Let-7 in Cardiovascular Diseases, Heart Development and Cardiovascular Differentiation from Stem Cells
Abstract
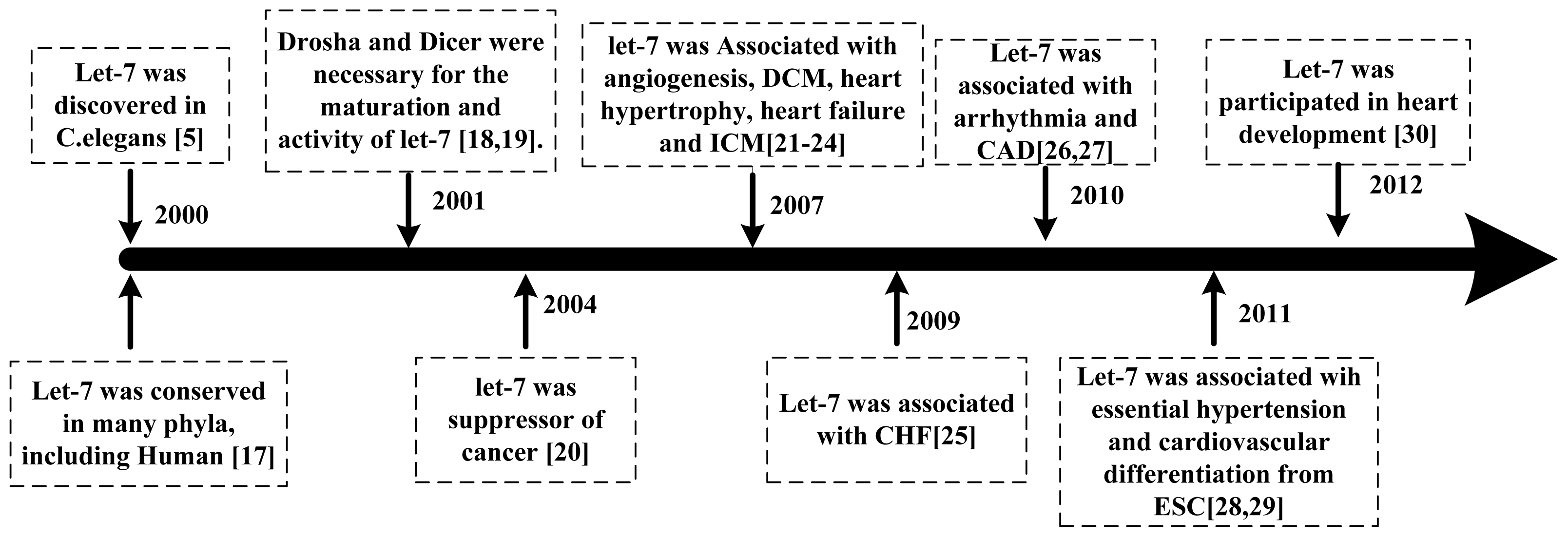
:1. Introduction
2. Let-7 in Cardiovascular Diseases
2.1. Let-7 in Cardiac Hypertrophy, Heart Development and Cardiac Fibrosis
2.2. Let-7 in Myocardial Infarction (MI) and Heart Failure
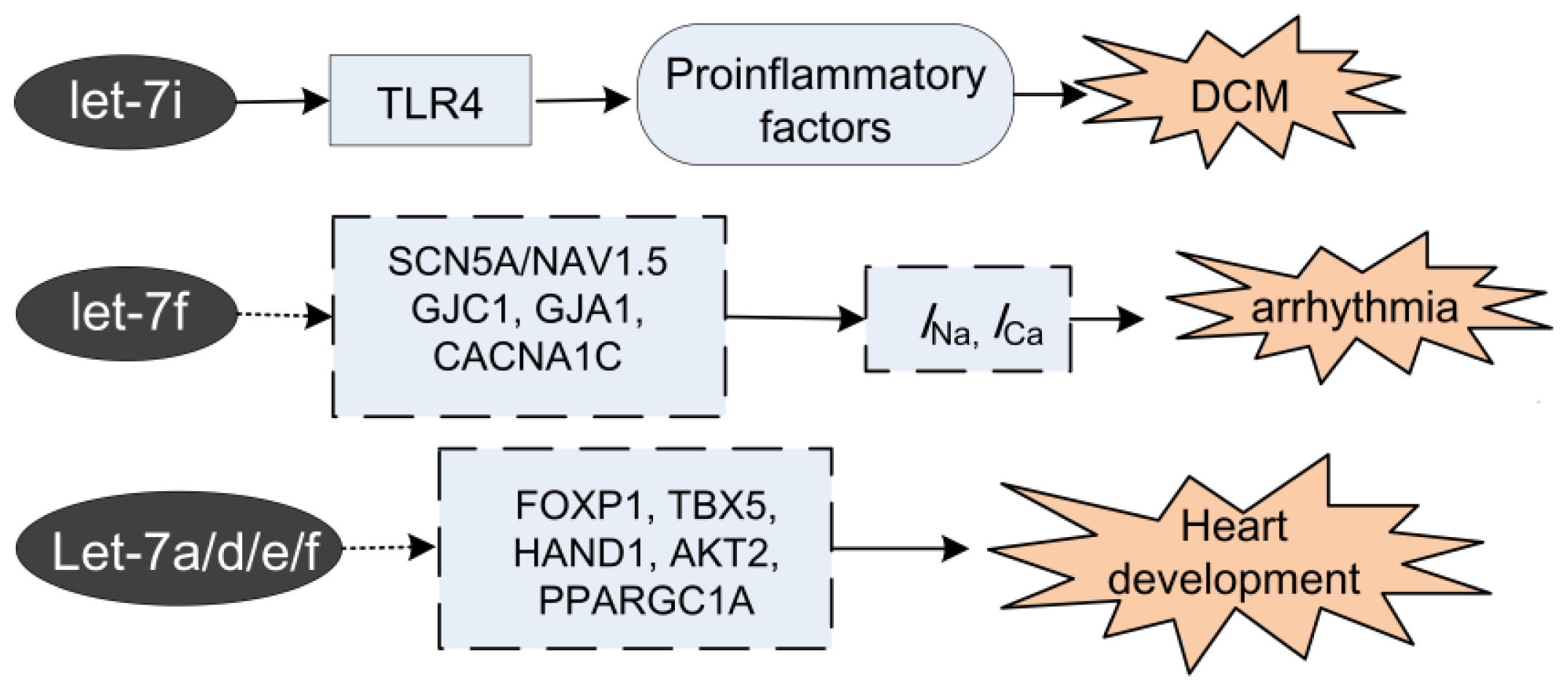
2.3. Let-7 in Dilated Cardiomyopathy (DCM)
2.4. Let-7 in Arrhythmia
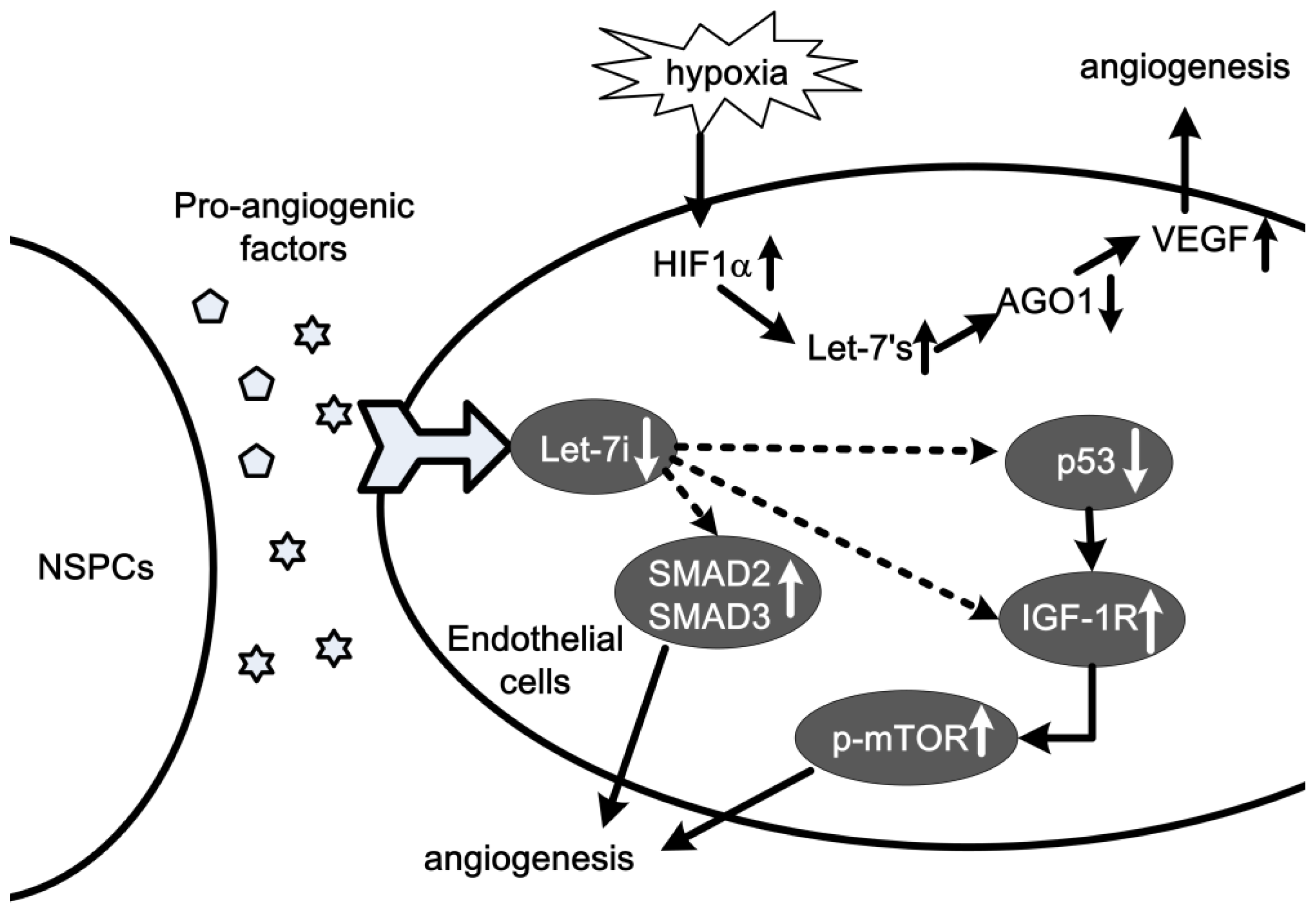
2.5. Let-7 in Angiogenesis
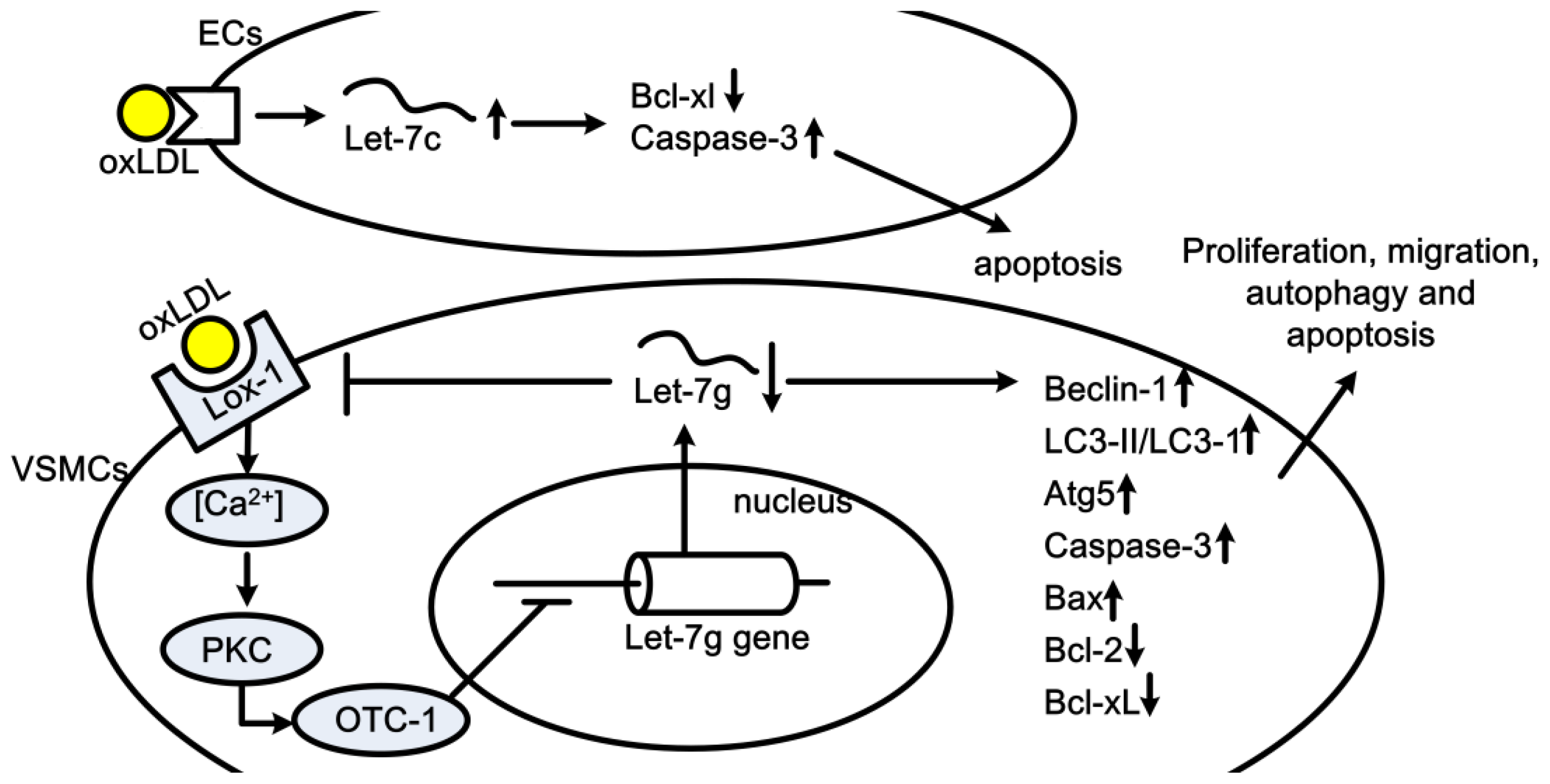
2.6. Let-7 in Atherogenesis and Coronary Artery Disease
2.7. Let-7 in Hypertension
3. Let-7 in Heart Development
4. Let-7 in Cardiovascular Differentiation of Embryonic Stem Cells
5. Circulating Let-7 as a Mediator of Intercellular Communication and as a Biomarker of Cardiovascular Disease
6. Conclusion and Perspective
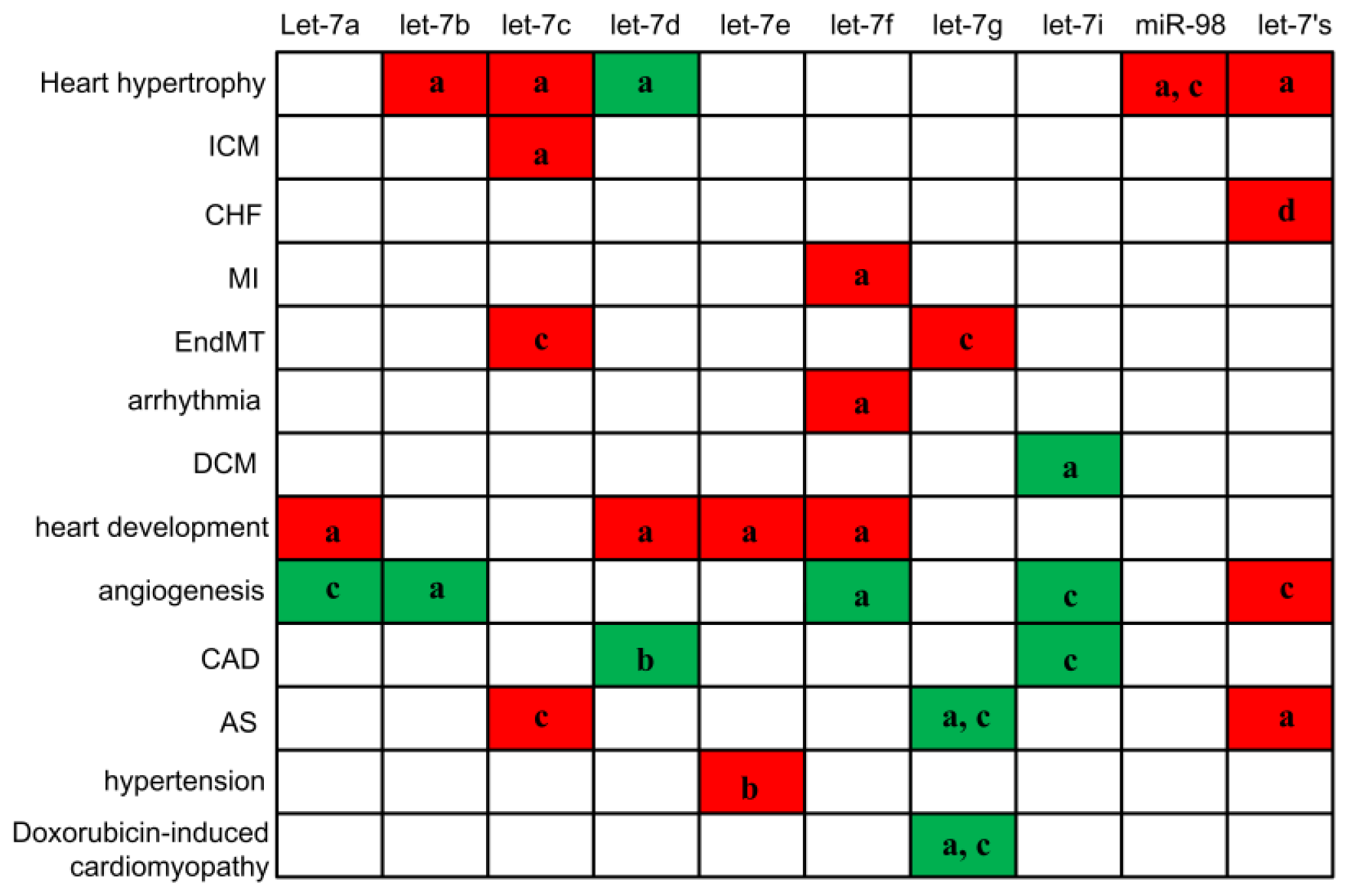
| Let-7 member | Up-/downregulation during cardiovascular diseases | Tissue or cell samples used | Predicted (in italic) or confirmed targeted genes | References |
|---|---|---|---|---|
| Let-7a | Heart development ↑ | Mouse fetal hearts | (FOXP1, TBX5, HAND1, AKT2, PPARGC1A) | [30] |
| Angiogenesis ↓ | Human endothelial cells | Nrp-2 | [53] | |
| Let-7b | Heart hypertrophy ↑ | Mouse heart | Undetected | [21] |
| Angiogenesis ↓ | Mouse ovary vessel | TIMP-1 c-Met | [54] | |
| Let-7c | Heart hypertrophy ↑ | Mouse heart | Undetected | [21] |
| ICM ↑ | Human left ventricular | Undetected | [22] | |
| EndMT ↑ | Mouse cardiac endothelial cells | Undetected | [40] | |
| AS ↑ | human endothelial cells | Bcl-xl | [62] | |
| Let-7e | Heart development ↑ | Mouse fetal hearts | (FOXP1, TBX5, HAND1, AKT2, PPARGC1A) | [30] |
| Hypertension ↑ | Human plasma samples (the origin is endothelial cells) | Undetected | [70] | |
| Let-7f | Arrhythmia ↑ | Rat hearts | Undetected | [26] |
| MI ↑ | Rat hearts | Undetected | [26] | |
| Heart development ↑ | Mouse fetal hearts | (FOXP1, TBX5, HAND1, AKT2, PPARGC1A) | [30] | |
| Angiogenesis ↓ | Human endothelial cells | thrombospondin-1, TSP-2 | [52] | |
| Let-7g | EndMT ↑ | Mouse cardiac endothelial cells | Undetected | [40] |
| AS ↓ | VSMCs, and mice aorta | Lectin-like LDL receptor 1 | [63] | |
| Doxorubicin-induced cardiomyopathy ↓ | Rat hearts and cardiac myocytes | Undetected | [42] | |
| Let-7i | DCM ↓ | Endomyocardial biopsy tissues | Toll like receptor 4 | [24] |
| Angiogenesis ↓ | Human endothelial cells | IGF-1R | [57] | |
| CAD ↓ | THP-1 cells and patient blood monocytes | Toll like receptor 4 | [64] | |
| MiR-98 | Heart hypertrophy ↑ | Mouse heart, and cardiac myocyte | Thioredoxin 1 | [34,35] |
| Let-7’s | Heart hypertrophy ↑ | Mouse heart | Undetected | [34] |
| CHF ↑ | Bioinformatics studies | Undetected | [25] | |
| Atherosclerotic AAA ↑ | Human aortic aneurysm | Undetected | [65] | |
| Angiogenesis ↑ | Human endothelial cells | Argonaute 1 | [11] | |
| ESC differentiation to myocardiac cells ↑ | ESCs and ESC-derived cardiomyocytes | Undetected | [28,29,71] | |
| iPS differentiation to cardiomyocytes ↓ | iPS cells and iPS-derived cardiomyocytes | Undetected | [73] |
Acknowledgments
Conflicts of Interest
References
- Bartel, D.P. MicroRNAs: Genomics, biogenesis, mechanism, and function. Cell 2004, 116, 281–297. [Google Scholar]
- Pushparaj, P.N.; Aarthi, J.J.; Kumar, S.D.; Manikandan, J. RNAi and RNAa—The yin and yang of RNAome. Bioinformatics 2008, 2, 235–237. [Google Scholar]
- Lewis, B.P.; Burge, C.B.; Bartel, D.P. Conserved seed pairing, often flanked by adenosines, indicates that thousands of human genes are microRNA targets. Cell 2005, 120, 15–20. [Google Scholar]
- Lee, R.C.; Feinbaum, R.L.; Ambros, V. The C. elegans heterochronic gene lin-4 encodes small RNAs with antisense complementarity to lin-14. Cell 1993, 75, 843–854. [Google Scholar]
- Reinhart, B.J.; Slack, F.J.; Basson, M.; Pasquinelli, A.E.; Bettinger, J.C.; Rougvie, A.E.; Horvitz, H.R.; Ruvkun, G. The 21-nucleotide let-7 RNA regulates developmental timing in Caenorhabditis elegan s. Nature 2000, 403, 901–906. [Google Scholar]
- MiRBase. Available online: http://www.mirbase.org/cgi-bin/browse.pl?org=hsa8 (accessed on 14 September 2013).
- Roush, S.; Slack, F.J. The let-7 family of microRNAs. Trends Cell Biol 2008, 18, 505–516. [Google Scholar]
- Boyerinas, B.; Park, S.M.; Hau, A.; Murmann, A.E.; Peter, M.E. The role of let-7 in cell differentiation and cancer. Endocr. Relat. Cancer 2010, 17, F19–F36. [Google Scholar]
- Charles, D.J.; Aurora, E.K.; Giovanni, S. The let-7 microRNA represses cell proliferation pathways in human cells. Cancer Res 2007, 67, 7713–7722. [Google Scholar]
- Ding, Z.; Wang, X.; Schnackenberg, L.; Khaidakov, M.; Liu, S.; Singla, S.; Dai, Y.; Mehta, J.L. Regulation of autophagy and apoptosis in response to ox-LDL in vascular smooth muscle cells, and the modulatory effects of the microRNA hsa-let-7g. Int. J. Cardiol 2013. [Google Scholar] [CrossRef]
- Chen, Z.; Lai, T.C.; Jan, Y.H.; Lin, F.M.; Wang, W.C.; Xiao, H.; Wang, Y.T.; Sun, W.; Cui, X.; Li, Y.S.; et al. Hypoxia-responsive miRNAs target argonaute 1 to promote angiogenesis. J. Clin. Investig 2013, 123, 1057–1067. [Google Scholar]
- Mamoru, S.; Yoshitaka, M.; Yuji, T.; Tsuyoshi, T.; Motoyuki, N. A cellular microRNA, let-7i, is a novel biomarker for clinical outcome in patients with dilated cardiomyopathy. J. Card. Fail 2011, 17, 923–929. [Google Scholar]
- Lagos-Quintana, M.; Rauhut, R.; Yalcin, A.; Meyer, J.; Lendeckel, W.; Tuschl, T. Identification of tissue-specific miRNAs from mouse. Curr. Biol 2002, 12, 735–739. [Google Scholar]
- Rao, P.K.; Toyama, T.; Chiang, H.R.; Gupta, S.; Bauer, M.; Medvid, R.; Reinhardt, F.; Liao, R.; Krieger, M.; Jaenisch, R.; et al. Loss of cardiac microRNA-mediated regulation leads to dilated cardiomyopathy and heart failure. Circ. Res 2009, 105, 585–594. [Google Scholar]
- Ji, R.; Cheng, Y.; Yue, J.; Yang, J.; Liu, X.; Chen, H.; Dean, D.B.; Zhang, C. MicroRNA expression signature and antisense-mediated depletion reveal an essential role of microRNA in vascular neointimal lesion formation. Circ. Res 2007, 100, 1579–1588. [Google Scholar]
- McCall, M.N.; Kent, O.A.; Yu, J.; Fox-Talbot, K.; Zaiman, A.L.; Halushka, M.K. MicroRNA profiling of diverse endothelial cell types. BMC Med. Genomics 2011, 4, 78–91. [Google Scholar]
- Pasquinelli, A.E.; Reinhart, B.J.; Slack, F.; Martindale, M.Q.; Kuroda, M.I.; Maller, B.; Hayward, D.C.; Ball, E.E.; Degnan, B.; Muller, P.; et al. Conservation of the sequence and temporal expression of let-7 heterochronic regulatory RNA. Nature 2000, 408, 86–89. [Google Scholar]
- Hutvagner, G.; Mclachlan, J.; Pasquenelli, A.E.; Balint, E.; Tuschl, T.; Zamore, P.D. A cellular function for the RNA-interference enzyme Dicer in the maturation of the let-7 small temporal RNA. Science 2001, 293, 834–838. [Google Scholar]
- Grishok, A.; Pasquinelli, A.E.; Conte, D.; Li, N.; Parrish, S.; Ha, I.; Baillie, D.L.; Fire, A.; Ruvkun, G.; Mello, C.C. Genes and mechanisms related to RNA interference regulate expression of the small temporal RNAs that control C. elegans developmental timing. Cell 2001, 106, 23–34. [Google Scholar]
- Takamizawa, J.; Konish, H.; Yanagisawa, K.; Tomida, S.; Osada, H.; Endoh, H.; Harano, T.; Yatabe, Y.; Nagino, M.; Nimura, Y.; et al. Reduced expression of the let-7 microRNAs in human lung cancers in association with shortened postoperative survival. Cancer Res 2004, 64, 3752–3756. [Google Scholar]
- Sayed, D.; Hong, C.; Chen, I.Y.; Lypowy, J.; Abdellatif, M. MicroRNAs play an essential role in the development of cardiac hypertrophy. Circ. Res 2007, 100, 416–424. [Google Scholar]
- Ikeda, S.; Kong, S.W.; Lu, J.; Bisping, E.; Zhang, H.; Allen, P.D.; Golub, T.R.; Pieske, B.; Pu, W.T. Altered microRNA expression in human heart disease. Physiol. Genomics 2007, 31, 367–373. [Google Scholar]
- Thum, T.; Galuppo, P.; Wolf, C.; Fiedler, J.; Kneitz, S.; van Laake, L.W.; Doevendans, P.A.; Mummery, C.L.; Borlak, J.; Haverich, A.; et al. MicroRNAs in the human heart: A clue to fetal gene reprogramming in heart failure. Circulation 2007, 116, 258–267. [Google Scholar]
- Chen, X.M.; Splinter, P.L.; O’Hara, S.P.; LaRusso, N.F. A cellular micro-RNA, let-7i, regulates Toll-like receptor 4 expression and contributes to cholangiocyte immune responses against Cryptosporidium parvum infection. J. Biol. Chem 2007, 282, 28929–28938. [Google Scholar]
- Shi, G.; Cui, Q.; Zhang, Y. MicroRNA set: A novel way to uncover the potential black box of chronic heart failure in microrna microarray analysis. J. Comput. Sci. Syst. Biol 2009, 2, 240–246. [Google Scholar]
- Luo, X.; Zhang, H.; Xiao, J.; Wang, Z. Regulation of human cardiac ion channel genes by microRNAs: Theoretical perspective and pathophysiological implications. Cell Physiol. Biochem 2010, 25, 571–586. [Google Scholar]
- Fichtlscherer, S.; Rosa, S.D.; Fox, H.; Schwietz, T.; Fischer, A.; Liebetrau, C.; Weber, M.; Hamm, C.W.; Roxe, T.; Muller-Ardogan, M.; et al. Circulating microRNAs in patients with coronary artery disease. Circ. Res 2010, 107, 677–684. [Google Scholar]
- Fu, J.D.; Rushing, S.N.; Lieu, D.K.; Chan, C.W.; Kong, C.W.; Geng, L.; Wilson, K.D.; Chiamvimonvat, N.; Boheler, K.R.; Wu, J.C.; et al. Distinct roles of microRNA-1 and −499 in ventricular specification and functional maturation of human embryonic stem cell-derived cardiomyocytes. PLoS One 2011, 6, e27417. [Google Scholar]
- Melton, C.; Judson, R.L.; Blelloch, R. Opposing microRNA families regulate self-renewal in mouse embryonic stem cells. Nature 2010, 463, 621–626. [Google Scholar]
- Cao, L.; Kong, L.P.; Yu, Z.B.; Han, S.P.; Bai, Y.F.; Zhu, J.; Hu, X.; Zhu, C.; Zhu, S.; Guo, X.R. MicroRNA expression profiling of the developing mouse heart. Int. J. Mol. Med 2012, 30, 1095–1104. [Google Scholar]
- Topkara, V.K.; Mann, D.L. Role of microRNAs in cardiac remodeling and heart failure. Cardiovasc. Drug Ther 2011, 25, 171–182. [Google Scholar]
- Cordes, K.R.; Srivastava, D.; Ivey, K.N. MicroRNAs in cardiac development. Pediatr. Cardiol 2010, 31, 349–356. [Google Scholar]
- Care, A.; Catalucci, D.; Felicetti, F.; Bonci, D.; Addario, A.; Gallo, P.; Bang, M.L.; Segnalini, P.; Gu, Y.; Dalton, N.D.; et al. MicroRNA-133 controls cardiac hypertrophy. Nat. Med 2007, 13, 613–618. [Google Scholar]
- Yang, Y.; Ago, T.; Zhai, P.; Abdellatif, M.; Sadoshima, J. Thioredoxin 1 negatively regulates angiotensin II-induced cardiac hypertrophy through upregulation of miR-98/let-7. Circ. Res 2011, 108, 305–313. [Google Scholar]
- Sun, H.; Wang, Y. Restriction of big hearts by a small RNA. Circ. Res 2011, 108, 274–276. [Google Scholar]
- Oka, S.; Ago, T.; Kitazono, T.; Zablocki, D.; Sadoshima, J. The role of redox modulation of class II histone deacetylases in mediating pathological cardiac hypertrophy. J. Mol. Med 2009, 87, 785–791. [Google Scholar]
- Ago, T.; Sadoshima, J. Thioredoxin1 as a negative regulator of cardiac hypertrophy. Antioxid. Redox Signal 2007, 9, 679–687. [Google Scholar]
- Arciniegas, E.; Frid, M.G.; Douglas, I.S.; Stenmark, K.R. Perspectives on endothelial-to-mesenchymal transition: Potential contribution to vascular remodeling in chronic pulmonary hypertension. Am. J. Physiol. Lung C 2007, 293, L1–L8. [Google Scholar]
- Zeisberg, M.E.; Tarnavski, O.; Zeisberg, M.; Dorfman, A.L.; McMullen, J.R.; Gustafsson, E.; Chandraker, A.; Yuan, X.; Pu, W.T.; Roberts, A.B.; et al. Endothelial-to-mesenchymal transition contributes to cardiac fibrosis. Nat. Med 2007, 13, 952–961. [Google Scholar]
- Ghosh, A.K.; Nagpal, V.; Covington, J.W.; Michaels, M.A.; Vaughan, D.E. Molecular basis of cardiac endothelial-to-mesenchymal transition (EndMT): Differential expression of microRNAs during EndMT. Cell Signal 2012, 24, 1031–1036. [Google Scholar]
- Satoh, M.; Nakamura, M.; Akatsu, T.; Shimoda, Y.; Segawa, I.; Hiramori, K. Toll-like receptor 4 is expressed with enteroviral replication in myocardium from patients with dilated cardiomyopathy. Lab. Investig 2004, 84, 173–181. [Google Scholar]
- Fu, J.; Peng, C.; Wang, W.; Jin, H.; Tang, Q.; Wei, X. Let-7g is involved in doxorubicin induced myocardial injury. Environ. Toxicol. Pharmacol 2012, 33, 312–317. [Google Scholar]
- Marbán, E. Cardiac channelopathies. Nature 2002, 415, 213–218. [Google Scholar]
- Yang, B.; Lin, H.; Xiao, J.; Lu, Y.; Luo, X.; Li, B.; Zhang, Y.; Xu, C.; Bai, Y.; Wang, H.; et al. The muscle-specific microRNA miR-1 regulates cardiac arrhythmogenic potential by targeting GJA1 and KCNJ2. Nat. Med 2007, 13, 486–491. [Google Scholar]
- Luo, X.; Lin, H.; Pan, Z.; Xiao, J.; Zhang, Y.; Lu, Y.; Yang, B.; Wang, Z. Downregulation of miRNA-1/miRNA-133 contributes to re-expression of pacemaker channel genes HCN2 and HCN4 in hypertrophic heart. J. Biol. Chem 2008, 283, 20045–20052. [Google Scholar]
- Xiao, J.; Luo, X.; Lin, H.; Zhang, Y.; Lu, Y.; Wang, N.; Zhang, Y.; Yang, B.; Wang, Z. MicroRNA miR-133 represses HERG K+ channel expression contributing to QT prolongation in diabetic hearts. J. Biol. Chem 2007, 282, 12363–12367. [Google Scholar]
- Dews, M.; Homayouni, A.; Yu, D.; Murphy, D.; Sevignani, C.; Wentzel, E.; Furth, E.E.; Lee, W.M.; Enders, G.H.; Mendell, J.T.; et al. Augmentation of tumor angiogenesis by a Myc-activated microRNA cluster. Nat. Genet 2006, 38, 1060–1065. [Google Scholar]
- Fish, J.E.; Santoro, M.M.; Morton, S.U.; Yu, S.; Yeh, R.F.; Wythe, J.D.; Ivey, K.N.; Bruneau, B.G.; Stainier, D.Y.R.; Srivastava, D. MiR-126 regulates angiogenic signaling and vascular integrity. Dev. Cell 2008, 15, 272–284. [Google Scholar]
- Wurdinger, T.; Tannous, B.A.; Saydam, O.; Skog, J.; Grau, S.; Soutschek, J.; Weissleder, R.; Breakefield, X.O.; Krichevsky, A.M. MiR-296 regulates growth factor receptor overexpression in angiogenic endothelial cells. Cancer Cell 2008, 14, 382–393. [Google Scholar]
- Lee, D.Y.; Deng, Z.; Wang, C.H.; Yang, B.B. MicroRNA-378 promotes cell survival, tumor growth, and angiogenesis by targeting SuFu and Fus-1 expression. Proc. Natl. Acad. Sci. USA 2007, 104, 20350–20355. [Google Scholar]
- Wang, S.; Olson, E.N. AngiomiRs-key regulators of angiogenesis. Curr. Opin. Genet. Dev 2009, 19, 205–211. [Google Scholar]
- Kuehbacher, A.; Urbich, C.; Zeiher, A.M.; Dimmeler, S. Role of dicer and drosha for endothelial microRNA expression and angiogenesis. Circ. Res 2007, 101, 59–68. [Google Scholar]
- Suárez, Y.; Fernández-Hernando, C.; Pober, J.S.; Sessa, W.C. Dicer dependent microRNAs regulate gene expression and functions in human endothelial cells. Circ. Res 2007, 100, 1164–1173. [Google Scholar]
- Otsuka, M.; Zheng, M.; Hayashi, M.; Lee, J.D.; Yoshino, O.; Lin, S.; Han, J. Impaired microRNA processing causes corpus luteum insufficiency and infertility in mice. J. Clin. Investig 2008, 118, 1944–1954. [Google Scholar]
- Bae, O.N.; Wang, J.M.; Baek, S.H.; Wang, Q.; Yuan, H.; Chen, A.F. Oxidative stress-mediated thrombospondin-2 upregulation impairs bone marrow-derived angiogenic cell function in diabetes mellitus. Arterioscler. Thromb. Vasc 2013, 33, 1920–1927. [Google Scholar]
- Poliseno, L.; Tuccoli, A.; Mariani, L.; Evangelista, M.; Citti, L.; Woods, K.; Mercatanti, A.; Hammond, S.; Rainaldi, G. MicroRNAs modulate the angiogenic properties of HUVECs. Blood 2006, 108, 3068–3071. [Google Scholar]
- Roitbak, T.; Bragina, O.; Padilla, J.L.; Pickett, G.G. The role of microRNAs in neural stem cell supported endothelial morphogenesis. Vasc. Cell 2011, 3, 1–15. [Google Scholar]
- Roitbak, T.; Li, L.; Cunningham, L.A. Neural stem/progenitor cells promote endothelial cell morphogenesis and protect endothelial cells against ischemia via HIF-1α-regulated VEGF signaling. J. Cereb. Blood Flow Metab 2008, 28, 1530–1542. [Google Scholar]
- Falk, E. Pathogenesis of atherosclerosis. J. Am. Coll. Cardiol 2006, 47, C7–C12. [Google Scholar]
- Cuaz-Pérolin, C.; Jguirim, I.; Lariqauderie, G.; Jlassi, A.; Furman, C.; Moreau, M.; Chapman, M.J.; Fruchart, J.C.; Slimane, M.N.; Mezdour, H.; Rouis, M. Apolipoprotein E knockout mice over-expressing human tissue inhibitor of metalloproteinase I are protected against aneurysm formation but not against atherosclerotic plaque development. J. Vasc. Res 2006, 43, 493–501. [Google Scholar]
- Ross, R. Atherosclerosis—An inflammatory disease. N. Engl. J. Med 1999, 340, 115–126. [Google Scholar]
- Qin, B.; Xiao, B.; Liang, D.; Li, Y.; Jiang, T.; Yang, H. MicroRNA let-7c inhibits Bcl-xl expression and regulates ox-LDL-induced endothelial apoptosis. BMB Rep 2012, 45, 464–469. [Google Scholar]
- Chen, K.C.; Hsieh, I.C.; Hsi, E.; Wang, Y.S.; Dai, C.Y.; Chou, W.W.; Juo, S.H. Negative feedback regulation between microRNA let-7g and the oxLDL receptor LOX-1. J. Cell Sci 2011, 124, 4115–4124. [Google Scholar]
- Satoh, M.; Tabuchi, T.; Minami, Y.; Takahashi, Y.; Itoh, T.; Nakamura, M. Expression of let-7i is associated with Toll-like receptor 4 signal in coronary artery disease: Effect of statins on let-7i and Toll-like receptor 4 signal. Immunobiology 2012, 217, 533–539. [Google Scholar]
- Kin, K.; Miyagawa, S.; Fukushima, S.; Shirakawa, Y.; Torikai, K.; Shimamura, K.; Daimon, T.; Kawahara, Y.; Kuratani, T.; Sawa, Y. Tissue- and plasma-specific microRNA signatures for atherosclerotic abdominal aortic aneurysm. J. Am. Heart Assoc 2012, 1, e000745. [Google Scholar]
- Satoh, M.; Shimoda, Y.; Maesawa, C.; Akatsu, T.; Ishikawa, Y.; Minami, Y.; Hiramori, K.; Nakamura, M. Activated Toll-like receptor 4 in monocytes is associated with heart failure after acute myocardial infarction. Int. J. Cardiol 2006, 109, 226–234. [Google Scholar]
- Synetos, A.; Toutouzas, K.; Stathogiannis, K.; Latsios, G.; Tsiamis, E.; Tousoulis, D.; Stefanadis, C. MicroRNAs in arterial hypertension. Curr. Top. Med. Chem 2013, 13, 1527–1532. [Google Scholar]
- Eskildsen, T.V.; Jeppesen, P.L.; Schneider, M.; Nossent, A.Y.; Sandberg, M.B.; Hansen, P.B.; Jensen, C.H.; Hansen, M.L.; Marcussen, N.; Rasmussen, L.M.; et al. Angiotensin II regulates microRNA-132/-212 in hypertensive rats and humans. Int. J. Mol. Sci 2013, 14, 11190–11207. [Google Scholar]
- Jackson, K.L.; Marques, F.Z.; Watson, A.M.; Palma-Rigo, K.; Nguyen-Huu, T.P.; Morris, B.J.; Charchar, F.J.; Davern, P.J.; Head, G.A. A novel interaction between sympathetic overactivity and aberrant regulation of renin by miR-181a in BPH/2J genetically hypertensive mice. Hypertension 2013. [Google Scholar] [CrossRef]
- Li, S.; Zhu, J.; Zhang, W.; Chen, Y.; Zhang, K.; Popescu, L.M.; Ma, X.; Lau, W.B.; Rong, R.; Yu, X.; et al. Signature microRNA expression profile of essential hypertension and its novel link to human cytomegalovirus infection. Circulation 2011, 124, 175–184. [Google Scholar]
- Wong, S.S.Y.; Ritner, C.; Ramachandran, S.; Aurigui, J.; Pitt, C.; Chandra, P.; Ling, V.B.; Yabut, O.; Bernstein, H.S. MiR-125b promotes early germ layer specification through Lin28/let-7d and preferential differentiation of mesoderm in human embryonic stem cells. PLoS One 2012, 7, e36121. [Google Scholar]
- Pasha, Z.; Haider, H.K.; Ashraf, M. Efficient non-viral reprogramming of myoblasts to stemness with a single small molecule to generate cardiac progenitor cells. Plos One 2011, 6, e23667. [Google Scholar]
- Ahmed, R.P.H.; Haider, H.K.; Buccini, S.; Li, L.; Jiang, S.; Ashraf, M. Reprogramming of skeletal myoblasts for induction of pluripotency for tumor-free cardiomyogenesis in the infarcted heart. Circ. Res 2011, 109, 60–70. [Google Scholar]
- Chen, X.; Liang, H.; Zhang, J.; Zen, K.; Zhang, C.Y. Secreted microRNAs: A new form of intercellular communication. Trends Cell Biol 2012, 22, 125–132. [Google Scholar]
- Ogawa, R.; Tanaka, C.; Sato, M.; Nagasaki, H.; Sugimura, K.; Okumura, K.; Nakagawa, Y.; Aoki, N. Adipocyte-derived microvesicles contain RNA that is transported into macrophages and might be secreted into blood circulation. Biochem. Biophys. Res. Commun 2010, 398, 723–729. [Google Scholar]
- Muller, G.; Schneider, M.; Biemer-Daub, G.; Weid, S. Microvesicles released from rat adipocytes and harboring glycosylphosphatidylinositol-anchored proteins transfer RNA stimulating lipid synthesis. Cell Signal 2011, 23, 1207–1223. [Google Scholar]
- Deddens, J.C.; Colijn, J.M.; Oerlemans, M.I.; Pasterkamp, G.; Chamuleau, S.A.; Doevendans, P.A.; Sluijter, J.P. Circulating microRNAs as novel biomarkers for the early diagnosis of acute coronary syndrome. J. Cardiovasc. Transl. Res 2013. [Google Scholar] [CrossRef]
- Chen, Y.J.; Zhu, J.M.; Wu, H.; Fan, J.; Zhou, J.; Hu, J.; Yu, Q.; Liu, T.T.; Yang, L.; Wu, C.L.; et al. Circulating microRNAs as a fingerprint for liver cirrhosis. PLoS One 2013, 8, e66577. [Google Scholar]
- Wei, C.; Henderson, H.; Spradley, C.; Li, L.; Kim, I.K.; Kumar, S.; Hong, N.; Arroliga, A.C.; Gupta, S. Circulating miRNAs as potential marker for pulmonary hypertension. PLoS One 2013, 8, e64396. [Google Scholar]
- Long, G.; Wang, F.; Duan, Q.; Yang, S.; Chen, F.; Gong, W.; Yang, X.; Wang, Y.; Chen, C.; Wang, D.W. Circulating miR-30a, miR-195 and let-7b associated with acute myocardial infarction. PLoS One 2012, 7, e50926. [Google Scholar]
- Li, T.; Cao, H.; Zhuang, J.; Wan, J.; Guan, M.; Yu, B.; Li, X.; Zhang, W. Identification of miR-130a, miR-27b and miR-210 as serum biomarkers for atherosclerosis obliterans. Clin. Chim. Acta 2010, 412, 66–70. [Google Scholar]
© 2013 by the authors; licensee MDPI, Basel, Switzerland This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution license (http://creativecommons.org/licenses/by/3.0/).
Share and Cite
Bao, M.-H.; Feng, X.; Zhang, Y.-W.; Lou, X.-Y.; Cheng, Y.; Zhou, H.-H. Let-7 in Cardiovascular Diseases, Heart Development and Cardiovascular Differentiation from Stem Cells. Int. J. Mol. Sci. 2013, 14, 23086-23102. https://doi.org/10.3390/ijms141123086
Bao M-H, Feng X, Zhang Y-W, Lou X-Y, Cheng Y, Zhou H-H. Let-7 in Cardiovascular Diseases, Heart Development and Cardiovascular Differentiation from Stem Cells. International Journal of Molecular Sciences. 2013; 14(11):23086-23102. https://doi.org/10.3390/ijms141123086
Chicago/Turabian StyleBao, Mei-Hua, Xing Feng, Yi-Wen Zhang, Xiao-Ya Lou, Yu Cheng, and Hong-Hao Zhou. 2013. "Let-7 in Cardiovascular Diseases, Heart Development and Cardiovascular Differentiation from Stem Cells" International Journal of Molecular Sciences 14, no. 11: 23086-23102. https://doi.org/10.3390/ijms141123086
APA StyleBao, M.-H., Feng, X., Zhang, Y.-W., Lou, X.-Y., Cheng, Y., & Zhou, H.-H. (2013). Let-7 in Cardiovascular Diseases, Heart Development and Cardiovascular Differentiation from Stem Cells. International Journal of Molecular Sciences, 14(11), 23086-23102. https://doi.org/10.3390/ijms141123086




