Molecular Cloning, Structural Analysis and Tissue Expression of Protein Phosphatase 3 Catalytic Subunit Alpha Isoform (PPP3CA) Gene in Tianfu Goat Muscle
Abstract
:1. Introduction
2. Results and Discussion
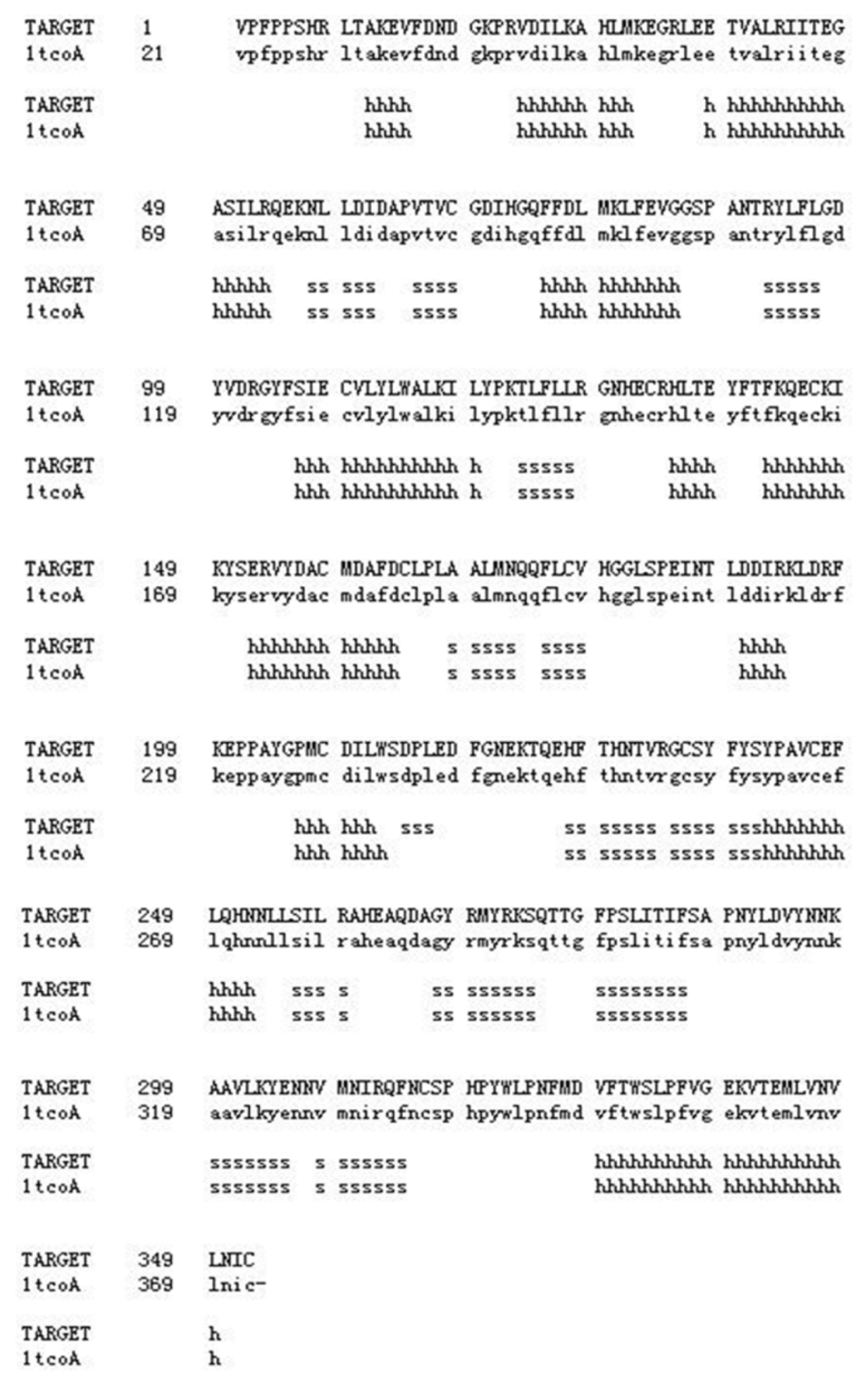
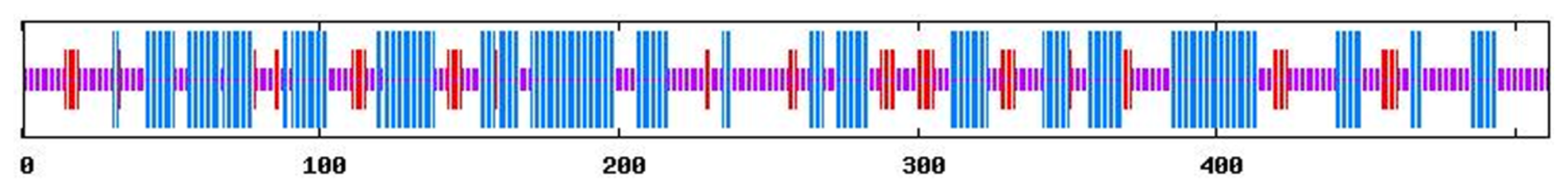
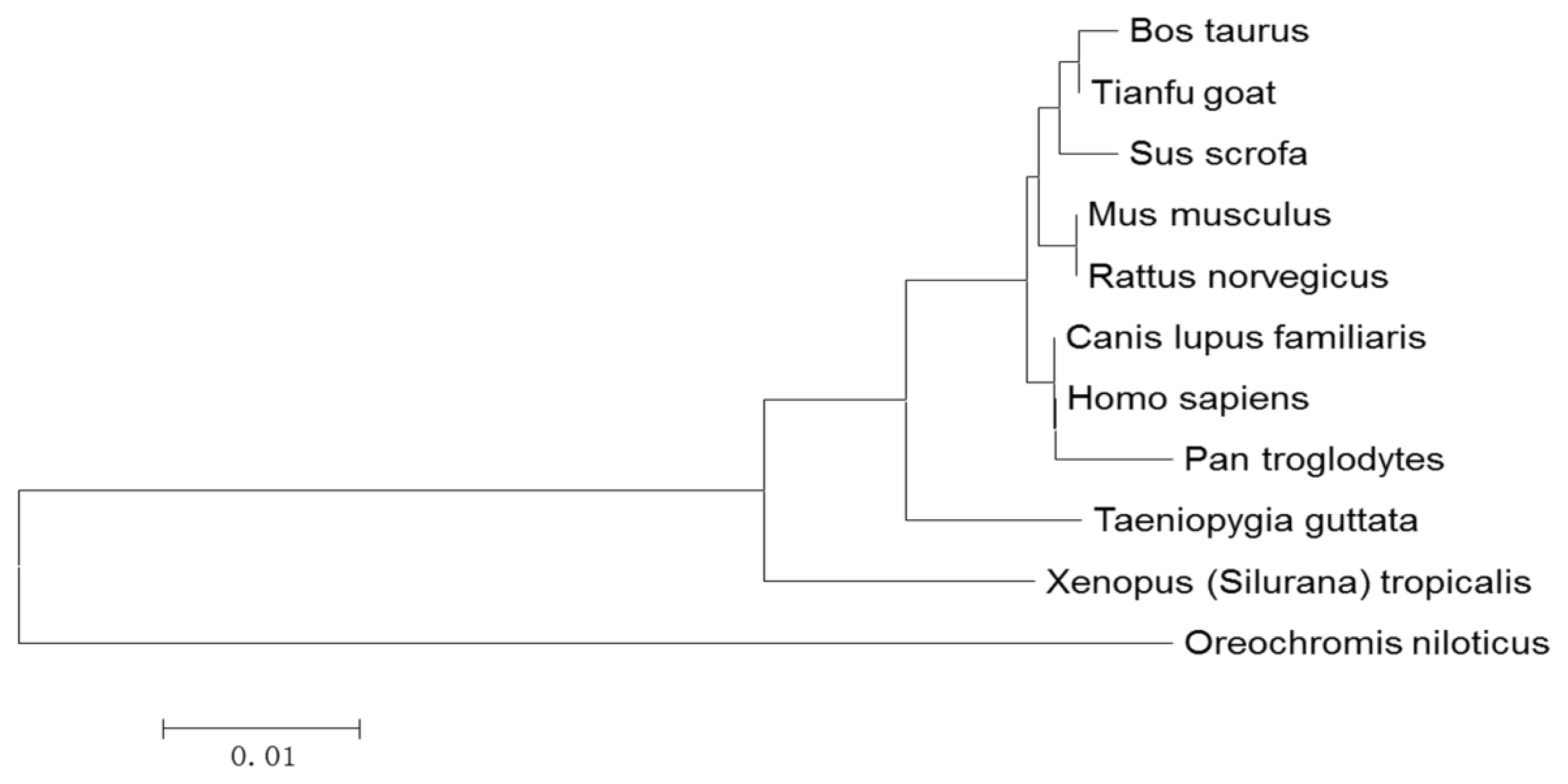
2.1. Summary of Characteristics of Tianfu Goat PPP3CA Sequences and Structures
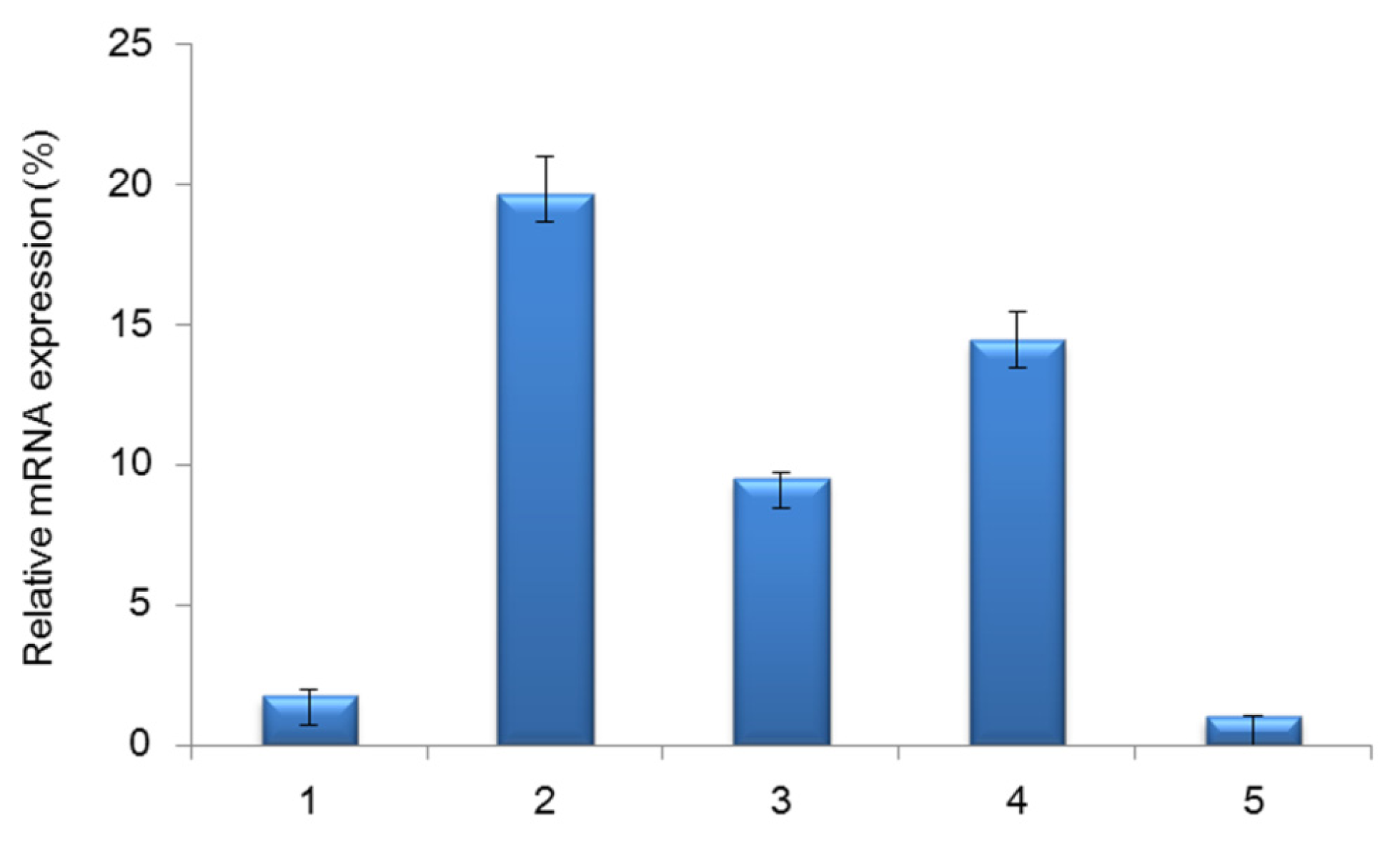
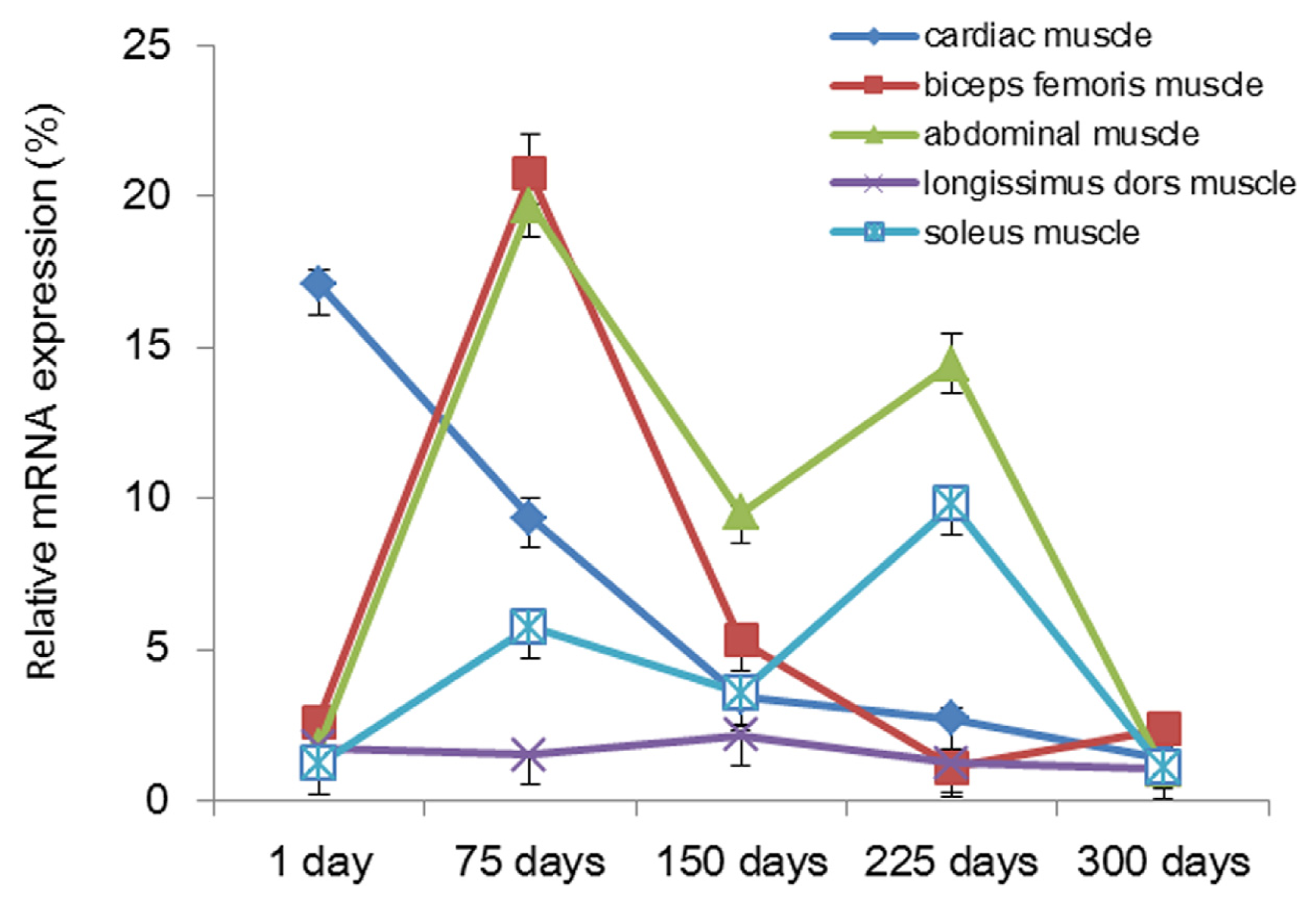
2.2. Analysis of mRNA Expression of Tianfu Goat PPP3CA in Different Muscle
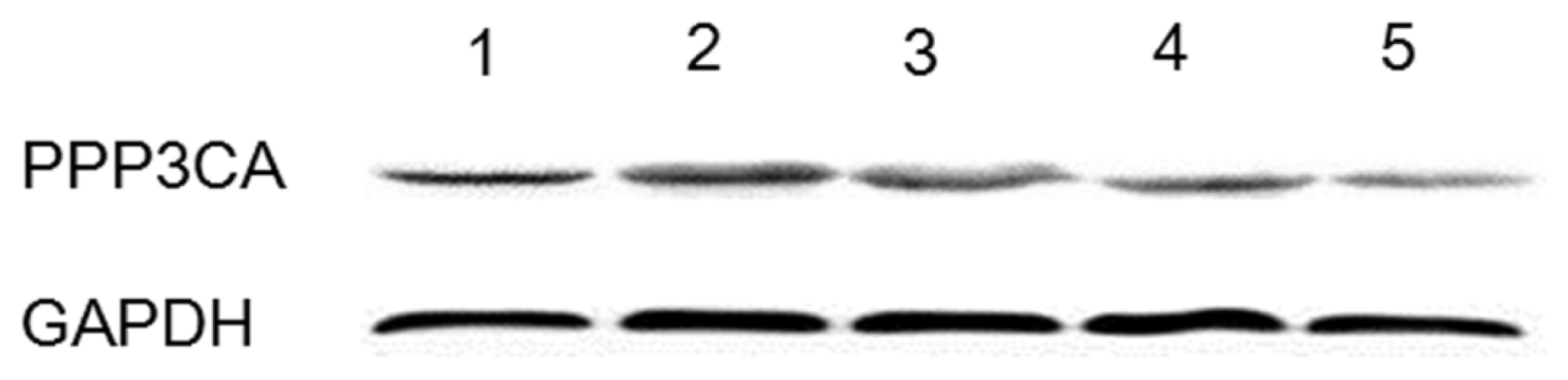
2.3. Western Blotting Analysis of Protein Expression of Tianfu Goat PPP3CA
3. Experimental Section
3.1. Animals and Sample Collection
3.2. Total RNA Isolation and Synthesis of cDNA
3.3. Cloning of the Tianfu Goat PPP3CA Gene
3.4. Sequence Analysis
3.5. RT-qPCR Analysis of PPP3CA Gene mRNA Expression
3.6. Western Blotting Analyses of PPP3CA Protein Expression
3.7. Statistical Analysis
4. Conclusions
Supplementary Information
ijms-15-02346-s001.pdfAcknowledgments
Conflicts of Interest
- Author ContributionsL.W. and J.M. designed the experiment under the supervision of G.X.; L.W., J.M. and N.W. performed the experiments and analyzed the data with the advice of G.X. and D.W.; L.W. and J.M. discussed the results; G.X. and D.W. gave conceptual advice; W.L. wrote the manuscript; all authors commented on the manuscript at all stages.
References
- Gentry, J.; McGlone, J.; Blanton, J.; Miller, M. Impact of spontaneous exercise on performance, meat quality, and muscle fiber characteristics of growing/finishing pigs. J. Anim. Sci 2002, 80, 2833–2839. [Google Scholar]
- Karlsson, A.H.; Klont, R.E.; Fernandez, X. Skeletal muscle fibres as factors for pork quality. Livest. Prod. Sci 1999, 60, 255–269. [Google Scholar]
- Klont, R.; Brocks, L.; Eikelenboom, G. Muscle fibre type and meat quality. Meat Sci 1998, 49, S219–S229. [Google Scholar]
- Lefaucheur, L.; Milan, D.; Ecolan, P.; Le Callennec, C. Myosin heavy chain composition of different skeletal muscles in Large White and Meishan pigs. J. Anim. Sci 2004, 82, 1931–1941. [Google Scholar]
- Argüello, A.; López-Fernández, J.L.; Rivero, J.L.L. Limb myosin heavy chain isoproteins and muscle fiber types in the adult goat (Capra hircus). Anat. Rec 2001, 264, 284–293. [Google Scholar]
- Ryu, Y.; Kim, B. The relationship between muscle fiber characteristics, postmortem metabolic rate, and meat quality of pig longissimus dorsi muscle. Meat Sci 2005, 71, 351–357. [Google Scholar]
- Rusnak, F.; Mertz, P. Calcineurin: Form and function. Physiol. Rev 2000, 80, 1483–1521. [Google Scholar]
- Olson, E.N.; Williams, R.S. Remodeling muscles with calcineurin. Bioessays 2000, 22, 510–519. [Google Scholar]
- Serrano, A.L.; Murgia, M.; Pallafacchina, G.; Calabria, E.; Coniglio, P.; Lømo, T.; Schiaffino, S. Calcineurin controls nerve activity-dependent specification of slow skeletal muscle fibers but not muscle growth. Proc. Natl. Acad. Sci. USA 2001, 98, 13108–13113. [Google Scholar]
- Chin, E.R.; Olson, E.N.; Richardson, J.A.; Yang, Q.; Humphries, C.; Shelton, J.M.; Wu, H.; Zhu, W.; Bassel-Duby, R.; Williams, R.S. A calcineurin-dependent transcriptional pathway controls skeletal muscle fiber type. Genes Dev 1998, 12, 2499–2509. [Google Scholar]
- Schiaffino, S.; Serrano, A. Calcineurin signaling and neural control of skeletal muscle fiber type and size. Trends Pharmacol. Sci 2002, 23, 569–575. [Google Scholar]
- Friday, B.B.; Horsley, V.; Pavlath, G.K. Calcineurin activity is required for the initiation of skeletal muscle differentiation. J. Cell Biol 2000, 149, 657–666. [Google Scholar]
- Da Costa, N.; Edgar, J.; Ooi, P.-T.; Su, Y.; Meissner, J.D.; Chang, K.-C. Calcineurin differentially regulates fast myosin heavy chain genes in oxidative muscle fibre type conversion. Cell Tissue Res 2007, 329, 515–527. [Google Scholar]
- Mitchell, P.O.; Mills, S.T.; Pavlath, G.K. Calcineurin differentially regulates maintenance and growth of phenotypically distinct muscles. Am. J. Physiol. Cell Physiol 2002, 282, C984–C992. [Google Scholar]
- Westerblad, H.; Allen, D. Changes of myoplasmic calcium concentration during fatigue in single mouse muscle fibers. J. Gen. Physiol 1991, 98, 615–635. [Google Scholar]
- Fiems, L.O. Double muscling in cattle: Genes, husbandry, carcasses and meat. Animals 2012, 2, 472–506. [Google Scholar]
- Guerini, D. Calcineurin: Not just a simple protein phosphatase. Biochem. Biophys. Res. Commun 1997, 235, 271–275. [Google Scholar]
- Parsons, S.A.; Millay, D.P.; Wilkins, B.J.; Bueno, O.F.; Tsika, G.L.; Neilson, J.R.; Liberatore, C.M.; Yutzey, K.E.; Crabtree, G.R.; Tsika, R.W. Genetic loss of calcineurin blocks mechanical overload-induced skeletal muscle fiber type switching but not hypertrophy. J. Biol. Chem 2004, 279, 26192–26200. [Google Scholar]
- Parsons, S.A.; Wilkins, B.J.; Bueno, O.F.; Molkentin, J.D. Altered skeletal muscle phenotypes in calcineurin Aα and Aβ gene-targeted mice. Mol. Cell Biol 2003, 23, 4331–4343. [Google Scholar]
- Kuno, T.; Mukai, H.; Ito, A.; Chang, C.D.; Kishima, K.; Saito, N.; Tanaka, C. Distinct cellular expression of calcineurin Aα and Aβ in rat brain. J. Neurochem 1992, 58, 1643–1651. [Google Scholar]
- Blough, E.; Dineen, B.; Esser, K. Extraction of nuclear proteins from striated muscle tissue. Biotechniques 1999, 26, 202–206. [Google Scholar]
- Naya, F.J.; Mercer, B.; Shelton, J.; Richardson, J.A.; Williams, R.S.; Olson, E.N. Stimulation of slow skeletal muscle fiber gene expression by calcineurin in vivo. J. Biol. Chem. 2000, 275, 4545–4548. [Google Scholar]
- Tůmová, L.; Petr, J.; Žalmanová, T.; Chmelíková, E.; Kott, T.; Tichovská, H.; Kučerová-Chrpová, V.; Hošková, K.; Jílek, F. Calcineurin expression and localisation during porcine oocyte growth and meiotic maturation. Anim. Reprod. Sci 2013, 141, 154–163. [Google Scholar]
- Klee, C.B.; Ren, H.; Wang, X. Regulation of the calmodulin-stimulated protein phosphatase, calcineurin. J. Biol. Chem 1998, 273, 13367–13370. [Google Scholar]
- Kincaid, R.L.; Nightingale, M.S.; Martin, B.M. Characterization of a cDNA clone encoding the calmodulin-binding domain of mouse brain calcineurin. Proc. Natl. Acad. Sci. USA 1988, 85, 8983–8987. [Google Scholar]
- Depreux, F.; Scheffler, J.; Grant, A.; Bidwell, C.; Gerrard, D. Molecular cloning and characterization of porcine calcineurin-α subunit expression in skeletal muscle. J. Anim. Sci 2010, 88, 562–571. [Google Scholar]
- Wang, M.; Yi, H.; Guerini, D.; Klee, C.; McBride, O. Calcineurin A α (PPP3CA), calcineurin A β (PPP3CB) and calcineurin B (PPP3R1) are located on human chromosomes 4, 10q21 → q22 and 2p16 → p15 respectively. Cytogenet. Genome Res 1996, 72, 236–241. [Google Scholar]
- Pette, D.; Staron, R.S. Myosin isoforms, muscle fiber types, and transitions. Microsc. Res. Tech 2000, 50, 500–509. [Google Scholar]
- McCoard, S.; McNabb, W.; Peterson, S.; McCutcheon, S.; Harris, P. Muscle growth, cell number, type and morphometry in single and twin fetal lambs during mid to late gestation. Reprod. Fertil. Dev 2001, 12, 319–327. [Google Scholar]
- Tauveron, I.; Larbaud, D.; Champredon, C.; Debras, E.; Tesseraud, S.; Bayle, G.; Bonnet, Y.; Thieblot, P.; Grizard, J. Effect of hyperinsulinemia and hyperaminoacidemia on muscle and liver protein synthesis in lactating goats. Am. J. Phys. Endocrinol. Metab 1994, 267, E877–E885. [Google Scholar]
- Yan, Z.; Booth, F.W. Cytochrome c promoter activity in soleus and white vastus lateralis muscles in rats. J. Appl. Phys 1998, 85, 973–978. [Google Scholar]
- Selvakumar, P.; Lakshmikuttyamma, A.; Anderson, D.H.; Sharma, R.K. Molecular cloning, expression, purification and characterization of calcineurin from bovine cardiac muscle. Biochimie 2005, 87, 975–983. [Google Scholar]
- Xu, H.; Xu, G.; Wang, D.; Ma, J.; Wan, L. Molecular cloning, sequence identification and expression analysis of novel caprine MYLPF gene. Mol. Biol. Rep 2013, 40, 2565–2572. [Google Scholar]



| Primer | Sequence fragment | Product length (bp) | Application |
|---|---|---|---|
| PPP3CA-F | GAACCACCTGCTTATGGACCTAT | 244 | Expression |
| PPP3CA-R | AAGGGAAGCCTGTTGTTTGG | 244 | Expression |
| GAPDH-F | GTCACCAACTGGGACGACA | 118 | RT-qPCR |
| GAPDH-R | AGGCGTACAGGGACAGCA | 118 | RT-qPCR |
© 2014 by the authors; licensee MDPI, Basel, Switzerland This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution license (http://creativecommons.org/licenses/by/3.0/).
Share and Cite
Wan, L.; Ma, J.; Xu, G.; Wang, D.; Wang, N. Molecular Cloning, Structural Analysis and Tissue Expression of Protein Phosphatase 3 Catalytic Subunit Alpha Isoform (PPP3CA) Gene in Tianfu Goat Muscle. Int. J. Mol. Sci. 2014, 15, 2346-2358. https://doi.org/10.3390/ijms15022346
Wan L, Ma J, Xu G, Wang D, Wang N. Molecular Cloning, Structural Analysis and Tissue Expression of Protein Phosphatase 3 Catalytic Subunit Alpha Isoform (PPP3CA) Gene in Tianfu Goat Muscle. International Journal of Molecular Sciences. 2014; 15(2):2346-2358. https://doi.org/10.3390/ijms15022346
Chicago/Turabian StyleWan, Lu, Jisi Ma, Gangyi Xu, Daihua Wang, and Nianlu Wang. 2014. "Molecular Cloning, Structural Analysis and Tissue Expression of Protein Phosphatase 3 Catalytic Subunit Alpha Isoform (PPP3CA) Gene in Tianfu Goat Muscle" International Journal of Molecular Sciences 15, no. 2: 2346-2358. https://doi.org/10.3390/ijms15022346




