EGFR Expression and KRAS and BRAF Mutational Status in Intestinal-Type Sinonasal Adenocarcinoma
Abstract
:1. Introduction
2. Results and Discussion
3. Experimental Section
3.1. Study Population
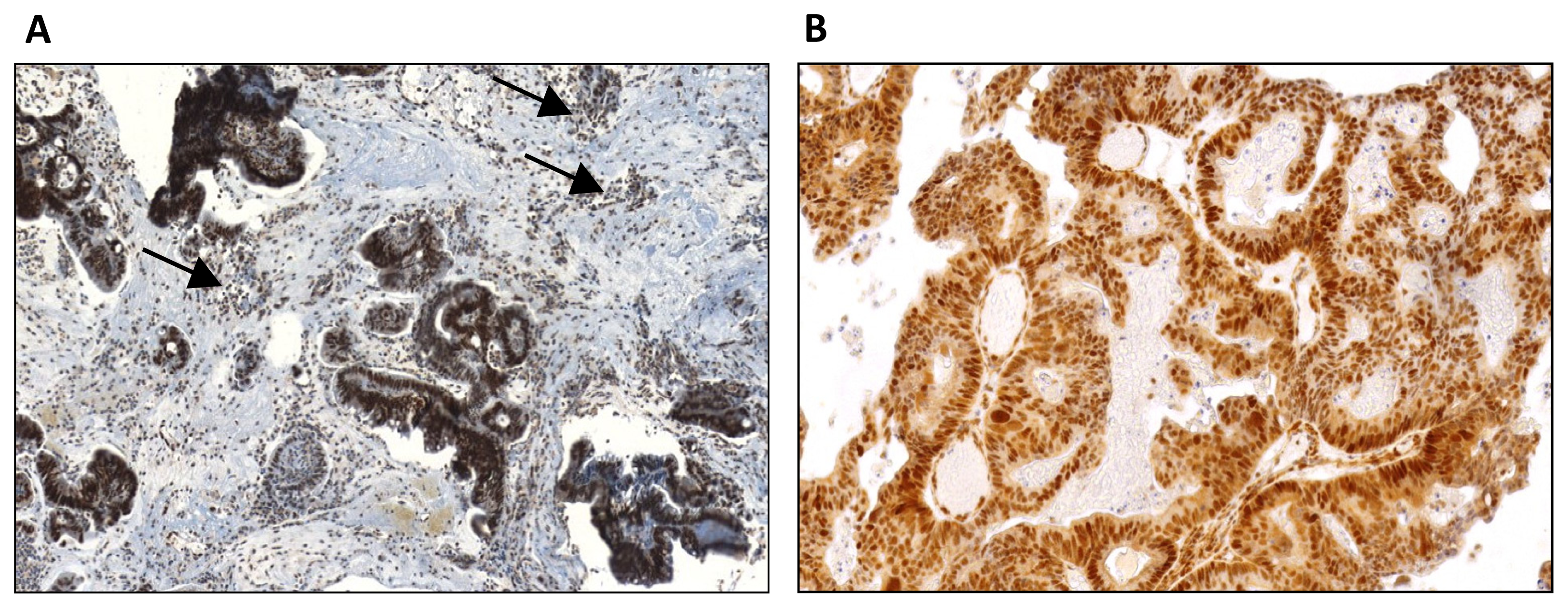
3.2. EGFR and hMLH1 and hMSH2 Expression Determination
3.3. KRAS and BRAF Analysis
- KRAS-F: 5′-GGCCTGCTGAAAATGACTGAA-3′;
- KRAS-R: 5′-ATTAGCTGTATCGTCAAGGCACTC-3′;
- BRAF-F: 5′-ATGAAGACCTCACAGTAAAAAJAGG-3′;
- BRAF-R: 5′-AGCAGCATCTTAGGGCCAAA-3′.
3.4. Statistical Analysis
4. Conclusions
Conflict of Interest
References
- 't Mannetje, A.; Kogevinas, M.; Luce, D.; Demers, P.A.; Bégin, D.; Bolm-Audorff, U.; Comba, P.; Gérin, M.; Hardell, L.; Hayes, R.B.; et al. Sinonasal cancer, occupation, and tobacco smoking in European women and men. Am. J. Ind. Med 1999, 36, 101–107. [Google Scholar]
- Turner, J.H.; Reh, D.D. Incidence and survival in patients with sinonasal cancer: A historical analysis of population-based data. Head Neck 2012, 34, 877–885. [Google Scholar]
- Barnes, L. Intestinal-type adenocarcinoma of the nasal cavity and paranasal sinuses. Am. J. Surg. Pathol 1986, 10, 192–202. [Google Scholar]
- Van Cutsem, E.; Köhne, C.-H.; Láng, I.; Folprecht, G.; Nowacki, M.P.; Cascinu, S.; Shchepotin, I.; Maurel, J.; Cunningham, D.; Tejpar, S.; et al. Cetuximab plus irinotecan, fluorouracil, and leucovorin as first-line treatment for metastatic colorectal cancer: Updated analysis of overall survival according to tumor KRAS and BRAF mutation status. J. Clin. Oncol 2011, 29, 2011–2019. [Google Scholar]
- Llorente, J.L.; Pérez-Escuredo, J.; Alvarez-Marcos, C.; Suárez, C.; Hermsen, M. Genetic and clinical aspects of wood dust related intestinal-type sinonasal adenocarcinoma: A review. Eur. Arch. Otorhinolaryngol 2009, 266, 1–7. [Google Scholar]
- Pérez-Escuredo, J.; Martínez, J.G.; Vivanco, B.; Marcos, C.Á.; Suárez, C.; Llorente, J.L.; Hermsen, M.A. Wood dust-related mutational profile of TP53 in intestinal-type sinonasal adenocarcinoma. Hum. Pathol. 2012, 43, 1894–1901. [Google Scholar]
- WHO Histological classification of tumors of the nasal cavity and paranasal sinuses. Available online: http://www.iarc.fr/en/publications/pdfs-online/pat-gen/bb9/bb9-chap1.pdf accessed on 27 February 2013.
- Franchi, A.; Fondi, C.; Paglierani, M.; Pepi, M.; Gallo, O.; Santucci, M. Epidermal growth factor receptor expression and gene copy number in sinonasal intestinal type adenocarcinoma. Oral Oncol 2009, 45, 835–838. [Google Scholar]
- Spano, J.-P.; Lagorce, C.; Atlan, D.; Milano, G.; Domont, J.; Benamouzig, R.; Attar, A.; Benichou, J.; Martin, A.; Morere, J.F.; et al. Impact of EGFR expression on colorectal cancer patient prognosis and survival. Ann. Oncol 2005, 16, 102–108. [Google Scholar]
- Saif, M.W. Colorectal cancer in review: The role of the EGFR pathway. Expert Opin. Invest. Drugs 2010, 19, 357–369. [Google Scholar]
- Goldstein, N.S.; Armin, M. Epidermal growth factor receptor immunohistochemical reactivity in patients with American Joint Committee on Cancer Stage IV colon adenocarcinoma: Implications for a standardized scoring system. Cancer 2001, 92, 1331–1346. [Google Scholar]
- Ooi, A.; Takehana, T.; Li, X.; Suzuki, S.; Kunitomo, K.; Iino, H.; Fujii, H.; Takeda, Y.; Dobashi, Y. Protein overexpression and gene amplification of HER-2 and EGFR in colorectal cancers: An immunohistochemical and fluorescent in situ hybridization study. Mod. Pathol 2004, 17, 895–904. [Google Scholar]
- Pai, R.K.; Jayachandran, P.; Koong, A.C.; Chang, D.T.; Kwok, S.; Ma, L.; Arber, D.A.; Balise, R.R.; Tubbs, R.R.; Shadrach, B.; et al. BRAF-mutated, microsatellite-stable adenocarcinoma of the proximal colon: An aggressive adenocarcinoma with poor survival, mucinous differentiation, and adverse morphologic features. Am. J. Surg. Pathol 2012, 36, 744–752. [Google Scholar]
- Borràs, E.; Jurado, I.; Hernan, I.; Gamundi, M.J.; Dias, M.; Martí, I.; Mañé, B.; Arcusa, À.; Agúndez, J.A.G.; Blanca, M.; et al. EGFR mutations by high resolution melting analysis and ultra-deep pyrosequencing. BMC Cancer 2011, 11, 406. [Google Scholar]
- Normanno, N.; Pinto, C.; Castiglione, F.; Bardelli, A.; Gambacorta, M.; Botti, G.; Nappi, O.; Siena, S.; Ciardiello, F.; Taddei, G.; et al. KRAS mutations testing in colorectal carcinoma patients in Italy: From guidelines to external quality assessment. PLoS One 2011, 6, e29146. [Google Scholar]
- Frattini, M.; Perrone, F.; Suardi, S.; Balestra, D.; Caramuta, S.; Colombo, F.; Licitra, L.; Cantù, G.; Pierotti, M.A.; Pilotti, S. Phenotype-genotype correlation: Challenge of intestinal-type adenocarcinoma of the nasal cavity and paranasal sinuses. Head Neck 2006, 28, 909–915. [Google Scholar]
- Yom, S.S.; Rashid, A.; Rosenthal, D.I.; Elliott, D.D.; Hanna, E.Y.; Weber, R.S.; El-Naggar, A.K. Genetic analysis of sinonasal adenocarcinoma phenotypes: Distinct alterations of histogenetic significance. Mod. Pathol 2005, 18, 315–319. [Google Scholar]
- López, F.; García Inclán, C.; Pérez-Escuredo, J.; Alvarez Marcos, C.; Scola, B.; Suárez, C.; Llorente, J.L.; Hermsen, M.A. KRAS and BRAF mutations in sinonasal cancer. Oral Oncol 2012, 48, 692–697. [Google Scholar]
- Saber, A.T.; Nielsen, L.R.; Dictor, M.; Hagmar, L.; Mikoczy, Z.; Wallin, H. K-ras mutations in sinonasal adenocarcinomas in patients occupationally exposed to wood or leather dust. Cancer Lett 1998, 126, 59–65. [Google Scholar]
- Wu, T.T.; Barnes, L.; Bakker, A.; Swalsky, P.A.; Finkelstein, S.D. K-ras-2 and p53 genotyping of intestinal-type adenocarcinoma of the nasal cavity and paranasal sinuses. Mod. Pathol 1996, 9, 199–204. [Google Scholar]
- Pérez, P.; Dominguez, O.; González, S.; González, S.; Triviño, A.; Suárez, C. Ras gene mutations in ethmoid sinus adenocarcinoma: Prognostic implications. Cancer 1999, 86, 255–264. [Google Scholar]
- Yokota, T. Are KRAS/BRAF mutations potent prognostic and/or predictive biomarkers in colorectal cancers? Anticancer Agents Med. Chem 2012, 12, 163–171. [Google Scholar]
- Nash, G.M.; Gimbel, M.; Cohen, A.M.; Zeng, Z.S.; Ndubuisi, M.I.; Nathanson, D.R.; Ott, J.; Barany, F.; Paty, P.B. KRAS mutation and microsatellite instability: Two genetic markers of early tumor development that influence the prognosis of colorectal cancer. Ann. Surg. Oncol 2010, 17, 416–424. [Google Scholar]
- Samowitz, W.S.; Curtin, K.; Schaffer, D.; Robertson, M.; Leppert, M.; Slattery, M.L. Slattery, Relationship of Ki-ras mutations in colon cancers to tumor location, stage, and survival: A population-based study. Cancer Epidemiol. Biomarkers Prev 2000, 9, 1193–1197. [Google Scholar]
- Belly, R.T.; Rosenblatt, J.D.; Steinmann, M.; Toner, J.; Sun, J.; Shehadi, J.; Peacock, J.L.; Raubertas, R.F.; Jani, N.; Ryan, C.K. Detection of mutated K12-ras in histologically negative lymph nodes as an indicator of poor prognosis in stage II colorectal cancer. Clin. Colorectal Cancer 2001, 1, 110–116. [Google Scholar]
- Inoue, Y.; Saigusa, S.; Iwata, T.; Okugawa, Y.; Toiyama, Y.; Tanaka, K.; Uchida, K.; Mohri, Y.; Kusunoki, M. The prognostic value of KRAS mutations in patients with colorectal cancer. Oncol. Rep 2012, 28, 1579–1584. [Google Scholar]
- Andreyev, H.J.; Norman, A.R.; Cunningham, D.; Oates, J.; Dix, B.R.; Iacopetta, B.J.; Young, J.; Walsh, T.; Ward, R.; Hawkins, N.; et al. Kirsten Ras mutations in patients with colorectal cancer: The “RASCAL II” study. Br. J. Cancer 2001, 85, 692–696. [Google Scholar]

| Characteristics | Number of patients (%) |
|---|---|
| Sex | |
| Male | 42 (98) |
| Female | 1 (2) |
| Age (years) | |
| Range | 39–80 |
| Median | 66 |
| Stage | |
| T1 | 1 (2) |
| T2 | 16 (37) |
| T3 | 9 (21) |
| T4 | 17 (40) |
| N | |
| Yes | 3 (7) |
| No | 39 (91) |
| NA | 1 (2) |
| M | |
| Yes | 6 (14) |
| No | 36 (84) |
| NA | 1 (2) |
| Histological type | |
| Papillary | 3 (7) |
| Colonic | 25 (58.1) |
| Solid | 5 (11.6) |
| Mucinous | 10 (23.3) |
| Wood dust exposure | |
| Yes | 26 (60) |
| No | 9 (21) |
| NA | 8 (19) |
| Recurrence | |
| Yes | 24 (56) |
| No | 17 (40) |
| NA | 2 (4) |
| Outcome | |
| Alive | 16 (37) |
| Died of disease | 16 (37) |
| Died of other causes | 9 (21) |
| NA | 2 (5) |
| Case | T | N | M | Wood exposure | Histology | EGFR overexpression | MLH1/MSH2 expression | KRAS exon 2 | BRAF exon 15 |
|---|---|---|---|---|---|---|---|---|---|
| 1 | T4 | No | No | No | Solid | No | Yes | Wild-type | Wild-type |
| 2 | T2 | No | No | Yes | Colonic | No | Yes | Not amplified | Not amplified |
| 3 | T4 | Yes | Yes | NA | Solid | No | Yes | Wild-type | Wild-type |
| 4 | T4 | No | No | No | Colonic | Yes | Yes | Mutation | Wild-type |
| 5 | T4 | No | No | Na | Mucinous | No | Yes | Not amplified | Not amplified |
| 6 | T2 | No | No | Yes | Colonic | No | Yes | Mutation | Wild-type |
| 7 | T2 | No | No | Yes | Colonic | No | Yes | Mutation | Wild-type |
| 8 | T4 | No | No | Yes | Colonic | No | Yes | Not amplified | Not amplified |
| 9 | T4 | No | Yes | No | Mucinous | No | Yes | Not amplified | Not amplified |
| 10 | T2 | No | No | No | Colonic | Yes | Yes | Wild-type | Wild-type |
| 11 | T2 | No | No | No | Mucinous | No | Yes | Wild-type | Wild-type |
| 12 | T2 | No | No | Yes | Colonic | No | Yes | Wild-type | Wild-type |
| 13 | T2 | No | No | NA | Colonic | Yes | Yes | Mutation | Wild-type |
| 14 | T2 | No | No | Yes | Papillary | No | Yes | Mutation | Wild-type |
| 15 | T3 | No | No | Yes | Colonic | No | Yes | Mutation | Wild-type |
| 16 | T3 | No | No | Yes | Colonic | No | Yes | Wild-type | Wild-type |
| 17 | T3 | No | No | No | Mucinous | No | Yes | Not amplified | Not amplified |
| 18 | T2 | No | No | Yes | Colonic | No | Yes | Wild-type | Wild-type |
| 19 | T2 | Yes | No | Yes | Solid | No | Yes | Mutation | Wild-type |
| 20 | T2 | No | No | Yes | Colonic | No | Yes | Not amplified | Not amplified |
| 21 | T4 | No | No | Yes | Mucinous | No | Yes | Not amplified | Not amplified |
| 22 | T2 | No | No | Yes | Colonic | No | Yes | Wild-type | Mutation |
| 23 | T2 | No | No | Yes | Mucinous | No | Yes | Not amplified | Wild-type |
| 24 | T3 | No | No | Yes | Mucinous | No | Yes | Mutation | Wild-type |
| 25 | T3 | No | No | Yes | Colonic | Yes | Yes | Not amplified | Not amplified |
| 26 | T2 | No | No | Yes | Papillary | Yes | Yes | Not amplified | Not amplified |
| 27 | T2 | Yes | Yes | Yes | Mucinous | No | Yes | Mutation | Wild-type |
| 28 | T4 | No | No | No | Colonic | Yes | Yes | Wild-type | Wild-type |
| 29 | T4 | No | No | No | Colonic | Yes | Yes | Wild-type | Wild-type |
| 30 | T3 | No | No | Yes | Colonic | Yes | Yes | Wild-type | Wild-type |
| 31 | T4 | No | No | NA | Solid | No | Yes | Wild-type | Wild-type |
| 32 | T1 | No | No | NA | Colonic | No | Yes | Wild-type | Wild-type |
| 33 | T4 | No | Yes | Yes | Colonic | Yes | Yes | Wild-type | Wild-type |
| 34 | T4 | No | No | Yes | Mucinous | No | Yes | Mutation | Wild-type |
| 35 | T4 | No | Yes | NA | Solid | No | No evaluable | Not amplified | Not amplified |
| 36 | T4 | No | No | Yes | Colonic | Yes | Yes | Wild-type | Wild-type |
| 37 | T3 | No | No | Yes | Colonic | Yes | Yes | Not amplified | Not amplified |
| 38 | T2 | No | No | NA | Colonic | No | Yes | Mutation | Wild-type |
| 39 | T4 | No | Yes | Yes | Colonic | Yes | Yes | Not amplified | Not amplified |
| 40 | T3 | No | No | Yes | Colonic | No | Yes | Not amplified | Not amplified |
| 41 | T4 | NA | NA | NA | Mucinous | No | Yes | Mutation | Wild-type |
| 42 | T3 | No | No | Yes | Colonic | Yes | Yes | Not amplified | Not amplified |
| 43 | T4 | No | No | No | Papillary | No | Yes | Wild-type | Wild-type |
© 2013 by the authors; licensee MDPI, Basel, Switzerland. This article is an open-access article distributed under the terms and conditions of the Creative Commons Attribution license (http://creativecommons.org/licenses/by/3.0/).
Share and Cite
Szablewski, V.; Solassol, J.; Poizat, F.; Larrieux, M.; Crampette, L.; Mange, A.; Bascoul-Mollevi, C.; Costes, V. EGFR Expression and KRAS and BRAF Mutational Status in Intestinal-Type Sinonasal Adenocarcinoma. Int. J. Mol. Sci. 2013, 14, 5170-5181. https://doi.org/10.3390/ijms14035170
Szablewski V, Solassol J, Poizat F, Larrieux M, Crampette L, Mange A, Bascoul-Mollevi C, Costes V. EGFR Expression and KRAS and BRAF Mutational Status in Intestinal-Type Sinonasal Adenocarcinoma. International Journal of Molecular Sciences. 2013; 14(3):5170-5181. https://doi.org/10.3390/ijms14035170
Chicago/Turabian StyleSzablewski, Vanessa, Jérôme Solassol, Flora Poizat, Marion Larrieux, Louis Crampette, Alain Mange, Caroline Bascoul-Mollevi, and Valérie Costes. 2013. "EGFR Expression and KRAS and BRAF Mutational Status in Intestinal-Type Sinonasal Adenocarcinoma" International Journal of Molecular Sciences 14, no. 3: 5170-5181. https://doi.org/10.3390/ijms14035170




