Specific siRNA Targeting Receptor for Advanced Glycation End Products (RAGE) Decreases Proliferation in Human Breast Cancer Cell Lines
Abstract
:1. Introduction
2. Results and Discussion
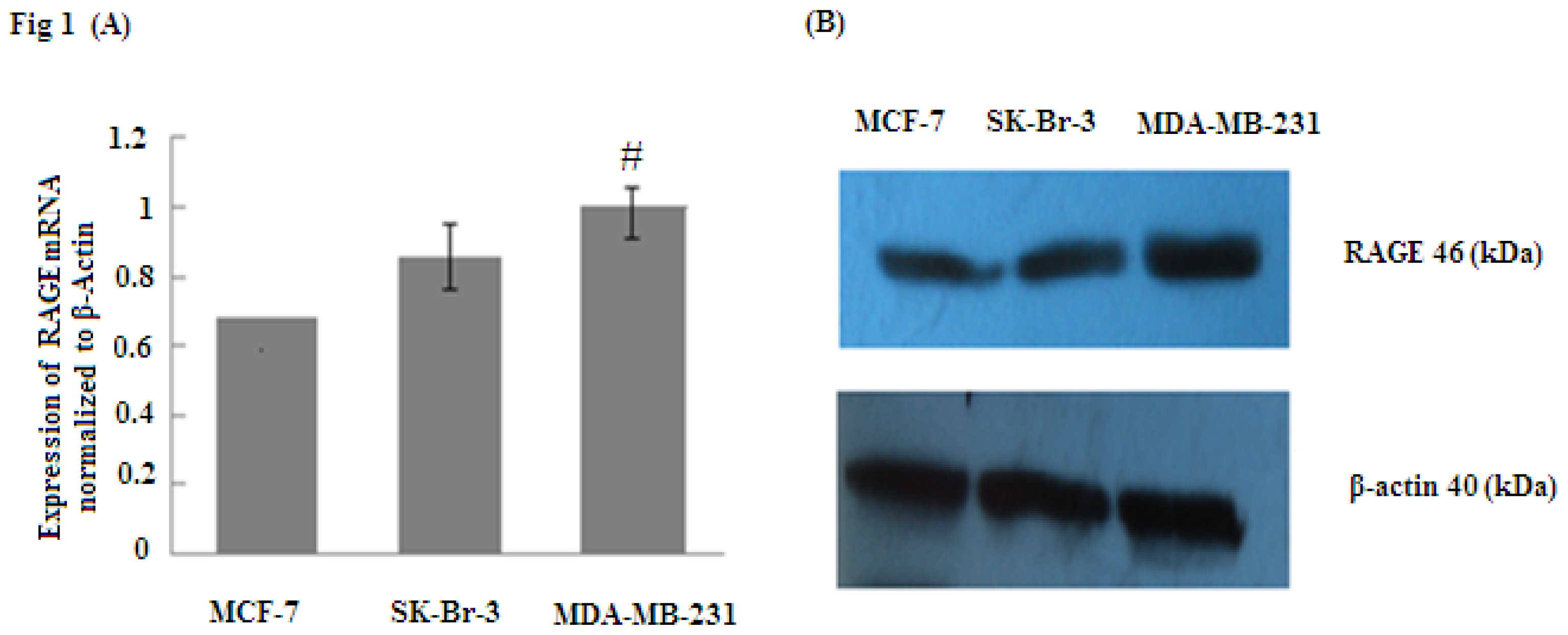
2.1. RAGE Expression in Different Sub-Types of Breast Cancer Cell Lines
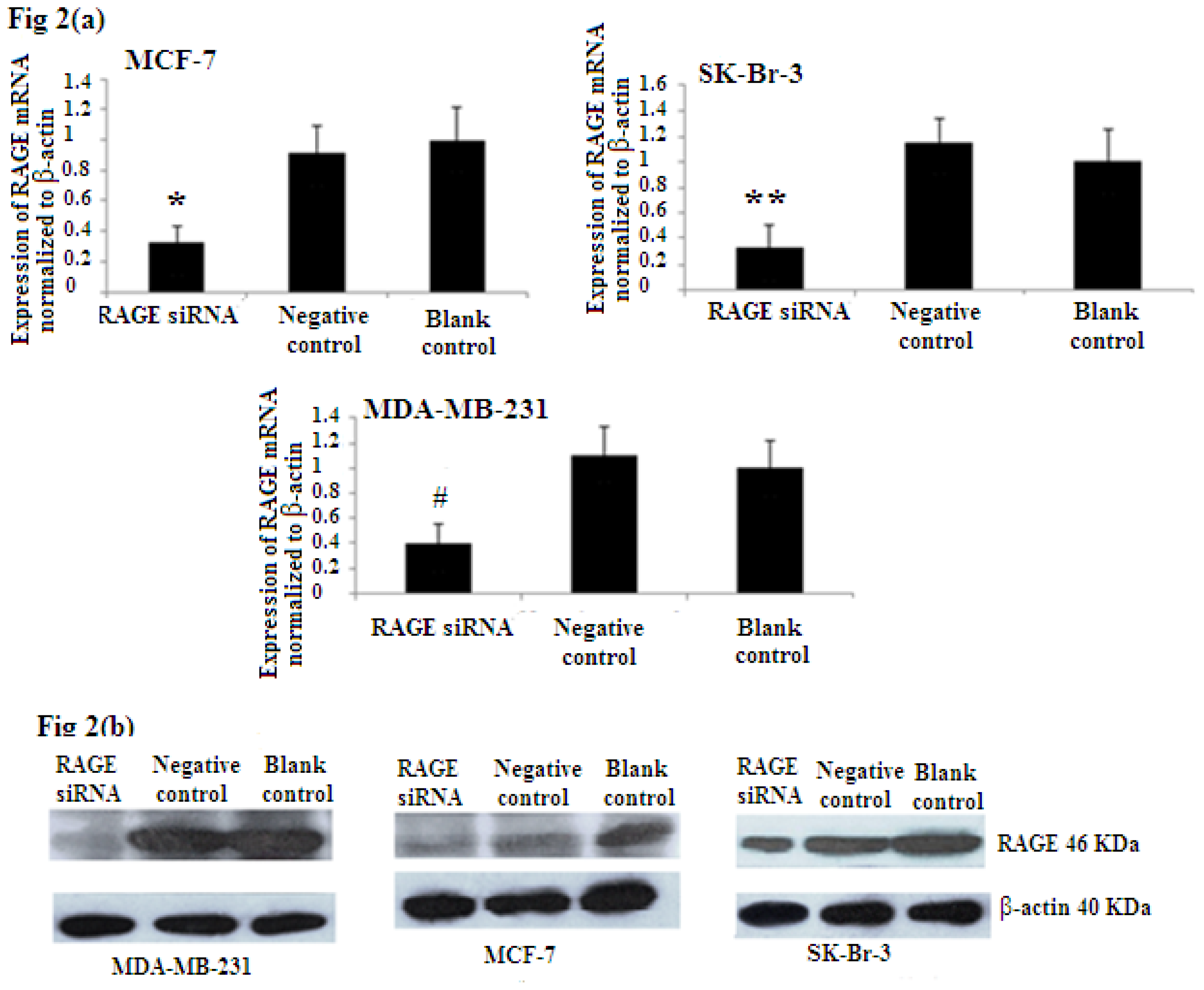
2.2. Efficiency of RAGE Gene Knockdown
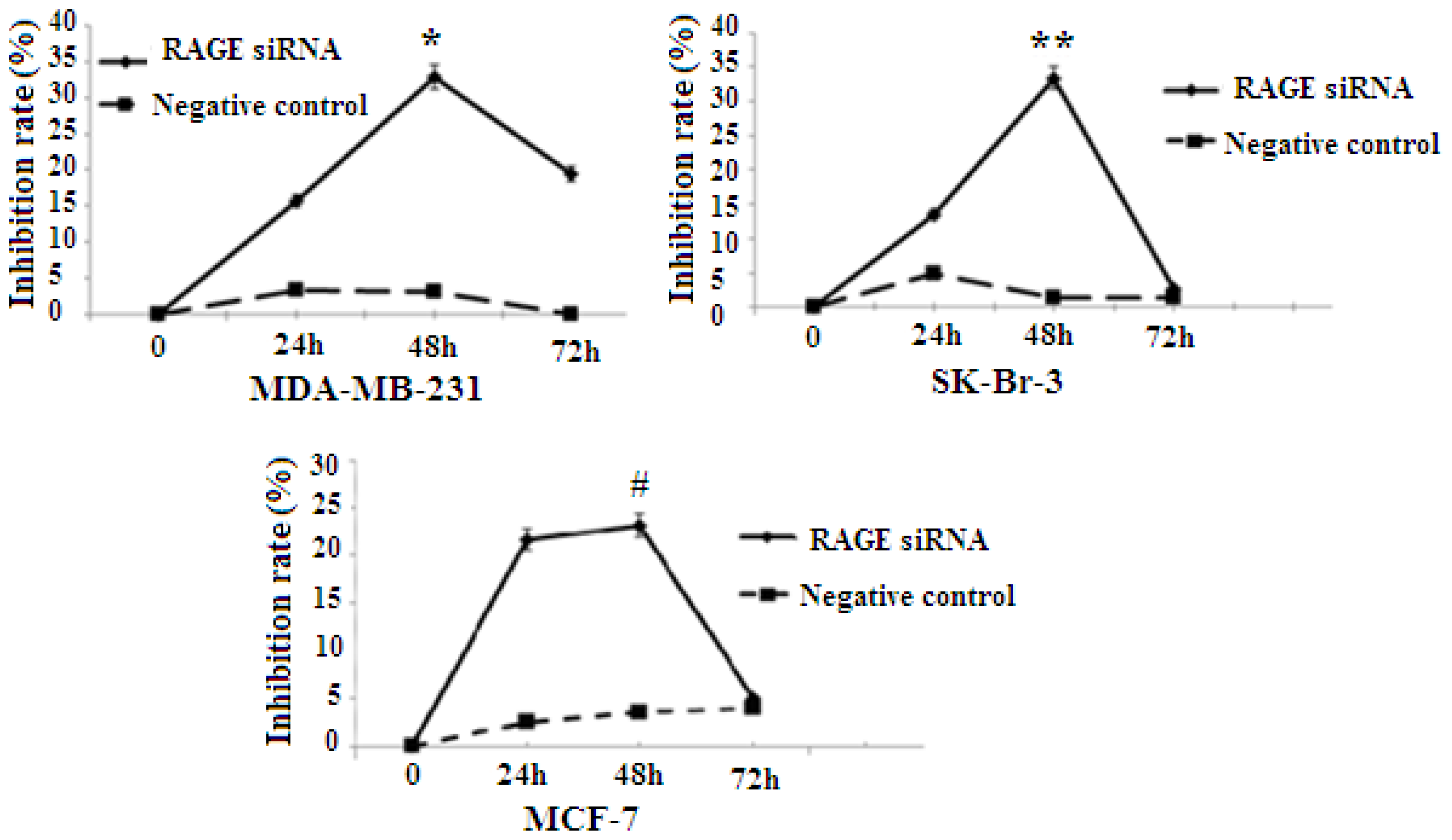
2.3. RAGE siRNA Decreases Viability in Breast Cancer
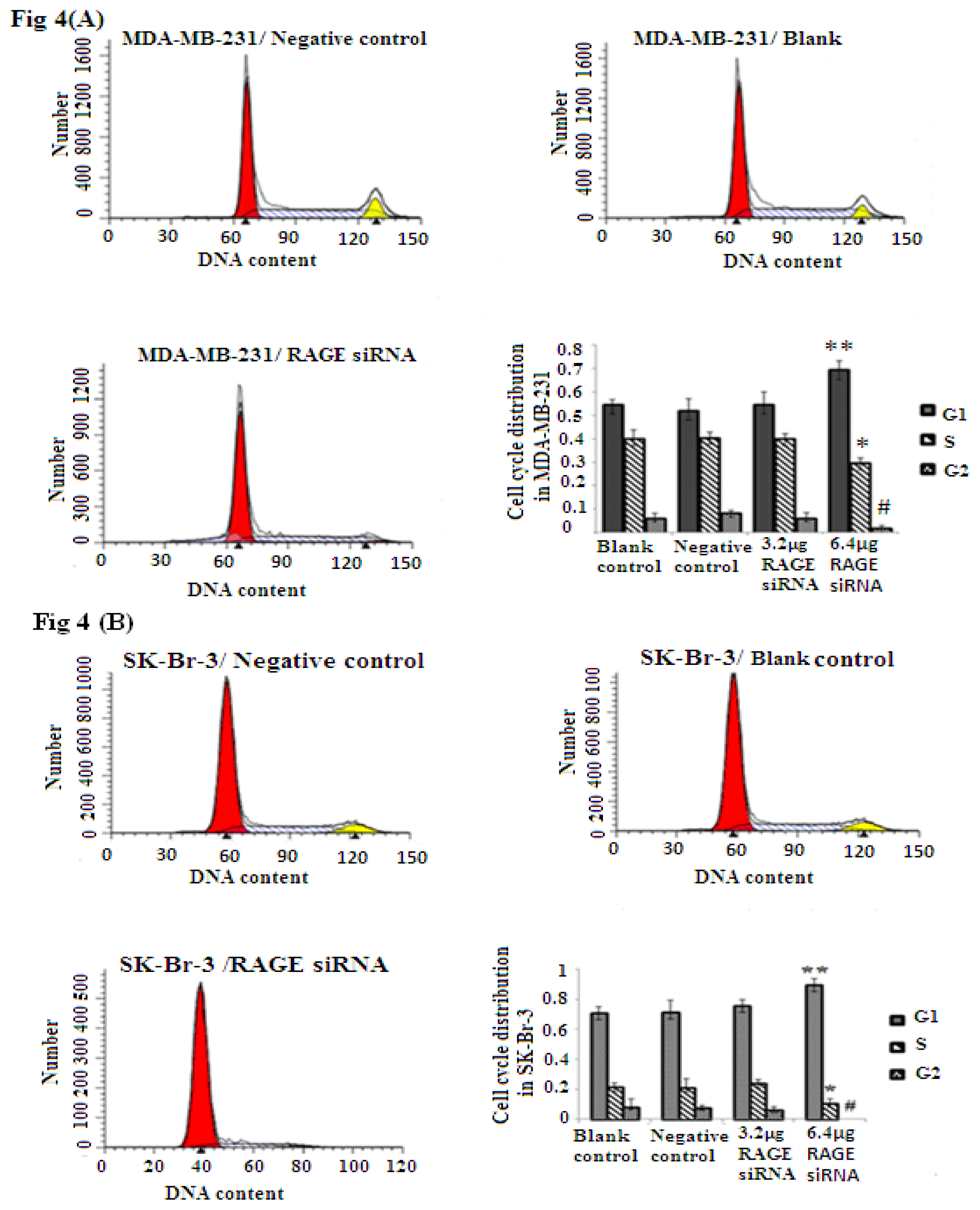
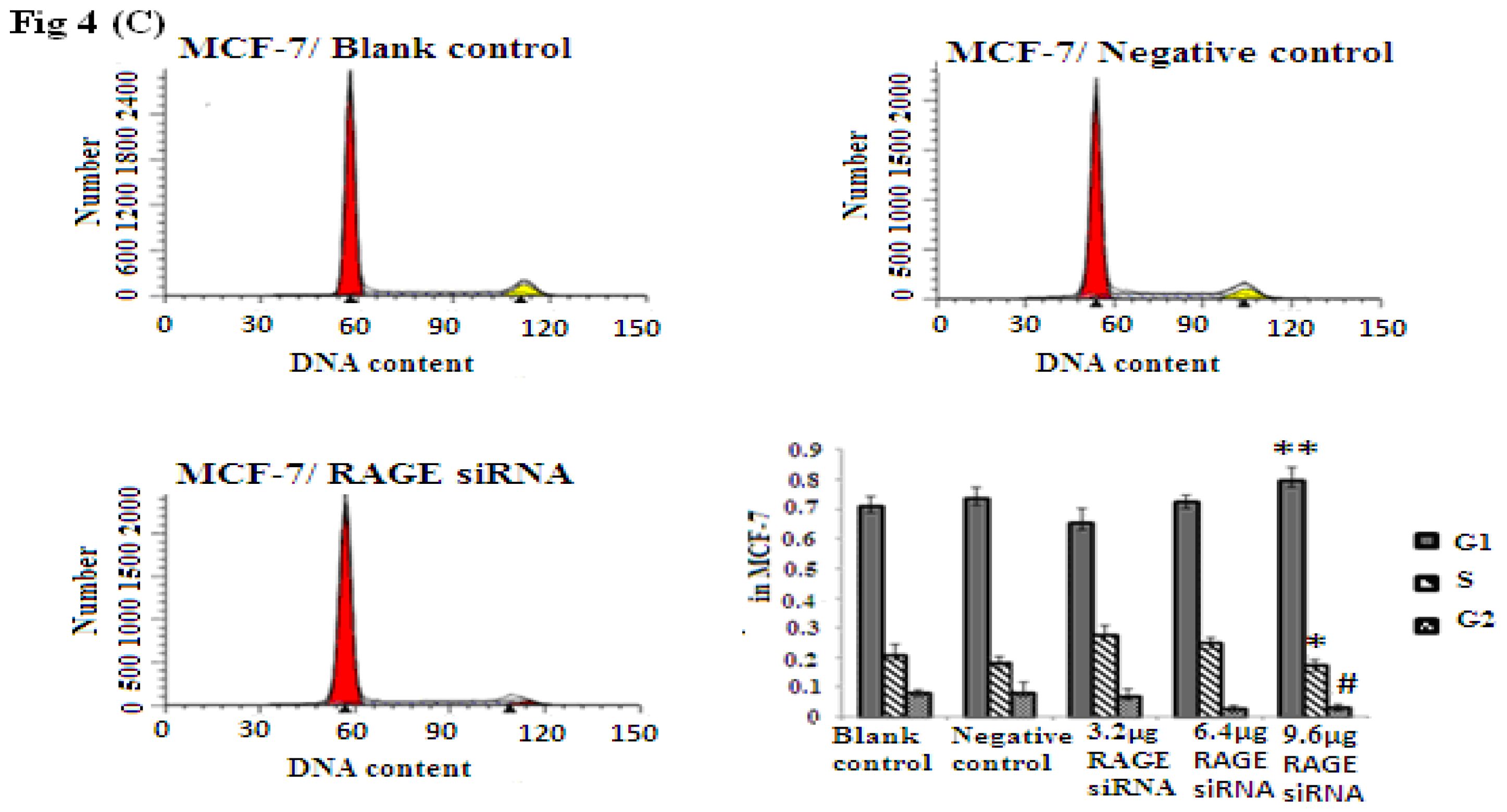
2.4. RAGE siRNA Induces G1 Arrest in Breast Cancer Cell Lines
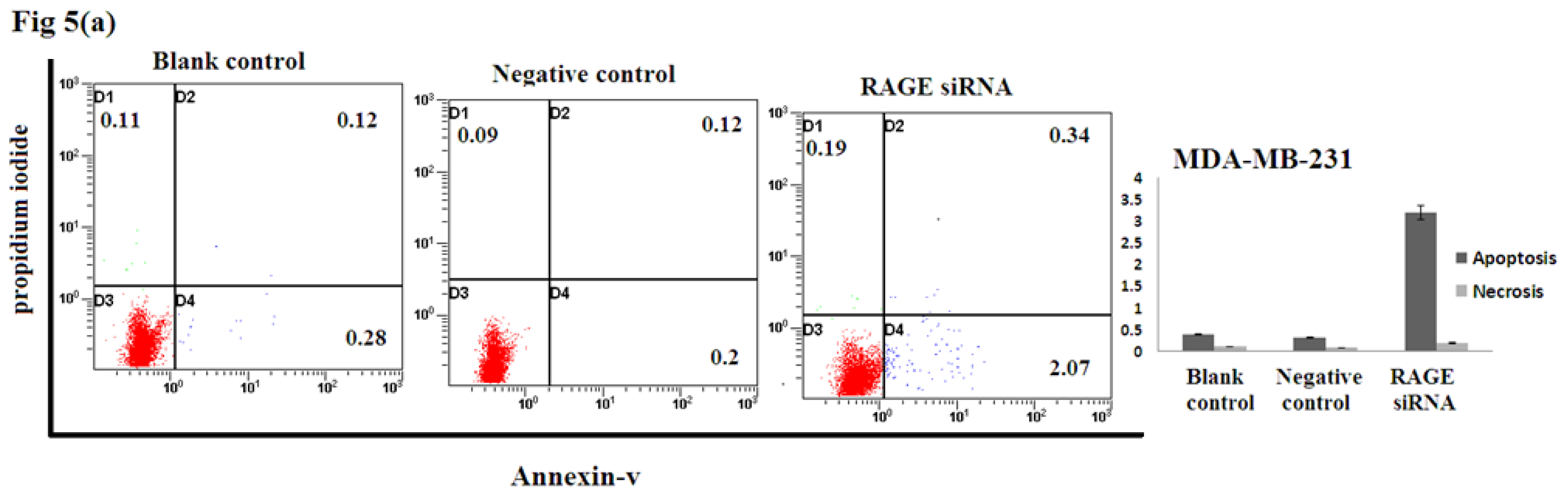
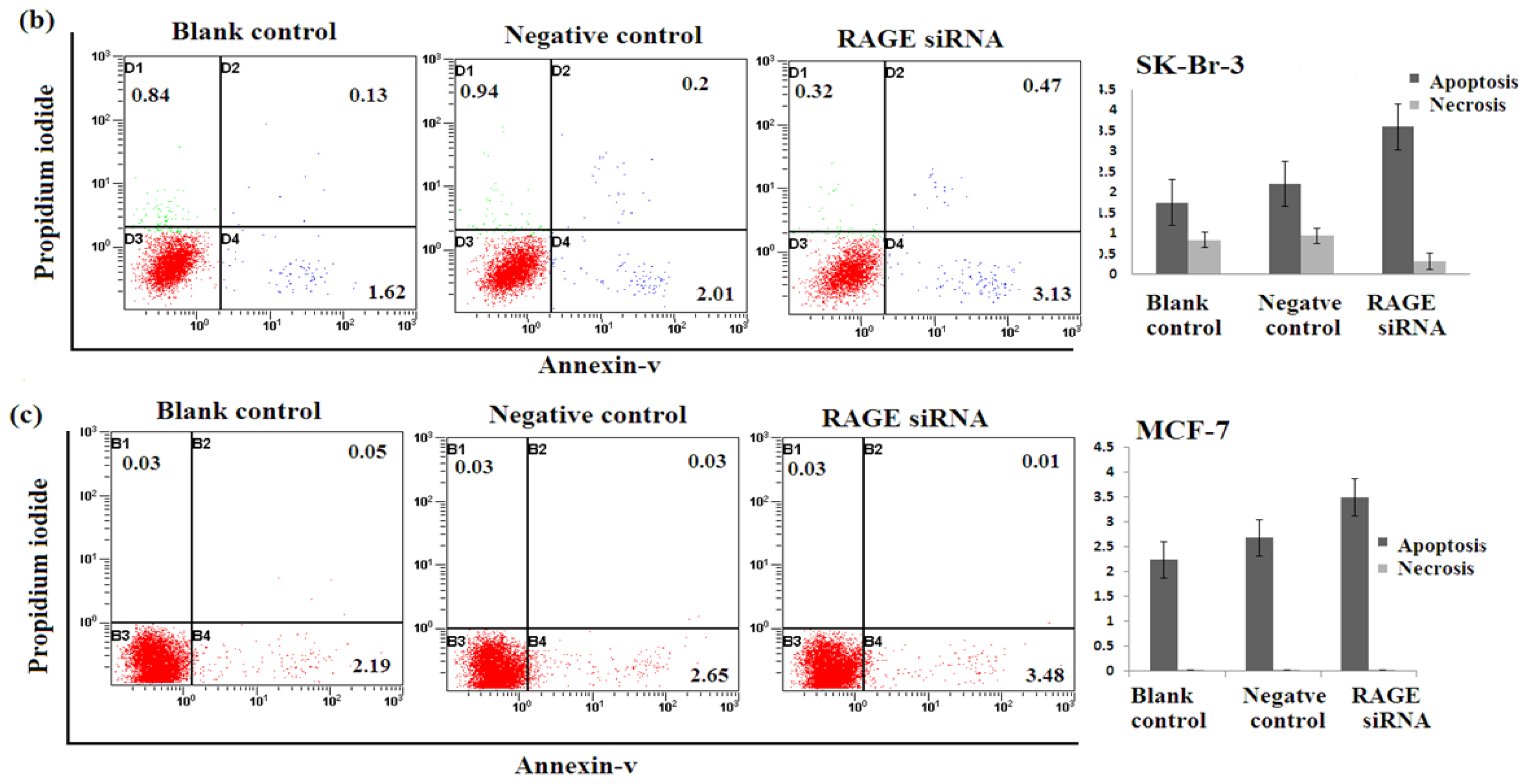
2.5. Silencing RAGE Induces Negligible Apoptosis
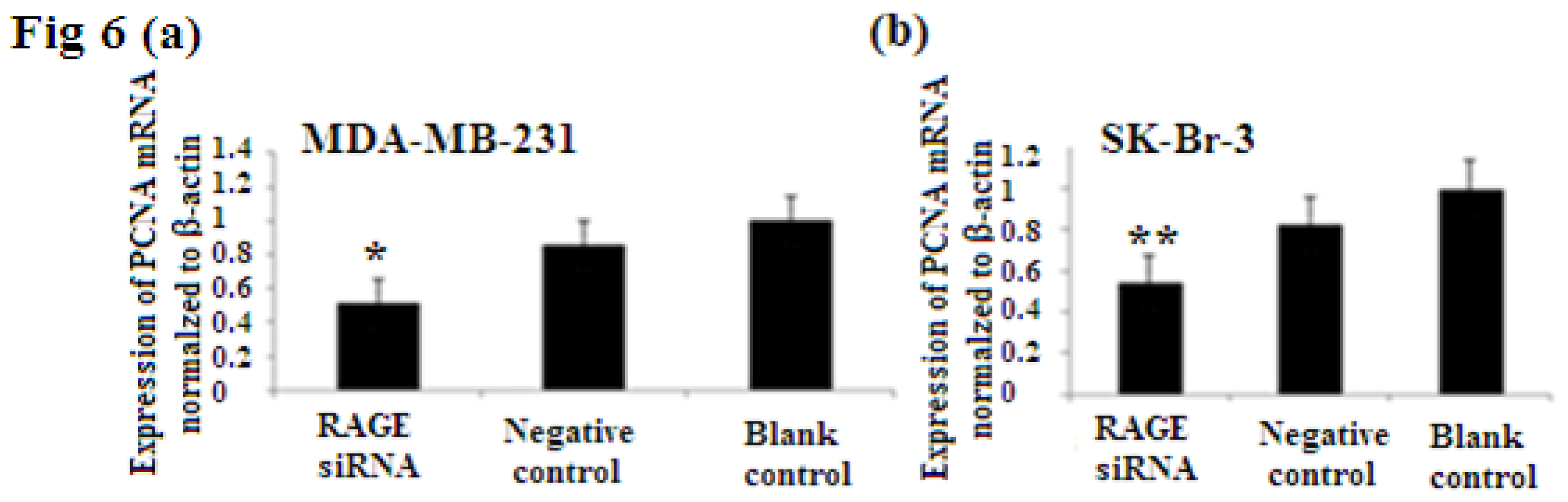
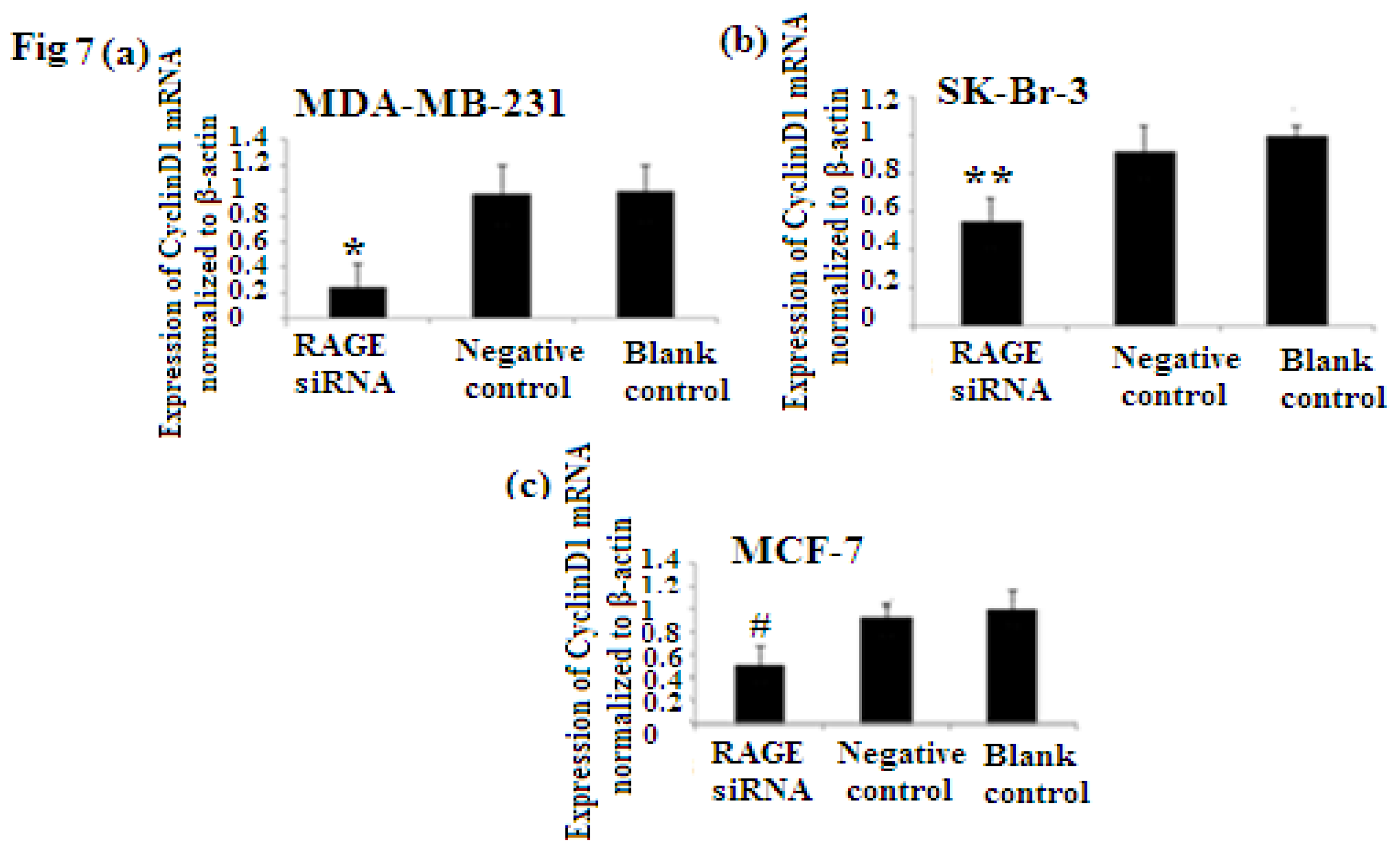
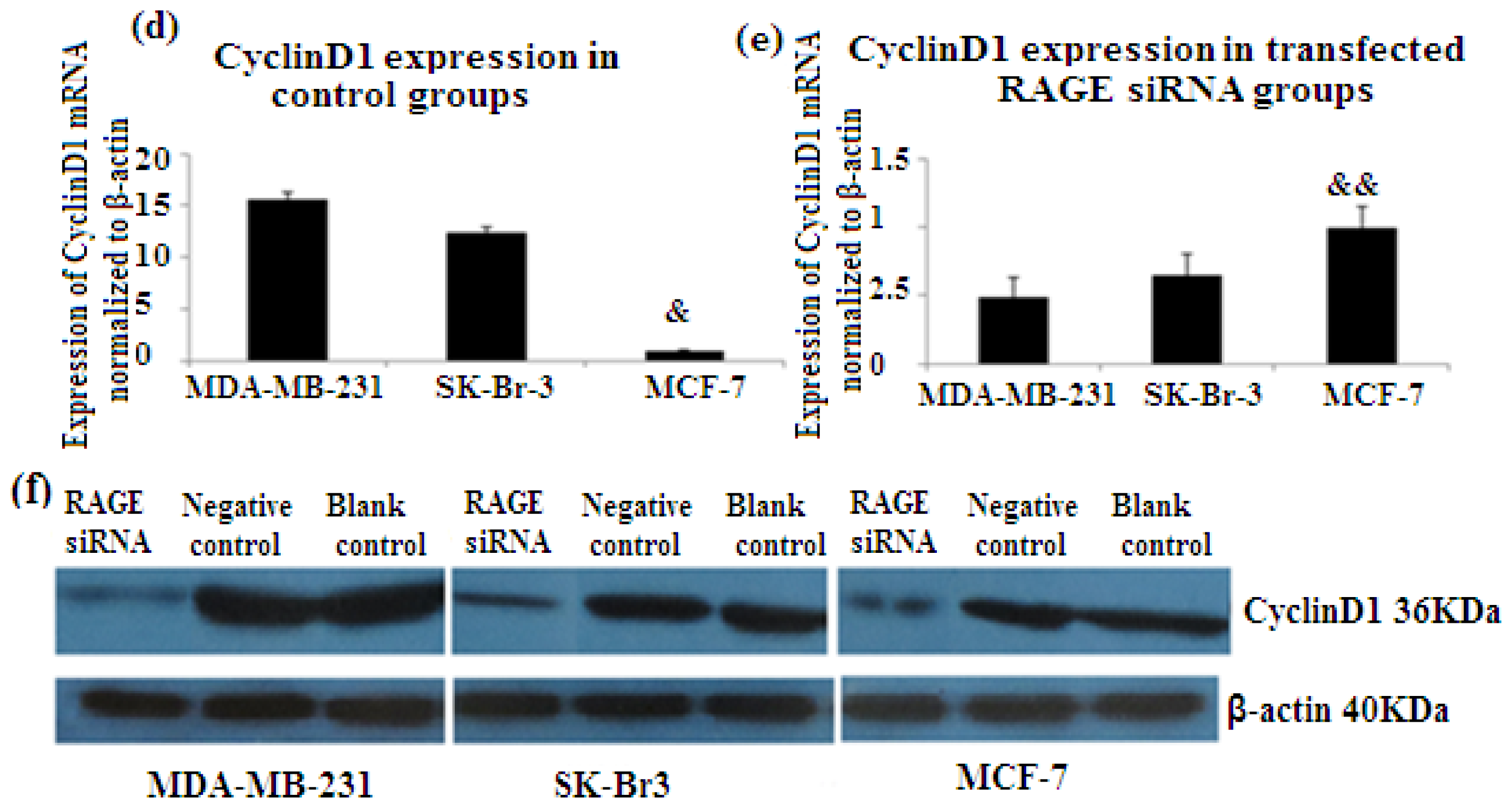
2.6. RAGE siRNA Decreases Expression of PCNA and CyclinD1 mRNA
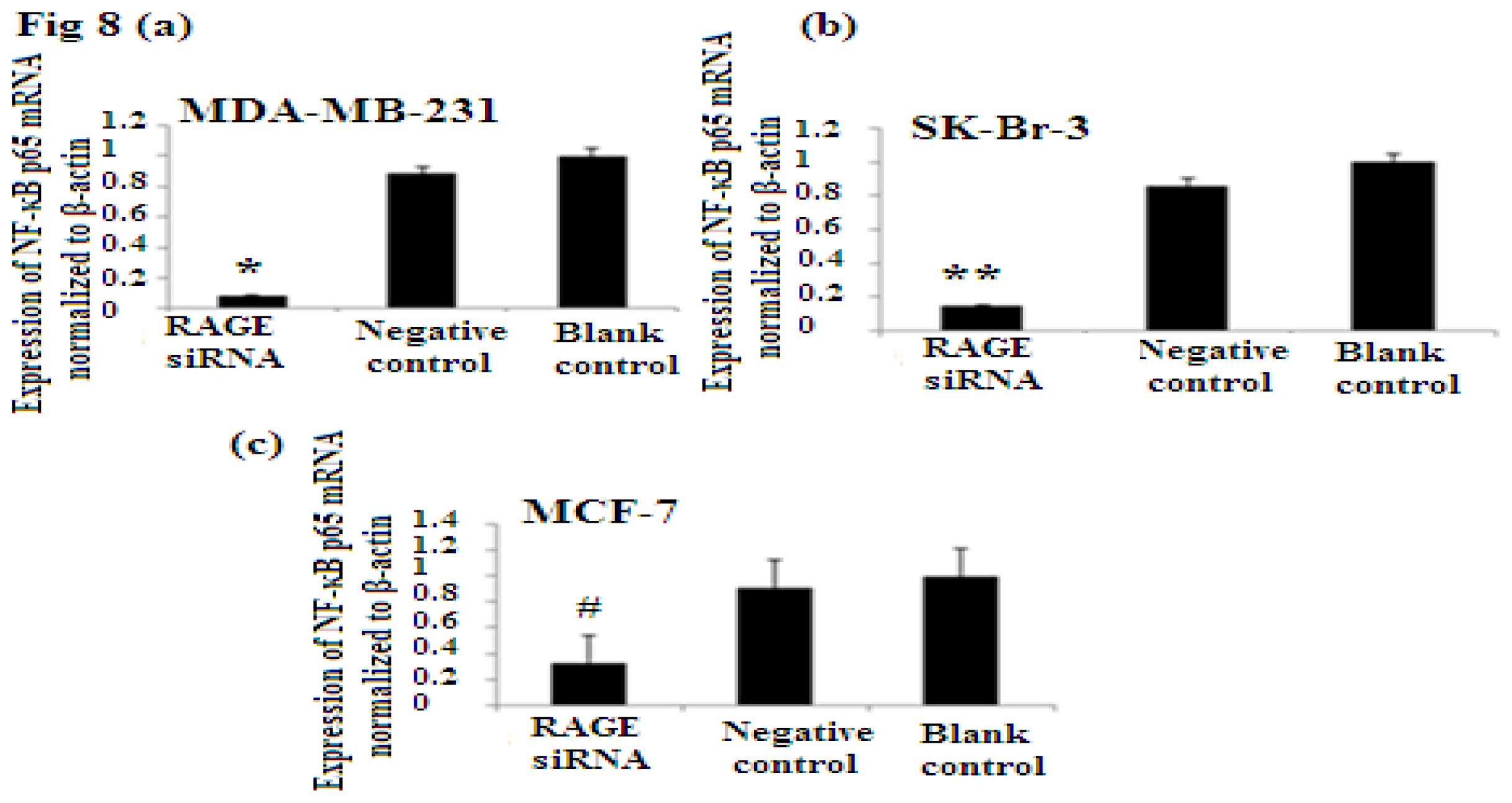
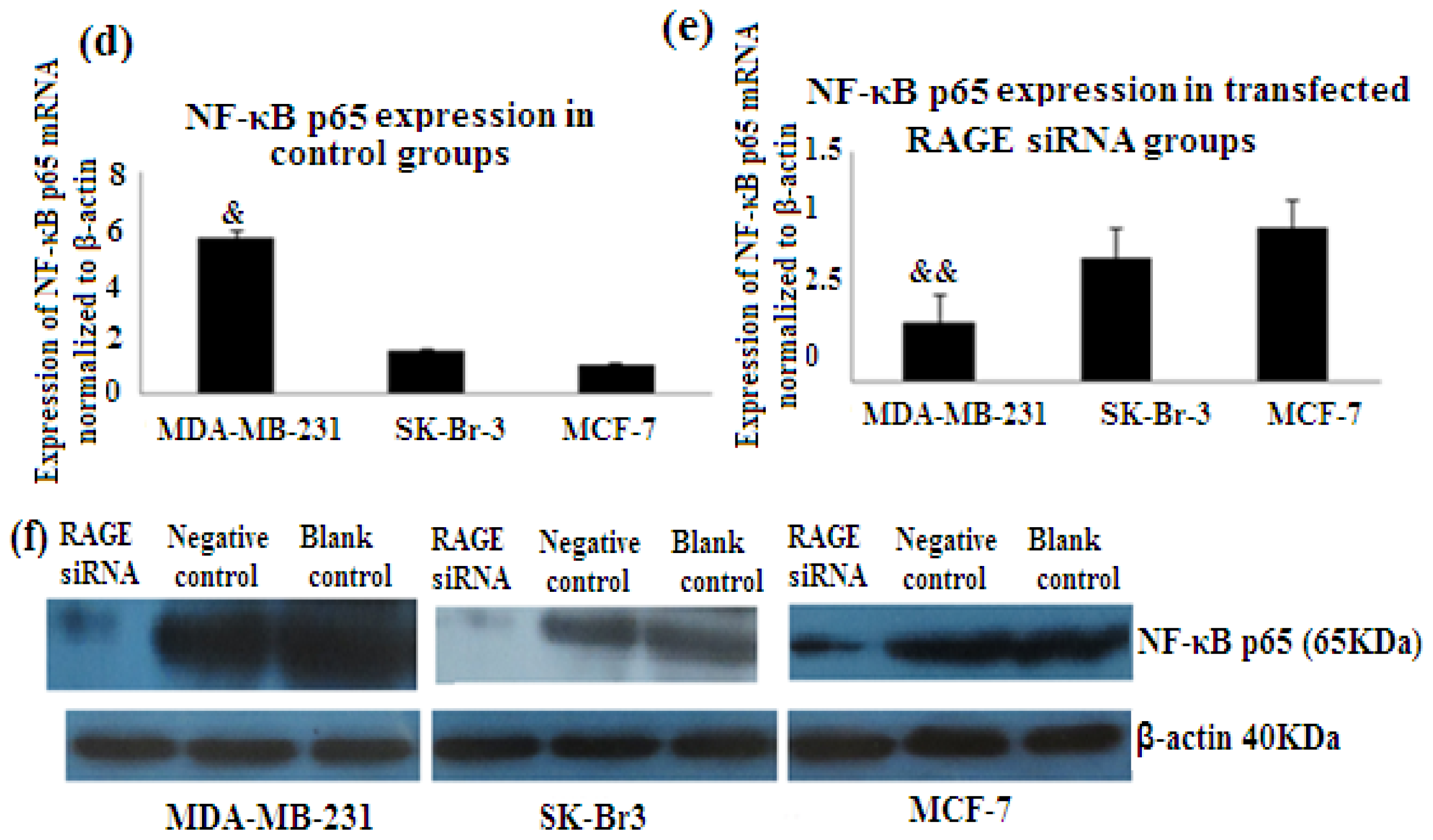
2.7. RAGE siRNA Decreases Expression of NF-κB p65
3. Discussion
4. Experimental Section
4.1. Cell Culture
4.2. siRNA Transfection
4.3. Quantitative Real Time PCR
4.4. Western Blot
4.5. Cell Viability Assay (MTT)
4.6. Cell Cycle Analysis
4.7. Apoptosis Assay
4.8. Statistical Analysis
5. Conclusions
Acknowledgments
References
- Poellinger, A. Near-infrared imaging of breast cancer using optical contrast agents. J. Biophotonics 2012, 5, 815–826. [Google Scholar]
- Smolarek, A.K.; Suh, N. Chemopreventive activity of vitamin E in breast cancer. A focus on ã- and ä-Tocopherol. Nutrients 2011, 3, 962–986. [Google Scholar]
- Beckmann, M.W.; Niederacher, D.; Schnurch, H.G.; Gusterson, B.A.; Bender, H.G. Multistep carcinogenesis of breast cancer and tumor heterogeneity. J. Mol. Med 1997, 75, 429–439. [Google Scholar]
- Lerebours, F.; Lidereau, R. Molecular alterations in sporadic breast cancer. Crit. Rev. Oncol. Hematol 2002, 44, 121–141. [Google Scholar]
- Balmain, A.; Gray, J.; Ponder, B. The genetics and genomics of cancer. Nat. Genet 2003, 33, 238– 244. [Google Scholar]
- Cynthia, O.; Paschal, W.; Debu, T. Oncogenes and tumor suppressor genes in breast cancer: Potential diagnostic and therapeutic applications. Oncologist 2004, 9, 361–377. [Google Scholar]
- DeNardo, D.G.; Coussens, L.M. Inflammation and breast cancer: Balancing immune response—Crosstalk between adaptive and innate immune cells during breast cancer progression. Breast Cancer Res 2007, 9, 212. [Google Scholar]
- Blows, F.M.; Driver, K.E.; Schmidt, M.K.; Broeks, A.; van Leeuwen, F.E.; Wesseling, J.; Cheang, M.C.; Gelmon, K.; Nielsen, T.O.; Blomqvist, C.; et al. Subtyping of breast cancer by immunohistochemistry to investigate a relationship between subtype and short and long term survival. A collaborative analysis of data for 10,159 cases from 12 studies. PLoS Med 2010, 7, e1000279. [Google Scholar]
- Badve, S.; Dabbs, D.J.; Schnitt, S.J.; Baehner, F.L.; Decker, T.; Eusebi, V.; Fox, S.B.; Ichihara, S.; Jacquemier, J.; Lakhani, S.R.; et al. Basal-like and triple-negative breast cancers. A critical review with an emphasis on the implications for pathologists and oncologists. Mod. Pathol 2011, 24, 157–167. [Google Scholar]
- Rakha, E.A.; Ellis, I.O.; Reis-Filho, J.S. Immunohistochemical heterogeneity of breast carcinomas negative for estrogen receptors, progesterone receptors and Her2/neu (basal-like breast carcinomas). Mod. Pathol 2008, 21, 1060–1061. [Google Scholar]
- Subhawong, A.P.; Subhawong, T.; Nassar, H.; Kouprina, N.; Begum, S.; Vang, R.H.; Westra, W.; Argani, P. Most basal-like breast carcinomas demonstrate the same Rb-/p16+ immunophenotype as the HPV-related poorly differentiated squamous cell carcinomas which they resemble morphologically. Am. J. Surg. Pathol 2009, 33, 163–175. [Google Scholar]
- Reis-Filho, J.S.; Tutt, A.N. Triple negative tumours: A critical review. Histopathology 2008, 52, 108–118. [Google Scholar]
- Nielsen, T.O.; Hsu, F.D.; Jensen, K.; Cheang, M.; Karaca, G.; Hu, Z.; Hernandez-Boussard, T.; Livasy, C.; Cowan, D.; Dressler, L.; et al. Immunohistochemical and clinical characterization of the basal like subtype of invasive breast carcinoma. Clin. Cancer Res 2004, 10, 5367–5374. [Google Scholar]
- Abd El-Rehim, D.M.; Pinder, S.E.; Paish, C.E.; Bell, J.; Blamey, R.W.; Robertson, J.F.; Nicholson, R.I.; Ellis, I.O. Expression of luminal and basal cytokeratins in human breast carcinoma. J. Pathol 2004, 203, 661–671. [Google Scholar]
- Fan, C.; Oh, D.S.; Wessels, L.; Weigelt, B.; Nuyten, D.S.; Nobel, A.B.; van’t Veer, L.J.; Perou, C.M. Concordance among gene-expression-based predictors for breast cancer. N. Engl. J. Med 2006, 355, 560–569. [Google Scholar]
- Peppercorn, J.; Perou, C.M.; Carey, L.A. Molecular subt-ypes in breast cancer evaluation and management: Divide and conquer. Cancer Invest 2008, 26, 1–10. [Google Scholar]
- Tan, D.S.; Marchío, C.; Jones, R.L.; Savage, K.; Smith, I.E.; Dowsett, M.; Reis-Filho, J.S. Triple negative breast cancer: Molecular profiling and prognostic impact in adjuvant anthracycline-treated patients. Breast Cancer Res. Treat 2008, 111, 27–44. [Google Scholar]
- Foulkes, W.D.; Smith, I.E.; Reis-Filho, J.S. Triple-negative breast cancer. N. Engl. J. Med 2010, 363, 1938–1948. [Google Scholar]
- Liedtke, C.; Mazouni, C.; Hess, K.R.; André, F.; Tordai, A.; Mejia, J.A.; Symmans, W.F.; Gonzalez-Angulo, A.M.; Hennessy, B.; Green, M.; et al. Response to neoadjuvant therapy and long term survival in patients with triple-negative breast cancer. J. Clin. Oncol 2008, 26, 1275–1281. [Google Scholar]
- Owens, M.A.; Horten, B.C.; da Silva, M.M. HER2 amplification ratios by fluorescence in situ hybridization and correlation with immunohistochemistry in a cohort of 6556 breast cancer tissues. Clin. Breast Cancer 2004, 5, 63–69. [Google Scholar]
- Baselga, J.; Swain, S.M. Novel anticancer targets: Revisiting ERBB2 and discovering ERBB3. Nat. Rev. Cancer 2009, 9, 463–475. [Google Scholar]
- Srikrishna, G.; Huttunen, H.J.; Johansson, L.; Weigle, B.; Yamaguchi, Y.; Rauvala, H.; Freeze, H.H. N-glycans on the receptor for advanced glycan end products influence amphoterin binding and neurite out growth. J. Neurochem 2002, 80, 998–1008. [Google Scholar]
- Al-Madhagi, Y.; Yan, H.; Rong-Rong, Z.; Guan-Sheng, H.; Mei-Fang, X.; Zhe-Bing, H.; Chao-Jun, D.; Wei, T.; Dao-Lin, T.; Xue-Gong, F. The role of receptor for advanced glycation end products (RAGE) in the proliferation of hepatocellular carcinoma. Int. J. Mol. Sci 2012, 13, 5982–5997. [Google Scholar]
- Logsdon, C.D.; Fuentes, M.K.; Huang, E.H.; Arumugam, T. RAGE and RAGE ligands in cancer. Curr. Mol. Med 2007, 7, 777–789. [Google Scholar]
- Bierhaus, A.; Nawroth, P.P. Multiple levels of regulation determine the role of the receptor for AGE (RAGE) as common soil in inflammation, immune responses and diabetes mellitus and its complications. Diabetologia 2009, 52, 2251–2263. [Google Scholar]
- Kojro, E.; Postina, R. Regulated proteolysis of RAGE and AbetaPP as possible link between type 2 diabetes mellitus and Alzheimer’s disease. J. Alzheimers. Dis 2009, 16, 865–878. [Google Scholar]
- Brett, J.; Schmidt, A.M.; Yan, S.D.; Zou, Y.S.; Weidman, E.; Pinsky, D.; Nowygrod, R.; Neeper, M.; Przysiecki, C.; Shaw, A. Survey of the distribution of a newly characterized receptor for advanced glycation end products in tissues. Am. J. Pathol 1993, 143, 1699–1712. [Google Scholar]
- Clynes, R.; Moser, B.; Yan, S.F.; Ramasamy, R.; Herold, K.; Schmidt, A.M. Receptor for AGE (RAGE): Weaving tangled webs within the inflammatory response. Curr. Mol. Med 2007, 7, 743–751. [Google Scholar]
- Yan, S.F.; Ramasamy, R.; Schmidt, A.M. Receptor for AGE (RAGE) and its ligands-cast into leading roles in diabetes and the inflammatory response. J. Mol. Med 2009, 87, 235–247. [Google Scholar]
- Chen, X.; Walker, D.G.; Schmidt, A.M.; Arancio, O.; Lue, L.F.; Yan, S.D. RAGE: A potential target for Abeta-mediated cellular perturbation in Alzheimer’s disease. Curr. Mol. Med 2007, 7, 735–742. [Google Scholar]
- Liguo, L.; Kaihua, Z.; Zhongkai, S.; Guozhong, W.; Guodong, D. Receptor for advanced glycation end products (RAGE) partially mediates HMGB1-ERKs activation in clear cell renal cell carcinoma. J. Cancer Res. Clin. Oncol 2012, 138, 11–22. [Google Scholar]
- Kiyokazu, H.; Shinichi, U.; Kazuhiro, A.; Fumitake, K.; Masahiko, S.; Ikuro, M.; Masahiro, H.; Shoji, N.; Takashi, A. Novel function of the receptor for advanced glycation end-products (RAGE) in association with tumorigenesis and tumor differentiation of HCC. Ann. Surg. Oncol 2008, 15, 923–933. [Google Scholar]
- Yamamoto, Y.; Kato, I.; Doi, T.; Yonekura, H.; Ohashi, S.; Takeuchi, M.; Watanabe, T.; Yamagishi, S.; Sakurai, S.; Takasawa, S.; et al. Development and prevention of advanced diabetic nephropathy in RAGE-overexpressing mice. J. Clin. Invest 2001, 108, 261–268. [Google Scholar]
- Wang, H.; Bloom, O.; Zhang, M.; Vishnubhakat, J.M.; Ombrellino, M.; Che, J.; Frazier, A.; Yang, H.; Ivanova, S.; Borovikova, L.; et al. HMG-1 as a late mediator of endotoxin lethality in mice. Science 1999, 285, 248–251. [Google Scholar]
- Liliensiek, B.; Weigand, M.A.; Bierhaus, A.; Nicklas, W.; Kasper, M.; Hofer, S.; Plachky, J.; Gröne, H.J.; Kurschus, F.C.; Schmidt, A.M.; et al. Receptor for advanced glycation end products (RAGE) regulates sepsis but not the adaptive immune response. J. Clin. Invest 2004, 113, 1641–1650. [Google Scholar]
- Hudson, B.I.; Kalea, A.Z.; Arriero, M.D.; Harja, E.; Boulanger, E.; D’Agati, V.; Schmidt, A.M. Interaction of the RAGE cytoplasmic domain with diaphanous-1 is required for ligand-stimulated cellular migration through activation of Rac1 and Cdc42. J. Biol. Chem 2008, 283, 34457–34468. [Google Scholar]
- Demling, N.; Ehrhardt, C.; Kasper, M.; Laue, M.; Knels, L.; Rieber, E. Promotion of cell adherence and spreading: A novel function of RAGE, the highly selective differentiation marker of human alveolar epithelial type I cells. Cell Tissue Res 2006, 323, 475–488. [Google Scholar]
- Ramasamy, R.; Yan, S.F.; Schmidt, A.M. Arguing for the motion. Yes, RAGE is a receptor for advanced glycation end products. Mol. Nutr. Food Res 2007, 51, 1111–1115. [Google Scholar]
- Brezniceanu, M.L.; Volp, K.; Bosser, S.; Solbach, C.; Lichter, P.; Joos, S.; Zornig, M. HMGB1 inhibits cell death in yeast and mammalian cells and is abundantly expressed in human breast carcinoma. FASEB J 2003, 17, 1295–1297. [Google Scholar]
- Flohr, A.M.; Rogalla, P.; Meiboom, M.; Borrmann, L.; Krohn, M.; Thode-Halle, B.; Bullerdiek, J. Variation of HMGB1 expression in breast cancer. Anticancer Res 2001, 21, 3881–3885. [Google Scholar]
- Lum, H.K.; Lee, K.L. The human HMGB1 promoter is modulated by a silencer and an enhancer-containing intron. Biochim. Biophys. Acta 2001, 1520, 79–84. [Google Scholar]
- Hsieh, H.L.; Schafer, B.W.; Sasaki, N.; Heizmann, C.W. Expression analysis of S100 proteins and RAGE in human tumors using tissue microarrays. Biochem. Biophys. Res. Commun 2003, 307, 375–381. [Google Scholar]
- Taguchi, A.; Blood, D.C.; del Toro, G.; Canet, A.; Lee, D.C.; Qu, W.; Tanji, N.; Lu, Y.; Lalla, E.; Fu, C.; et al. Blockade of RAGE amphoterin signaling suppresses tumor growth and metastases. Nature 2000, 405, 354–360. [Google Scholar]
- Kuniyasu, H.; Oue, N.; Wakikawa, A.; Shigieshi, H.; Matsutani, N.; Kuraoka, K.; Ito, R.; Yokozaki, H.; Yasui, W. Expression of receptors for advanced glycation end-products (RAGE) is closely associated with the invasive and metastatic activity of gastric cancer. J. Pathol 2002, 196, 163–170. [Google Scholar]
- Kuniyasu, H.; Chihara, Y.; Kondo, H. Differential effects between amphoterin and advanced glycation end products on colon cancer cells. Int. J. Cancer 2003, 104, 722–727. [Google Scholar]
- Kuniyasu, H.; Chihara, Y.; Takahashi, T. Co-Expression of receptor for advanced glycation end products and the ligand amphoterin associates closely with metastasis of colorectal cancer. Oncol. Rep 2003, 10, 445–448. [Google Scholar]
- Hirata, K.; Takada, M.; Suzuki, Y.; Kuroda, Y. Expression of receptor for advanced glycation end products (RAGE) in human biliary cancer cells. Hepatogastroenterology 2003, 50, 1205–1207. [Google Scholar]
- Takada, M.; Koizumi, T.; Toyama, H.; Suzuki, Y.; Kuroda, Y. Differential expression of RAGE in human pancreatic carcinoma cells. Hepatogastroenterology 2001, 48, 1577–1578. [Google Scholar]
- Ishiguro, H.; Nakaigawa, N.; Miyoshi, Y.; Fujinami, K.; Kubota, Y.; Uemura, H. Receptor for advanced glycation end products (RAGE) and its ligand, amphoterin are overexpressed and associated with prostate cancer development. Prostate 2005, 64, 92–100. [Google Scholar]
- Karin, M.; Ben-Neriah, Y. Phosphorylation meets ubiquitination: The control of NF-κB activity. Annu. Rev. Immunol 2000, 18, 621–663. [Google Scholar]
- Cheng, Q.; Cant, C.A.; Moll, T.; Hofer-Warbinek, R.; Wagner, E.; Birnstiel, M.L.; Bach, F.H.; de Martin, R. NK-kappa B subunit-specific regulation of the I kappa B alpha promoter. J. Biol. Chem 1994, 269, 13551–13557. [Google Scholar]
- Yeh, C.H.; Sturgis, L.; Haidacher, J.; Zhang, X.N.; Sherwood, S.J.; Bjercke, R.J.; Juhasz, O.; Crow, M.T.; Tilton, R.G.; Denner, L. Requirement for p38 and p44/p42 mitogen-activated protein kinases in RAGE mediated nuclear factor-κB transcriptional activation and cytokine secretion. Diabetes 2001, 50, 1495–1504. [Google Scholar]
- Huttunen, H.J.; Fages, C.; Rauvala, H. Receptor for advanced glycation end products (RAGE)-mediated neurite outgrowth and activation of NF-κB require the cytoplasmic domain of the receptor but different downstream signaling pathways. J. Biol. Chem 1999, 274, 19919–19924. [Google Scholar]
- Gilmore, T.D. Introduction to NF-κB: Players, pathways, perspectives. Oncogene 2006, 25, 6680–6684. [Google Scholar]
- Brasier, A.R. The NF-κB regulatory network. Cardiovasc. Toxicol 2006, 6, 111–130. [Google Scholar]
- Perkins, N.D. Integrating cell-signalling pathways with NF-κB and IKK function. Nat. Rev. Mol. Cell Biol 2007, 8, 49–62. [Google Scholar]
- Tian, B.; Brasier, A.R. Identification of a nuclear factor NF-κB -dependent gene network. Recent Prog. Horm. Res 2003, 58, 95–130. [Google Scholar]
- Jones, R.L.; Rojo, F.G.; A’Hern, R.; Villena, N.; Corominas, J.; Servitja, S.; Corominas, J.M.; Servitja, S.; Smith, I.E.; Rovira, A.; et al. Nuclear NF-κB p65 expression and response to neoadjuvant chemotherapy in breast cancer. J. Clin. Pathol 2011, 64, 130–135. [Google Scholar]
- Gramantieri, L.; Trere, D.; Chieco, P.; Lacchini, M.; Giovannini, C.; Piscaglia, F.; Cavallari, A.; Bolondi, L. In human hepatocellular carcinoma in cirrhosis proliferating cell nuclear antigen (PCNA) is involved in cell proliferation and cooperates with P21 in DNA repair. J Hepatol 2003, 39, 997–1003. [Google Scholar]
- Daolin, T.; Rui, K.; Herbert, J.Z.; Michael, T.L. High-mobility group box 1 and cancer. Biochim. Biophys. Acta 2010, 1799, 131–140. [Google Scholar]
- Hinz, M.; Krappmann, D.; Eichten, A.; Heder, A.; Scheidereit, C.; Strauss, M. NF-κB function in growth control, Regulation of cyclin D1 expression and G0/G1-to-S-Phase transition. Mol. Cell Biol 1999, 19, 2690–2698. [Google Scholar]

© 2013 by the authors; licensee MDPI, Basel, Switzerland This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution license (http://creativecommons.org/licenses/by/3.0/).
Share and Cite
Radia, A.-M.; Yaser, A.-M.; Ma, X.; Zhang, J.; Yang, C.; Dong, Q.; Rong, P.; Ye, B.; Liu, S.; Wang, W. Specific siRNA Targeting Receptor for Advanced Glycation End Products (RAGE) Decreases Proliferation in Human Breast Cancer Cell Lines. Int. J. Mol. Sci. 2013, 14, 7959-7978. https://doi.org/10.3390/ijms14047959
Radia A-M, Yaser A-M, Ma X, Zhang J, Yang C, Dong Q, Rong P, Ye B, Liu S, Wang W. Specific siRNA Targeting Receptor for Advanced Glycation End Products (RAGE) Decreases Proliferation in Human Breast Cancer Cell Lines. International Journal of Molecular Sciences. 2013; 14(4):7959-7978. https://doi.org/10.3390/ijms14047959
Chicago/Turabian StyleRadia, AL-Madhagi, AL-Madhagi Yaser, Xiaoqian Ma, Juan Zhang, Cejun Yang, Qiong Dong, Pengfei Rong, Bin Ye, Sheng Liu, and Wei Wang. 2013. "Specific siRNA Targeting Receptor for Advanced Glycation End Products (RAGE) Decreases Proliferation in Human Breast Cancer Cell Lines" International Journal of Molecular Sciences 14, no. 4: 7959-7978. https://doi.org/10.3390/ijms14047959



