Leucyl-tRNA Synthetase Regulates Lactation and Cell Proliferation via mTOR Signaling in Dairy Cow Mammary Epithelial Cells
Abstract
:1. Introduction
2. Results and Discussion
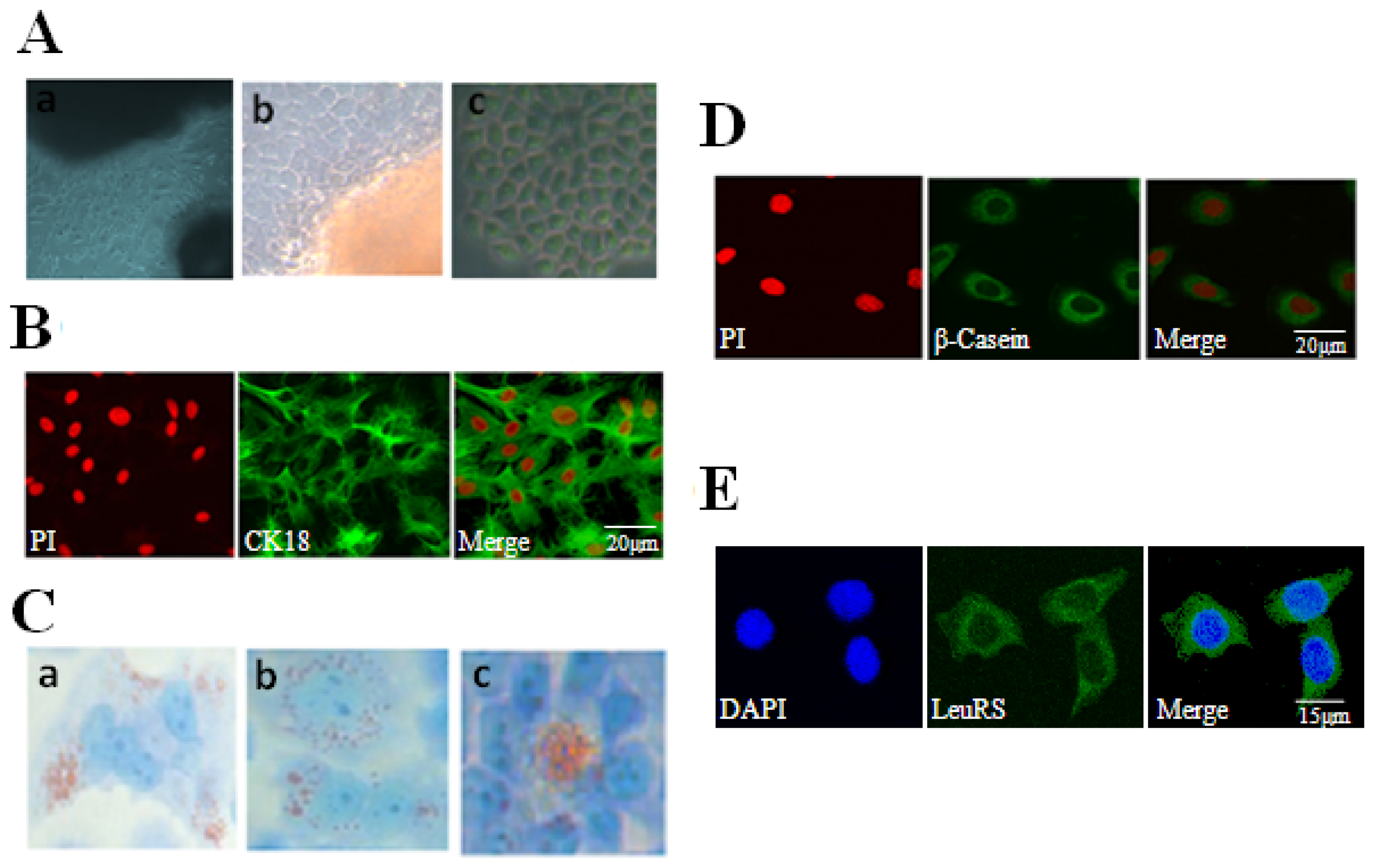
2.1. Localization and Expression of LeuRS in Mammary Gland Tissues
2.2. Culture and Identification of DCMECs and Cytolocalization of LeuRS
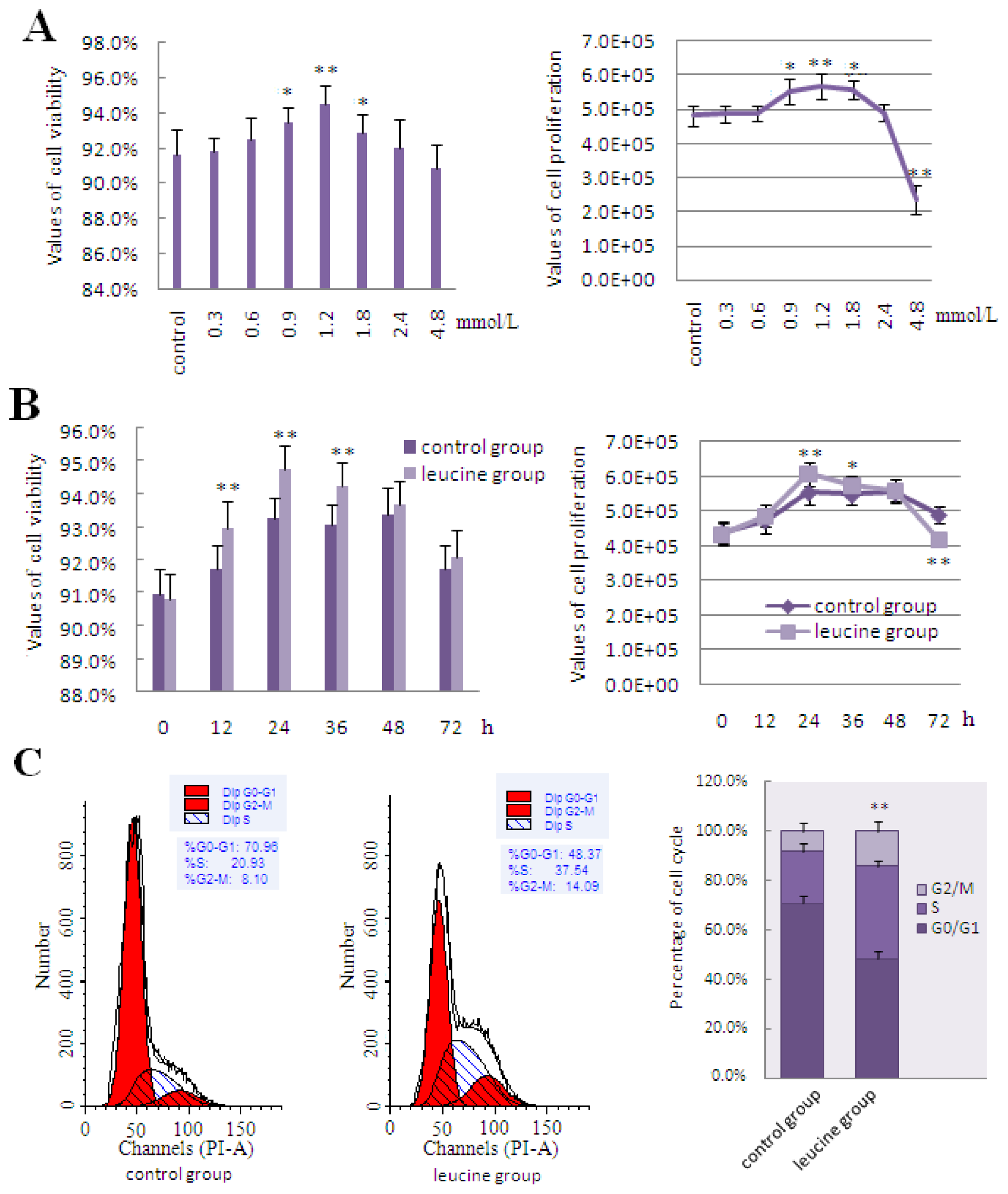
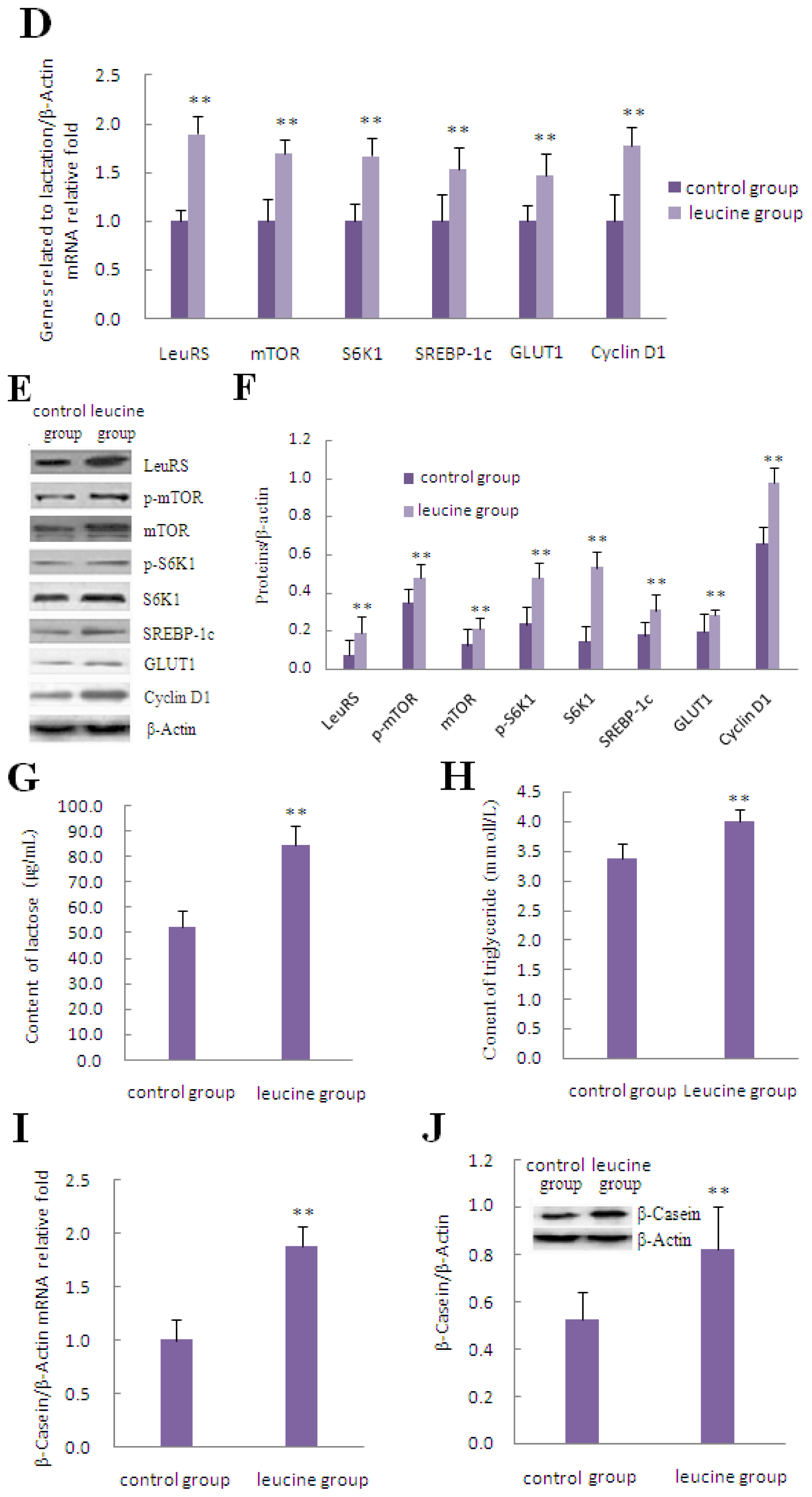
2.3. The Effect of Leucine on LeuRS to Regulate Cell Growth and Proteins in mTOR Signaling in DCMECs
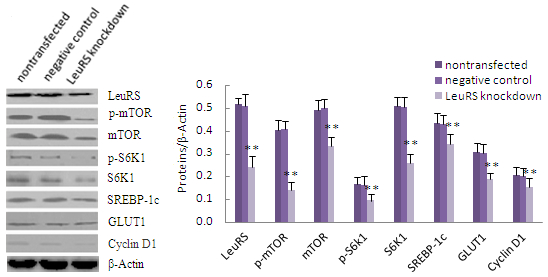
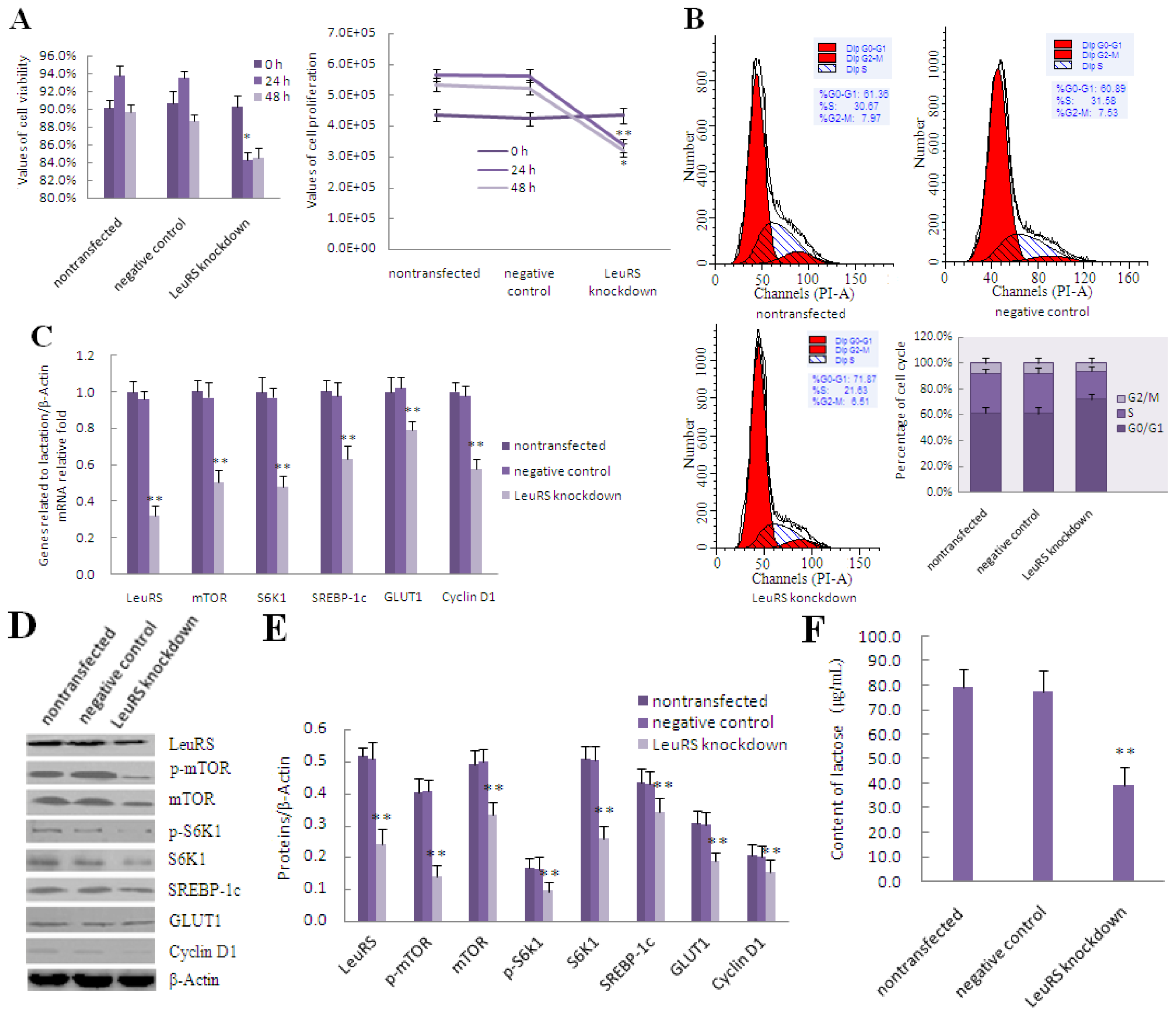
2.4. LeuRS Knockdown Reduced Cell Growth, the Expression of Lactation-Associated Proteins, and Milk Synthesis
2.5. Discussion
3. Experimental Section
3.1. Cell Preparation and Treatments
3.2. Cell Viability and Cell Proliferation Assay
3.3. Oil Red O Staining
3.4. Analysis of Cell Cycle Progression by Flow Cytometry
3.5. Small Interfering RNA Transfection
3.6. Immunohistochemistry
3.7. Immunofluorescence
3.8. RNA Extraction and Quantitative Real-Time PCR
3.9. Western Blotting Analysis
3.10. Quantitation of Secreted Triglyceride and Lactose in the Culture Medium
3.11. Statistical Analysis
4. Conclusions
Acknowledgments
Conflicts of Interest
- Author ContributionsLina Wang developed the conception and design of the study, analyzed data and co-drafting of the manuscript; Ye-Lin analyzed data and co-drafting of the manuscript; Yanjie Bian performed quantitation of secreted triglyceride and lactose experimental work; Lili Liu participated in collection of the literature review; Li Shao and Lin Lin participated in collection of the experimental samples; Bo Qu performed inimmunohistochemistry experimental work and analyzed data; Feng Zhao performed immunofluorescence experimental work; Xuejun Gao analyzed data and revision of the manuscript; Qingzhang Li suggested the research idea, supervised the project and revision of the manuscript.
References
- Herbein, J.H.; Aiello, R.J.; Eckler, L.I.; Pearson, R.E.; Akers, R.M. Glucagon, insulin, growth hormone, and glucose concentrations in blood plasma of lactating dairy cows. J. Dairy Sci 1985, 68, 320–325. [Google Scholar]
- Schmidt, G.H. Effect of insulin on yield and composition of milk of dairy cows. J. Dairy Sci 1966, 9, 381–385. [Google Scholar]
- Mepham, T.B. Amino acid utilization by lactating mammary gland. J. Dairy Sci 1982, 65, 87–98. [Google Scholar]
- Mackle, T.R.; Dwyer, D.A.; Ingvartsen, K.L.; Chouinard, P.Y.; Ross, D.A.; Bauman, D.E. Evaluation of whole blood and plasma in the interorgan supply of free amino acids for the mammary gland of lactating dairy cows. J. Dairy Sci 2000, 83, 1300–1309. [Google Scholar]
- Bauman, D.E.; Mather, I.H.; Wall, R.J.; Lock, A.L. Major advances associated with the biosynthesis of milk. J. Dairy Sci 2006, 89, 1235–1243. [Google Scholar]
- Jenkins, T.C.; McGuire, M.A. Major advances in nutrition: Impact on milk composition. J. Dairy Sci 2006, 89, 2–10. [Google Scholar]
- Neville, M.C.; McFadden, T.B.; Forsyth, I. Hormonal regulation of mammary differentiation and milk secretion. J. Mammary Gland Biol. Neoplasia 2002, 7, 49–66. [Google Scholar]
- Caron, E.; Ghosh, S.; Matsuoka, Y.; Ashton-Beaucage, D.; Therrien, M.; Lemieux, S.; Perreault, C.; Roux, P.P.; Kitano, H. A comprehensive map of the mTOR signaling network. Mol. Syst. Biol 2010, 108, 1–14. [Google Scholar]
- Cho, C.H. Frontier of epilepsy research-mTOR signaling pathway. Exp. Mol. Med 2011, 43, 231–274. [Google Scholar]
- Jewell, J.L.; Guan, K.L. Nutrient signaling to mTOR and cell growth. Trends Biochem. Sci 2013, 38, 233–242. [Google Scholar]
- Sancak, Y.; Bar-Peled, L.; Zoncu, R.; Markhard, A.L.; Nada, S.; Sabatini, D.M. Ragulator-Rag complex targets mTORC1 to the lysosomal surface and is necessary for its activation by amino acids. Cell 2010, 141, 290–303. [Google Scholar]
- Yuan, H.X.; Xiong, Y.; Guan, K.L. Nutrient sensing, metabolism, and cell growth control. Mol. Cell 2013, 49, 379–387. [Google Scholar]
- Schadewaldt, P.; Wendel, U. Metabolism of branched-chain amino acids in maple syrup urine disease. Eur. J. Pediatr 1997, 156, 62–66. [Google Scholar]
- Proud, C.G. Amino acids and mTOR signalling in anabolic function. Biochem. Soc. Trans 2007, 35, 1187–1190. [Google Scholar]
- Stipanuk, M.H. Leucine and protein synthesis: mTOR and beyond. Nutr. Rev 2007, 65, 122–129. [Google Scholar]
- Kim, M.S.; Wu, K.Y.; Auyeung, V.; Chen, Q.; Gruppuso, P.A.; Phornphutkul, C. Leucine restriction inhibits chondrocyte proliferation and differentiation through mechanisms both dependent and independent of mTOR signaling. Am. J. Physiol. Endocrinol. Metab 2009, 296, 1374–1382. [Google Scholar]
- Durán, R.V.; Hall, M.N. Leucyl-tRNA synthetase: Double duty in amino acid sensing. Cell Res 2012, 68, 1207–1209. [Google Scholar]
- Bonfils, G.; Jaquenoud, M.; Bontron, S.; Ostrowicz, C.; Ungermann, C.; de Virgilio, C. Leucyl-tRNA synthetase controls TORC1 via the EGO complex. Mol. Cell 2012, 2, 105–110. [Google Scholar]
- Han, J.M.; Jeong, S.J.; Park, M.C.; Kim, G.; Kwon, N.H.; Kim, H.K.; Ha, S.H.; Ryu, S.H.; Kim, S. Leucyl-tRNA synthetase is an intracellular leucine sensor for the mTORC1-signaling pathway. Cell 2012, 149, 410–424. [Google Scholar]
- Charles, H.S. Cell Adhesion in manlmary gland biology and neoplasia. J. Mammary Gland Biol. Neoplasia 2003, 8, 375–381. [Google Scholar]
- Naylor, M.J.; Oakes, S.R.; Gardiner-Garden, M.; Harris, J.; Blazek, K.; Ho, T.W.; Li, F.C.; Wynick, D.; Walker, A.M.; Ormandy, C.J. Transcriptional changes underlying the secretory activation phase of mammary gland development. Mol. Endocrinol 2005, 19, 1868–1883. [Google Scholar]
- Wu, C.; Luan, H.; Wang, S.; Zhang, X.; Wang, R.; Jin, L.; Guo, P.; Chen, X. Modulation of lipogenesis and glucose consumption in HepG2 cells and C2C12 myotubes by sophoricoside. Molecules 2013, 18, 15624–15635. [Google Scholar]
- Fujita, S.; Dreyer, H.C.; Drummond, M.J.; Glynn, E.L.; Cadenas, J.G.; Yoshizawa, F.; Volpi, E.; Rasmussen, B.B. Nutrient signaling in the regulation of human muscle protein synthesis. J. Physiol 2007, 582, 813–823. [Google Scholar]
- Pasiakos, S.M.; McClung, H.L.; McClung, J.P.; Margolis, L.M.; Andersen, N.E.; Cloutier, G.J.; Pikosky, M.A.; Rood, J.C.; Fielding, R.A.; Young, A.J. Leucine-enriched essential amino acid supplementation during moderate steady state exercise enhances postexercise muscle protein synthesis. Am. J. Clin. Nutr 2011, 94, 809–818. [Google Scholar]
- Kimball, S.R.; Jefferson, L.S. Signaling pathways and molecular mechanisms through which branched-chain amino acids mediate translational control of protein synthesis. J. Nutr 2006, 136, 227S–231S. [Google Scholar]
- Avruch, J.; Long, X.; Ortiz-Vega, S.; Rapley, J.; Papageorgiou, A.; Dai, N. Amino acid regulation of TOR complex 1. Am. J. Physiol. Endocrinol. Metab 2009, 296, E592–E602. [Google Scholar]
- Nicklin, P.; Bergman, P.; Zhang, B.; Triantafellow, E.; Wang, H.; Nyfeler, B.; Yang, H.; Hild, M.; Kung, C.; Wilson, C.; et al. Bidirectional transport of amino acids regulates mTOR and autophagy. Cell 2009, 136, 521–534. [Google Scholar]
- Wang, X.; Proud, C.G. Nutrient control of TORC1, a cell-cycle regulator. Trends Cell Biol 2009, 19, 260–267. [Google Scholar]
- Ktistakis, N.T.; Manifava, M.; Schoenfelder, P.; Rotondo, S. How phosphoinositide 3-phosphate controls growth downstream of amino acids and autophagy downstream of amino acid withdrawal. Biochem. Soc. Trans 2012, 40, 37–43. [Google Scholar]
- Gulati, P.; Gaspers, L.D.; Dann, S.G.; Joaquin, M.; Nobukuni, T.; Natt, F.; Kozma, S.C.; Thomas, A.P.; Thomas, G. Amino acids activate mTOR complex 1 via Ca2+/CaM signaling to hVps34. Cell Metab 2008, 3, 747–755. [Google Scholar]
- Um, S.H.; D’Alessio, D.; Thomas, G. Nutrient overload, insulin resistance, and ribosomal protein S6 kinase 1, S6K1. Cell Metab 2006, 3, 393–402. [Google Scholar]
- Dann, S.G.; Selvaraj, A.; Thomas, G. mTOR Complex1-S6K1 signaling: At the crossroads of obesity, diabetes and cancer. Trends Mol. Med 2007, 13, 252–259. [Google Scholar]
- Sancak, Y.; Peterson, T.R.; Shaul, Y.D.; Lindquist, R.A.; Thoreen, C.C.; Bar-Peled, L.; Sabatini, D.M. The Rag GTPases bind raptor and mediate amino acid signaling to mTORC1. Science 2008, 320, 1496–1501. [Google Scholar]
- Kim, E.; Goraksha-Hicks, P.; Li, L.; Neufeld, T.P.; Guan, K.L. Regulation of TORC1 by Rag GTPases in nutrient response. Nat. Cell Biol 2008, 10, 935–945. [Google Scholar]
- Rhoads, E.R.; Grudzien-Nogalska, E. Translational regulation of milk protein synthesis at secretory activation. J. Mammary Gland Biol. Neoplasia 2007, 12, 283–292. [Google Scholar]
- Burgos, S.A.; Dai, M.; Cant, J.P. Nutrient availability and lactogenic hormones regulate mammary protein synthesis through the mammalian target of rapamycin signalling pathway. J. Dairy Sci 2010, 93, 153–161. [Google Scholar]
- Byfield, M.P.; Murray, J.T.; Backer, J.M. hVps34 is a nutrient-regulated lipid kinase required for activation of p70 S6 kinase. J. Biol. Chem 2005, 280, 33076–33082. [Google Scholar]
- Nobukuni, T.; Joaquin, M.; Roccio, M.; Dann, S.G.; Kim, S.Y.; Gulati, P.; Byfield, M.P.; Backer, J.M.; Natt, F.; Bos, J.L.; et al. Amino acids mediate mTOR/raptor signaling through activation of class 3 phosphatidylinositol 3OH-kinase. Proc. Natl. Acad. Sci. USA 2005, 102, 14238–14243. [Google Scholar]
- Santos, S.J.; Aupperlee, M.D.; Xie, J.; Durairaj, S.; Miksicek, R.; Conrad, S.E.; Leipprandt, J.R.; Tan, Y.S.; Schwartz, R.C.; Haslam, S.Z. Progesterone receptor A-regulated gene expression in mammary organoid cultures. J. Steroid Biochem. Mol. Biol 2009, 115, 161–172. [Google Scholar]
- Bionaz, M.; Loor, J.J. Gene networks driving bovine mammary protein synthesis during the lactation cycle. Bioinform. Biol. Insights 2011, 5, 83–98. [Google Scholar]
- Lin, X.; Luo, J.; Zhang, L.; Wang, W.; Gou, D. MiR-103 Controls Milk fat accumulation in goat (Capra hircus) mammary gland during lactation. PLoS One 2013, 11, e79258. [Google Scholar]
- Nishitani, S.; Matsumura, T.; Fujitani, S.; Sonaka, I.; Miura, Y.; Yagasaki, K. Leucine promotes glucose uptake in skeletal muscles of rats. Biochem. Biophys. Res. Commun 2002, 295, 693–696. [Google Scholar]
- Burnol, A.F.; Letirque, A.; Loizeau, M.; Postic, C.; Girard, J. Glucose transporter expression in rat mammary gland. Biochem. J 1990, 270, 277–279. [Google Scholar]
- Camps, M.; Vilaro, S.; Testar, X.; Palacín, M.; Zorzano, A. High and polarized expression of GLUT1 glucose transporters inepithelial cells from mammary gland: Acute down regulation of GLUT1 carriers by weaning. Endocrinology 1994, 134, 924–934. [Google Scholar]
- Bionaz, M.; Loor, J.J. Gene networks driving bovine milk fat synthesis during the lactation cycle. BMC Gen 2008, 9, 83–98. [Google Scholar]
- Yecies, J.L.; Zhang, H.H.; Menon, S.; Liu, S.; Yecies, D.; Lipovsky, A.I.; Gorgun, C.; Kwiatkowski, D.J.; Hotamisligil, G.S.; Lee, C.H.; et al. Akt stimulates hepatic SREBP1c and lipogenesis through parallel mTORC1-dependent and independent pathways. Cell Metab 2011, 14, 21–32. [Google Scholar]
- Bakan, I.; Laplante, M. Connecting mTORC1 signaling to SREBP-1 activation. Curr. Opin. Lipidol 2012, 23, 226–234. [Google Scholar]
- Tong, H.L.; Gao, X.J.; Li, Q.Z. Metabolic regulation of mammary gland epithelial cells of dairy cow by galactopoietic compound isolated from Vaccariae segetalis. Agric. Sci. China 2011, 10, 1106–1116. [Google Scholar]
- Li, H.-M.; Wang, C.-M.; Li, Q.-Z.; Gao, X.-J. MiR-15a decreases bovine mammary epithelial cell viability and lactation and regulates growth hormone receptor expression. Molecules 2012, 17, 12037–12048. [Google Scholar]
- Kampa-Schittenhelm, K.M.; Heinrich, M.C.; Akmut, F.; Rasp, K.H.; Illing, B.; Döhner, H.; Döhner, K.; Schittenhelm, M.M. Cell cycle-dependent activity of the novel dual PI3K-MTORC1/2 inhibitor NVP-BGT226 in acute leukemia. Mol. Cancer 2013, 12. [Google Scholar] [CrossRef]
- Cui, Y.-J.; Li, Q.-Z.H. Expression and functions of fibroblast growth factor 10 in the mouse mammary gland. Int. J. Mol. Sci 2013, 14, 4094–4105. [Google Scholar]
- Chan, C.-B.; Liu, X.; Ensslin, M.A.; Dillehay, D.L.; Ormandy, C.J.; Sohn, P.; Serra, R.; Ye, K. PIKE-A is required for prolactin-mediated STAT5a activation in mammary gland development. EMBO J 2010, 29, 956–968. [Google Scholar]
- Lu, L.M.; Li, Q.Z.; Huang, J.G.; Gao, X.J. Proteomic and functional analyses reveal MAPK1 regulates milk protein synthesis. Molecules 2013, 18, 263–275. [Google Scholar]
- Huang, Y.-L.; Zhao, F.; Luo, C.-C.; Zhang, X.; Si, Y.; Sun, Z.; Zhang, L.; Li, Q.-Z.; Gao, X.-J. SOCS3-Mediated blockade reveals major contribution of JAK2/STAT5 signaling pathway to lactation and proliferation of dairy cow mammary epithelial cellsin vitro. Molecules 2013, 18, 12987–13002. [Google Scholar]
- Huang, J.-G.; Gao, X.-J.; Li, Q.-Z.; Lu, L.-M.; Liu, R.; Luo, C.-C.; Wang, J.-L.; Qiao, B.; Jin, X. Proteomic analysis of the nuclear phosphorylated proteins in bovine mammary epithelial cells treated with estrogen. In Vitro Cell. Dev. Biol. Anim 2012, 48, 449–457. [Google Scholar]
- Luo, Z.-D.; Ma, L.-Q.; Zhao, Z.-G.; He, H.-B.; Yang, D.-C.; Feng, X.-L.; Ma, S.-T.; Chen, X.-P.; Zhu, T.-Q.; Cao, T.-B.; et al. TRPV1 activation improves exercise endurance and energy metabolism through PGC-1α upregulation in mice. Cell Res 2011, 205, 551–564. [Google Scholar]


© 2014 by the authors; licensee MDPI, Basel, Switzerland This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution license (http://creativecommons.org/licenses/by/3.0/).
Share and Cite
Wang, L.; Lin, Y.; Bian, Y.; Liu, L.; Shao, L.; Lin, L.; Qu, B.; Zhao, F.; Gao, X.; Li, Q. Leucyl-tRNA Synthetase Regulates Lactation and Cell Proliferation via mTOR Signaling in Dairy Cow Mammary Epithelial Cells. Int. J. Mol. Sci. 2014, 15, 5952-5969. https://doi.org/10.3390/ijms15045952
Wang L, Lin Y, Bian Y, Liu L, Shao L, Lin L, Qu B, Zhao F, Gao X, Li Q. Leucyl-tRNA Synthetase Regulates Lactation and Cell Proliferation via mTOR Signaling in Dairy Cow Mammary Epithelial Cells. International Journal of Molecular Sciences. 2014; 15(4):5952-5969. https://doi.org/10.3390/ijms15045952
Chicago/Turabian StyleWang, Lina, Ye Lin, Yanjie Bian, Lili Liu, Li Shao, Lin Lin, Bo Qu, Feng Zhao, Xuejun Gao, and Qingzhang Li. 2014. "Leucyl-tRNA Synthetase Regulates Lactation and Cell Proliferation via mTOR Signaling in Dairy Cow Mammary Epithelial Cells" International Journal of Molecular Sciences 15, no. 4: 5952-5969. https://doi.org/10.3390/ijms15045952