The Association of Il28b Genotype with the Histological Features of Chronic Hepatitis C Is HCV Genotype Dependent
Abstract
:Materials and Methods
1. Introduction
2. Results
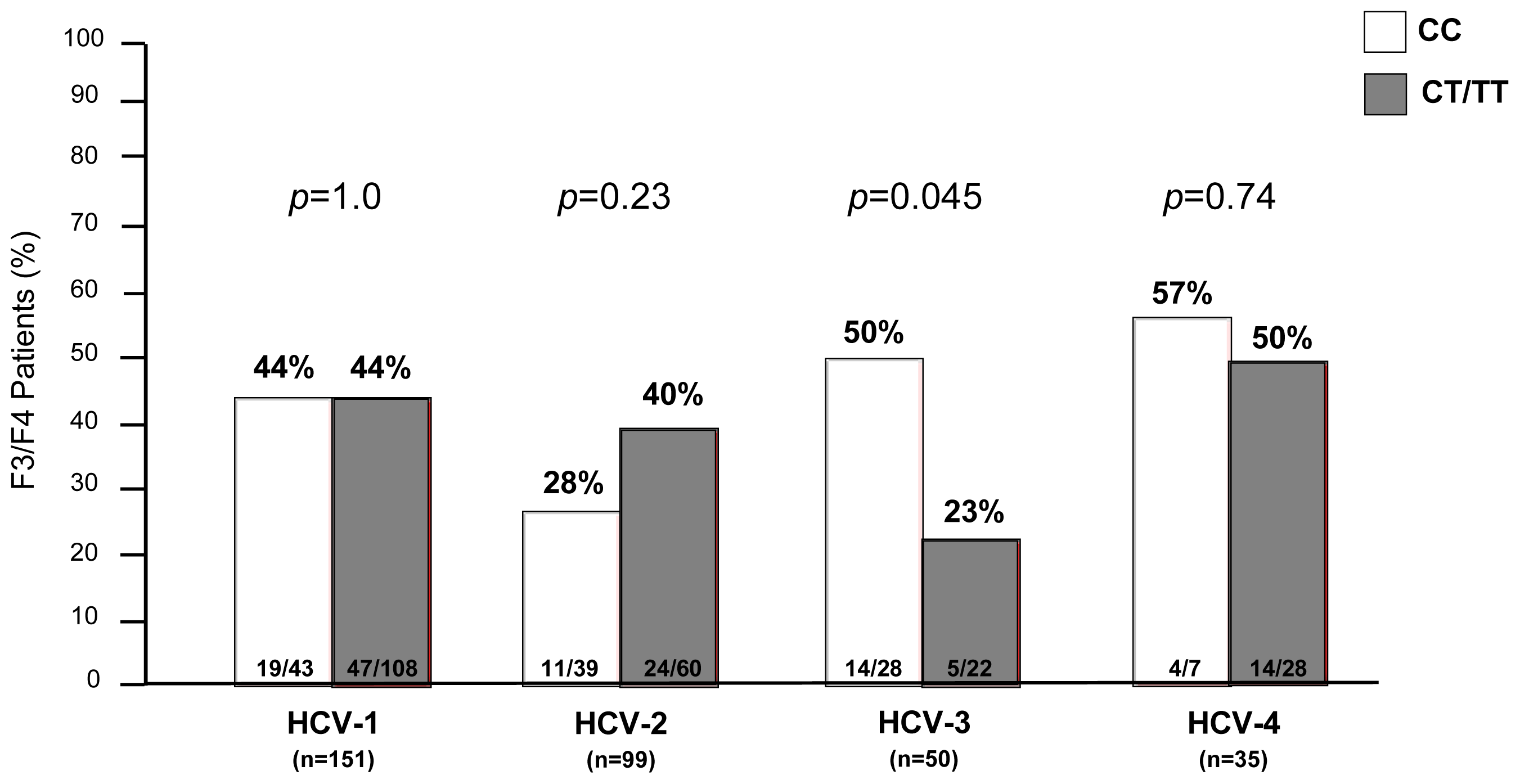
2.1. Fibrosis
2.2. Inflammation
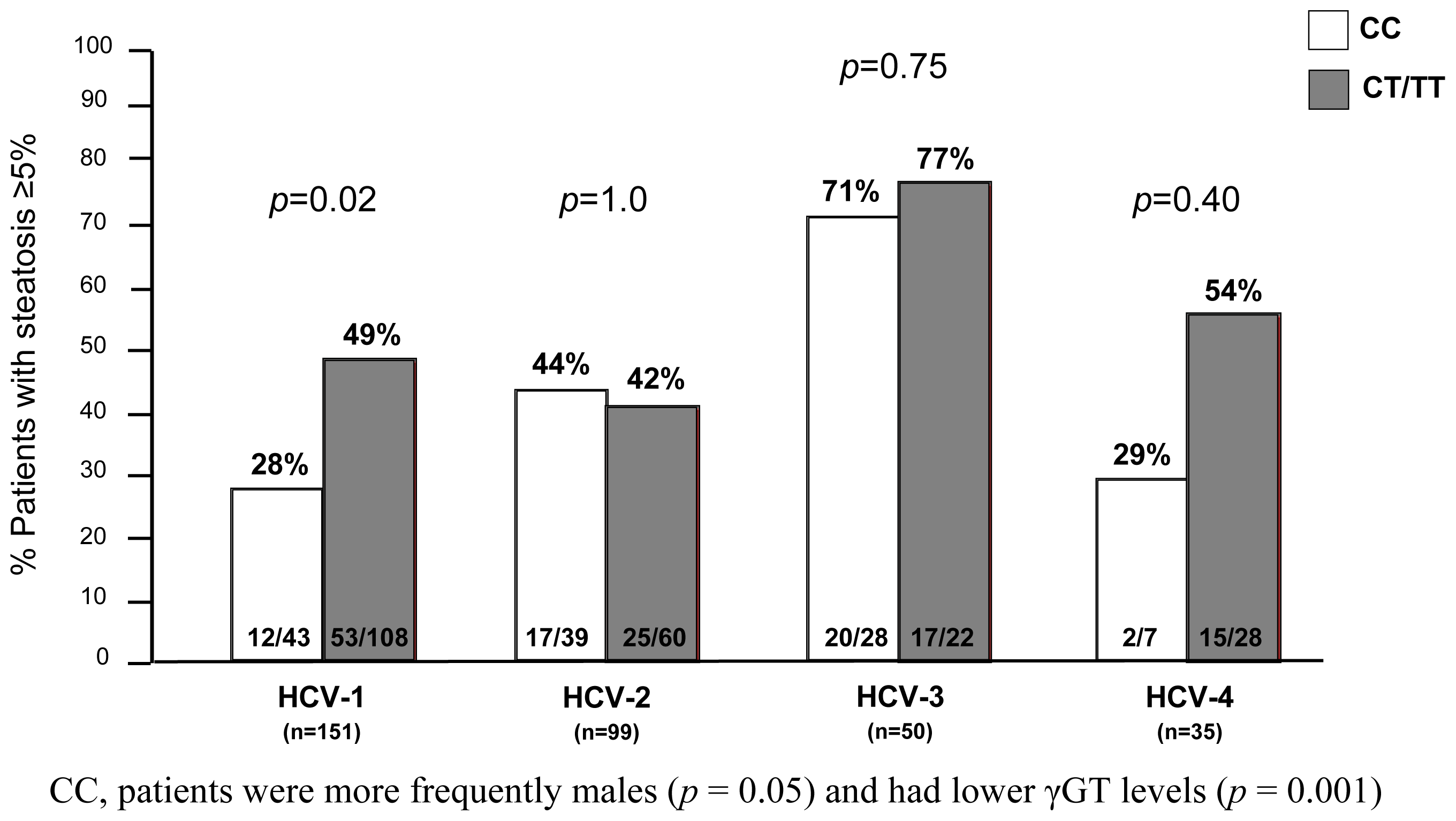
2.3. Steatosis
3. Discussion
4. Materials and Methods
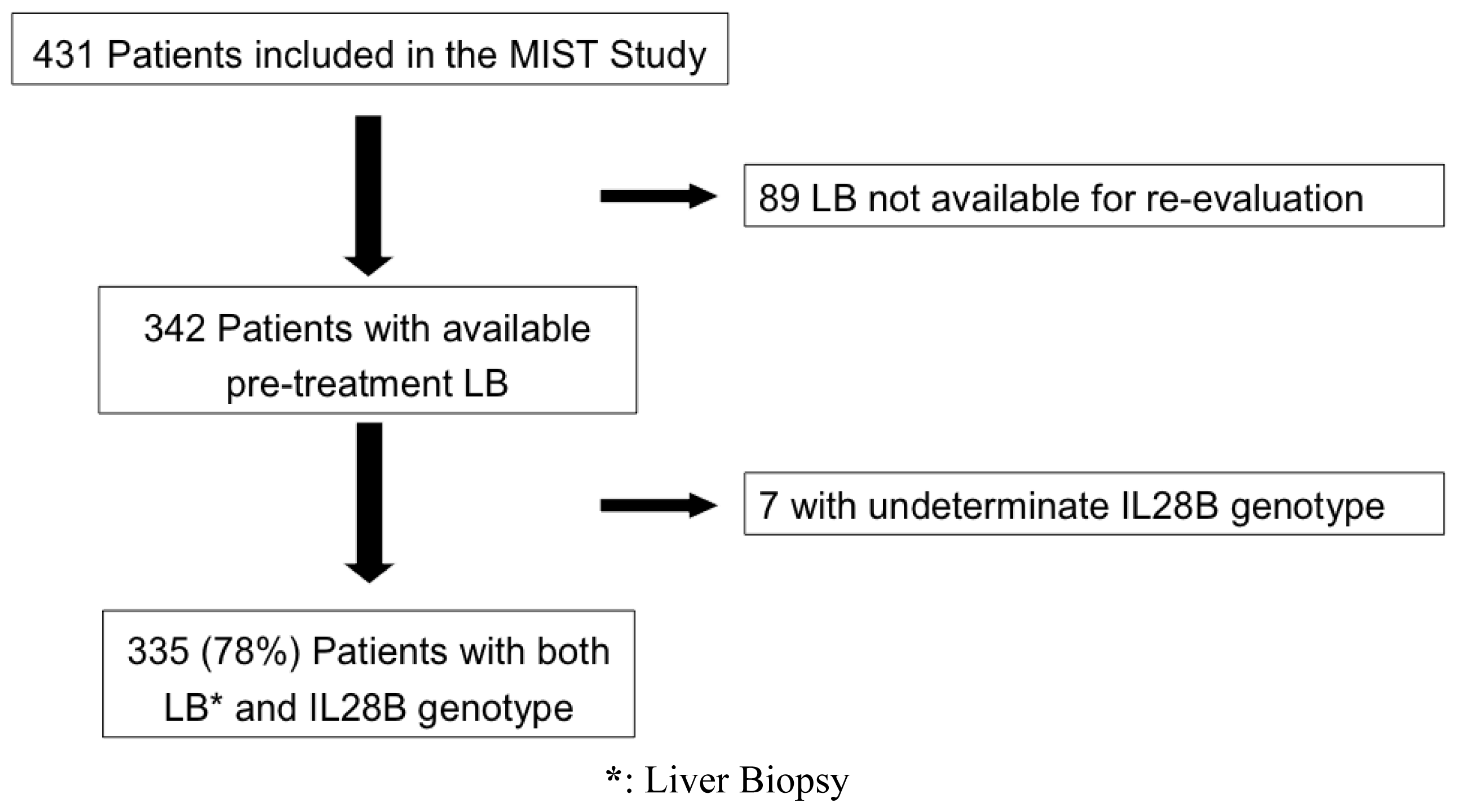
4.1. Patient Population
4.2. Laboratory Assessment
4.3. IL28B Genotyping
4.4. Histological Assessment
4.5. Statistical Analysis
5. Conclusions
Acknowledgments
Conflicts of Interest
- Author ContributionsStudy Conception and Design: Roberta D’Ambrosio, Alessio Aghemo, Massimo Colombo. Acquisition of data: Roberta D’Ambrosio and Stella De Nicola. Liver Biopsies Reading: Roberta D’Ambrosio and Guido Ronchi. IL28B Genotyping; Raffaele De Francesco, Enrico Galmozzi and Cristina Cheroni. Analysis and Interpretation of data: Roberta D’Ambrosio, Alessio Aghemo, Pietro Lampertico. Statistical Analysis: Paul J Clark and Pietro Lampertico. Drafting of the Manuscript: Roberta D’Ambrosio and Alessio Aghemo. Critical Revision: Roberta D’Ambrosio, Alessio Aghemo, Maria Grazia Rumi, Pietro Lampertico, Massimo Colombo.
References
- Thomas, D.L.; Thio, C.L.; Martin, M.P.; Qi, Y.; Ge, D.; O’hUigin, C.; Kidd, J.; Kidd, K.; Khakoo, S.I.; Alexander, G.; et al. Genetic variation in IL28B and spontaneous clearance of hepatitis C virus. Nature 2009, 461, 798–801. [Google Scholar]
- Rauch, A.; Kutalik, Z.; descombes, P.; Cai, T.; Iulio, J.D.; Mueller, T.; Bochud, M.; Battegay, M.; Bernasconi, E.; Borovicka, J.; et al. Genetic variation in IL28B is associated with chronic hepatitis C and treatment failure: A genome-wide association study. Gastroenterology 2010, 138, 1338–1345. [Google Scholar]
- Ge, D.; fellay, J.; Thompson, A.J.; Simon, J.S.; Shianna, K.V.; Urban, T.J.; Heinzen, E.L.; Qiu, P.; Bertelsen, A.H.; Muir, A.J.; et al. Genetic variation in IL28B predicts hepatitis C treatment-induced viral clearance. Nature 2009, 461, 399–401. [Google Scholar]
- Suppiah, V.; Moldovan, M.; Ahlenstiel, G.; Berg, T.; Weltman, M.; Abate, M.L.; Bassendine, M.; Spengler, U.; Dore, G.J.; Powell, E.; et al. IL28B is associated with response to chronic hepatitis C interferon-alpha and ribavirin therapy. Nat. Genet 2009, 41, 1100–1104. [Google Scholar]
- Tanaka, Y.; Nishida, N.; Sugiyama, M.; Kurosaki, M.; Matsuura, K.; Sakamoto, N.; Nakagawa, M.; Korenaga, M.; Hino, K.; Hige, S.; et al. Genome-wide association of IL28B with response to pegylated interferon-alpha and ribavirin therapy for chronic hepatitis C. Nat. Genet 2009, 41, 1105–1109. [Google Scholar]
- Thompson, A.J.; Muir, A.J.; Sulkowski, M.S.; Ge, D.; Fellay, J.; Shianna, K.V.; Urban, T.; Afdhal, N.H.; Jacobson, I.M.; Esteban, R.; et al. Interleukin-28B polymorphism improves viral kinetics and is the strongest pretreatment predictor of sustained virological response in genotype 1 hepatitis C virus. Gastroenterology 2010, 139, 120–129. [Google Scholar]
- Mangia, A.; Thompson, A.J.; Santoro, R.; Piazzolla, V.; Tillmann, H.L.; Patel, K.; Shianna, K.V.; Mottola, L.; Petruzzellis, D.; Bacca, D.; et al. An IL28B polymorphism determines treatment response of hepatitis C virus genotype 2 or 3 patients who do not achieve a rapid virological response. Gastroenterology 2010, 139, 821–827. [Google Scholar]
- De Nicola, S.; Aghemo, A.; Rumi, M.G.; Galmozzi, E.; Valenti, L.; Soffredini, R.; de Francesco, R.; Prati, G.M.; D’Ambrosio, R.; Cheroniet, C.; et al. Interleukin 28B polymorphism predicts pegylated interferon plus ribavirin treatment outcome in chronic hepatitis C genotype 4. Hepatology 2012, 55, 336–342. [Google Scholar]
- Asselah, T.; de Muynck, S.; Broët, P.; Masliah-Planchon, J.; Blanluet, M.; Bièche, I.; Lapalus, M.; Martinot-Peignoux, M.; Lada, O.; Estrabaud, E.; et al. IL28B polymorphism is associated with treatment response in patients with genotype 4 chronic hepatitis C. J. Hepatol 2012, 56, 527–532. [Google Scholar]
- Noureddin, M.; Wright, E.C.; Alter, H.J.; Clark, S.; Thomas, E.; Chen, R.; Zhao, X.; Conry-Cantilena, C.; Kleiner, D.E.; Liang, T.J.; et al. Association of IL28B genotype with fibrosis progression and clinical outcomes in patients with chronic hepatitis C: A longitudinal analysis. Hepatology 2013, 58, 1548–1557. [Google Scholar]
- D’Ambrosio, R.; Aghemo, A.; Rumi, M.G.; Ronchi, G.; Donato, M.F.; Paradis, V.; Colombo, M.; Bedossa, P. A morphometrical and immunoistochemical study to assess the benefit of a sustained virological response in hepatitis C virus patients with cirrhosis. Hepatology 2012, 56, 532–543. [Google Scholar]
- Golden-Mason, L.; bambha, K.M.; Cheng, L.; Howell, C.D.; Taylor, M.W.; Clark, P.J.; Afdhal, N.; Rosen, H.R. On behalf of the Virahep-C study group. Natural killer inhibitory receptor expression associated with treatment failure and interleukin-28B genotype in patients with chronic hepatitis C. Hepatology 2011, 54, 1559–1569. [Google Scholar]
- Naggie, S.; Osinusi, A.; Katsounas, A.; Lempicki, R.; Herrmann, E.; Thompson, A.J.; Clark, P.J.; Patel, K.; Muir, A.J.; McHutchison, J.G.; et al. Dysregulation of innate immunity in hepatitis C virus genotype 1 IL28B-unfavorable genotype patients: impaired viral kinetics and therapeutic response. Hepatology 2012, 56, 444–454. [Google Scholar]
- Thompson, A.J.; Clark, P.J.; Zhu, M.; Zhu, Q.; Ge, D.; Sulkowski, M.S.; Muir, A.J.; Tillmann, H.L.; Patel, K.; Naggie, S.; et al. Genome wide-association study identifies IL28B polymorphism to be associated with baseline ALT and hepatic necro-inflammatory activity in chronic hepatitis C patients enrolled in the IDEAL study. Hepatology 2010, 52, 1220A. [Google Scholar]
- Rembeck, K.; Alsio, A.; Christensen, P.B.; Färkkilä, M.; Langeland, N.; Buhl, M.R.; Pedersen, C.; Mørch, K.; Westin, J.; Lindh, M.; et al. Impact of IL28B-related single nucleotide polymorphisms on liver histopathology in chronic hepatitis C genotype 2 and 3. PLoS One 2012, 7. [Google Scholar] [CrossRef]
- Bochud, P.Y.; Bibert, S.; Kutalik, Z.; Patin, E.; Guergnon, J.; Nalpas, B.; Goossens, N.; Kuske, L.; Müllhaupt, B.; Gerlach, T.; et al. IL28B alleles associated with poor HCV clearance protect against inflammation and fibrosis in patients infected with non-1 HCV genotypes. Hepatology 2012, 55, 1038–1047. [Google Scholar]
- Check, P.J.; Thompson, A.J.; Vock, D.M.; Kratz, L.E.; Tolun, A.A.; Muir, A.J.; McHutchison, J.G.; Subramanian, M.; Millington, D.M.; Kelley, R.I.; et al. Hepatitis C virus selectively perturbs the distal cholesterol synthesis pathway in agenotype-specific manner. Hepatology 2012, 56, 49–56. [Google Scholar]
- Bochud, P.Y.; Cai, T.; Overbeck, K.; Bochud, M.; Dufour, J.F.; Müllhaupt, B.; Borovicka, J.; Heim, M.; Moradpour, D.; Cerny, A.; et al. Genotype 3 is associated with accelerated fibrosis progression in chronic hepatitis C. J. Hepatol 2009, 51, 655–666. [Google Scholar]
- Tillmann, H.L.; Patel, K.; Muir, A.J.; Guy, C.D.; Li, J.H.; Lao, X.Q.; Thompson, A.; Clark, P.J.; Gardner, S.D.; McHutchison, J.G.; et al. Beneficial IL28B genotype associated with lower frequency of hepatic Steatosis in patients with chronic hepatitis C. J. Hepatol 2011, 55, 1195–1200. [Google Scholar]
- Rousselet, M.C.; Michalak, S.; Duprè, F.; Croué, A.; Bedossa, P.; Saint-André, J.P.; Calès, P. Sources of variability in histological scoring of chronic viral hepatitis. Hepatology 2005, 41, 257–264. [Google Scholar]
- Rumi, M.G.; Aghemo, A.; Prati, G.M.; D’Ambrosio, R.; Donato, M.F.; Soffredini, R.; del Ninno, E.; Russo, A.; Colombo, M. Randomized study of peginterferon-alpha2a plus ribavirin and peginterferon-alpha2b plus ribavirin in chronic hepatitis C. Gastroenterology 2010, 138, 108–115. [Google Scholar]
- Aghemo, A.; Prati, G.M.; Rumi, M.G.; Soffredini, R.; D’Ambrosio, R.; Orsi, E.; de Nicola, S.; Degasperi, E.; Grancini, V.; Colombo, M. Sustained virological response prevents the development of insulin resistance in patients with chronic hepatitis C. Hepatology 2012, 56. [Google Scholar] [CrossRef]
- Bedossa, P. Intraobserver and interobserver variations in liver biopsy interpretation in patients with chronic hepatitis C. Hepatology 1994, 20, 15–20. [Google Scholar]
- Bedossa, P.; Poynard, T. An alghoritm for the grading of activity in chronic hepatitis C. Hepatology 1996, 24, 289–293. [Google Scholar]
- Ishak, K.; Baptista, A.; Bianchi, L.; Callea, F.; de Groote, J.; Denk, F.G.; Desmet, V.; Korb, G.; MacSween, R.N.M. Histological grading and staging of chronic hepatitis. J. Hepatol 1995, 22, 696–699. [Google Scholar]
| Characteristics | Overall (N = 335) | IL28B CC (N = 117) | IL28B CT/TT (N = 218) | p-value |
|---|---|---|---|---|
| Males ∘ | 197 (59) | 74 (63) | 123 (56) | 0.25 |
| Age, years * | 54 (21–73) | 55 (26–72) | 24 (21–73) | 0.42 |
| Body Mass Index * | 25 (16–36) | 25 (16–35) | 25 (18–36) | 0.91 |
| HCV genotype ∘ | ||||
| 1 | 151 (45) | 43 (37) | 108 (50) | 0.0008 |
| 2 | 99 (30) | 39 (33) | 60 (27) | |
| 3 | 50 (15) | 28 (24) | 22 (10) | |
| 4 | 35 (10) | 7 (6) | 28 (13) | |
| HCV-RNA, IU/mL ** | 774, 308 | 914, 086 | 743, 038 | 0.69 |
| ALT, U/L * | 101 (9–691) | 113 (22–691) | 96 (9–486) | 0.042 |
| γGT, U/L * | 59 (11–633) | 46 (12–633) | 67 (11–502) | < 0.0001 |
| Cholesterol, mg/dL * | 172 (60–320) | 174 (81–265) | 170 (60–320) | 0.08 |
| Tryglicerides, mg/dL * | 92 (40–380) | 86 (40–340) | 95 (65–380) | 0.13 |
| F3/F4 ∘ | 138 (41) | 48 (41) | 90 (41) | 1.0 |
| A2/A3 ∘ | 167 (50) | 59 (50) | 108 (46) | 0.9 |
| HCV genotype | Histological features | Univariate analysis | Multivariate analysis | OR (95% CI) |
|---|---|---|---|---|
| HCV–1 | Portal inflammation | Age | Age | 1.04 (1.01–1.07) |
| IL28B CC | IL28B CC | 3.24 (1.23–8.51) | ||
| Steatosis | Age | γGT | 1.02 (1.01–1.03) | |
| Body weight | Body Mass Index | 1.17 (1.03–1.33) | ||
| Body Mass Index | HOMA Index | 2.10 (1.03–4.27) | ||
| AST | Tryglicerides | 1.01 (1.00–1.02) | ||
| γGT | - | - | ||
| IL28B CT/TT | - | - | ||
| Diabetes | - | - | ||
| Glucose | - | - | ||
| HOMA Index | - | - | ||
| Tryglicerides | - | - | ||
| HCV–2 | Lobular inflammation | AST | AST | 1.01 (1.00–1.01) |
| ALT | - | - | ||
| γGT | - | - | ||
| IL28B CC | - | - | ||
| HCV–3 | Fibrosis | AST | AST | 1.07 (1.02–1.11) |
| ALT | - | - | ||
| IL28B CC | - | - | ||
| Glucose | - | - | ||
| HOMA Index | - | - | ||
| Grading | HCV-1 | HCV-2 | HCV-3 | HCV-4 | ||||
|---|---|---|---|---|---|---|---|---|
| CC (n = 43) | CT/TT (n = 108) | CC (n = 39) | CT/TT (n = 60) | CC (n = 28) | CT/TT (n = 22) | CC (n = 7) | CT/TT (n = 28) | |
| METAVIR A ≥ 2 | 49% | 50% | 54% | 48% | 43% | 45% | 71% | 54% |
| Ishak | ||||||||
| Piecemeal ≥ 3 | 23% | 13% | 28% | 18% | 14% | 10% | 14% | 18% |
| Portal ≥ 3 | 86% | 63% | 69% | 65% | 50% | 55% | 71% | 46% |
| Lobular ≥ 3 | 19% | 19% | 44% | 23% | 21% | 36% | 71% | 29% |
| Confluent ≥ 3 | 23% | 5% | 18% | 7% | 14% | 5% | 0 | 14% |
© 2014 by the authors; licensee MDPI, Basel, Switzerland This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution license (http://creativecommons.org/licenses/by/3.0/).
Share and Cite
D'Ambrosio, R.; Aghemo, A.; De Francesco, R.; Rumi, M.G.; Galmozzi, E.; De Nicola, S.; Cheroni, C.; Clark, P.J.; Ronchi, G.; Lampertico, P.; et al. The Association of Il28b Genotype with the Histological Features of Chronic Hepatitis C Is HCV Genotype Dependent. Int. J. Mol. Sci. 2014, 15, 7213-7224. https://doi.org/10.3390/ijms15057213
D'Ambrosio R, Aghemo A, De Francesco R, Rumi MG, Galmozzi E, De Nicola S, Cheroni C, Clark PJ, Ronchi G, Lampertico P, et al. The Association of Il28b Genotype with the Histological Features of Chronic Hepatitis C Is HCV Genotype Dependent. International Journal of Molecular Sciences. 2014; 15(5):7213-7224. https://doi.org/10.3390/ijms15057213
Chicago/Turabian StyleD'Ambrosio, Roberta, Alessio Aghemo, Raffaele De Francesco, Maria Grazia Rumi, Enrico Galmozzi, Stella De Nicola, Cristina Cheroni, Paul J. Clark, Guido Ronchi, Pietro Lampertico, and et al. 2014. "The Association of Il28b Genotype with the Histological Features of Chronic Hepatitis C Is HCV Genotype Dependent" International Journal of Molecular Sciences 15, no. 5: 7213-7224. https://doi.org/10.3390/ijms15057213
APA StyleD'Ambrosio, R., Aghemo, A., De Francesco, R., Rumi, M. G., Galmozzi, E., De Nicola, S., Cheroni, C., Clark, P. J., Ronchi, G., Lampertico, P., & Colombo, M. (2014). The Association of Il28b Genotype with the Histological Features of Chronic Hepatitis C Is HCV Genotype Dependent. International Journal of Molecular Sciences, 15(5), 7213-7224. https://doi.org/10.3390/ijms15057213




