Innovative Strategy for MicroRNA Delivery in Human Mesenchymal Stem Cells via Magnetic Nanoparticles
Abstract
:1. Introduction
2. Results and Discussion
2.1. Optimization of Transfection Complexes in hMSCs
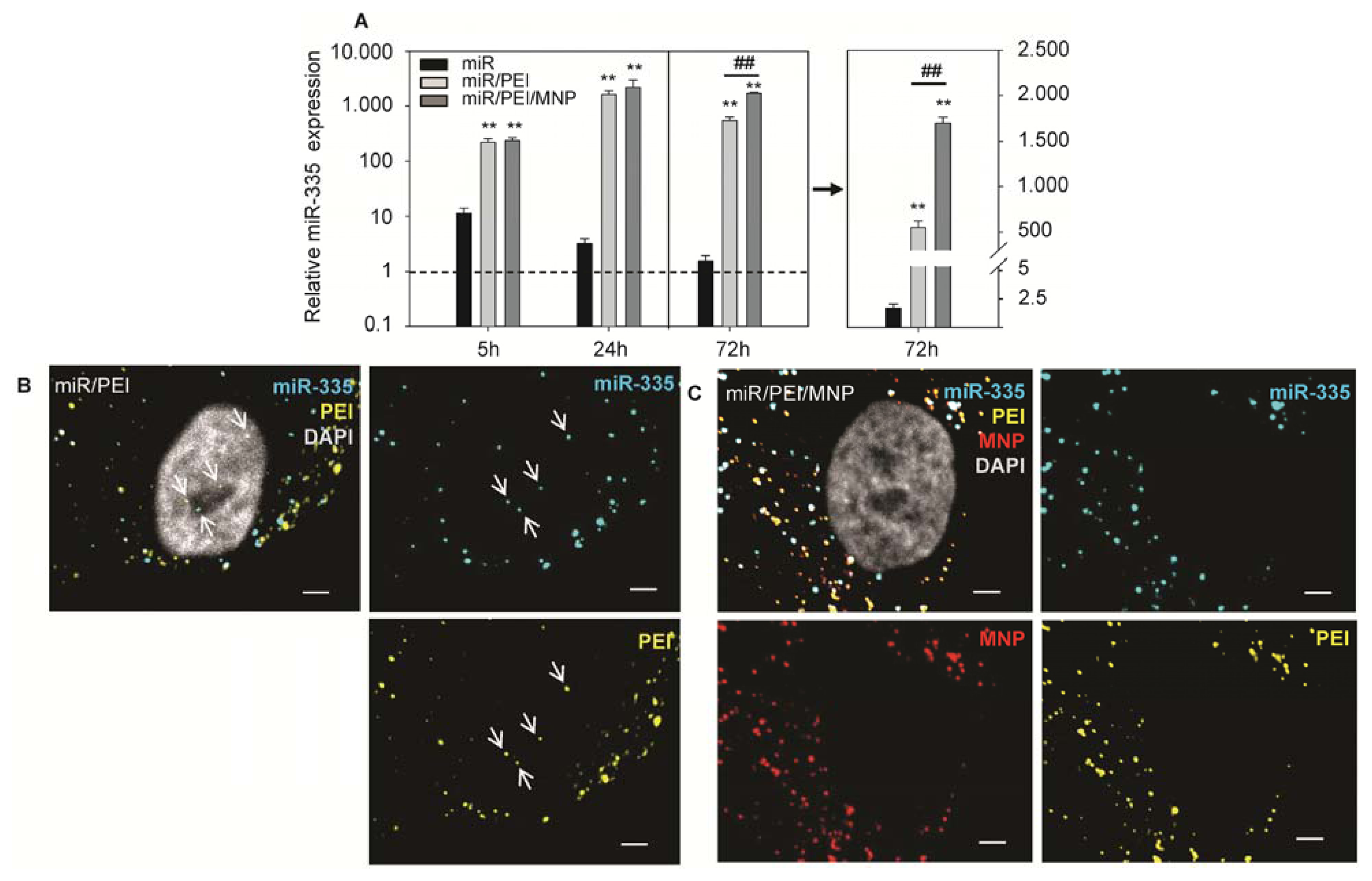
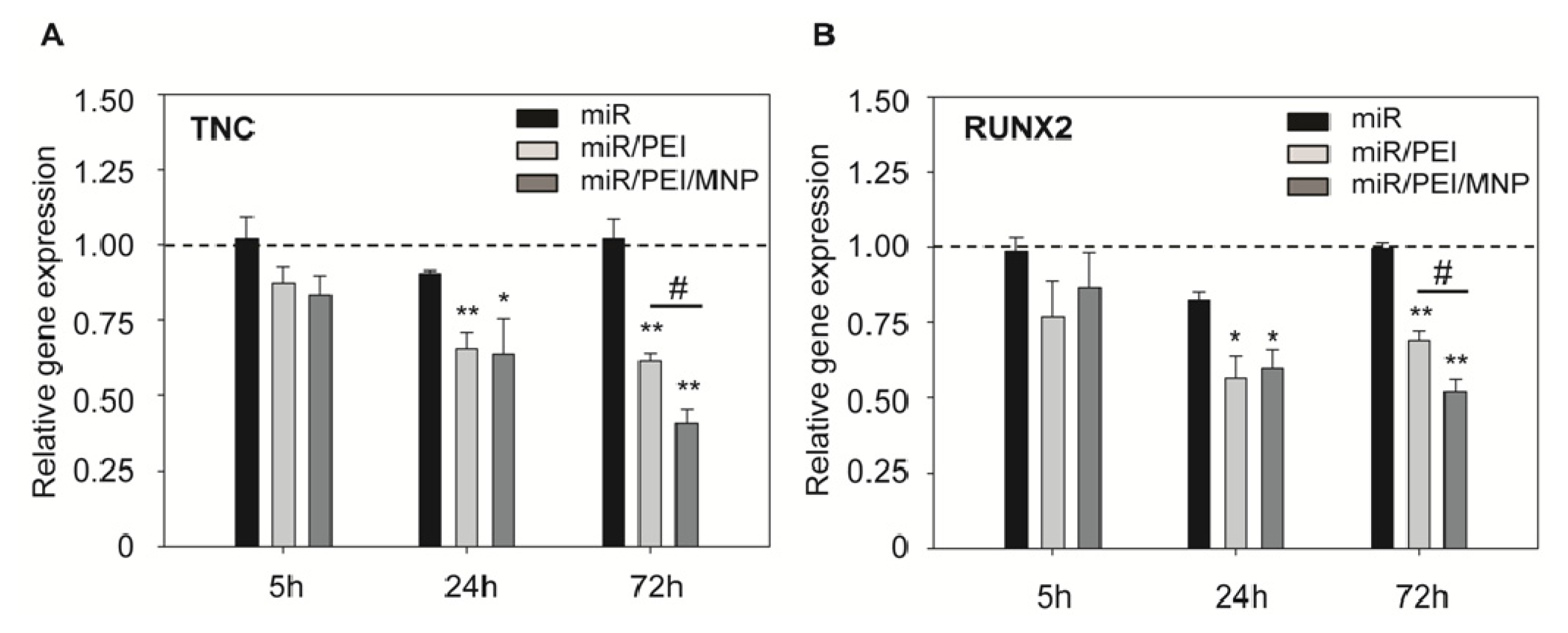
2.2. Monitoring of miR Processing in hMSCs over Time
3. Experimental Section
3.1. Culture of hMSCs
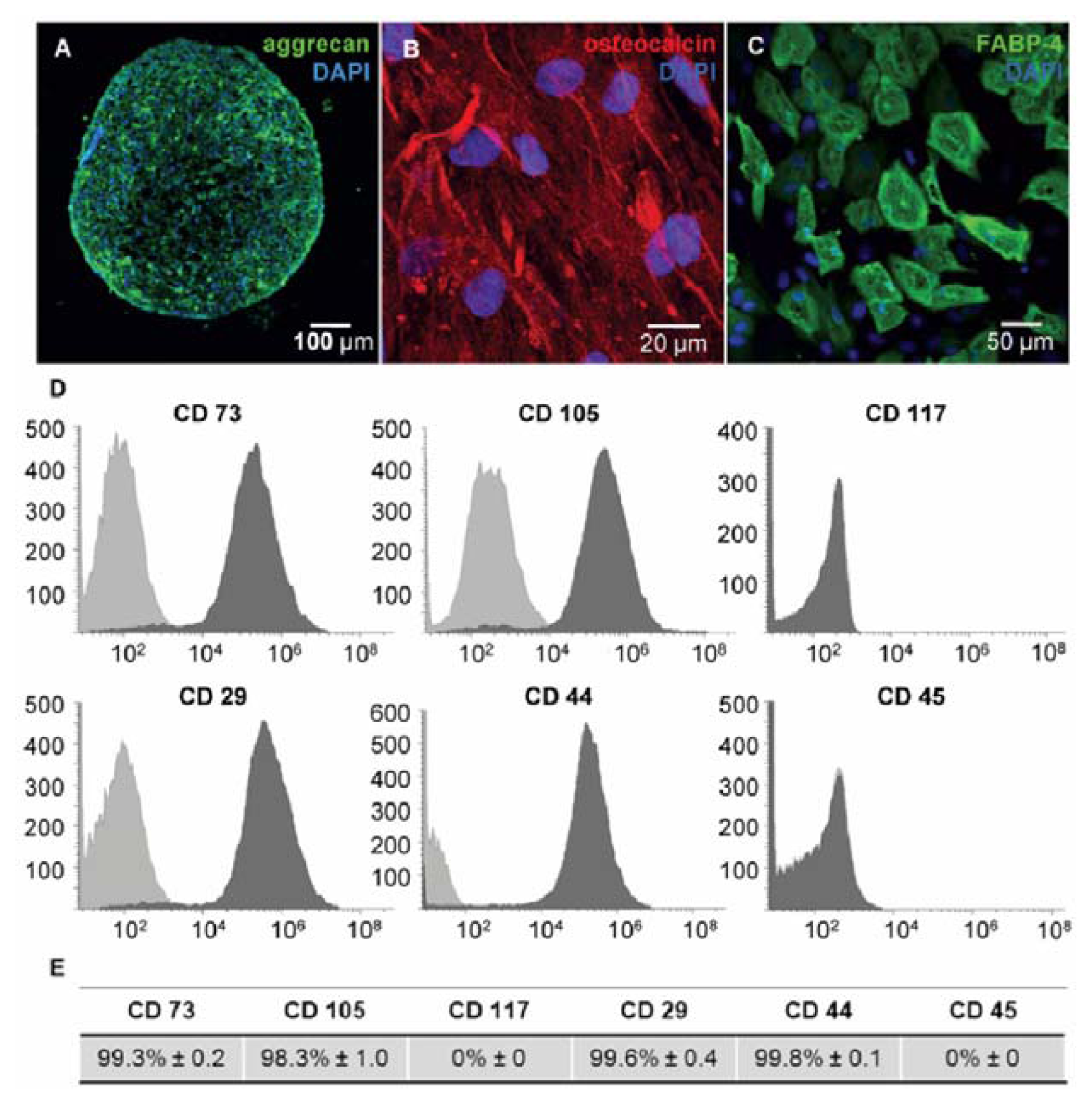
3.2. Immunophenotyping of hMSCs
3.3. Functional Characterization for hMSCs
3.4. Preparation and Characterization of Transfection Complexes
3.5. Transfection
3.6. Uptake Efficiency and Cytotoxicity
3.7. Fluorescent Labeling of Transfection Complexes
3.8. Confocal Laser Scanning Microscopy
3.9. Real-Time PCR
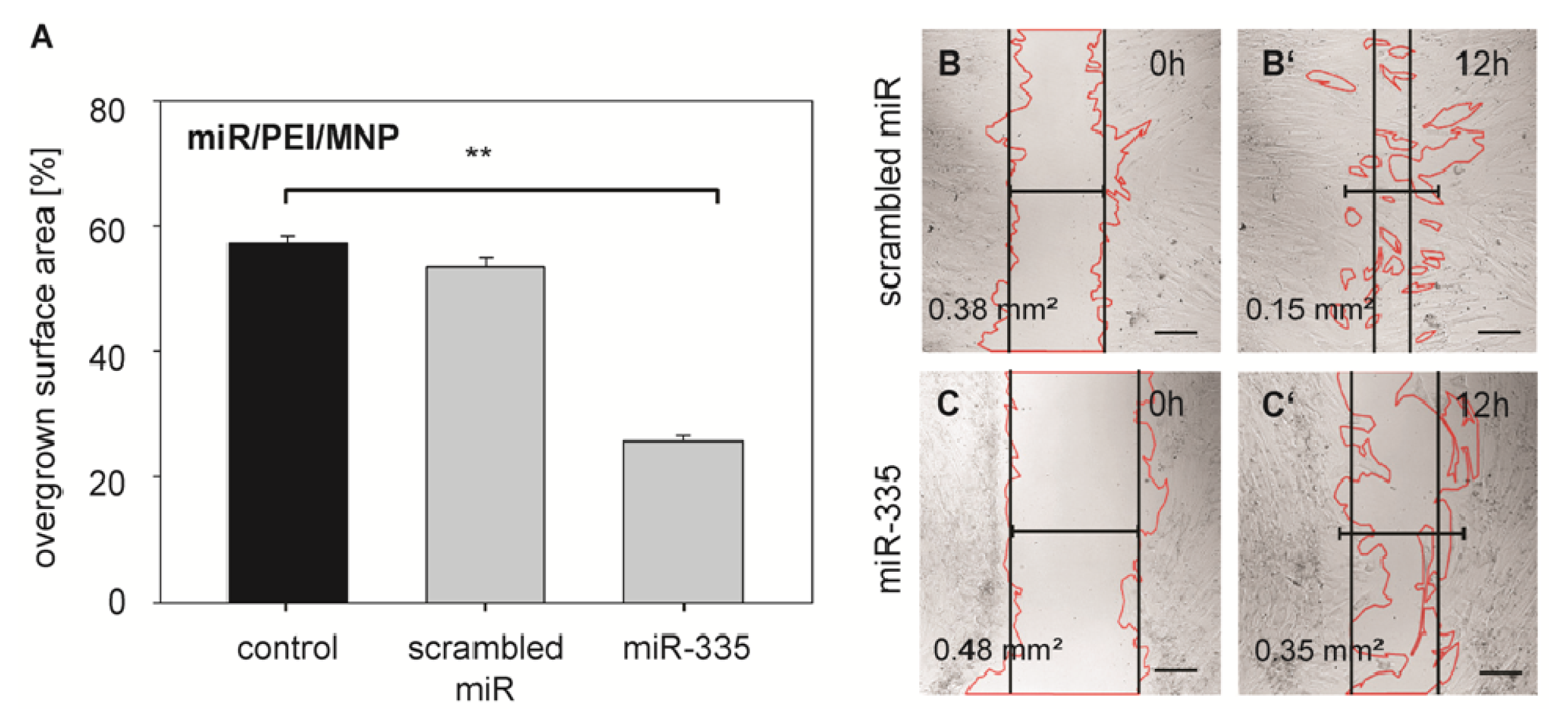
3.10. Wound Healing Assay
3.11. Statistical Analysis
4. Conclusions
Supplementary Information
ijms-14-10710-s001.zipAcknowledgments
Conflict of Interest
References
- Nesselmann, C.; Li, W.; Ma, N.; Steinhoff, G. Stem cell-mediated neovascularization in heart repair. Ther. Adv. Cardiovasc. Dis 2010, 4, 27–42. [Google Scholar]
- Wang, W.; Li, W.; Ou, L.; Flick, E.; Mark, P.; Nesselmann, C.; Lux, C.A.; Gatzen, H.H.; Kaminski, A.; Liebold, A.; et al. Polyethylenimine-mediated gene delivery into human bone marrow mesenchymal stem cells from patients. J. Cell. Mol. Med 2011, 15, 1989–1998. [Google Scholar]
- Iso, Y.; Spees, J.L.; Serrano, C.; Bakondi, B.; Pochampally, R.; Song, Y.H.; Sobel, B.E.; Delafontaine, P.; Prockop, D.J. Multipotent human stromal cells improve cardiac function after myocardial infarction in mice without long-term engraftment. Biochem. Biophys. Res. Commun 2007, 354, 700–706. [Google Scholar]
- Silva, G.V.; Litovsky, S.; Assad, J.A.; Sousa, A.L.; Martin, B.J.; Vela, D.; Coulter, S.C.; Lin, J.; Ober, J.; Vaughn, W.K.; et al. Mesenchymal stem cells differentiate into an endothelial phenotype, enhance vascular density, and improve heart function in a canine chronic ischemia model. Circulation 2005, 111, 150–156. [Google Scholar]
- Huang, F.; Zhu, X.; Hu, X.Q.; Fang, Z.F.; Tang, L.; Lu, X.L.; Zhou, S.H. Mesenchymal stem cells modified with miR-126 release angiogenic factors and activate Notch ligand Delta-like-4, enhancing ischemic angiogenesis and cell survival. Int. J. Mol. Med 2013, 31, 484–492. [Google Scholar]
- Chen, J.J.; Zhou, S.H. Mesenchymal stem cells overexpressing MiR-126 enhance ischemic angiogenesis via the AKT/ERK-related pathway. Cardiol. J 2011, 18, 675–681. [Google Scholar]
- Wen, Z.; Zheng, S.; Zhou, C.; Yuan, W.; Wang, J.; Wang, T. Bone marrow mesenchymal stem cells for post-myocardial infarction cardiac repair: microRNAs as novel regulators. J. Cell. Mol. Med 2012, 16, 657–671. [Google Scholar]
- Berezikov, E.; Guryev, V.; van de Belt, J.; Wienholds, E.; Plasterk, R.H.; Cuppen, E. Phylogenetic shadowing and computational identification of human microRNA genes. Cell 2005, 120, 21–24. [Google Scholar]
- Jakob, P.; Landmesser, U. Role of microRNAs in stem/progenitor cells and cardiovascular repair. Cardiovasc. Res 2012, 93, 614–622. [Google Scholar]
- Boulaiz, H.; Marchal, J.A.; Prados, J.; Melguizo, C.; Aranega, A. Non-viral and viral vectors for gene therapy. Cell. Mol. Biol 2005, 51, 3–22. [Google Scholar]
- Al-Dosari, M.S.; Gao, X. Nonviral gene delivery: Principle, limitations, and recent progress. AAPS J 2009, 11, 671–681. [Google Scholar]
- Godbey, W.T.; Wu, K.K.; Mikos, A.G. Poly(ethylenimine) and its role in gene delivery. J. Control. Release 1999, 60, 149–160. [Google Scholar]
- Pathak, A.; Patnaik, S.; Gupta, K.C. Recent trends in non-viral vector-mediated gene delivery. Biotechnol. J 2009, 4, 1559–1572. [Google Scholar]
- Godbey, W.T.; Mikos, A.G. Recent progress in gene delivery using non-viral transfer complexes. J. Control. Release 2001, 72, 115–125. [Google Scholar]
- Bieber, T.; Meissner, W.; Kostin, S.; Niemann, A.; Elsasser, H.P. Intracellular route and transcriptional competence of polyethylenimine-DNA complexes. J. Control. Release 2002, 82, 441–454. [Google Scholar]
- Li, W.; Ma, N.; Ong, L.L.; Kaminski, A.; Skrabal, C.; Ugurlucan, M.; Lorenz, P.; Gatzen, H.H.; Lutzow, K.; Lendlein, A.; et al. Enhanced thoracic gene delivery by magnetic nanobead-mediated vector. J. Gene Med 2008, 10, 897–909. [Google Scholar]
- Delyagina, E.; Li, W.; Ma, N.; Steinhoff, G. Magnetic targeting strategies in gene delivery. Nanomedicine 2011, 6, 1593–1604. [Google Scholar]
- Dobson, J. Gene therapy progress and prospects: Magnetic nanoparticle-based gene delivery. Gene Ther 2006, 13, 283–287. [Google Scholar]
- Muthana, M.; Scott, S.D.; Farrow, N.; Morrow, F.; Murdoch, C.; Grubb, S.; Brown, N.; Dobson, J.; Lewis, C.E. A novel magnetic approach to enhance the efficacy of cell-based gene therapies. Gene Ther 2008, 15, 902–910. [Google Scholar]
- Hofmann, A.; Wenzel, D.; Becher, U.M.; Freitag, D.F.; Klein, A.M.; Eberbeck, D.; Schulte, M.; Zimmermann, K.; Bergemann, C.; Gleich, B.; et al. Combined targeting of lentiviral vectors and positioning of transduced cells by magnetic nanoparticles. Proc. Natl. Acad. Sci. USA 2009, 106, 44–49. [Google Scholar]
- Scherer, F.; Anton, M.; Schillinger, U.; Henke, J.; Bergemann, C.; Kruger, A.; Gansbacher, B.; Plank, C. Magnetofection: Enhancing and targeting gene delivery by magnetic force in vitro and in vivo. Gene Ther 2002, 9, 102–109. [Google Scholar]
- Yiu, H.H.; McBain, S.C.; Lethbridge, Z.A.; Lees, M.R.; Dobson, J. Preparation and characterization of polyethylenimine-coated Fe3O4-MCM-48 nanocomposite particles as a novel agent for magnet-assisted transfection. J. Biomed. Mater. Res. Part A 2008, 92, 386–392. [Google Scholar]
- Del Pino, P.; Munoz-Javier, A.; Vlaskou, D.; Rivera Gil, P.; Plank, C.; Parak, W.J. Gene silencing mediated by magnetic lipospheres tagged with small interfering RNA. Nano Lett 2010, 10, 3914–3921. [Google Scholar]
- Lim, J.; Clements, M.A.; Dobson, J. Delivery of short interfering ribonucleic acid-complexed magnetic nanoparticles in an oscillating field occurs via caveolae-mediated endocytosis. PLoS One 2012, 7, e51350. [Google Scholar]
- Schillinger, U.; Brill, T.; Rudolph, C.; Huth, S.; Gersting, S.; Krötz, F.; Hirschberger, J.; Bergemann, C.; Plank, C. Advances in magnetofection-magnetically guided nucleic acid delivery. J. Magn. Magn. Mater 2005, 293, 501–508. [Google Scholar]
- McBain, S.C.; Griesenbach, U.; Xenariou, S.; Keramane, A.; Batich, C.D.; Alton, E.W.; Dobson, J. Magnetic nanoparticles as gene delivery agents: Enhanced transfection in the presence of oscillating magnet arrays. Nanotechnology 2008, 19, 405102. [Google Scholar]
- Pickard, M.; Chari, D. Enhancement of magnetic nanoparticle-mediated gene transfer to astrocytes by ‘magnetofection’: Effects of static and oscillating fields. Nanomedicine 2010, 5, 217–232. [Google Scholar]
- Delyagina, E.; Schade, A.; Scharfenberg, D.; Skorska, A.; Lux, C.; Li, W.; Steinhoff, G. Improved transfection in human mesenchymal stem cells: Effective intracellular release of pDNA by magnetic polyplexes. Nanomedicine 2013, in press. [Google Scholar]
- Tome, M.; Lopez-Romero, P.; Albo, C.; Sepulveda, J.C.; Fernandez-Gutierrez, B.; Dopazo, A.; Bernad, A.; Gonzalez, M.A. miR-335 orchestrates cell proliferation, migration and differentiation in human mesenchymal stem cells. Cell Death Differ 2011, 18, 985–995. [Google Scholar]
- Greco, S.; de Simone, M.; Colussi, C.; Zaccagnini, G.; Fasanaro, P.; Pescatori, M.; Cardani, R.; Perbellini, R.; Isaia, E.; Sale, P.; et al. Common micro-RNA signature in skeletal muscle damage and regeneration induced by Duchenne muscular dystrophy and acute ischemia. FASEB J 2009, 23, 3335–3346. [Google Scholar]
- Choi, Y.H.; Kurtz, A.; Stamm, C. Mesenchymal stem cells for cardiac cell therapy. Hum. Gene Ther 2011, 22, 3–17. [Google Scholar]
- Pittenger, M.F.; Martin, B.J. Mesenchymal stem cells and their potential as cardiac therapeutics. Circul. Res 2004, 95, 9–20. [Google Scholar]
- Grzelinski, M.; Urban-Klein, B.; Martens, T.; Lamszus, K.; Bakowsky, U.; Hobel, S.; Czubayko, F.; Aigner, A. RNA interference-mediated gene silencing of pleiotrophin through polyethylenimine-complexed small interfering RNAs in vivo exerts antitumoral effects in glioblastoma xenografts. Hum. Gene Ther 2006, 17, 751–766. [Google Scholar]
- Hobel, S.; Koburger, I.; John, M.; Czubayko, F.; Hadwiger, P.; Vornlocher, H.P.; Aigner, A. Polyethylenimine/small interfering RNA-mediated knockdown of vascular endothelial growth factor in vivo exerts anti-tumor effects synergistically with Bevacizumab. J. Gene Med 2010, 12, 287–300. [Google Scholar]
- Urban-Klein, B.; Werth, S.; Abuharbeid, S.; Czubayko, F.; Aigner, A. RNAi-mediated gene-targeting through systemic application of polyethylenimine (PEI)-complexed siRNA in vivo. Gene Ther 2005, 12, 461–466. [Google Scholar]
- Ibrahim, A.F.; Weirauch, U.; Thomas, M.; Grunweller, A.; Hartmann, R.K.; Aigner, A. MicroRNA replacement therapy for miR-145 and miR-33a is efficacious in a model of colon carcinoma. Cancer Res 2011, 71, 5214–5224. [Google Scholar]
- Kwok, A.; Hart, S.L. Comparative structural and functional studies of nanoparticle formulations for DNA and siRNA delivery. Nanomedicine 2011, 7, 210–219. [Google Scholar]
- Giouroudi, I.; Kosel, J. Recent progress in biomedical applications of magnetic nanoparticles. Recent Pat. Nanotechnol 2010, 4, 111–118. [Google Scholar]
- Von Gersdorff, K.; Sanders, N.N.; Vandenbroucke, R.; de Smedt, S.C.; Wagner, E.; Ogris, M. The internalization route resulting in successful gene expression depends on both cell line and polyethylenimine polyplex type. Mol. Ther 2006, 14, 745–753. [Google Scholar]
- Judge, A.; MacLachlan, I. Overcoming the innate immune response to small interfering RNA. Hum. Gene Ther 2008, 19, 111–124. [Google Scholar]
- Arsianti, M.; Lim, M.; Marquis, C.P.; Amal, R. Polyethylenimine based magnetic iron-oxide vector: The effect of vector component assembly on cellular entry mechanism, intracellular localization, and cellular viability. Biomacromolecules 2010, 11, 2521–2531. [Google Scholar]
- Treuel, L.; Jiang, X.; Nienhaus, G.U. New views on cellular uptake and trafficking of manufactured nanoparticles. J. R. Soc. Interface 2013, 10, 20120939. [Google Scholar]
- Prabha, S.; Zhou, W.Z.; Panyam, J.; Labhasetwar, V. Size-dependency of nanoparticle-mediated gene transfection: Studies with fractionated nanoparticles. Int. J. Pharm 2002, 244, 105–115. [Google Scholar]
- Boussif, O.; Lezoualc’h, F.; Zanta, M.A.; Mergny, M.D.; Scherman, D.; Demeneix, B.; Behr, J.P. A versatile vector for gene and oligonucleotide transfer into cells in culture and in vivo: Polyethylenimine. Proc. Natl. Acad. Sci. USA 1995, 92, 7297–7301. [Google Scholar]
- Kim, W.J.; Kim, S.W. Efficient siRNA delivery with non-viral polymeric vehicles. Pharm. Res 2009, 26, 657–666. [Google Scholar]
- Duman, O.; Tunç, S. Electrokinetic and rheological properties of Na-bentonite in some electrolyte solutions. Microporous Mesoporous Mater 2009, 117, 331–338. [Google Scholar]
- Mekhamer, W.K. The colloidal stability of raw bentonite deformed mechanically by ultrasound. J. Saudi Chem. Soc 2010, 14, 301–306. [Google Scholar]
- Ma, C.; Liu, Y.; He, L. MicroRNAs—Powerful repression comes from small RNAs. Sci. China Ser. C 2009, 52, 323–330. [Google Scholar]
- Clamme, J.P.; Krishnamoorthy, G.; Mely, Y. Intracellular dynamics of the gene delivery vehicle polyethylenimine during transfection: Investigation by two-photon fluorescence correlation spectroscopy. Biochim. Biophys. Acta 2003, 1617, 52–61. [Google Scholar]
- Tavazoie, S.F.; Alarcon, C.; Oskarsson, T.; Padua, D.; Wang, Q.; Bos, P.D.; Gerald, W.L.; Massague, J. Endogenous human microRNAs that suppress breast cancer metastasis. Nature 2008, 451, 147–152. [Google Scholar]
- Tsunoda, T.; Inada, H.; Kalembeyi, I.; Imanaka-Yoshida, K.; Sakakibara, M.; Okada, R.; Katsuta, K.; Sakakura, T.; Majima, Y.; Yoshida, T. Involvement of large tenascin-C splice variants in breast cancer progression. Am. J. Pathol 2003, 162, 1857–1867. [Google Scholar]
- Yoshida, T.; Yoshimura, E.; Numata, H.; Sakakura, Y.; Sakakura, T. Involvement of tenascin-C in proliferation and migration of laryngeal carcinoma cells. Virchows Arch 1999, 435, 496–500. [Google Scholar]
- Scholz, C.; Wagner, E. Therapeutic plasmid DNA versus siRNA delivery: Common and different tasks for synthetic carriers. J. Control. Release 2012, 161, 554–565. [Google Scholar]
- Lynch, J.; Fay, J.; Meehan, M.; Bryan, K.; Watters, K.M.; Murphy, D.M.; Stallings, R.L. MiRNA-335 suppresses neuroblastoma cell invasiveness by direct targeting of multiple genes from the non-canonical TGF-beta signalling pathway. Carcinogenesis 2012, 33, 976–985. [Google Scholar]
- Gaebel, R.; Furlani, D.; Sorg, H.; Polchow, B.; Frank, J.; Bieback, K.; Wang, W.; Klopsch, C.; Ong, L.L.; Li, W.; et al. Cell origin of human mesenchymal stem cells determines a different healing performance in cardiac regeneration. PLoS One 2011, 6, e15652. [Google Scholar]



© 2013 by the authors; licensee MDPI, Basel, Switzerland This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution license ( http://creativecommons.org/licenses/by/3.0/).
Share and Cite
Schade, A.; Delyagina, E.; Scharfenberg, D.; Skorska, A.; Lux, C.; David, R.; Steinhoff, G. Innovative Strategy for MicroRNA Delivery in Human Mesenchymal Stem Cells via Magnetic Nanoparticles. Int. J. Mol. Sci. 2013, 14, 10710-10726. https://doi.org/10.3390/ijms140610710
Schade A, Delyagina E, Scharfenberg D, Skorska A, Lux C, David R, Steinhoff G. Innovative Strategy for MicroRNA Delivery in Human Mesenchymal Stem Cells via Magnetic Nanoparticles. International Journal of Molecular Sciences. 2013; 14(6):10710-10726. https://doi.org/10.3390/ijms140610710
Chicago/Turabian StyleSchade, Anna, Evgenya Delyagina, Dorothee Scharfenberg, Anna Skorska, Cornelia Lux, Robert David, and Gustav Steinhoff. 2013. "Innovative Strategy for MicroRNA Delivery in Human Mesenchymal Stem Cells via Magnetic Nanoparticles" International Journal of Molecular Sciences 14, no. 6: 10710-10726. https://doi.org/10.3390/ijms140610710




