]
- Harris, W.J.; Huang, X.; Lynch, J.T.; Spencer, G.J.; Hitchin, J.R.; Li, Y.; Ciceri, F.; Blaser, J.G.; Greystoke, B.F.; Jordan, A.M.; et al. The Histone demethylase KDM1A sustains the oncogenic potential of MLL-AF9 leukemia stem cells. Cancer Cell 2012, 21, 473–487. [Google Scholar] [CrossRef]
- Wissmann, M.; Yin, N.; Muller, J.M.; Greschik, H.; Fodor, B.D.; Jenuwein, T.; Vogler, C.; Schneider, R.; Gunther, T.; Buettner, R.; et al. Cooperative demethylation by JMJD2C and LSD1 promotes androgen receptor-dependent gene expression. Nat. Cell Biol. 2007, 9, 347–353. [Google Scholar] [CrossRef]
- Bennani-Baiti, I.M.; Machado, I.; Llombart-Bosch, A.; Kovar, H. Lysine-specific demethylase 1 (LSD1/KDM1A/AOF2/BHC110) is expressed and is an epigenetic drug target in chondrosarcoma, Ewing’s sarcoma, osteosarcoma, and rhabdomyosarcoma. Hum. Pathol. 2012, 43, 1300–1307. [Google Scholar] [CrossRef]
- Lv, T.; Yuan, D.; Miao, X.; Lv, Y.; Zhan, P.; Shen, X.; Song, Y. Over-expression of LSD1 promotes proliferation, migration and invasion in non-small cell lung cancer. PLoS One 2012, 7, e35065. [Google Scholar]
- Lim, S.; Janzer, A.; Becker, A.; Zimmer, A.; Schule, R.; Buettner, R.; Kirfel, J. Lysine-specific demethylase 1 (LSD1) is highly expressed in ER-negative breast cancers and a biomarker predicting aggressive biology. Carcinogenesis 2010, 31, 512–520. [Google Scholar] [CrossRef]
- Kauffman, E.C.; Robinson, B.D.; Downes, M.J.; Powell, L.G.; Lee, M.M.; Scherr, D.S.; Gudas, L.J.; Mongan, N.P. Role of androgen receptor and associated lysine-demethylase coregulators, LSD1 and JMJD2A, in localized and advanced human bladder cancer. Mol. Carcinog. 2011, 50, 931–944. [Google Scholar] [CrossRef]
- Metzger, E.; Wissmann, M.; Yin, N.; Muller, J.M.; Schneider, R.; Peters, A.H.; Gunther, T.; Buettner, R.; Schule, R. LSD1 demethylates repressive histone marks to promote androgen-receptor-dependent transcription. Nature 2005, 437, 436–439. [Google Scholar]
- Schulte, J.H.; Lim, S.; Schramm, A.; Friedrichs, N.; Koster, J.; Versteeg, R.; Ora, I.; Pajtler, K.; Klein-Hitpass, L.; Kuhfittig-Kulle, S.; et al. Lysine-specific demethylase 1 is strongly expressed in poorly differentiated neuroblastoma: Implications for therapy. Cancer Res. 2009, 69, 2065–2071. [Google Scholar]
- Magerl, C.; Ellinger, J.; Braunschweig, T.; Kremmer, E.; Koch, L.K.; Holler, T.; Buttner, R.; Luscher, B.; Gutgemann, I. H3K4 dimethylation in hepatocellular carcinoma is rare compared with other hepatobiliary and gastrointestinal carcinomas and correlates with expression of the methylase Ash2 and the demethylase LSD1. Hum. Pathol. 2010, 41, 181–189. [Google Scholar] [CrossRef]
- Wang, Y.; Zhang, H.; Chen, Y.; Sun, Y.; Yang, F.; Yu, W.; Liang, J.; Sun, L.; Yang, X.; Shi, L.; et al. LSD1 is a subunit of the NuRD complex and targets the metastasis programs in breast cancer. Cell 2009, 138, 660–672. [Google Scholar] [CrossRef]
- Zeng, J.; Ge, Z.; Wang, L.; Li, Q.; Wang, N.; Bjorkholm, M.; Jia, J.; Xu, D. The histone demethylase RBP2 Is overexpressed in gastric cancer and its inhibition triggers senescence of cancer cells. Gastroenterology 2010, 138, 981–992. [Google Scholar]
- Van Zutven, L.J.; Onen, E.; Velthuizen, S.C.; van Drunen, E.; von Bergh, A.R.; van den Heuvel-Eibrink, M.M.; Veronese, A.; Mecucci, C.; Negrini, M.; et al. Identification of NUP98 abnormalities in acute leukemia: JARID1A (12p13) as a new partner gene. Genes Chromosomes Cancer 2006, 45, 437–446. [Google Scholar] [CrossRef]
- Barrett, A.; Santangelo, S.; Tan, K.; Catchpole, S.; Roberts, K.; Spencer-Dene, B.; Hall, D.; Scibetta, A.; Burchell, J.; Verdin, E.; et al. Breast cancer associated transcriptional repressor PLU-1/JARID1B interacts directly with histone deacetylases. Int. J. Cancer 2007, 121, 265–275. [Google Scholar] [CrossRef]
- Liggins, A.P.; Lim, S.H.; Soilleux, E.J.; Pulford, K.; Banham, A.H. A panel of cancer-testis genes exhibiting broad-spectrum expression in haematological malignancies. Cancer Immun. 2010, 10, 8. [Google Scholar]
- Lu, P.J.; Sundquist, K.; Baeckstrom, D.; Poulsom, R.; Hanby, A.; Meier-Ewert, S.; Jones, T.; Mitchell, M.; Pitha-Rowe, P.; Freemont, P.; Taylor-Papadimitriou, J. A novel gene (PLU-1) containing highly conserved putative DNA/chromatin binding motifs is specifically up-regulated in breast cancer. Biol. Chem. 1999, 274, 15633–15645. [Google Scholar]
- Yamane, K.; Tateishi, K.; Klose, R.J.; Fang, J.; Fabrizio, L.A.; Erdjument-Bromage, H.; Taylor-Papadimitriou, J.; Tempst, P.; Zhang, Y. PLU-1 is an H3K4 demethylase involved in transcriptional repression and breast cancer cell proliferation. Mol. Cell 2007, 25, 801–812. [Google Scholar]
- Xiang, Y.; Zhu, Z.; Han, G.; Ye, X.; Xu, B.; Peng, Z.; Ma, Y.; Yu, Y.; Lin, H.; Chen, A.P.; et al. JARID1B is a histone H3 lysine 4 demethylase up-regulated in prostate cancer. Proc. Natl. Acad. Sci. USA 2007, 104, 19226–19231. [Google Scholar]
- Kim, J.; Shin, S.; Subramaniam, M.; Bruinsma, E.; Kim, T.D.; Hawse, J.R.; Spelsberg, T.C.; Janknecht, R. Histone demethylase JARID1B/KDM5B is a corepressor of TIEG1/KLF10. Biochem. Biophys. Res. Commun. 2010, 401, 412–416. [Google Scholar]
- Catchpole, S.; Spencer-Dene, B.; Hall, D.; Santangelo, S.; Rosewell, I.; Guenatri, M.; Beatson, R.; Scibetta, A.G.; Burchell, J.M.; Taylor-Papadimitriou, J. PLU-1/JARID1B/KDM5B is required for embryonic survival and contributes to cell proliferation in the mammary gland and in ER+ breast cancer cells. Int. J. Oncol. 2011, 38, 1267–1277. [Google Scholar]
- Ma, X.D.; Han, H.D.; Huang, Y.Q.; Zou, Y. The effects of JARID1B siRNA on proliferation and apoptosis in HL-60 cell. Zhonghua Xue Ye Xue Za Zhi 2012, 33, 392–396. [Google Scholar]
- Niu, X.; Zhang, T.; Liao, L.; Zhou, L.; Lindner, D.J.; Zhou, M.; Rini, B.; Yan, Q.; Yang, H. The von Hippel-Lindau tumor suppressor protein regulates gene expression and tumor growth through histone demethylase JARID1C. Oncogene 2012, 31, 776–786. [Google Scholar]
- Smith, J.A.; White, E.A.; Sowa, M.E.; Powell, M.L.; Ottinger, M.; Harper, J.W.; Howley, P.M. Genome-wide siRNA screen identifies SMCX, EP400, and Brd4 as E2-dependent regulators of human papillomavirus oncogene expression. Proc. Natl. Acad. Sci. USA 2010, 107, 3752–3757. [Google Scholar]
- Varela, I.; Tarpey, P.; Raine, K.; Huang, D.; Ong, C.K.; Stephens, P.; Davies, H.; Jones, D.; Lin, M.L.; Teague, J.; et al. Exome sequencing identifies frequent mutation of the SWI/SNF complex gene PBRM1 in renal carcinoma. Nature 2011, 469, 539–542. [Google Scholar]
- He, J.; Nguyen, A.T.; Zhang, Y. KDM2b/JHDM1b, an H3K36me2-specific demethylase, is required for initiation and maintenance of acute myeloid leukemia. Blood 2011, 117, 3869–3880. [Google Scholar] [CrossRef]
- Nakamura, S.; Tan, L.; Nagata, Y.; Takemura, T.; Asahina, A.; Yokota, D.; Yagyu, T.; Shibata, K.; Fujisawa, S.; Ohnishi, K. JmjC-domain containing histone demethylase 1B-mediated p15(Ink4b) suppression promotes the proliferation of leukemic progenitor cells through modulation of cell cycle progression in acute myeloid leukemia. Mol. Carcinog. 2011. [Google Scholar]
- Suzuki, T.; Minehata, K.; Akagi, K.; Jenkins, N.A.; Copeland, N.G. Tumor suppressor gene identification using retroviral insertional mutagenesis in Blm-deficient mice. EMBO J. 2006, 25, 3422–3431. [Google Scholar]
- Frescas, D.; Guardavaccaro, D.; Bassermann, F.; Koyama-Nasu, R.; Pagano, M. JHDM1B/FBXL10 is a nucleolar protein that represses transcription of ribosomal RNA genes. Nature 2007, 450, 309–313. [Google Scholar]
- Cloos, P.A.; Christensen, J.; Agger, K.; Maiolica, A.; Rappsilber, J.; Antal, T.; Hansen, K.H.; Helin, K. The putative oncogene GASC1 demethylates tri- and dimethylated lysine 9 on histone H3. Nature 2006, 442, 307–311. [Google Scholar]
- Italiano, A.; Attias, R.; Aurias, A.; Perot, G.; Burel-Vandenbos, F.; Otto, J.; Venissac, N.; Pedeutour, F. Molecular cytogenetic characterization of a metastatic lung sarcomatoid carcinoma: 9p23 Neocentromere and 9p23-p24 amplification including JAK2 and JMJD2C. Cancer Genet. Cytogenet. 2006, 167, 122–130. [Google Scholar]
- Helias, C.; Struski, S.; Gervais, C.; Leymarie, V.; Mauvieux, L.; Herbrecht, R.; Lessard, M. Polycythemia vera transforming to acute myeloid leukemia and complex abnormalities including 9p homogeneously staining region with amplification of MLLT3, JMJD2C, JAK2, and SMARCA2. Cancer Genet. Cytogenet. 2008, 180, 51–55. [Google Scholar] [CrossRef]
- Nacheva, E.P.; Brazma, D.; Virgili, A.; Howard-Reeves, J.; Chanalaris, A.; Gancheva, K.; Apostolova, M.; Valganon, M.; Mazzullo, H.; Grace, C. Deletions of immunoglobulin heavy chain and T cell receptor gene regions are uniquely associated with lymphoid blast transformation of chronic myeloid leukemia. BMC Genomics 2010, 11, 41. [Google Scholar]
- Min, D.J.; Licht, J.D. Partners in crime: Genes within an amplicon collude to globally deregulate chromatin in lymphoma. Cancer Cell 2010, 18, 539–541. [Google Scholar]
- Rui, L.; Emre, N.C.; Kruhlak, M.J.; Chung, H.J.; Steidl, C.; Slack, G.; Wright, G.W.; Lenz, G.; Ngo, V.N.; Shaffer, A.L.; et al. Cooperative epigenetic modulation by cancer amplicon genes. Cancer Cell 2010, 18, 590–605. [Google Scholar] [CrossRef]
- Liu, G.; Bollig-Fischer, A.; Kreike, B.; van de Vijver, M.J.; Abrams, J.; Ethier, S.P.; Yang, Z.Q. Genomic amplification and oncogenic properties of the GASC1 histone demethylase gene in breast cancer. Oncogene 2009, 28, 4491–4500. [Google Scholar]
- Wu, J.; Liu, S.; Liu, G.; Dombkowski, A.; Abrams, J.; Martin-Trevino, R.; Wicha, M.S.; Ethier, S.P.; Yang, Z.Q. Identification and functional analysis of 9p24 amplified genes in human breast cancer. Oncogene 2012, 31, 333–341. [Google Scholar]
- Fischer, U.; Wullich, B.; Sattler, H.P.; Gottert, E.; Zang, K.D.; Meese, E. DNA amplifications on chromosomes 7, 9 and 12 in glioblastoma detected by reverse chromosome painting. Eur. J. Cancer 1994, 30A, 1124–1127. [Google Scholar]
- Haque, A.; Banik, N.L.; Ray, S.K. Molecular alterations in glioblastoma: Potential targets for immunotherapy. Prog. Mol. Biol. Transl. Sci. 2011, 98, 187–234. [Google Scholar]
- Margareto, J.; Leis, O.; Larrarte, E.; Pomposo, I.C.; Garibi, J.M.; Lafuente, J.V. DNA copy number variation and gene expression analyses reveal the implication of specific oncogenes and genes in GBM. Cancer Investig. 2009, 27, 541–548. [Google Scholar]
- Vinatzer, U.; Gollinger, M.; Mullauer, L.; Raderer, M.; Chott, A.; Streubel, B. Mucosa-associated lymphoid tissue lymphoma: Novel translocations including rearrangements of ODZ2, JMJD2C, and CNN3. Clin. Cancer Res. 2008, 14, 6426–6431. [Google Scholar] [CrossRef]
- Ehrbrecht, A.; Muller, U.; Wolter, M.; Hoischen, A.; Koch, A.; Radlwimmer, B.; Actor, B.; Mincheva, A.; Pietsch, T.; Lichter, P.; et al. Comprehensive genomic analysis of desmoplastic medulloblastomas: Identification of novel amplified genes and separate evaluation of the different histological components. J. Pathol. 2006, 208, 554–563. [Google Scholar] [CrossRef]
- Yang, Z.Q.; Imoto, I.; Fukuda, Y.; Pimkhaokham, A.; Shimada, Y.; Imamura, M.; Sugano, S.; Nakamura, Y.; Inazawa, J. Identification of a novel gene, GASC1, within an amplicon at 9p23-24 frequently detected in esophageal cancer cell lines. Cancer Res. 2000, 60, 4735–4739. [Google Scholar]
- Chen, Z.; Wang, L.; Wang, Q.; Li, W. Histone modifications and chromatin organization in prostate cancer. Epigenomics 2010, 2, 551–560. [Google Scholar] [CrossRef]
- Shin, S.; Janknecht, R. Activation of androgen receptor by histone demethylases JMJD2A and JMJD2D. Biochem. Biophys. Res. Commun. 2007, 359, 742–746. [Google Scholar] [CrossRef]
- Chang, P.C.; Fitzgerald, L.D.; Hsia, D.A.; Izumiya, Y.; Wu, C.Y.; Hsieh, W.P.; Lin, S.F.; Campbell, M.; Lam, K.S.; Luciw, P.A.; et al. Histone demethylase JMJD2A regulates Kaposi's sarcoma-associated herpesvirus replication and is targeted by a viral transcriptional factor. J. Virol. 2011, 85, 3283–3293. [Google Scholar]
- Li, B.X.; Zhang, M.C.; Luo, C.L.; Yang, P.; Li, H.; Xu, H.M.; Xu, H.F.; Shen, Y.W.; Xue, A.M.; Zhao, Z.Q. Effects of RNA interference-mediated gene silencing of JMJD2A on human breast cancer cell line MDA-MB-231 in vitro. J. Exp. Clin. Cancer Res. 2011, 30, 90. [Google Scholar] [CrossRef]
- Kim, T.D.; Shin, S.; Berry, W.L.; Oh, S.; Janknecht, R. The JMJD2A demethylase regulates apoptosis and proliferation in colon cancer cells. J. Cell. Biochem. 2011, 113, 368–1376. [Google Scholar]
- Anderton, J.A.; Bose, S.; Vockerodt, M.; Vrzalikova, K.; Wei, W.; Kuo, M.; Helin, K.; Christensen, J.; Rowe, M.; Murray, P.G.; et al. The H3K27me3 demethylase, KDM6B, is induced by Epstein-Barr virus and over-expressed in Hodgkin's Lymphoma. Oncogene 2011, 30, 2037–2043. [Google Scholar]
- Martinelli, P.; Bonetti, P.; Sironi, C.; Pruneri, G.; Fumagalli, C.; Raviele, P.R.; Volorio, S.; Pileri, S.; Chiarle, R.; McDuff, F.K.; et al. The lymphoma-associated NPM-ALK oncogene elicits a p16INK4a/pRb-dependent tumor-suppressive pathway. Blood 2011, 117, 6617–6626. [Google Scholar]
- Pereira, F.; Barbachano, A.; Silva, J.; Bonilla, F.; Campbell, M.J.; Munoz, A.; Larriba, M.J. KDM6B/JMJD3 histone demethylase is induced by vitamin D and modulates its effects in colon cancer cells. Hum. Mol. Genet. 2011, 20, 4655–4665. [Google Scholar]
- Dalgliesh, G.L.; Furge, K.; Greenman, C.; Chen, L.; Bignell, G.; Butler, A.; Davies, H.; Edkins, S.; Hardy, C.; Latimer, C.; et al. Systematic sequencing of renal carcinoma reveals inactivation of histone modifying genes. Nature 2010, 463, 360–363. [Google Scholar]
- Gui, Y.; Guo, G.; Huang, Y.; Hu, X.; Tang, A.; Gao, S.; Wu, R.; Chen, C.; Li, X.; Zhou, L.; et al. Frequent mutations of chromatin remodeling genes in transitional cell carcinoma of the bladder. Nat. Genet. 2011, 43, 875–878. [Google Scholar]
- Mar, B.G.; Bullinger, L.; Basu, E.; Schlis, K.; Silverman, L.B.; Dohner, K.; Armstrong, S.A. Sequencing histone-modifying enzymes identifies UTX mutations in acute lymphoblastic leukemia. Leukemia 2012, 26, 1881–1883. [Google Scholar]
- McDevitt, M.A. Clinical applications of epigenetic markers and epigenetic profiling in myeloid malignancies. Semin. Oncol. 2012, 39, 109–122. [Google Scholar]
- Muramatsu, H.; Makishima, H.; Maciejewski, J.P. Chronic myelomonocytic leukemia and atypical chronic myeloid leukemia: Novel pathogenetic lesions. Semin. Oncol. 2012, 39, 67–73. [Google Scholar]
- Van Haaften, G.; Dalgliesh, G.L.; Davies, H.; Chen, L.; Bignell, G.; Greenman, C.; Edkins, S.; Hardy, C.; O’Meara, S.; Teague, J.; et al. Somatic mutations of the histone H3K27 demethylase gene UTX in human cancer. Nat. Genet. 2009, 41, 521–523. [Google Scholar]
- Shi, Y.J.; Matson, C.; Lan, F.; Iwase, S.; Baba, T.; Shi, Y. Regulation of LSD1 histone demethylase activity by its associated factors. Mol. Cell 2005, 19, 857–864. [Google Scholar]
- Yang, M.; Gocke, C.B.; Luo, X.; Borek, D.; Tomchick, D.R.; Machius, M.; Otwinowski, Z.; Yu, H. Structural basis for CoREST-dependent demethylation of nucleosomes by the human LSD1 histone demethylase. Mol. Cell 2006, 23, 377–387. [Google Scholar]
- Baron, R.; Binda, C.; Tortorici, M.; McCammon, J.A.; Mattevi, A. Molecular mimicry and ligand recognition in binding and catalysis by the histone demethylase LSD1-CoREST complex. Structure 2011, 19, 212–220. [Google Scholar]
- Adamo, A.; Sese, B.; Boue, S.; Castano, J.; Paramonov, I.; Barrero, M.J.; Izpisua Belmonte, J.C. LSD1 regulates the balance between self-renewal and differentiation in human embryonic stem cells. Nat. Cell Biol. 2011, 13, 652–659. [Google Scholar]
- Nicholson, T.B.; Su, H.; Hevi, S.; Wang, J.; Bajko, J.; Li, M.; Valdez, R.; Loureiro, J.; Cheng, X.; Li, E.; et al. Defective heart development in hypomorphic LSD1 mice. Cell Res. 2011. [Google Scholar]
- Sun, G.; Ye, P.; Murai, K.; Lang, M.F.; Li, S.; Zhang, H.; Li, W.; Fu, C.; Yin, J.; Wang, A.; et al. miR-137 Forms a regulatory loop with nuclear receptor TLX and LSD1 in neural stem cells. Nat. Commun. 2011, 2, 529. [Google Scholar] [CrossRef]
- Choi, J.; Jang, H.; Kim, H.; Kim, S.T.; Cho, E.J.; Youn, H.D. Histone demethylase LSD1 is required to induce skeletal muscle differentiation by regulating myogenic factors. Biochem. Biophys. Res. Commun. 2010, 401, 327–332. [Google Scholar]
- Hino, S.; Sakamoto, A.; Nagaoka, K.; Anan, K.; Wang, Y.; Mimasu, S.; Umehara, T.; Yokoyama, S.; Kosai, K.; Nakao, M. FAD-dependent lysine-specific demethylase-1 regulates cellular energy expenditure. Nat. Commun. 2012, 3, 758. [Google Scholar]
- Janzer, A.; Lim, S.; Fronhoffs, F.; Niazy, N.; Buettner, R.; Kirfel, J. Lysine-specific demethylase 1 (LSD1) and histone deacetylase 1 (HDAC1) synergistically repress proinflammatory cytokines and classical complement pathway components. Biochem. Biophys. Res. Commun. 2012, 421, 665–670. [Google Scholar]
- Sprussel, A.; Schulte, J.H.; Weber, S.; Necke, M.; Handschke, K.; Thor, T.; Pajtler, K.W.; Schramm, A.; Konig, K.; Diehl, L.; et al. Lysine-specific demethylase 1 restricts hematopoietic progenitor proliferation and is essential for terminal differentiation. Leukemia 2012. [Google Scholar]
- Stavropoulos, P.; Blobel, G.; Hoelz, A. Crystal structure and mechanism of human lysine-specific demethylase-1. Nat. Struct. Mol. Biol. 2006, 13, 626–632. [Google Scholar]
- Mimasu, S.; Umezawa, N.; Sato, S.; Higuchi, T.; Umehara, T.; Yokoyama, S. Structurally designed trans-2-phenylcyclopropylamine derivatives potently inhibit histone demethylase LSD1/KDM1. Biochemistry 2010, 49, 6494–6503. [Google Scholar]
- Huang, Y.; Marton, L.J.; Woster, P.M.; Casero, R.A. Polyamine analogues targeting epigenetic gene regulation. Essays Biochem. 2009, 46, 95–110. [Google Scholar]
- Yu, V.; Fisch, T.; Long, A.M.; Tang, J.; Lee, J.H.; Hierl, M.; Chen, H.; Yakowec, P.; Schwandner, R.; Emkey, R. High-throughput TR-FRET assays for identifying inhibitors of LSD1 and JMJD2C histone lysine demethylases. J. Biomol. Screen. 2012, 17, 27–38. [Google Scholar]
- Yang, M.; Culhane, J.C.; Szewczuk, L.M.; Gocke, C.B.; Brautigam, C.A.; Tomchick, D.R.; Machius, M.; Cole, P.A.; Yu, H. Structural basis of histone demethylation by LSD1 revealed by suicide inactivation. Nat. Struct. Mol. Biol. 2007, 14, 535–539. [Google Scholar]
- Chen, Y.; Jie, W.; Yan, W.; Zhou, K.; Xiao, Y. Lysine-specific histone demethylase 1 (LSD1): A potential molecular target for tumor therapy. Crit. Rev. Eukaryot. Gene Expr. 2012, 22, 53–59. [Google Scholar]
- Karytinos, A.; Forneris, F.; Profumo, A.; Ciossani, G.; Battaglioli, E.; Binda, C.; Mattevi, A. A novel mammalian flavin-dependent histone demethylase. Biol. Chem. 2009, 284, 17775–17782. [Google Scholar]
- Fang, R.; Barbera, A.J.; Xu, Y.; Rutenberg, M.; Leonor, T.; Bi, Q.; Lan, F.; Mei, P.; Yuan, G.C.; Lian, C.; et al. Human LSD2/KDM1b/AOF1 regulates gene transcription by modulating intragenic H3K4me2 methylation. Mol. Cell 2010, 39, 222–233. [Google Scholar] [CrossRef]
- Van Essen, D.; Zhu, Y.; Saccani, S. A feed-forward circuit controlling inducible NF-kappaB target gene activation by promoter histone demethylation. Mol. Cell 2010, 39, 750–760. [Google Scholar]
- Ciccone, D.N.; Su, H.; Hevi, S.; Gay, F.; Lei, H.; Bajko, J.; Xu, G.; Li, E.; Chen, T. KDM1B is a histone H3K4 demethylase required to establish maternal genomic imprints. Nature 2009, 461, 415–418. [Google Scholar]
- Lin, S.L. Concise review: Deciphering the mechanism behind induced pluripotent stem cell generation. Stem Cells 2011, 29, 1645–1649. [Google Scholar]
- Binda, C.; Valente, S.; Romanenghi, M.; Pilotto, S.; Cirilli, R.; Karytinos, A.; Ciossani, G.; Botrugno, O.A.; Forneris, F.; Tardugno, M.; et al. Biochemical, structural, and biological evaluation of tranylcypromine derivatives as inhibitors of histone demethylases LSD1 and LSD2. J. Am. Chem. Soc. 2010, 132, 6827–6833. [Google Scholar]
- Alvarez-Saavedra, M.; Antoun, G.; Yanagiya, A.; Oliva-Hernandez, R.; Cornejo-Palma, D.; Perez-Iratxeta, C.; Sonenberg, N.; Cheng, H.Y. miRNA-132 Orchestrates chromatin remodeling and translational control of the circadian clock. Hum. Mol. Genet. 2011, 20, 731–751. [Google Scholar]
- DiTacchio, L.; Le, H.D.; Vollmers, C.; Hatori, M.; Witcher, M.; Secombe, J.; Panda, S. Histone lysine demethylase JARID1a activates CLOCK-BMAL1 and influences the circadian clock. Science 2011, 333, 1881–1885. [Google Scholar]
- Christensen, J.; Agger, K.; Cloos, P.A.; Pasini, D.; Rose, S.; Sennels, L.; Rappsilber, J.; Hansen, K.H.; Salcini, A.E.; Helin, K. RBP2 belongs to a family of demethylases, specific for tri-and dimethylated lysine 4 on histone 3. Cell 2007, 128, 1063–1076. [Google Scholar]
- Lee, C.H.; Alpert, B.O.; Sankaranarayanan, P.; Alter, O. GSVD comparison of patient-matched normal and tumor aCGH profiles reveals global copy-number alterations predicting glioblastoma multiforme survival. PLoS One 2012, 7, e30098. [Google Scholar]
- Sharma, S.V.; Lee, D.Y.; Li, B.; Quinlan, M.P.; Takahashi, F.; Maheswaran, S.; McDermott, U.; Azizian, N.; Zou, L.; Fischbach, M.A.; et al. A chromatin-mediated reversible drug-tolerant state in cancer cell subpopulations. Cell 2010, 141, 69–80. [Google Scholar] [CrossRef]
- Frank, S.A.; Rosner, M.R. Nonheritable cellular variability accelerates the evolutionary processes of cancer. PLoS Biol. 2012, 10, e1001296. [Google Scholar]
- Pointon, J.J.; Harvey, D.; Karaderi, T.; Appleton, L.H.; Farrar, C.; Wordsworth, B.P. The histone demethylase JARID1A is associated with susceptibility to ankylosing spondylitis. Genes Immun. 2011, 12, 395–398. [Google Scholar]
- Zhao, M.; Liang, G.; Wu, X.; Wang, S.; Zhang, P.; Su, Y.; Yin, H.; Tan, Y.; Zhang, J.; Lu, Q. Abnormal epigenetic modifications in peripheral blood mononuclear cells from patients with alopecia areata. Br. J. Dermatol. 2012, 166, 226–273. [Google Scholar]
- Liefke, R.; Oswald, F.; Alvarado, C.; Ferres-Marco, D.; Mittler, G.; Rodriguez, P.; Dominguez, M.; Borggrefe, T. Histone demethylase KDM5A is an integral part of the core Notch-RBP-J repressor complex. Genes Dev. 2010, 24, 590–601. [Google Scholar]
- Chicas, A.; Kapoor, A.; Wang, X.; Aksoy, O.; Evertts, A.G.; Zhang, M.Q.; Garcia, B.A.; Bernstein, E.; Lowe, S.W. H3K4 demethylation by Jarid1a and Jarid1b contributes to retinoblastoma-mediated gene silencing during cellular senescence. Proc. Natl. Acad. Sci. USA 2012, 109, 8971–8976. [Google Scholar]
- Dey, B.K.; Stalker, L.; Schnerch, A.; Bhatia, M.; Taylor-Papidimitriou, J.; Wynder, C. The histone demethylase KDM5b/JARID1b plays a role in cell fate decisions by blocking terminal differentiation. Mol. Cell. Biol. 2008, 28, 5312–5327. [Google Scholar]
- Schmitz, S.U.; Albert, M.; Malatesta, M.; Morey, L.; Johansen, J.V.; Bak, M.; Tommerup, N.; Abarrategui, I.; Helin, K. Jarid1b targets genes regulating development and is involved in neural differentiation. EMBO J. 2011, 30, 4586–4600. [Google Scholar]
- Scibetta, A.G.; Santangelo, S.; Coleman, J.; Hall, D.; Chaplin, T.; Copier, J.; Catchpole, S.; Burchell, J.; Taylor-Papadimitriou, J. Functional analysis of the transcription repressor PLU-1/JARID1B. Mol. Cell. Biol. 2007, 27, 7220–7235. [Google Scholar]
- Tarantino, C.; Paolella, G.; Cozzuto, L.; Minopoli, G.; Pastore, L.; Parisi, S.; Russo, T. miRNA 34a, 100, and 137 Modulate differentiation of mouse embryonic stem cells. FASEB J. 2010, 24, 3255–3263. [Google Scholar]
- Kristensen, L.H.; Nielsen, A.L.; Helgstrand, C.; Lees, M.; Cloos, P.; Kastrup, J.S.; Helin, K.; Olsen, L.; Gajhede, M. Studies of H3K4me3 demethylation by KDM5B/Jarid1B/PLU1 reveals strong substrate recognition in vitro and identifies 2,4-pyridine-dicarboxylic acid as an in vitro and in cell inhibitor. FEBS J. 2012, 279, 1905–1914. [Google Scholar]
- Akbarian, S.; Huang, H.S. Epigenetic regulation in human brain-focus on histone lysine methylation. Biol. Psychiatry 2009, 65, 198–203. [Google Scholar]
- Xu, J.; Andreassi, M. Reversible histone methylation regulates brain gene expression and behavior. Horm. Behav. 2011, 59, 383–392. [Google Scholar]
- Abidi, F.E.; Holloway, L.; Moore, C.A.; Weaver, D.D.; Simensen, R.J.; Stevenson, R.E.; Rogers, R.C.; Schwartz, C.E. Mutations in JARID1C are associated with X-linked mental retardation, short stature and hyperreflexia. J. Med. Genet. 2008, 45, 787–793. [Google Scholar]
- Jensen, L.R.; Bartenschlager, H.; Rujirabanjerd, S.; Tzschach, A.; Numann, A.; Janecke, A.R.; Sporle, R.; Stricker, S.; Raynaud, M.; Nelson, J.; et al. A distinctive gene expression fingerprint in mentally retarded male patients reflects disease-causing defects in the histone demethylase KDM5C. Pathogenetics 2010, 3, 2. [Google Scholar] [CrossRef]
- Santos-Reboucas, C.B.; Fintelman-Rodrigues, N.; Jensen, L.R.; Kuss, A.W.; Ribeiro, M.G.; Campos, M., Jr.; Santos, J.M.; Pimentel, M.M. A novel nonsense mutation in KDM5C/JARID1C gene causing intellectual disability, short stature and speech delay. Neurosci. Lett. 2011, 498, 67–71. [Google Scholar] [CrossRef]
- Adegbola, A.; Gao, H.; Sommer, S.; Browning, M. A novel mutation in JARID1C/SMCX in a patient with autism spectrum disorder (ASD). Am. J. Med. Genet. A 2008, 146A, 505–511. [Google Scholar]
- Upadhyay, A.K.; Rotili, D.; Han, J.W.; Hu, R.; Chang, Y.; Labella, D.; Zhang, X.; Yoon, Y.S.; Mai, A.; Cheng, X. An analog of BIX-01294 selectively inhibits a family of histone H3 lysine 9 Jumonji demethylases. J. Mol. Biol. 2012, 416, 319–327. [Google Scholar]
- He, J.; Kallin, E.M.; Tsukada, Y.; Zhang, Y. The H3K36 demethylase Jhdm1b/Kdm2b regulates cell proliferation and senescence through p15(Ink4b). Nat. Struct. Mol. Biol. 2008, 15, 1169–1175. [Google Scholar]
- Tzatsos, A.; Paskaleva, P.; Lymperi, S.; Contino, G.; Stoykova, S.; Chen, Z.; Wong, K.K.; Bardeesy, N. Lysine-specific demethylase 2B (KDM2B)-let-7-enhancer of zester homolog 2 (EZH2) pathway regulates cell cycle progression and senescence in primary cells. Biol. Chem. 2011, 286, 33061–33069. [Google Scholar]
- Wang, T.; Chen, K.; Zeng, X.; Yang, J.; Wu, Y.; Shi, X.; Qin, B.; Zeng, L.; Esteban, M.A.; Pan, G.; et al. The histone demethylases Jhdm1a/1b enhance somatic cell reprogramming in a vitamin-C-dependent manner. Cell Stem Cell 2011, 9, 575–587. [Google Scholar] [CrossRef]
- Bredel, M.; Bredel, C.; Juric, D.; Harsh, G.R.; Vogel, H.; Recht, L.D.; Sikic, B.I. Functional network analysis reveals extended gliomagenesis pathway maps and three novel MYC-interacting genes in human gliomas. Cancer Res. 2005, 65, 8679–8689. [Google Scholar]
- Freije, W.A.; Castro-Vargas, F.E.; Fang, Z.; Horvath, S.; Cloughesy, T.; Liau, L.M.; Mischel, P.S.; Nelson, S.F. Gene expression profiling of gliomas strongly predicts survival. Cancer Res. 2004, 64, 6503–6510. [Google Scholar]
- Lizcano, F.; Romero, C.; Vargas, D. Regulation of adipogenesis by nuclear receptor PPARgamma is modulated by the histone demethylase JMJD2C. Genet. Mol. Biol. 2011, 34, 19–24. [Google Scholar]
- Pollard, P.J.; Loenarz, C.; Mole, D.R.; McDonough, M.A.; Gleadle, J.M.; Schofield, C.J.; Ratcliffe, P.J. Regulation of Jumonji-domain-containing histone demethylases by hypoxia-inducible factor (HIF)-1alpha. Biochem. J. 2008, 416, 387–394. [Google Scholar]
- Loh, Y.H.; Zhang, W.; Chen, X.; George, J.; Ng, H.H. Jmjd1a and Jmjd2c histone H3 Lys 9 demethylases regulate self-renewal in embryonic stem cells. Genes Dev. 2007, 21, 2545–2557. [Google Scholar]
- Katoh, Y.; Katoh, M. Comparative integromics on JMJD2A, JMJD2B and JMJD2C: Preferential expression of JMJD2C in undifferentiated ES cells. Int. J. Mol. Med. 2007, 20, 269–273. [Google Scholar]
- Kantojarvi, K.; Onkamo, P.; Vanhala, R.; Alen, R.; Hedman, M.; Sajantila, A.; Nieminen-von Wendt, T.; Jarvela, I. Analysis of 9p24 and 11p12–13 regions in autism spectrum disorders: rs1340513 In the JMJD2C gene is associated with ASDs in Finnish sample. Psychiatr. Genet. 2010, 20, 102–108. [Google Scholar]
- Ponnaluri, V.K.; Vavilala, D.T.; Mukherji, M. Studies on substrate specificity of Jmjd2a-c histone demethylases. Biochem. Biophys. Res. Commun. 2011, 405, 588–592. [Google Scholar]
- Gray, S.G.; Iglesias, A.H.; Lizcano, F.; Villanueva, R.; Camelo, S.; Jingu, H.; Teh, B.T.; Koibuchi, N.; Chin, W.W.; Kokkotou, E.; et al. Functional characterization of JMJD2A, a histone deacetylase- and retinoblastoma-binding protein. Biol. Chem. 2005, 280, 28507–28518. [Google Scholar] [CrossRef]
- Klose, R.J.; Yamane, K.; Bae, Y.; Zhang, D.; Erdjument-Bromage, H.; Tempst, P.; Wong, J.; Zhang, Y. The transcriptional repressor JHDM3A demethylates trimethyl histone H3 lysine 9 and lysine 36. Nature 2006, 442, 312–316. [Google Scholar]
- Verrier, L.; Escaffit, F.; Chailleux, C.; Trouche, D.; Vandromme, M. A new isoform of the histone demethylase JMJD2A/KDM4A is required for skeletal muscle differentiation. PLoS Genet. 2011, 7, e1001390. [Google Scholar]
- Mallette, F.A.; Mattiroli, F.; Cui, G.; Young, L.C.; Hendzel, M.J.; Mer, G.; Sixma, T.K.; Richard, S. RNF8- and RNF168-dependent degradation of KDM4A/JMJD2A triggers 53BP1 recruitment to DNA damage sites. EMBO J. 2012, 31, 1865–1878. [Google Scholar]
- Zhang, Q.J.; Chen, H.Z.; Wang, L.; Liu, D.P.; Hill, J.A.; Liu, Z.P. The histone trimethyllysine demethylase JMJD2A promotes cardiac hypertrophy in response to hypertrophic stimuli in mice. J. Clin. Invest. 2011, 121, 2447–2456. [Google Scholar]
- Couture, J.F.; Collazo, E.; Ortiz-Tello, P.A.; Brunzelle, J.S.; Trievel, R.C. Specificity and mechanism of JMJD2A, a trimethyllysine-specific histone demethylase. Nat. Struct. Mol. Biol. 2007, 14, 689–695. [Google Scholar]
- Huang, Y.; Fang, J.; Bedford, M.T.; Zhang, Y.; Xu, R.M. Recognition of histone H3 lysine-4 methylation by the double tudor domain of JMJD2A. Science 2006, 312, 748–751. [Google Scholar]
- Ng, S.S.; Kavanagh, K.L.; McDonough, M.A.; Butler, D.; Pilka, E.S.; Lienard, B.M.; Bray, J.E.; Savitsky, P.; Gileadi, O.; von Delft, F.; et al. Crystal structures of histone demethylase JMJD2A reveal basis for substrate specificity. Nature 2007, 448, 87–91. [Google Scholar]
- zboyaci, M.; Gursoy, A.; Erman, B.; Keskin, O. Molecular recognition of H3/H4 histone tails by the tudor domains of JMJD2A: A comparative molecular dynamics simulations study. PLoS One 2011, 6, e14765. [Google Scholar]
- Sanli, D.; Keskin, O.; Gursoy, A.; Erman, B. Structural cooperativity in histone H3 tail modifications. Protein Sci. 2011, 20, 1982–1990. [Google Scholar]
- Rose, N.R.; Woon, E.C.; Kingham, G.L.; King, O.N.; Mecinovic, J.; Clifton, I.J.; Ng, S.S.; Talib-Hardy, J.; Oppermann, U.; McDonough, M.A.; et al. Selective inhibitors of the JMJD2 histone demethylases: Combined nondenaturing mass spectrometric screening and crystallographic approaches. J. Med. Chem. 2010, 53, 1810–1818. [Google Scholar]
- Hutchinson, S.E.; Leveridge, M.V.; Heathcote, M.L.; Francis, P.; Williams, L.; Gee, M.; Munoz-Muriedas, J.; Leavens, B.; Shillings, A.; Jones, E.; et al. Enabling lead discovery for histone lysine demethylases by high-throughput RapidFire mass spectrometry. J. Biomol. Screen. 2012, 17, 39–48. [Google Scholar] [CrossRef]
- Xiang, Y.; Zhu, Z.; Han, G.; Lin, H.; Xu, L.; Chen, C.D. JMJD3 is a histone H3K27 demethylase. Cell Res. 2007, 17, 850–857. [Google Scholar]
- Hong, S.; Cho, Y.W.; Yu, L.R.; Yu, H.; Veenstra, T.D.; Ge, K. Identification of JmjC domain-containing UTX and JMJD3 as histone H3 lysine 27 demethylases. Proc. Natl. Acad. Sci. USA 2007, 104, 18439–18444. [Google Scholar]
- Issaeva, I.; Zonis, Y.; Rozovskaia, T.; Orlovsky, K.; Croce, C.M.; Nakamura, T.; Mazo, A.; Eisenbach, L.; Canaani, E. Knockdown of ALR (MLL2) reveals ALR target genes and leads to alterations in cell adhesion and growth. Mol. Cell. Biol. 2007, 27, 1889–1903. [Google Scholar]
- Lee, M.G.; Villa, R.; Trojer, P.; Norman, J.; Yan, K.P.; Reinberg, D.; di Croce, L.; Shiekhattar, R. Demethylation of H3K27 regulates polycomb recruitment and H2A ubiquitination. Science 2007, 318, 447–450. [Google Scholar]
- Vernimmen, D.; Lynch, M.D.; de Gobbi, M.; Garrick, D.; Sharpe, J.A.; Sloane-Stanley, J.A.; Smith, A.J.; Higgs, D.R. Polycomb eviction as a new distant enhancer function. Genes Dev. 2011, 25, 1583–1588. [Google Scholar]
- Dahle, O.; Kumar, A.; Kuehn, M.R. Nodal signaling recruits the histone demethylase Jmjd3 to counteract polycomb-mediated repression at target genes. Sci. Signal. 2010, 3, ra48. [Google Scholar]
- Kim, S.W.; Yoon, S.J.; Chuong, E.; Oyolu, C.; Wills, A.E.; Gupta, R.; Baker, J. Chromatin and transcriptional signatures for Nodal signaling during endoderm formation in hESCs. Dev. Biol. 2011, 357, 492–504. [Google Scholar]
- Ohtani, K.; Vlachojannis, G.J.; Koyanagi, M.; Boeckel, J.N.; Urbich, C.; Farcas, R.; Bonig, H.; Marquez, V.E.; Zeiher, A.M.; Dimmeler, S. Epigenetic regulation of endothelial lineage committed genes in pro-angiogenic hematopoietic and endothelial progenitor cells. Circ. Res. 2011, 109, 1219–1229. [Google Scholar]
- Sola, S.; Xavier, J.M.; Santos, D.M.; Aranha, M.M.; Morgado, A.L.; Jepsen, K.; Rodrigues, C.M. p53 interaction with JMJD3 results in its nuclear distribution during mouse neural stem cell differentiation. PLoS One 2011, 6, e18421. [Google Scholar]
- Yasui, T.; Hirose, J.; Tsutsumi, S.; Nakamura, K.; Aburatani, H.; Tanaka, S. Epigenetic regulation of osteoclast differentiation: Possible involvement of Jmjd3 in the histone demethylation of Nfatc1. J. Bone Miner. Res. 2011, 26, 2665–2671. [Google Scholar]
- Satoh, T.; Takeuchi, O.; Vandenbon, A.; Yasuda, K.; Tanaka, Y.; Kumagai, Y.; Miyake, T.; Matsushita, K.; Okazaki, T.; Saitoh, T.; et al. The Jmjd3-Irf4 axis regulates M2 macrophage polarization and host responses against helminth infection. Nat. Immunol. 2010, 11, 936–944. [Google Scholar] [CrossRef]
- Zhang, Q.; Long, H.; Liao, J.; Zhao, M.; Liang, G.; Wu, X.; Zhang, P.; Ding, S.; Luo, S.; Lu, Q. Inhibited expression of hematopoietic progenitor kinase 1 associated with loss of jumonji domain containing 3 promoter binding contributes to autoimmunity in systemic lupus erythematosus. J. Autoimmun. 2011, 37, 180–189. [Google Scholar]
- Ciavatta, D.J.; Yang, J.; Preston, G.A.; Badhwar, A.K.; Xiao, H.; Hewins, P.; Nester, C.M.; Pendergraft, W.F., 3rd; Magnuson, T.R.; Jennette, J.C.; et al. Epigenetic basis for aberrant upregulation of autoantigen genes in humans with ANCA vasculitis. J. Clin. Invest. 2010, 120, 3209–3219. [Google Scholar]
- Lin, T.Y.; Cheng, Y.C.; Yang, H.C.; Lin, W.C.; Wang, C.C.; Lai, P.L.; Shieh, S.Y. Loss of the candidate tumor suppressor BTG3 triggers acute cellular senescence via the ERK-JMJD3-p16INK4a signaling axis. Oncogene 2011, 31, 3287–3297. [Google Scholar]
- Svotelis, A.; Bianco, S.; Madore, J.; Huppe, G.; Nordell-Markovits, A.; Mes-Masson, A.M.; Gevry, N. H3K27 demethylation by JMJD3 at a poised enhancer of anti-apoptotic gene BCL2 determines ERalpha ligand dependency. EMBO J. 2011, 30, 3947–3961. [Google Scholar]
- Mulji, A.; Haslam, C.; Brown, F.; Randle, R.; Karamshi, B.; Smith, J.; Eagle, R.; Munoz-Muriedas, J.; Taylor, J.; Sheikh, A.; Bridges, A.; et al. Configuration of a high-content imaging platform for hit identification and pharmacological assessment of JMJD3 demethylase enzyme inhibitors. J. Biomol. Screen. 2012, 17, 108–120. [Google Scholar] [CrossRef]
- Sengoku, T.; Yokoyama, S. Structural basis for histone H3 Lys 27 demethylation by UTX/KDM6A. Genes Dev. 2011, 25, 2266–2277. [Google Scholar]
- Lee, S.; Lee, J.W.; Lee, S.K. UTX, a histone H3-lysine 27 demethylase, acts as a critical switch to activate the cardiac developmental program. Dev. Cell 2012, 22, 25–37. [Google Scholar] [CrossRef]
- Yap, D.B.; Walker, D.C.; Prentice, L.M.; McKinney, S.; Turashvili, G.; Mooslehner-Allen, K.; de Algara, T.R.; Fee, J.; de Tassigny, X.; Colledge, W.H.; et al. Mll5 is required for normal spermatogenesis. PLoS One 2011, 6, e27127. [Google Scholar]
- Liu, J.; Mercher, T.; Scholl, C.; Brumme, K.; Gilliland, D.G.; Zhu, N. A functional role for the histone demethylase UTX in normal and malignant hematopoietic cells. Exp. Hematol. 2012, 40, 487–498. [Google Scholar]
- Seenundun, S.; Rampalli, S.; Liu, Q.C.; Aziz, A.; Palii, C.; Hong, S.; Blais, A.; Brand, M.; Ge, K.; Dilworth, F.J. UTX mediates demethylation of H3K27me3 at muscle-specific genes during myogenesis. EMBO J. 2010, 29, 1401–1411. [Google Scholar]
- Wang, J.K.; Tsai, M.C.; Poulin, G.; Adler, A.S.; Chen, S.; Liu, H.; Shi, Y.; Chang, H.Y. The histone demethylase UTX enables RB-dependent cell fate control. Genes Dev. 2010, 24, 327–332. [Google Scholar]
- Terashima, M.; Ishimura, A.; Yoshida, M.; Suzuki, Y.; Sugano, S.; Suzuki, T. The tumor suppressor Rb and its related Rbl2 genes are regulated by Utx histone demethylase. Biochem. Biophys. Res. Commun. 2010, 399, 238–244. [Google Scholar]
- Tsai, M.C.; Wang, J.K.; Chang, H.Y. Tumor suppression by the histone demethylase UTX. Cell Cycle 2010, 9, 2043–2044. [Google Scholar]
- Verbrugge, I.; Johnstone, R.W.; Bots, M. Promises and challenges of anticancer drugs that target the epigenome. Epigenomics 2011, 3, 547–565. [Google Scholar]
- Wu, Y.; Steinbergs, N.; Murray-Stewart, T.; Marton, L.J.; Casero, R.A. Oligoamine analogues in combination with 2-difluoromethylornithine synergistically induce re-expression of aberrantly silenced tumour-suppressor genes. Biochem. J. 2012, 442, 693–701. [Google Scholar]
- Gooden, D.M.; Schmidt, D.M.; Pollock, J.A.; Kabadi, A.M.; McCafferty, D.G. Facile synthesis of substituted trans-2-arylcyclopropylamine inhibitors of the human histone demethylase LSD1 and monoamine oxidases A and B. Bioorg. Med. Chem. Lett. 2008, 18, 3047–3051. [Google Scholar]
- Ueda, R.; Suzuki, T.; Mino, K.; Tsumoto, H.; Nakagawa, H.; Hasegawa, M.; Sasaki, R.; Mizukami, T.; Miyata, N. Identification of cell-active lysine specific demethylase 1-selective inhibitors. J. Am. Chem. Soc. 2009, 131, 17536–17537. [Google Scholar]
- Willmann, D.; Lim, S.; Wetzel, S.; Metzger, E.; Jandausch, A.; Wilk, W.; Jung, M.; Forne, I.; Imhof, A.; Janzer, A.; et al. Impairment of prostate cancer cell growth by a selective and reversible LSD1 inhibitor. Int. J. Cancer 2012. [Google Scholar]
- Ortega-Muñoz, A.; Castro-Palomino Laria, J.; Fyfe, M.C.T. Lysine Specific Demethylase-1 Inhibitors and Their Use. WO 2011035941A1, 31 March 2011. [Google Scholar]
- Lokken, A.A.; Zeleznik-Le, N.J. Breaking the LSD1/KDM1A addiction: Therapeutic targeting of the epigenetic modifier in AML. Cancer Cell 2012, 21, 451–453. [Google Scholar]
- Schenk, T.; Chen, W.C.; Gollner, S.; Howell, L.; Jin, L.; Hebestreit, K.; Klein, H.U.; Popescu, A.C.; Burnett, A.; Mills, K.; et al. Inhibition of the LSD1 (KDM1A) demethylase reactivates the all-trans-retinoic acid differentiation pathway in acute myeloid leukemia. Nat. Med. 2012, 18, 605–611. [Google Scholar]
- McDonough, M.A.; McNeill, L.A.; Tilliet, M.; Papamicael, C.A.; Chen, Q.Y.; Banerji, B.; Hewitson, K.S.; Schofield, C.J. Selective inhibition of factor inhibiting hypoxia-inducible factor. J. Am. Chem. Soc. 2005, 127, 7680–7681. [Google Scholar]
- Sakurai, M.; Rose, N.R.; Schultz, L.; Quinn, A.M.; Jadhav, A.; Ng, S.S.; Oppermann, U.; Schofield, C.J.; Simeonov, A. A miniaturized screen for inhibitors of Jumonji histone demethylases. Mol. Biosyst. 2010, 6, 357–364. [Google Scholar]
- Chang, K.H.; King, O.N.; Tumber, A.; Woon, E.C.; Heightman, T.D.; McDonough, M.A.; Schofield, C.J.; Rose, N.R. Inhibition of histone demethylases by 4-carboxy-2,2'-bipyridyl compounds. ChemMedChem 2011, 6, 759–764. [Google Scholar]
- King, O.N.; Li, X.S.; Sakurai, M.; Kawamura, A.; Rose, N.R.; Ng, S.S.; Quinn, A.M.; Rai, G.; Mott, B.T.; Beswick, P.; et al. Quantitative high-throughput screening identifies 8-hydroxyquinolines as cell-active histone demethylase inhibitors. PLoS One 2010, 5, e15535. [Google Scholar]
- Ulucan, O.; Keskin, O.; Erman, B.; Gursoy, A. A comparative molecular dynamics study of methylation state specificity of JMJD2A. PLoS One 2011, 6, e24664. [Google Scholar]
- Kong, X.; Ouyang, S.; Liang, Z.; Lu, J.; Chen, L.; Shen, B.; Li, D.; Zheng, M.; Li, K.K.; Luo, C.; et al. Catalytic mechanism investigation of lysine-specific demethylase 1 (LSD1): A computational study. PLoS One 2011, 6, e25444. [Google Scholar]
- Szewczuk, L.M.; Culhane, J.C.; Yang, M.; Majumdar, A.; Yu, H.; Cole, P.A. Mechanistic analysis of a suicide inactivator of histone demethylase LSD1. Biochemistry 2007, 46, 6892–6902. [Google Scholar]
- Culhane, J.C.; Wang, D.; Yen, P.M.; Cole, P.A. Comparative analysis of small molecules and histone substrate analogues as LSD1 lysine demethylase inhibitors. J. Am. Chem. Soc. 2010, 132, 3164–3176. [Google Scholar]
© 2012 by the authors; licensee MDPI, Basel, Switzerland. This article is an open-access article distributed under the terms and conditions of the Creative Commons Attribution license (http://creativecommons.org/licenses/by/3.0/).
Share and Cite
Lizcano, F.; Garcia, J. Epigenetic Control and Cancer: The Potential of Histone Demethylases as Therapeutic Targets. Pharmaceuticals 2012, 5, 963-990. https://doi.org/10.3390/ph5090963
Lizcano F, Garcia J. Epigenetic Control and Cancer: The Potential of Histone Demethylases as Therapeutic Targets. Pharmaceuticals. 2012; 5(9):963-990. https://doi.org/10.3390/ph5090963
Chicago/Turabian StyleLizcano, Fernando, and Jeison Garcia. 2012. "Epigenetic Control and Cancer: The Potential of Histone Demethylases as Therapeutic Targets" Pharmaceuticals 5, no. 9: 963-990. https://doi.org/10.3390/ph5090963
APA StyleLizcano, F., & Garcia, J. (2012). Epigenetic Control and Cancer: The Potential of Histone Demethylases as Therapeutic Targets. Pharmaceuticals, 5(9), 963-990. https://doi.org/10.3390/ph5090963




