Non-invasive Optical Biosensor for Probing Cell Signaling
Abstract
:1. Introduction
2. Optical Biosensors
3. Theories for Living Cell Sensing with Optical Biosensor
4. High Throughput Optical Biosensor Systems
5. Probing Cell Signaling with Optical Biosensor
5.1 Cell-surface interactions
5.2 Cells under un-stimulated conditions
5.3 Cellular responses upon stimulation
5.4 Characteristics of endogenous GPCR DMR signals
5.5 Cellular mechanisms of GPCR ligand-induced DMR signals
5.6 Applications of optical biosensors in cell signaling
6. Conclusion
References
- Leatherbarrow, R.J.; Edwards, P.R. Analysis of molecular recognition using optical biosensors. Curr. Opin. Chem. Biol. 1999, 3, 544–547. [Google Scholar]
- Cooper, M.A. Optical biosensors in drug discovery. Nature Rev. Drug Discov. 2002, 1, 515–528. [Google Scholar]
- Ramsden, J.J.; Li, S.-Y.; Prenosil, J.E.; Heinzle, E. Kinetics of adhesion and spreading of animan cells. Biotechnol. Bioeng. 1994, 43, 939–945. [Google Scholar]
- Giebel, K.-F.; Bechinger, C.; Herminghaus, S.; Riedel, M.; Leiderer, P.; Weiland, U.; Bastmeyer, M. Imaging of cell/substrate contacts of living cells with surface plasmon resonance microscopy. Biophys. J. 1999, 76, 509–516. [Google Scholar]
- Horath, R.; Pedersen, H.C.; Skivesen, N.; Selmeczi, D.; Larsen, N.B. Monitoring of living cell attachment and spreading using reverse symmetry waveguide sensing. Appl. Phys. Lett. 2005, 86, e071101. [Google Scholar]
- Quinn, J.G.; O'Neill, S.; Doyle, A.; McAtamney, C.; Diamond, D.; MacCraith, B.D.; O'Kennedy, R. Development and application of surface plasmon resonance-based biosensors for the detection of cell-ligand interactions. Anal. Biochem. 2000, 281, 135–143. [Google Scholar]
- Hide, M.; Tsutsui, T.; Sato, H.; Nishimura, T.; Morimoto, K.; Yamamoto, S.; Yoshizato, K. Real-time analysis of ligand-induced cell surface and intracellular reactions of living mast cells using a surface plasmon resonance-based biosensor. Anal. Biochem. 2002, 302, 28–37. [Google Scholar]
- Voros, J.; Graf, R.; Kenausis, G.L.; Bruinink, A.; Mayer, J.; Textor, M.; Wintermantel, E.; Spencer, N.D. Feasibility study of an online toxicological sensor based on the optical waveguide technique. Biosensors & Bioelectronics 2000, 15, 423–429. [Google Scholar]
- Fang, Y.; Ferrie, A.M.; Fontaine, N.H.; Yuen, P.K. Characteristics of dynamic mass redistribution of EGF receptor signaling in living cells measured with label free optical biosensors. Anal. Chem. 2005, 77, 5720–5725. [Google Scholar]
- Fang, Y.; Li, G.; Peng, J. Optical biosensor provides insights for bradykinin B2 receptor signaling in A431 cells. FEBS Lett. 2005, 579, 6365–6374. [Google Scholar]
- Fang, Y.; Ferrie, A.M.; Li, G. Probing cytoskeleton modulation by optical biosensors. FEBS Lett. 2005, 579, 4175–4180. [Google Scholar]
- Fang, Y.; Ferrie, A.M.; Li, G.; Fontaine, N.H. Cellular functions of cholesterol probed with optical biosensors”. Biochim. Biophys. Acta 2006, 1763, 254–261. [Google Scholar]
- Fang, Y.; Ferrie, A.M.; Fontaine, N.H.; Mauro, J.; Balakrishnan, J. Resonant waveguide grating biosensor for living cell sensing. Biophys. J. 2006, 91, 1925–1940. [Google Scholar]
- Li, G.; Ferrie, A.M.; Fang, Y. Label-free profiling of endogenous G protein-coupled receptors using a cell-based high throughput screening technology. J. Assoc. Lab. Automat. 2006, 11, 181–187. [Google Scholar]
- Fang, Y.; Li, G.; Ferrie, A.M. Non-invasive optical biosensor for assaying endogenous G protein-coupled receptors in adherent cells. J. Pharmacol. Toxicol. Methods 2007, 55, 314–322. [Google Scholar]
- Fang, Y.; Ferrie, A.M. Optical biosensor differentiates signaling of endogenous PAR1 and PAR2 in A431 cells. BMC Cell Biol. 2007, 8, 24:1–24:12. [Google Scholar]
- Fang, Y.; Ferrie, A.M.; Lahiri, J. Systems biology and pharmacology of β2 adrenergic receptors in A431. In Trends in Signal Transduction Research; Meyers, J.N., Ed.; Nova Science Publishers Inc.: New York, 2007; pp. 145–171. [Google Scholar]
- Fang, Y. Label-free cell-based assays with optical biosensors in drug discovery. Assays and Drug Development Technologies 2006, 4, 583–595. [Google Scholar]
- Cooper, M.A. Optical biosensors: where next and how soon. Drug Discov. Today 2006, 11, 1061–1067. [Google Scholar]
- Rich, R.L.; Myszka, D.G. Higher-throughput, label-free, real-time molecular interaction analysis. Anal. Biochem. 2007, 361, 1–6. [Google Scholar]
- Tiefenthaler, K.; Lukosz, W. Sensitivity of grating couplers as integrated-optical chemical sensors. J. Opt. Soc. Am. B 1989, 6, 209–220. [Google Scholar]
- Barer, R.; Joseph, S. Refractometry of living cells. Part I. basic principles. Quart. J. Microsc. Science 1954, 95, 399–423. [Google Scholar]
- De Feijter, J.A.; Benjamins, J.; Veer, F.A. Ellipsometry as a tool to study the adsorption behavior of synthetic and biopolymers at the air-water interface. Biopolymers 1978, 17, 1759–1772. [Google Scholar]
- Brandenburg, A.; Krauter, R.; Kunzel, C.; Stefan, M.; Schulte, H. Interferometric sensor for detection of surface-bound bioreactions. Appl. Optics 2000, 39, 6396–6405. [Google Scholar]
- Burridge, K.; Fath, K.; Kelly, T.; Nuckolls, G.; Turner, C. Focal adhesions: transmembrane junctions between the extracellular matrix and the cytoskeleton. Annu. Rev. Cell Biol. 1988, 4, 487–525. [Google Scholar]
- Chen, W.T.; Singer, S.J. Immunoelectron microscopic studies of the sites of cell-substratum and cell-cell contacts in cultured fibroblasts. J. Cell Boil. 1982, 95, 205–222. [Google Scholar]
- Discher, D.E.; Janmey, P.; Wang, Y. Tissue cells feel and respond to the stiffness of their substrate. Science 2005, 310, 1139–1143. [Google Scholar]
- Kato, M.; Mrksich, M. Rewiring cell adhesion. J. Am. Chem. Soc. 2006, 126, 6504–6505. [Google Scholar]
- Giaever, I.; Keese, C.R. Micromotion of mammalian cells measured electrically. Proc. Natl. Acad. Sci. USA 1991, 88, 7896–7900. [Google Scholar]
- Hudder, A.; Nathanson, L.; Deutscher, M.P. Organization of mammalian cytoplasm. Mol. Cell Biol. 2003, 23, 9318–9326. [Google Scholar]
- Sato, T.K.; Overduin, M.; Emr, S.D. Location, location, location: membrane targeting directed by PX domains. Science 2001, 294, 1881–1885. [Google Scholar]
- Kholodenko, B.N. Four-dimensional organization of protein kinase signaling cascades: the roles of diffusion, endocytosis and molecular motors. J. Exper. Biol. 2003, 206, 2073–2082. [Google Scholar]
- Aldridge, B.B.; Burke, J.M.; Lauffenburger, D.A.; Sorger, P.K. Physicochemical modelling of cell signalling pathways. Nat. Cell Biol. 2006, 8, 1195–1203. [Google Scholar]
- Wiley, H.S. Trafficking of the ErbB receptors and its influence on signaling. Exper. Cell Res. 2003, 284, 78–88. [Google Scholar]
- Schnapp, B.J. Trafficking of signaling modules by kinesin motors. J. Cell Sci. 2003, 116, 2125–2135. [Google Scholar]
- Lefkowitz, R.J.; Whalen, E.J. β-arrestins: traffic cops of cell signaling. Curr. Opinion Cell Biol. 2004, 16, 162–168. [Google Scholar]
- Fredriksson, R.; Schioth, H.B. The repertoire of G-protein-coupled receptors in fully sequenced genomes. Mol. Pharmacol. 2005, 67, 1414–1425. [Google Scholar]
- Fang, Y.; Lahiri, J.; Picard, L. G protein-coupled receptor microarrays for drug discovery. Drug Discov. Today 2003, 8, 755–761. [Google Scholar]
- van Baal, J.; de Widt, J.; Divecha, N.; van Blitterswijk, W.J. Translocation of diacylglycerol kinase θ from cytosol to plasma membrane in response to activation of G protein-coupled receptors and protein kinase C. J. Biol. Chem. 2005, 280, 9870–9878. [Google Scholar]
- Degerman, E.; Belfrage, P.; Manganiello, V.C. Structure, localization, and regulation of cGMP-inhibited phosphodiesterase (PDE3). J. Biol. Chem. 1997, 272, 6823–6826. [Google Scholar]
- Fan, G.; Shumay, E.; Wang, H.; Malbon, C.C. The scaffold protein gravin (cAMP-dependent protein kinase-anchoring protein 250) binds the β2-adrenergic receptor via the receptor cytoplasmic Arg-329 to Leu-413 domain and provides a mobile scaffold during desensitization. J. Biol. Chem. 2001, 276, 24005–24014. [Google Scholar]
- Lin, F.; Wang, H.-Y.; Malbon, C.C. Gravin-mediated formation of signaling complexes in β2-adrenergic receptor desensitization and resensitization. J. Biol. Chem. 2000, 275, 19025–19034. [Google Scholar]
- Lu, Z.; Jiang, G.; Blume-Jensen, P.; Hunter, T. Epidermal growth factor-induced tumor cell invasion and metastasis initiated by dephosphorylation and downregulation of focal adhesion kinase. Mol. Cell Biol. 2001, 21, 4016–4031. [Google Scholar]
- Marceau, F.; Regoli, D. Bradykinin receptor ligands: therapeutic perspectives. Nat. Rev. Drug Discov. 2004, 3, 845–852. [Google Scholar]
- Liebmann, C.; Graness, A.; Ludwig, B.; Adomeit, A.; Boehmer, A.; Boehmer, F.-D.; Nürnberg, B.; Wetzker, R. Dual bradykinin B2 receptor signalling in A431 human epidermoid carcinoma cells: activation of protein kinase C Is counteracted by a Gs-mediated stimulation of the cyclic AMP pathway. Biochem. J. 1996, 313, 109–118. [Google Scholar]
- Ferguson, S.S.G. Evolving concepts in G protein-coupled receptor endocytosis: the role in receptor desensitization and signaling. Pharmacol. Rev. 2001, 53, 1–24. [Google Scholar]
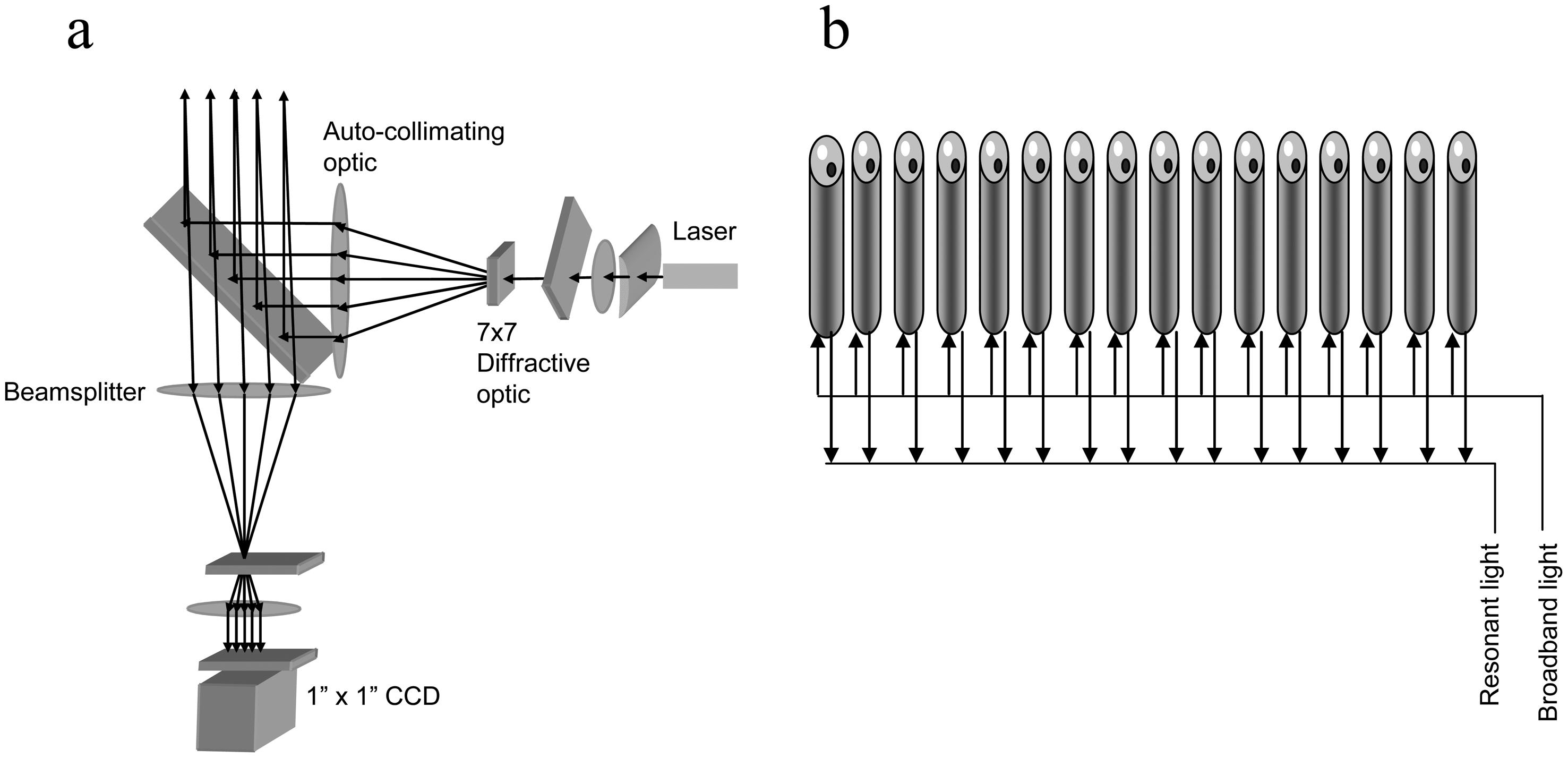
 ) in solution bind to the immobilized receptors. (b) RWG biosensor. The coupling of incident light into the waveguide is achieved by the diffraction grating. The intensity of reflected light is a function of the resonant wavelength. The binding of target molecules in a sample to the immobilized receptors results in a shift in the resonant wavelength. (c) Interferometric biosensor. A broadband light resource is directly incident at the solution-surface interface. Two types of reflected light waves originated from the two reflecting surfaces, the interface with the optical layer and the surface of bio-receptors, interact each other, leading to interference patterns. The binding of biomolecules to the bio-receptors alters the inference pattern.
) in solution bind to the immobilized receptors. (b) RWG biosensor. The coupling of incident light into the waveguide is achieved by the diffraction grating. The intensity of reflected light is a function of the resonant wavelength. The binding of target molecules in a sample to the immobilized receptors results in a shift in the resonant wavelength. (c) Interferometric biosensor. A broadband light resource is directly incident at the solution-surface interface. Two types of reflected light waves originated from the two reflecting surfaces, the interface with the optical layer and the surface of bio-receptors, interact each other, leading to interference patterns. The binding of biomolecules to the bio-receptors alters the inference pattern.
 ) in solution bind to the immobilized receptors. (b) RWG biosensor. The coupling of incident light into the waveguide is achieved by the diffraction grating. The intensity of reflected light is a function of the resonant wavelength. The binding of target molecules in a sample to the immobilized receptors results in a shift in the resonant wavelength. (c) Interferometric biosensor. A broadband light resource is directly incident at the solution-surface interface. Two types of reflected light waves originated from the two reflecting surfaces, the interface with the optical layer and the surface of bio-receptors, interact each other, leading to interference patterns. The binding of biomolecules to the bio-receptors alters the inference pattern.
) in solution bind to the immobilized receptors. (b) RWG biosensor. The coupling of incident light into the waveguide is achieved by the diffraction grating. The intensity of reflected light is a function of the resonant wavelength. The binding of target molecules in a sample to the immobilized receptors results in a shift in the resonant wavelength. (c) Interferometric biosensor. A broadband light resource is directly incident at the solution-surface interface. Two types of reflected light waves originated from the two reflecting surfaces, the interface with the optical layer and the surface of bio-receptors, interact each other, leading to interference patterns. The binding of biomolecules to the bio-receptors alters the inference pattern.


© 2007 by MDPI ( http://www.mdpi.org). Reproduction is permitted for noncommercial purposes.
Share and Cite
Fang, Y. Non-invasive Optical Biosensor for Probing Cell Signaling. Sensors 2007, 7, 2316-2329. https://doi.org/10.3390/s7102316
Fang Y. Non-invasive Optical Biosensor for Probing Cell Signaling. Sensors. 2007; 7(10):2316-2329. https://doi.org/10.3390/s7102316
Chicago/Turabian StyleFang, Ye. 2007. "Non-invasive Optical Biosensor for Probing Cell Signaling" Sensors 7, no. 10: 2316-2329. https://doi.org/10.3390/s7102316



