Overexpression of Arachis hypogaea AREB1 Gene Enhances Drought Tolerance by Modulating ROS Scavenging and Maintaining Endogenous ABA Content
Abstract
:1. Introduction
2. Results
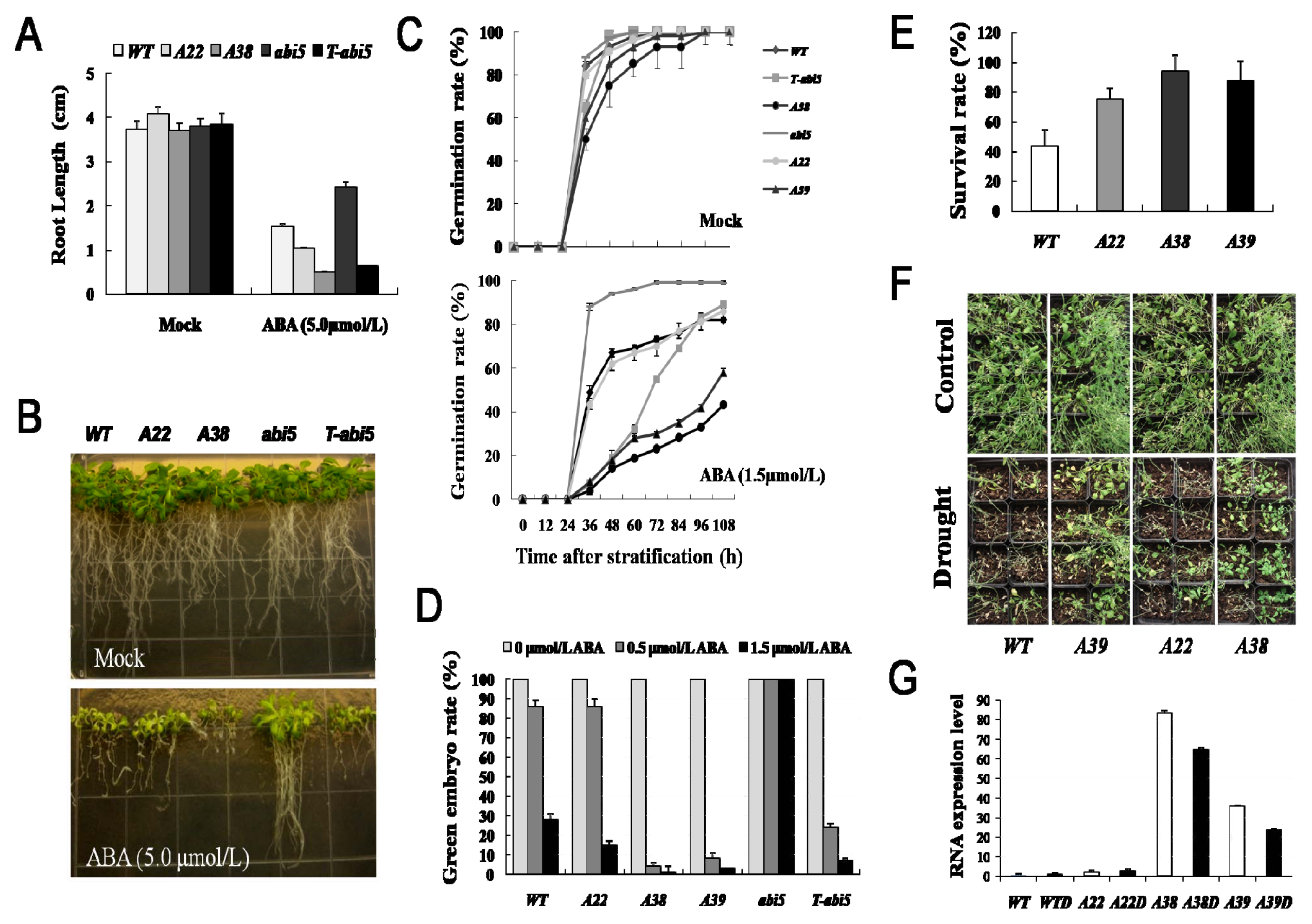
2.1. The AhAREB1 Overexpression Greatly Improves Drought Tolerance and ABA Sensitivity in Transgenic Arabidopsis Plants
2.2. Identification of Target Genes of AhAREB1 Using Genome-Wide Microarray Expression Analysis
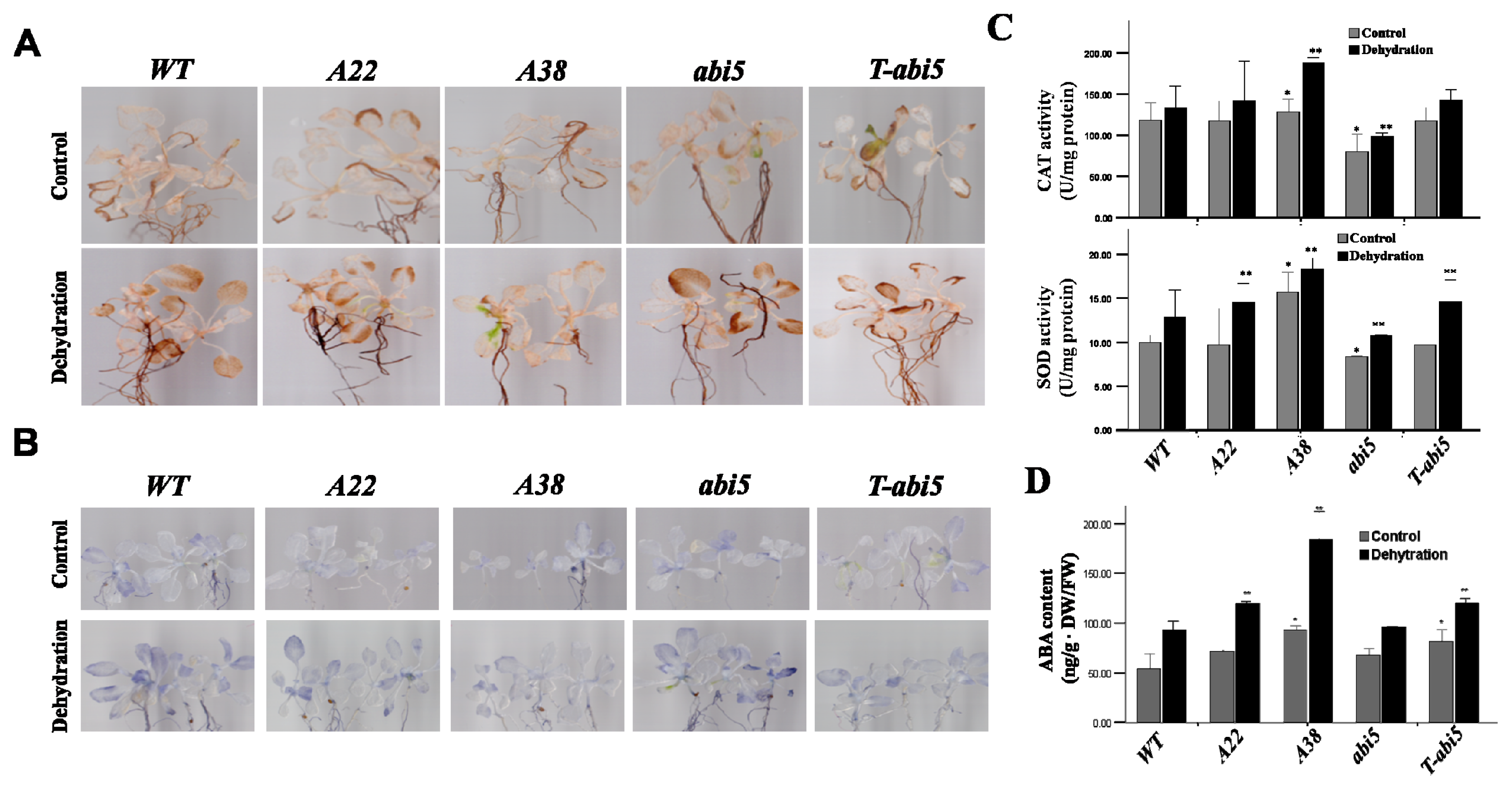
2.3. Effect of Constitutive Expression of AhAREB1 on ROS Levels and ABA Content in Arabidopsis thaliana
3. Discussion
4. Experimental Section
4.1. Plant Materials
4.2. Plasmid Construction and Arabidopsis Transformation
4.3. Assays of Seed Germination and Green Cotyledons; Growth of Roots
4.4. Drought Stress Tolerance Assays
4.5. Microarray Analysis
4.6. Quantitative PCR Assay
4.7. In Situ NBT Staining and Measurement of SOD Activity
4.8. In Situ DAB Staining and Measurement of CAT Activity
4.9. Quantification of ABA Levels
4.10. Subcellular Localization
5. Conclusions
Supplementary File 1
ijms-14-12827-s001.pdfAcknowledgements
Conflict of Interest
References
- Wan, X.R.; Li, L. Regulation of ABA level and water-stress tolerance of Arabidopsis by ectopic expression of a peanut 9-cis-epoxycarotenoid dioxygenase gene. Biochem. Biophys. Res. Commun 2006, 347, 1030–1038. [Google Scholar]
- Yamaguchi-Shinozaki, K.; Shinozaki, K. Transcriptional regulatory networks in cellular responses and tolerance to dehydration and cold stresses. Annu. Rev. Plant Biol 2006, 57, 781–803. [Google Scholar]
- Seki, M.; Umezawa, T.; Urano, K.; Shinozaki, K. Regulatory metabolic networks in drought stress responses. Curr. Opin. Plant Biol 2007, 10, 296–302. [Google Scholar]
- Huh, S.U.; Lee, S.B.; Kim, H.H.; Paek, K.H. ATAF2, a NAC transcription factor, binds to the promoter and regulates NIT2 gene expression involved in auxin biosynthesis. Mol. Cells 2012. [Google Scholar] [CrossRef]
- Sreenivasulu, N.; Harshavardhan, V.T.; Govind, G.; Seiler, C.; Kohli, A. Contrapuntal role of ABA: Does it mediate stress tolerance or plant growth retardation under long-term drought stress? Gene 2012, 506, 265–273. [Google Scholar]
- Bensmihen, S.; Rippa, S.; Lambert, G.; Jublot, D.; Pautot, V.; Granier, F.; Giraudat, J.; Parcy, F. The homologous ABI5 and EEL transcription factors function antagonistically to fine-tune gene expression during late embryogenesis. Plant Cell 2002, 14, 1391–1403. [Google Scholar]
- Zheng, Y.; Schumaker, K.S.; Guo, Y. Sumoylation of transcription factor MYB30 by the small ubiquitin-like modifier E3 ligase SIZ1 mediates abscisic acid response in Arabidopsis thaliana. Proc. Natl. Acad. Sci. USA 2012, 109, 12822–12827. [Google Scholar]
- Kim, S.; Kang, J.Y.; Cho, D.I.; Park, J.H.; Kim, S.Y. ABF2, an ABRE-binding bZIP factor, is an essential component of glucose signaling and its overexpression affects multiple stress tolerance. Plant J 2004, 40, 75–87. [Google Scholar]
- Sakuma, Y.; Maruyama, K.; Qin, F.; Osakabe, Y.; Shinozaki, K. Dual function of an Arabidopsis transcription factor DREB2A in water-stress-responsive and heat-stress-responsive gene expression. Proc. Natl. Acad. Sci. USA 2006, 103, 18822–18827. [Google Scholar]
- Narusaka, Y.; Nakashima, K.; Shinwari, Z.K.; Sakuma, Y.; Furihata, T.; Abe, H.; Narusaka, M.; Shinozaki, K.; Yamaguchi-Shinozaki, K. Interaction between two cis-acting elements, ABRE and DRE, in ABA-dependent expression of Arabidopsis rd29A gene in response to dehydration and high-salinity stresses. Plant. J 2003, 34, 137–148. [Google Scholar]
- Kang, J.Y.; Choi, H.I.; Im, M.Y.; Kim, S.Y. Arabidopsis basic leucine zipper proteins that mediate stress-responsive abscisic acid signaling. Plant. Cell 2002, 14, 343–357. [Google Scholar]
- Kim, J.B.; Kang, J.Y.; Kim, S.Y. Overexpression of a transcription factor regulating ABA-responsive gene expression confers multiple stress tolerance. Plant Biotechnol. J 2004, 2, 459–466. [Google Scholar]
- Fujita, Y.; Fujita, M.; Satoh, R.; Maruyama, K.; Parvez, M.M.; Seki, M.; Hiratsu, K.; Ohme-Takagi, M.; Shinozaki, K.; Yamaguchi-Shinozaki, K. AREB1 is a transcription activator of novel ABRE-dependent ABA signaling that enhances drought stress tolerance in Arabidopsis. Plant Cell 2005, 17, 3470–3488. [Google Scholar]
- Yoshida, T.; Fujita, Y.; Sayama, H.; Kidokoro, S.; Maruyama, K.; Mizoi, J.; Shinozaki, K.; Yamaguchi-Shinozaki, K. AREB1, AREB2, and ABF3 are master transcription factors that cooperatively regulate ABRE-dependent ABA signaling involved in drought stress tolerance and require ABA for full activation. Plant J 2010, 61, 672–685. [Google Scholar]
- Zhang, X.; Wang, L.; Meng, H.; Wen, H.; Fan, Y.; Zhao, J. Maize ABP9 enhances tolerance to multiple stresses in transgenic Arabidopsis by modulating ABA signaling and cellular levels of reactive oxygen species. Plant Mol. Biol 2011, 75, 365–378. [Google Scholar]
- Huang, X.S.; Liu, J.H.; Chen, X.J. Overexpression of PtrABF gene, a bZIP transcription factor isolated from Poncirus trifoliata, enhances dehydration and drought tolerance in tobacco via scavenging ROS and modulating expression of stress-responsive genes. BMC Plant Biol 2010, 10, 230. [Google Scholar]
- Rizhsky, L.; Hallak-Herr, E.; van Breusegem, F.; Rachmilevitch, S.; Barr, J.E.; Rodermel, S.; Inzé, D.; Mittler, R. Double antisense plants lacking ascorbate peroxidase and catalase are less sensitive to oxidative stress than single antisense plants lacking ascorbate peroxidase or catalase. Plant J 2002, 32, 329–342. [Google Scholar]
- Hong, L.; Hu, B.; Liu, X.; He, C.Y.; Yao, Y.; Li, X.L.; Li, L. Molecular cloning and expression analysis of a new stress-related AREB gene from Arachis hypogaea. Biologia. Plantarum 2012. [Google Scholar] [CrossRef]
- Yao, Y.; Liu, X.; Hong, L.; Li, T.F.; Zhu, M.K.; Li, L. Preliminary study on transcriptional activity of AhAREB1 protein. Chin. Sci. Tech. Pap. Online 2012, 5, 408–414. [Google Scholar]
- Liu, Q.; Kasuga, M.; Sakuma, Y.; Abe, H.; Miura, S.; Yamaguchi-Shinozaki, K.; Shinozaki, K. Two transcription factors, DREB1 and DREB2, with an EREBP/AP2 DNA binding domain separate two cellular signal transduction pathways in drought- and low-temperature-responsive gene expression, respectively, in Arabidopsis. Plant Cell 1998, 10, 1391–1406. [Google Scholar]
- Kasuga, M.; Liu, Q.; Miura, S.; Yamaguchi-Shinozaki, K.; Shinozaki, K. Improving plant drought, salt, and freezing tolerance by gene transfer of a single stress-inducible transcription factor. Nat. Biotechnol 1999, 17, 287–291. [Google Scholar]
- Gilmour, S.J.; Sebolt, A.M.; Salazar, M.P.; Everard, J.D.; Thomashow, M.F. Overexpression of the Arabidopsis CBF3 transcriptional activator mimics multiple biochemical changes associated with cold acclimation. Plant. Physiol 2000, 124, 1854–1865. [Google Scholar]
- Hobo, T.; Kowyama, Y.; Hattori, T. A bZIP factor, TRAB1, interacts with VP1 and mediates abscisic acid-induced transcription. Proc. Natl. Acad. Sci. USA 1999, 96, 15348–15353. [Google Scholar]
- Takahashi, S.; Katagiri, T.; Yamaguchi-Shinozaki, K.; ozaki, K. An Arabidopsis gene encoding a Ca2+-binding protein is induced by abscisic acid during dehydration. Plant Cell Physiol 2000, 41, 898–903. [Google Scholar]
- Mao, X.; Zhang, H.; Qian, X.; Li, A.; Zhao, G.; Jing, R. TaNAC2, a NAC-type wheat transcription factor conferring enhanced multiple abiotic stress tolerances in Arabidopsis. J. Exp. Bot 2012, 63, 2933–2946. [Google Scholar]
- Stone, S.L.; William, L.A.; Farmer, L.M.; Vierstra, R.D.; Callis, J. KEEP ON GOING, a RING E3 ligase essential for Arabidopsis growth and development, is involved in abscisic acid signaling. Plant Cell 2006, 18, 3415–3428. [Google Scholar]
- Wang, Y.; Li, L.; Ye, T.; Lu, Y.; Chen, X.; Wu, Y. The inhibitory effect of ABA on floral transition is mediated by ABI5 in Arabidopsis. J. Exp. Bot 2013, 64, 675–684. [Google Scholar]
- Brocard, I.M.; Lynch, T.J.; Finkelstein, R.R. Regulation and role of the Arabidopsis abscisic acid-insensitive 5 gene in abscisic acid, sugar, and stress response. Plant Physiol 2002, 129, 1533–1543. [Google Scholar]
- Levine, A.; Tenhaken, R.; Dixon, R.; Lamb, C. H2O2 from the oxidative burst orchestrates the plant hypersensitive disease resistance response. Cell 1994, 79, 583–593. [Google Scholar]
- Desikan, R.; Reynolds, A.; Hancock, J.T.; Neill, S.J. Harpin and hydrogen peroxide both initiate programmed cell death but have differential effects on defence gene expression in Arabidopsis suspension cultures. Biochem. J 1998, 330, 115–120. [Google Scholar]
- Liu, Y.; Ye, N.; Liu, R.; Chen, M.; Zhang, J. H2O2 mediates the regulation of ABA catabolism and GA biosynthesis in Arabidopsis seed dormancy and germination. J. Exp. Bot 2010, 61, 2979–2990. [Google Scholar]
- Zhu, D.; Scandalios, J.G. Differential accumulation of manganese-superoxide dismutase transcripts in maize in response to abscisic acid and high osmoticum. Plant Physiol 1994, 106, 173–178. [Google Scholar]
- Fryer, M.J.; Ball, L.; Oxborough, K.; Karpinski, S.; Mullineaux, P.M.; Baker, N.R. Control of ascorbate peroxidase 2 expression by hydrogen peroxide and leaf water status during excess light stress reveals a functional organisation of Arabidopsis leaves. Plant J 2003, 33, 691–705. [Google Scholar]
- Kwak, J.M.; Mori, I.C.; Pei, Z.M.; Leonhardt, N.; Torres, M.A.; Dangl, J.L.; Bloom, R.E.; Bodde, S.; Jones, J.D.; Schroeder, J.I. NADPH oxidase AtrbohD and AtrbohF genes function in ROS-dependent ABA signaling in Arabidopsis. EMBO J 2003, 22, 2623–2633. [Google Scholar]
- Jiang, M; Zhang, J. Water stress-induced abscisic acid accumulation triggers the increased generation of reactive oxygen species and up-regulates the activities of antioxidant enzymes in maize leaves. J. Exp. Bot 2002, 53, 2401–2410. [Google Scholar]
- Iuch, S.; Kobayashi, M.; Taji, T.; Naramoto, M.; Seki, M.; Kato, T.; Tabata, S.; Kakubari, Y.; Yamaguchi-Shinozaki, K.; Shinozaki, K. Regulation of drought tolerance by gene manipulation of 9-cis-epoxycarotenoid dioxygenase, a key enzyme in abscisic acid biosynthesis in Arabidopsis. Plant J 2001, 27, 325–333. [Google Scholar]
- Hauser, F.; Waadt, R.; Schroeder, J.I. Evolution of abscisic acid synthesis and signaling mechanisms. Curr. Biol 2011, 21, R346–R355. [Google Scholar]
- Qin, X.; Zeevaart, J.A. The 9-cis-epoxycarotenoid cleavage reaction is the key regulatory step of abscisic acid biosynthesis in water-stressed bean. Proc. Natl. Acad. Sci. USA 1999, 96, 15354–15361. [Google Scholar]
- Park, H.Y.; Seok, H.Y.; Park, B.K.; Kim, S.H.; Goh, C.H.; Lee, B.H.; Lee, C.H.; Moon, Y.H. Overexpression of Arabidopsis ZEP enhances tolerance to osmotic stress. Biochem. Biophys. Res. Commun 2008, 375, 80–85. [Google Scholar]
- Kushiro, T.; Okamoto, M; Nakabayashi, K.; Yamagishi, K.; Kitamura, S.; Asami, T.; Hirai, N.; Koshiba, T.; Kamiya, Y.; Nambara, E. The Arabidopsis cytochrome P450 CYP707A encodes ABA 8′-hydroxylases: Key enzymes in ABA catabolism. EMBO J 2004, 23, 1647–1656. [Google Scholar]
- Zhu, G.; Liu, Y; Ye, N.; Liu, R.; Zhang, J. Involvement of the abscisic acid catabolic gene CYP707A2 in the glucose-induced delay in seed germination and post-germination growth of Arabidopsis. Physiol. Plant 2011, 143, 375–384. [Google Scholar]
- Liu, Y.; Zhang, J. Rapid accumulation of NO regulates ABA catabolism and seed dormancy during imbibition in Arabidopsis. Plant Signal. Behav 2009, 4, 905–907. [Google Scholar]
- Saito, S.; Hirai, N.; Matsumoto, C.; Ohigashi, H.; Ohta, D.; Sakata, K.; Mizutani, M. Arabidopsis CYP707As encode (+)-abscisic acid 8′-hydroxylase, a key enzyme in the oxidative catabolism of abscisic acid. Plant Physiol 2004, 134, 1439–1449. [Google Scholar]
- Umezawa, T.; Okamoto, M.; Kushiro, T.; Nambara, E.; Oono, Y.; Seki, M.; Kobayashi, M.; Koshiba, T.; Kamiya, Y.; Shinozaki, K. CYP707A3, a major ABA 8′-hydroxylase involved in dehydration and rehydration response in Arabidopsis thaliana. Plant J 2006, 46, 171–182. [Google Scholar]
- Hu, B.; Hong, L.; Liu, X.; Xiao, S.N.; Lv, Y.; Li, L. Identification of different ABA biosynthesis sites at seedling and fruiting stages in Arachis hypogaea L following water stress. Plant Growth Regul 2013. [Google Scholar] [CrossRef]
- Hu, B.; Hong, L.; Liu, X.; Xiao, S.N.; Lv, Y.; Li, L. The higher expression and distribution of 9-cis-epoxycarotenoid dioxygenase1 (AhNCED1) from Arachis hypogaea L contribute to tolerance to water stress in a drought-tolerant cultivar. Acta Physiol. Plant 2013, 35, 1667–1674. [Google Scholar]
- Kang, J.; Hwang, J.U.; Lee, M.; Kim, Y.Y.; Assmann, S.M.; Martinoia, E.; Lee, Y. PDR-type ABC transporter mediates cellular uptake of the phytohormone abscisic acid. Proc. Natl. Acad. Sci. USA 2010, 107, 2355–2360. [Google Scholar] [Green Version]
- Liu, X.; Hong, L.; Li, X.Y.; Yao, Y.; Hu, B.; Li, L. Improved drought and salt tolerance in transgenic Arabidopsis overexpressing a NAC transcriptional factor from Arachis hypogaea. Biosci. Biotechnol. Biochem 2011, 75, 443–450. [Google Scholar]
- Hughes, T.R.; Mao, M.; Jones, A.R.; Burchard, J.; Marton, M.J.; Shannon, K.W.; Lefkowitz, S.M.; Ziman, M.; Schelter, J.M.; Meyer, M.R.; et al. Expression profiling using microarrays fabricated by an ink-jet oligonucleotide sythesizer. Nat. Biotechnol 2001, 19, 342–347. [Google Scholar]
- Cheng, Y.; Song, C. Hydrogen peroxide homeostasis and signaling in plant cells. Sci. China C Life Sci 2006, 49, 1–11. [Google Scholar]
- Arenas-Huertero, F.; Arroyo, A.; Zhou, L.; Sheen, J.; León, P. Analysis of Arabidopsis glucose insensitive mutants, gin5 and gin6, reveals a central role of the plant hormone ABA in the regulation of plant vegetative development by sugar. Genes Dev 2000, 14, 2085–2096. [Google Scholar]

© 2013 by the authors; licensee MDPI, Basel, Switzerland This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution license ( http://creativecommons.org/licenses/by/3.0/).
Share and Cite
Li, X.-Y.; Liu, X.; Yao, Y.; Li, Y.-H.; Liu, S.; He, C.-Y.; Li, J.-M.; Lin, Y.-Y.; Li, L. Overexpression of Arachis hypogaea AREB1 Gene Enhances Drought Tolerance by Modulating ROS Scavenging and Maintaining Endogenous ABA Content. Int. J. Mol. Sci. 2013, 14, 12827-12842. https://doi.org/10.3390/ijms140612827
Li X-Y, Liu X, Yao Y, Li Y-H, Liu S, He C-Y, Li J-M, Lin Y-Y, Li L. Overexpression of Arachis hypogaea AREB1 Gene Enhances Drought Tolerance by Modulating ROS Scavenging and Maintaining Endogenous ABA Content. International Journal of Molecular Sciences. 2013; 14(6):12827-12842. https://doi.org/10.3390/ijms140612827
Chicago/Turabian StyleLi, Xiao-Yun, Xu Liu, Yao Yao, Yi-Hao Li, Shuai Liu, Chao-Yong He, Jian-Mei Li, Ying-Ying Lin, and Ling Li. 2013. "Overexpression of Arachis hypogaea AREB1 Gene Enhances Drought Tolerance by Modulating ROS Scavenging and Maintaining Endogenous ABA Content" International Journal of Molecular Sciences 14, no. 6: 12827-12842. https://doi.org/10.3390/ijms140612827



