Discriminant Analysis of Raman Spectra for Body Fluid Identification for Forensic Purposes
Abstract
:1. Introduction
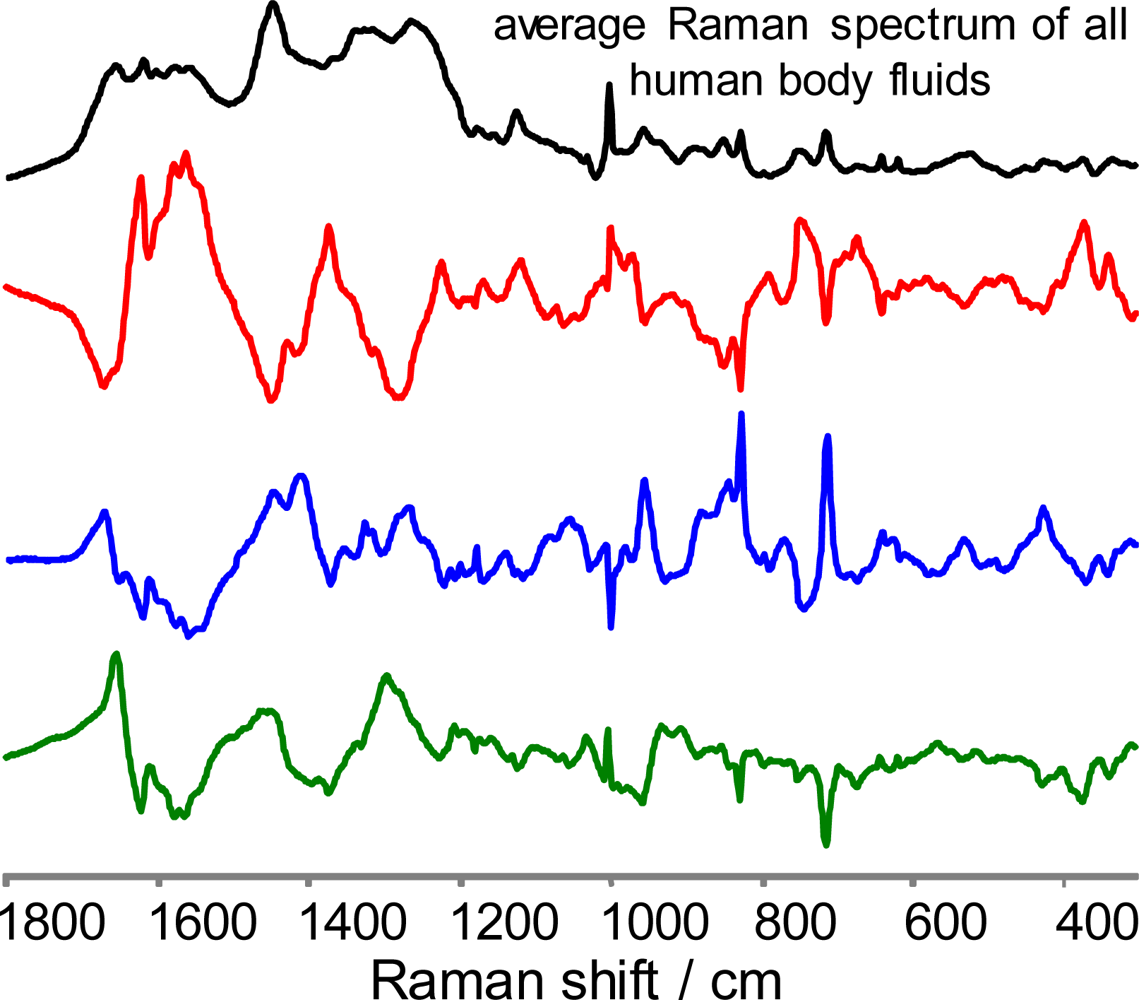
2. Multi-dimensional Raman spectroscopic signature for a single body fluid (review of data published recently)
2.1. Experimental
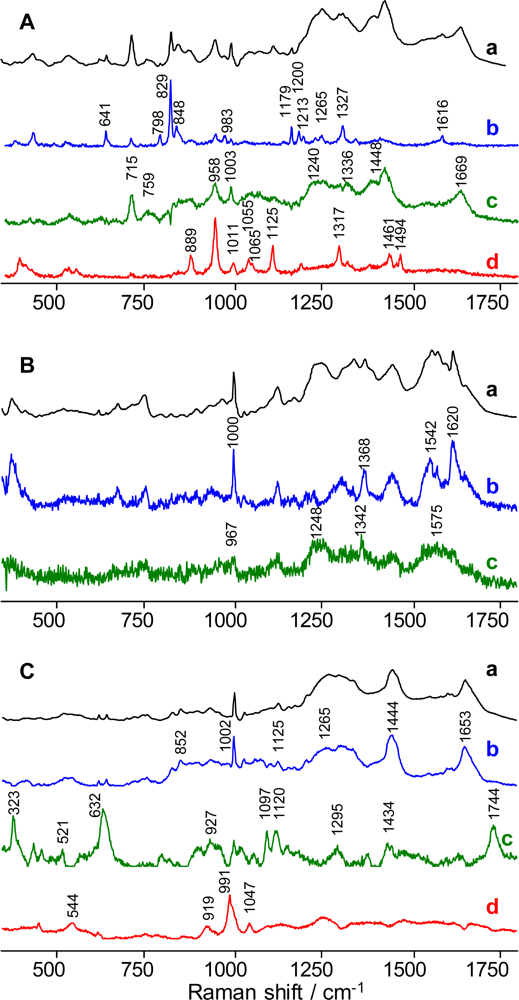
2.2. Raman Spectroscopic Signature of Semen
2.3. Raman Spectroscopic Signature of Blood
2.4. Raman Spectroscopic Signature of Saliva
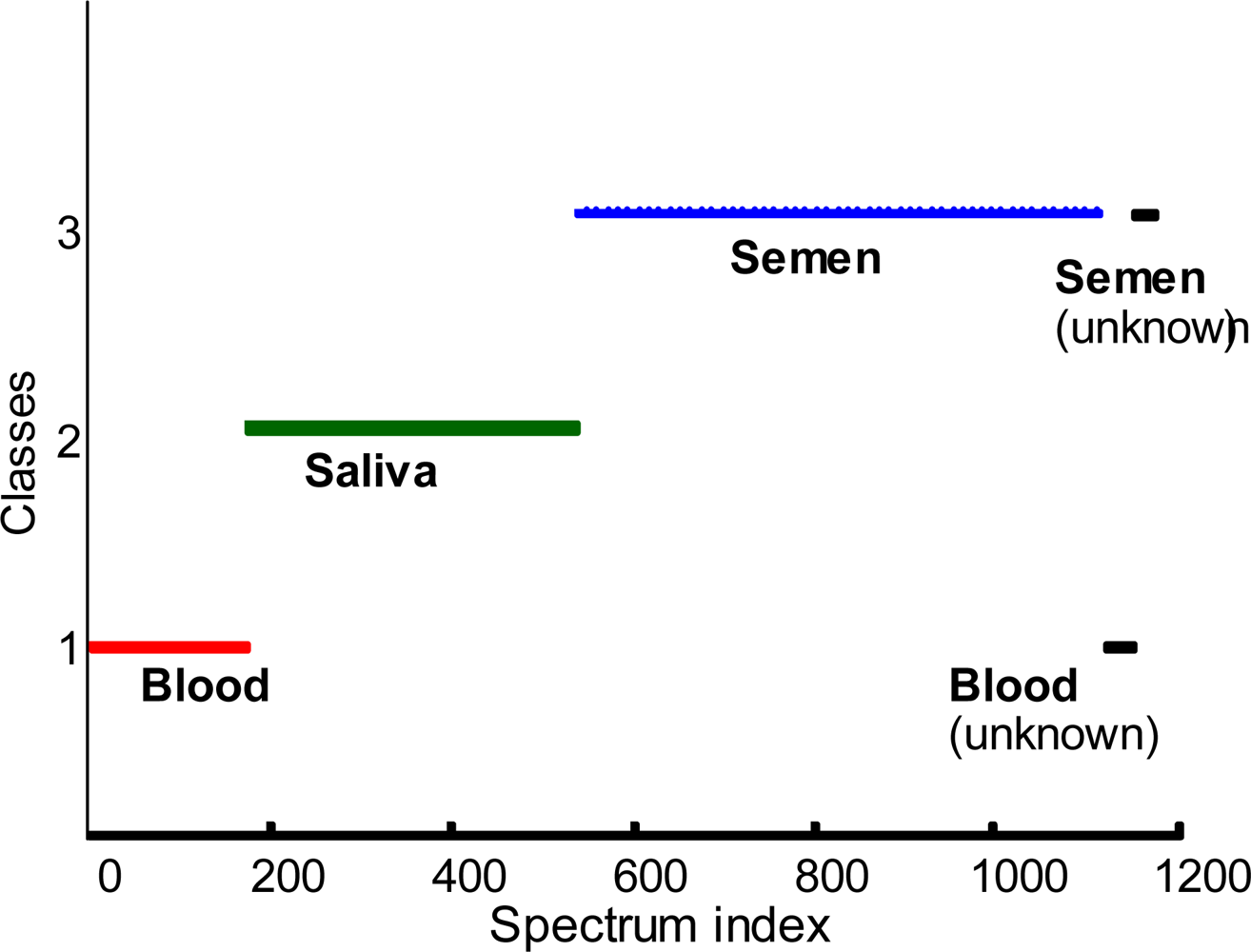
3. Identification of unknown body fluids
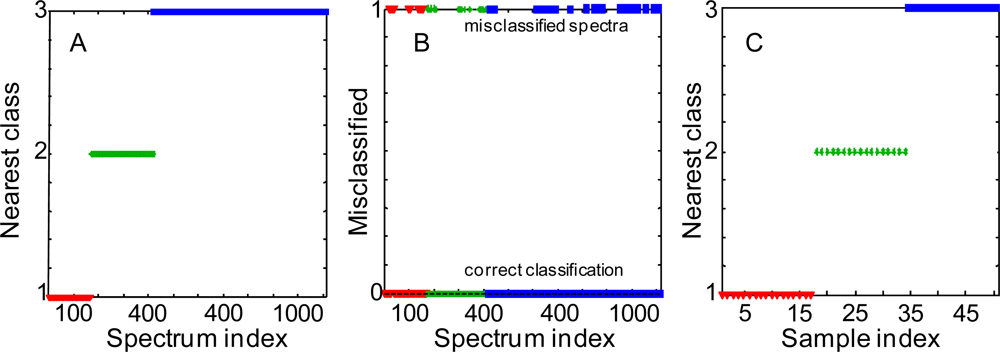
3.1. Soft Independent Modeling of Class Analogy
3.2. Linear Discriminant Analysis
3.3. Partial Least Squares Discriminant Analysis
4. Conclusions
Acknowledgments
References and Notes
- Greenfield, A.; Sloan, M.M. Identification of biological fluids and stains. In Forensic Science: an Introduction to Scientific and Investigative Techniques; CRC Press: Boca Ratom, FL, USA, 2005; pp. 261–271. [Google Scholar]
- Juusola, J.; Ballantyne, J. mRNA profiling for body fluid identification by multiplex quantitative RT-PCR. J. Forensic. Sci 2007, 52, 1252–1262. [Google Scholar]
- Crime Scene Investigation: A Guide for Law Enforcement; U.S. Department of Justice Office of Justice Programs: Washington, DC, USA, 2000.
- Soukos, N.S.; Crowley, K.; Bamberg, M.P.; Gillies, R.; Doukas, A.G.; Evans, R.; Kollias, N. A rapid method to detect dried saliva stains swabbed from human skin using fluorescence spectroscopy. Forensic. Sci. Int 2000, 114, 133–138. [Google Scholar]
- Powers, L.S.; Lloyd, C.R. Method and apparatus for detecting and imaging the presence of biological materials. US Patent 7186990,; March 6 2007. [Google Scholar]
- Estes, C.; Duncan, A.; Wade, B.; Lloyd, C.; Ellis, W., Jr.; Powers, L. Reagentless detection of microorganisms by intrinsic fluorescence. Biosens. Bioelectron 2003, 18, 511–519. [Google Scholar]
- Virkler, K.; Lednev, I.K. Raman spectroscopic signature of semen and its potential application to forensic body fluid identification. Forensic. Sci. Int 2009, 193, 56–62. [Google Scholar]
- Virkler, K.; Lednev, I.K. Forensic body fluid identification: The Raman spectroscopic signature of saliva. Analyst 2010. [Google Scholar] [CrossRef]
- Virkler, K.; Lednev, I.K. Raman spectroscopic signature of blood and its potential application to forensic body fluid identification. Anal. Bioanal. Chem 2009, 396, 525–534. [Google Scholar]
- Virkler, K.; Lednev, I.K. Raman spectroscopy offers great potential for the nondestructive confirmatory identification of body fluids. Forensic. Sci. Int 2008, 181, e1–5. [Google Scholar]
- Virkler, K.; Lednev, I.K. Analysis of body fluids for forensic purposes: from laboratory testing to non-destructive rapid confirmatory identification at a crime scene. Forensic. Sci. Int 2009, 188, 1–17. [Google Scholar]
- De Wael, K.; Lepot, L.; Gason, F.; Gilbert, B. In search of blood--detection of minute particles using spectroscopic methods. Forensic. Sci. Int 2008, 180, 37–42. [Google Scholar]
- Virkler, K.; Lednev, I.K. Blood species identification for forensic purposes using Raman spectroscopy combined with advanced statistical analysis. Anal. Chem 2009, 81, 7773–7777. [Google Scholar]
- Gebel, E. Species in a snap: Raman analysis of blood. Anal. Chem 2009, 81, 7862. [Google Scholar]
- Yu, N.T. Raman spectroscopy: a conformational probe in biochemistry. CRC Crit. Rev. Biochem 1977, 4, 229–280. [Google Scholar]
- Carey, P.R. Resonance Raman spectroscopy in biochemistry and biology. Q. Rev. Biophys 1978, 11, 309–370. [Google Scholar]
- Peticolas, W.L. Application of Raman spectroscopy to biological macromolecules. Biochimie 1975, 57, 417–428. [Google Scholar]
- Shashilov, V.; Lednev, I.K. Deep-UV Raman spectroscopy directly probes a fibrillation nucleus. Laser Focus World 2007, 43, 87–90. [Google Scholar]
- Shashilov, V.A.; Lednev, I.K. Two-dimensional correlation Raman spectroscopy for characterizing protein structure and dynamics. J. Raman Spectrosc 2009, 40, 1749–1758. [Google Scholar]
- Shashilov, V.; Xu, M.; Ermolenkov, V.V.; Fredriksen, L.; Lednev, I.K. Probing a fibrillation nucleus directly by deep ultraviolet Raman spectroscopy. J. Am. Chem. Soc 2007, 129, 6972–6973. [Google Scholar]
- Xu, M.; Shashilov, V.; Lednev, I.K. Probing the cross-beta core structure of amyloid fibrils by hydrogen-deuterium exchange deep ultraviolet resonance Raman spectroscopy. J. Am. Chem. Soc 2007, 129, 11002–11003. [Google Scholar]
- Xu, M.; Shashilov, V.A.; Ermolenkov, V.V.; Fredriksen, L.; Zagorevski, D.; Lednev, I.K. The first step of hen egg white lysozyme fibrillation, irreversible partial unfolding, is a two-state transition. Protein. Sci 2007, 16, 815–832. [Google Scholar]
- Shashilov, V.A.; Ermolenkov, V.V.; Lednev, I.K. Multiple bicyclic diamide-lutetium complexes in solution: chemometric analysis of deep-UV Raman spectroscopic data. Inorg. Chem 2006, 45, 3606–3612. [Google Scholar]
- Sikirzhytski, V.; Topilina, N.I.; Higashiya, S.; Welch, J.T.; Lednev, I.K. Genetic engineering combined with deep UV resonance Raman spectroscopy for structural characterization of amyloid-like fibrils. J. Am. Chem. Soc 2008, 130, 5852–5853. [Google Scholar]
- Shashilov, V.A.; Lednev, I.K. 2D correlation deep UV resonance raman spectroscopy of early events of lysozyme fibrillation: kinetic mechanism and potential interpretation pitfalls. J. Am. Chem. Soc 2008, 130, 309–317. [Google Scholar]
- Shashilov, V.A.; Xu, M.; Ermolenkov, V.V.; Lednev, I.K. Latent variable analysis of Raman spectra for structural characterization of proteins. J. Quant. Spectrosc. Radiat. Transfer 2006, 102, 46–61. [Google Scholar]
- Shashilov, V.A.; Lednev, I.K. Bayesian Extraction of Deep UV Resonance Raman Signature of Fibrillar Cross-β Sheet Core based on H-D Exchange Data; Knuth, K.H., Caticha, A., Giffin, A., Rodríguez, C.C., Center, J.L.J., Eds.; Saratoga Springs: New York, NY, USA, 2007; pp. 450–457. [Google Scholar]
- De Gelder, J.; Gussem, K.D.; Vandenabeele, P.; Moens, L. Reference database of Raman spectra of biological molecules. J. Raman. Spectrosc 2007, 38, 1133–1147. [Google Scholar]
- Johnson, C.R.; Ludwig, M.; Asher, S.A. Ultraviolet resonance Raman characterization of photochemical transients of phenol, tyrosine, and tryptophan. J. Am. Chem. Soc 1986, 108, 905–912. [Google Scholar]
- Chikishev, A.Y.; Lucassen, G.W.; Koroteev, N.I.; Otto, C.; Greve, J. Polarization sensitive coherent anti-Stokes Raman scattering spectroscopy of the amide I band of proteins in solutions. Biophys. J 1992, 63, 976–985. [Google Scholar]
- Berjot, M.; Marx, J.; Alix, A.J.P. Determination of the secondary structure of proteins from the Raman amide I band: The reference intensity profiles method. J. Raman. Spectrosc 2005, 18, 289–300. [Google Scholar]
- Mikhonin, A.V.; Bykov, S.V.; Myshakina, N.S.; Asher, S. A. Peptide secondary structure folding reaction coordinate: correlation between uv raman amide III frequency, Psi Ramachandran angle, and hydrogen bonding. J. Phys. Chem. B 2006, 110, 1928–1943. [Google Scholar]
- Carter, E.A.; Edwards, H.G.M. Biological applications of Raman spectroscopy. In Infrared and Raman Spectroscopy of Biological Materials; Hans-Ulrich, G., Yan, B., Eds.; Marcel Dekker, Inc: New York, NY, USA, 2001; Volume 24, pp. 421–475. [Google Scholar]
- Liang, J.; Cheng, Y.; Han, H. Study on the interaction of bovine serum albumin and CdTe quantum dots with spectroscopic techniques. J. Mol. Struct 2008, 892, 116–120. [Google Scholar]
- Ivanov, A.I.; Zhbankov, R.G.; Korolenko, E.A.; Korolik, E.V.; Meleshchenko, L.A.; Marchewka, M.; Ratajczak, H. Infrared, Raman spectroscopic studies of the structure of human serum albumin under various ligand loads. J. Appl. Spectrosc 1994, 60, 399–405. [Google Scholar]
- Edsall, J.T. Raman spectra of amino acids and related compounds. VI. Sarcosine, ethanolamine, choline, betaine, and betaine derivatives. J. Am. Chem. Soc 1943, 65, 1767–1770. [Google Scholar]
- Koyama, Y.; Toda, S.; Kyogoku, Y. Raman spectra and conformation of the glycerophosphorylcholine headgroup. Chem. Phys. Lipids 1977, 19, 74–92. [Google Scholar]
- Spiker, R.C., Jr.; Levin, I.W. Raman spectra and vibrational assignments for dipalmitoyl phosphatidylcholine and structurally related molecules. Biochim. Biophys. Acta 1975, 388, 361–373. [Google Scholar]
- Bertoluzza, A.; Fagnano, C.; Finelli, P.; Morelli, M.A.; Simoni, R.; Tosi, R. Raman and infrared spectra of spermidine and spermine and their hydrochlorides and phosphates as a basis for the study of the interactions between polyamines and nucleic acids. J. Raman. Spectrosc 1983, 14, 385–394. [Google Scholar]
- Eapen, A.; Joe, I.H.; Aruldhas, G. Vibrational and SERS spectra of spermine phosphate hexahydrate. Spectrosc. Lett 1997, 30, 751–770. [Google Scholar]
- Sato, H.; Chiba, H.; Tashiro, H.; Ozaki, Y. Excitation wavelength-dependent changes in Raman spectra of whole blood and hemoglobin: comparison of the spectra with 514.5-, 720-, and 1064-nm excitation. J. Biomed. Opt 2001, 6, 366–370. [Google Scholar]
- Venkatesh, B.; Ramasamy, S.; Mylrajan, M.; Asokan, R.; Manoharan, P.T.; Rifkind, J.M. Fourier transform Raman approach to structural correlation in hemoglobin derivatives. Spectrochim. Acta A Mol. Biomol. Spectrosc 1999, 55A, 1691–1697. [Google Scholar]
- Asher, S.A.; Vickery, L.E.; Schuster, T.M.; Sauer, K. Resonance Raman spectra of methemoglobin derivatives. Selective enhancement of axial ligand vibrations and lack of an effect of inositol hexaphosphate. Biochemistry 1977, 16, 5849–5856. [Google Scholar]
- Blood. In Encyclopaedia Britannica, 14th ed; Hutchins, R.M. (Ed.) Encyclopaedia Britannica, Inc: Chicago, IL, USA, 1970; Volume 3, p. 796.
- Adar, F.; Gouterman, M; Aronowitz, S. Fluorescence, resonance Raman, and radiationless decay in several hemoproteins. J. Phys. Chem 1976, 80, 2184–2191. [Google Scholar]
- Marx, J.; Hudry-Clergeon, G.; Capet-Antonini, F.; Bernard, L. Laser Raman spectroscopy study of bovine fibrinogen and fibrin. Biochim. Biophys. Acta 1979, 578, 107–115. [Google Scholar]
- Vanni, S. Raman chemical imaging provides rapid, non-invasive and reagentless biothreat detection. Proceedings of 19th Annual NDIA Security Symposium & Exhibition; ChemImage Corporation: Pittsburgh, PA, USA, 2003. [Google Scholar]
- Kalasinsky, K.S.; Hadfield, T.; Shea, A.A.; Kalasinsky, V.F.; Nelson, M.P.; Neiss, J.; Drauch, A.J.; Vanni, G.S.; Treado, P.J. Raman chemical imaging spectroscopy reagentless detection and identification of pathogens: signature development and evaluation. Anal. Chem 2007, 79, 2658–2673. [Google Scholar]
- FT-IR Raman spectrum of propyl acetate. Sigma-Aldrich Co: St. Louis, MO, USA, 2009.
- FT-IR Raman spectrum of dodecyl acetate. Sigma-Aldrich Co: St. Louis, MO, USA, 2009.
- FT-IR Raman spectrum of amyl acetate. Sigma-Aldrich Co: St. Louis, MO, USA, 2009.
- Barrett, T.W. Laser Raman spectra of mono-, oligo- and polysaccharides in solution. Spectrochim. Acta A Mol. Spectrosc 1981, 37, 233–239. [Google Scholar]
- Habuchi, O. Diversity and functions of glycosaminoglycan sulfotransferases. Biochim. Biophys. Acta 2000, 1474, 115–127. [Google Scholar]
- Walsh, M.C.; Brennan, L.; Malthouse, J.P.; Roche, H.M.; Gibney, M.J. Effect of acute dietary standardization on the urinary, plasma, and salivary metabolomic profiles of healthy humans. Am. J. Clin. Nutr 2006, 84, 531–539. [Google Scholar]
- Effros, R.M.; Casaburi, R.; Su, J.; Dunning, M.; Torday, J.; Biller, J.; Shaker, R. The effects of volatile salivary acids and bases on exhaled breath condensate pH. Am. J. Respir. Crit. Care Med 2006, 173, 386–392. [Google Scholar]
- Mariey, L.; Signolle, J.P.; Amiel, C.; Travert, J. Discrimination, classification, identification of microorganisms using FTIR spectroscopy and chemometrics. Vib. Spectrosc 2001, 26, 151–159. [Google Scholar]
- Coomans, D.; Smyth, C.; Lee, I.; Hancock, T.; Yang, J. Unsupervised data mining: introduction. In Comprehensive chemometrics: chemical and biochemical data analysis; Brown, S.D., Tauler, R., Walczak, B., Eds.; Elsevier B.V.: Amsterdam, the Netherlands, 2009; Volume 2, pp. 559–576. [Google Scholar]
- Orav, E.J.; Louis, T.A.; Palmer, R.H.; Wright, E.A. Variance components and their implications for statistical information in medical data. Stat. Med 1991, 10, 599–616. [Google Scholar]
- Beljebbar, A.; Bouche, O.; Diebold, M.D.; Guillou, P.J.; Palot, J.P.; Eudes, D.; Manfait, M. Identification of Raman spectroscopic markers for the characterization of normal and adenocarcinomatous colonic tissues. Crit. Rev. Oncol. Hematol 2009, 72, 255–264. [Google Scholar]
- Miller, D.J.; Wang, Y.; Kesidis, G. Emergent unsupervised clustering paradigms with potential application to bioinformatics. Front. Biosci 2008, 13, 677–690. [Google Scholar]
- Ester, M.; Kriegel, H.-P.; Xu, X. A density-based algorithm for discovering clusters in large spatial databases with noise; AAAI Press: Menlo Park, CA, USA, 1996; pp. 226–231. [Google Scholar]
- Krafft, C.; Steiner, G.; Beleites, C.; Salzer, R. Disease recognition by infrared and Raman spectroscopy. J. Biophotonics 2009, 2, 13–28. [Google Scholar]
- Ivosev, G.; Burton, L.; Bonner, R. Dimensionality reduction and visualization in principal component analysis. Anal. Chem 2008, 80, 4933–4944. [Google Scholar]
- Baxter, M.J. A review of supervised and unsupervised pattern recognition in archaeometry. Archaeometry 2006, 48, 671–694. [Google Scholar]
- Larranaga, P.; Calvo, B.; Santana, R.; Bielza, C.; Galdiano, J.; Inza, I.; Lozano, J.; Armananzas, R.; Santafe, G.; Perez, A.; Robles, V. Machine learning in bioinformatics. Briefings Bioinformatics 2006, 7, 86–112. [Google Scholar]
- Lavine, B.K.; Rayens, W.S. Statistical discriminant analysis. In Comprehensive chemometrics: chemical and biochemical data analysis; Brown, S.D., Tauler, R., Walczak, B., Eds.; Elsevier B.V.: Amsterdam, Netherlands, 2009; Volume 3, pp. 517–540. [Google Scholar]
- Coomans, D.; Massart, D. L.; Kaufman, L. Optimization by statistical linear discriminant analysis in analytical chemistry. Anal. Chim. Acta 1979, 112, 97–122. [Google Scholar]
- Huang, J.; Yuen, P.C.; Chen, W.-S.; Lai, J.H. Choosing parameters of kernel subspace LDA for recognition of face images under pose and illumination variations. IEEE Trans. Syst. Man. Cybern. Part B Cybern 2007, 37, 847–862. [Google Scholar]
- Yu, H.; Yang, J. A direct LDA algorithm for high-dimensional data with application to face recognition. Pattern. Recognit 2001, 34, 2067–2070. [Google Scholar]
- Loog, M.; Duin, R.P.W. Linear dimensionality reduction via a heteroscedastic extension of LDA: the Chernoff criterion. IEEE Trans. Pattern. Anal. Mach. Intell 2004, 26, 732–739. [Google Scholar]
- Zhu, M.; Martinez, A.M. Subclass discriminant analysis. IEEE Trans. Pattern. Anal. Mach. Intell 2006, 28, 1274–1286. [Google Scholar]
- Dunn, W.J.I.; Wold, S. SIMCA pattern recognition and classification. Meth. Princip. Med. Chem 1995, 2, 179–193. [Google Scholar]
- Goerlitz, L.; Menze, B.H.; Kelm, B.M.; Hamprecht, F.A. Processing spectral data. Surf. Interface Anal 2009, 41, 636–644. [Google Scholar]
- Everitt, B.S. Analysis of longitudinal data. Beyond MANOVA. Br. J. Psychiatry 1998, 172, 7–10. [Google Scholar]
- Barker, M.; Rayens, W. Partial least squares for discrimination. J. Chemom 2003, 17, 166–173. [Google Scholar]

| Modeled class | Blood | Semen | Saliva |
|---|---|---|---|
| Sensitivity (Cal) | 1.000 | 1.000 | 0.999 |
| Specificity (Cal) | 1.000 | 0.999 | 1.000 |
| Sensitivity (CV) | 1.000 | 1.000 | 0.999 |
| Specificity (CV) | 1.000 | 0.999 | 1.000 |
| Class. Err (Cal) | 0 | 0.000579 | 0.000721 |
| Class. Err (CV) | 0 | 0.000579 | 0.000721 |
| RMSEC | 0.0344 | 0.107 | 0.103 |
© 2010 by the authors; licensee Molecular Diversity Preservation International, Basel, Switzerland. This article is an open-access article distributed under the terms and conditions of the Creative Commons Attribution license ( http://creativecommons.org/licenses/by/3.0/).
Share and Cite
Sikirzhytski, V.; Virkler, K.; Lednev, I.K. Discriminant Analysis of Raman Spectra for Body Fluid Identification for Forensic Purposes. Sensors 2010, 10, 2869-2884. https://doi.org/10.3390/s100402869
Sikirzhytski V, Virkler K, Lednev IK. Discriminant Analysis of Raman Spectra for Body Fluid Identification for Forensic Purposes. Sensors. 2010; 10(4):2869-2884. https://doi.org/10.3390/s100402869
Chicago/Turabian StyleSikirzhytski, Vitali, Kelly Virkler, and Igor K. Lednev. 2010. "Discriminant Analysis of Raman Spectra for Body Fluid Identification for Forensic Purposes" Sensors 10, no. 4: 2869-2884. https://doi.org/10.3390/s100402869




