Blood microRNAs in Low or No Risk Ischemic Stroke Patients
Abstract
:1. Introduction
2. Results and Discussion
2.1. MicroRNAs That Show Common Expression in No Risk Ischemic Stroke
2.2. Potential Subtype Differentiation Using miRNAs
2.3. MicroRNAs Associated with Functional Outcome in Large Artery Stroke
3. Experimental Section
3.1. Standard Protocol Approvals, Registrations, and Patient Consents
3.2. Blood Collection and Total RNA Isolation
3.3 MicroRNA Microarray and Analysis
4. Conclusions
Supplementary Information
ijms-14-02072-s001.pdfAcknowledgement
References
- Amarenco, P.; Bogousslavsky, J.; Caplan, L.R.; Donnan, G.A.; Hennerici, M.G. Classification of stroke subtypes. Cerebrovasc. Dis 2008, 27, 493–501. [Google Scholar]
- Hatano, S. Experience from a multicentre stroke register: A preliminary report. Bull. World Health Organ 1976, 54, 541–553. [Google Scholar]
- Raichle, M.E. The pathophysiology of brain ischemia. Ann. Neurol 1983, 13, 2–10. [Google Scholar]
- Sacco, R.L.; Benjamin, E.J.; Broderick, J.P.; Dyken, M.; Easton, J.D.; Feinberg, W.M.; Goldstein, L.B.; Gorelick, P.B.; Howard, G.; Kittner, S.J.; et al. American Heart Association; Prevention Conference. IV. Prevention and Rehabilitation of Stroke. Risk Factors. In Stroke; 1997; Volume 28, pp. 1507–1517. [Google Scholar]
- Allegra, A.; Alonci, A.; Campo, S.; Penna, G.; Petrungaro, A.; Gerace, D.; Musolino, C. Circulating microRNAs: New biomarkers in diagnosis, prognosis and treatment of cancer (Review). Int. J. Oncol. 2012. [Google Scholar] [CrossRef]
- Satoh, J.I. Molecular network analysis of human microRNA targetome: From cancers to Alzheimer’s disease. BioData Min 2012, 5, 17. [Google Scholar]
- Saito, Y.; Saito, H. MicroRNAs in cancers and neurodegenerative disorders. Front Genet 2012, 3, 194. [Google Scholar]
- Jeyaseelan, K.; Lim, K.Y.; Armugam, A. MicroRNA expression in the blood and brain of rats subjected to transient focal ischemia by middle cerebral artery occlusion. Stroke 2008, 39, 959–966. [Google Scholar]
- Dharap, A.; Bowen, K.; Place, R.; Li, L.C.; Vemuganti, R. Transient focal ischemia induces extensive temporal changes in rat cerebral microRNAome. J. Cereb. Blood Flow Metab 2009, 29, 675–687. [Google Scholar]
- Lee, R.C.; Feinbaum, R.L.; Ambros, V. The C. elegans heterochronic gene lin-4 encodes small RNAs with antisense complementarity to lin-14. Cell 1993, 75, 843–854. [Google Scholar]
- Bartel, D.P. MicroRNAs: Genomics, biogenesis, mechanism, and function. Cell 2004, 116, 281–297. [Google Scholar]
- Stamova, B.; Xu, H.; Jickling, G.; Bushnell, C.; Tian, Y.; Ander, B.P.; Zhan, X.; Liu, D.; Turner, R.; Adamczyk, P.; et al. Gene expression profiling of blood for the prediction of ischemic stroke. Stroke 2010, 41, 2171–2177. [Google Scholar]
- Jickling, G.C.; Xu, H.; Stamova, B.; Ander, B.P.; Zhan, X.; Tian, Y.; Turner, R.J.; Mesias, M.; Verro, P.; Khoury, J.; et al. Signatures of cardioembolic and large-vessel ischemic stroke. Ann. Neurol. 2010, 68, 681–692. [Google Scholar]
- Jickling, G.C.; Stamova, B.; Ander, B.P.; Zhan, X.; Tian, Y.; Liu, D.; Xu, H.; Johnston, S.C.; Verro, P.; Sharp, F.R. Profiles of lacunar and nonlacunar stroke. Ann. Neurol 2011, 70, 477–485. [Google Scholar]
- Tan, K.S.; Armugam, A.; Sepramaniam, S.; Lim, K.Y.; Setyowati, K.D.; Wang, C.W.; Jeyaseelan, K. Expression profile of MicroRNAs in young stroke patients. PLoS One 2009, 4, e7689. [Google Scholar]
- Wu, P.; Zuo, X.; Ji, A. Stroke-induced microRNAs: The potential therapeutic role for stroke. Exp. Ther. Med 2012, 3, 571–576. [Google Scholar]
- Gan, C.S.; Wang, C.W.; Tan, K.S. Circulatory microRNA-145 expression is increased in cerebral ischemia. Genet. Mol. Res 2012, 11, 147–152. [Google Scholar]
- Zeng, L.; Liu, J.; Wang, Y.; Wang, L.; Weng, S.; Tang, Y.; Zheng, C.; Cheng, Q.; Chen, S.; Yang, G.Y. MicroRNA-210 as a novel blood biomarker in acute cerebral ischemia. Front. Biosci 2011, 3, 1265–1272. [Google Scholar]
- Weng, H.; Shen, C.; Hirokawa, G.; Ji, X.; Takahashi, R.; Shimada, K.; Kishimoto, C.; Iwai, N. Plasma miR-124 as a biomarker for cerebral infarction. Biomed. Res 2011, 32, 135–141. [Google Scholar]
- Quintavalle, M.; Condorelli, G.; Elia, L. Arterial remodeling and atherosclerosis: miRNAs involvement. Vascul. Pharmacol 2011, 55, 106–110. [Google Scholar]
- Shantikumar, S.; Caporali, A.; Emanueli, C. Role of miRNA in Diabetes and its Cardiovascular Complications. Cardiovasc. Res 2012, 93, 583–593. [Google Scholar]
- Van Empel, V.P.; De Windt, L.J.; Martins, P.A. Circulating miRNAs: Reflecting or Affecting Cardiovascular Disease? Curr. Hypertens. Rep. 2012. [Google Scholar] [CrossRef]
- Karolina, D.S.; Tavintharan, S.; Armugam, A.; Sepramaniam, S.; Pek, S.L.; Wong, M.T.; Lim, S.C.; Sum, C.F.; Jeyaseelan, K. Circulating miRNA Profiles in Patients with Metabolic Syndrome. J. Clin. Endocrinol. Metab. 2012. [Google Scholar] [CrossRef]
- Lewis, B.P.; Burge, C.B.; Bartel, D.P. Conserved seed pairing, often flanked by adenosines, indicates that thousands of human genes are microRNA targets. Cell 2005, 120, 15–20. [Google Scholar]
- Garcia, D.M.; Baek, D.; Shin, C.; Bell, G.W.; Grimson, A.; Bartel, D.P. Weak seed-pairing stability and high target-site abundance decrease the proficiency of lsy-6 and other microRNAs. Nat. Struct. Mol. Biol 2011, 18, 1139–1146. [Google Scholar]
- Zhang, L.; Sullivan, P.S.; Goodman, J.C.; Gunaratne, P.H.; Marchetti, D. MicroRNA-1258 suppresses breast cancer brain metastasis by targeting heparanase. Cancer Res 2011, 71, 645–654. [Google Scholar]
- Takahashi, H.; Matsumoto, H.; Kumon, Y.; Ohnishi, T.; Freeman, C.; Imai, Y.; Tanaka, J. Expression of heparanase in nestin-positive reactive astrocytes in ischemic lesions of rat brain after transient middle cerebral artery occlusion. Neurosci. Lett 2007, 417, 250–254. [Google Scholar]
- Tong, J.L.; Zhang, C.P.; Nie, F.; Xu, X.T.; Zhu, M.M.; Xiao, S.D.; Ran, Z.H. MicroRNA 506 regulates expression of PPAR alpha in hydroxycamptothecin-resistant human colon cancer cells. FEBS Lett 2011, 585, 3560–3568. [Google Scholar]
- Collino, M.; Aragno, M.; Mastrocola, R.; Benetti, E.; Gallicchio, M.; Dianzani, C.; Danni, O.; Thiemermann, C.; Fantozzi, R. Oxidative stress and inflammatory response evoked by transient cerebral ischemia/reperfusion: effects of the PPAR-alpha agonist WY14643. Free Radic. Biol. Med 2006, 41, 579–589. [Google Scholar]
- Rademakers, R.; Eriksen, J.L.; Baker, M.; Robinson, T.; Ahmed, Z.; Lincoln, S.J.; Finch, N.; Rutherford, N.J.; Crook, R.J.; Josephs, K.A.; et al. Common variation in the miR-659 binding-site of GRN is a major risk factor for TDP43-positive frontotemporal dementia. Hum. Mol. Genet. 2008, 17, 3631–3642. [Google Scholar]
- Ahmed, Z.; Mackenzie, I.R.; Hutton, M.L.; Dickson, D.W. Progranulin in frontotemporal lobar degeneration and neuroinflammation. J. Neuro. Inflamm 2007, 4, 7. [Google Scholar]
- Jiang, L.; Huang, Q.; Chang, J.; Wang, E.; Qiu, X. MicroRNA Hsa-miR-125a-5p induces apoptosis by activating p53 in lung cancer cells. Exp. Lung Res 2011, 37, 387–398. [Google Scholar]
- Broughton, B.R.; Reutens, D.C.; Sobey, C.G. Apoptotic mechanisms after cerebral ischemia. Stroke 2009, 40, e331–e339. [Google Scholar]
- Martinelli, R.; Nardelli, C.; Pilone, V.; Buonomo, T.; Liguori, R.; Castanò, I.; Castanò, I.; Buono, P.; Masone, S.; Persico, G.; et al. miR-519d overexpression is associated with human obesity. Obesity (Silver Spring) 2010, 18, 2170–2176. [Google Scholar]
- Zandbergen, F.; Plutzky, J. PPARalpha in atherosclerosis and inflammation. Biochim. Biophys. Acta 2007, 1771, 972–982. [Google Scholar]
- Meder, B.; Keller, A.; Vogel, B.; Haas, J.; Sedaghat-Hamedani, F.; Kayvanpour, E.; Just, S.; Borries, A.; Rudloff, J.; Leidinger, P.; et al. MicroRNA signatures in total peripheral blood as novel biomarkers for acute myocardial infarction. Basic Res. Cardiol 2011, 106, 13–23. [Google Scholar]
- Xu, H.; Tang, Y.; Liu, D.Z.; Ran, R.; Ander, B.P.; Apperson, M.; Liu, X.S.; Khoury, J.C.; Gregg, J.P.; Pancioli, A.; et al. Gene expression in peripheral blood differs after cardioembolic compared with large-vessel atherosclerotic stroke: biomarkers for the etiology of ischemic stroke. J. Cereb. Blood Flow Metab. 2008, 28, 1320–1328. [Google Scholar]
- Xia, H.F.; He, T.Z.; Liu, C.M.; Cui, Y.; Song, P.P.; Jin, X.H.; Ma, X. MiR-125b expression affects the proliferation and apoptosis of human glioma cells by targeting Bmf. Cell Physiol. Biochem 2009, 23, 347–358. [Google Scholar]
- Le, M.T.; Teh, C.; Shyh-Chang, N.; Xie, H.; Zhou, B.; Korzh, V.; Lodish, H.F.; Lim, B. MicroRNA-125b is a novel negative regulator of p53. Genes Dev 2009, 23, 862–876. [Google Scholar]
- Yang, H.; Kong, W.; He, L.; Zhao, J.J.; O’Donnell, J.D.; Wang, J.; Wenham, R.M.; Coppola, D.; Kruk, P.A.; Nicosia, S.V.; et al. MicroRNA expression profiling in human ovarian cancer: miR-214 induces cell survival and cisplatin resistance by targeting PTEN. Cancer Res. 2008, 68, 425–433. [Google Scholar]
- Zhang, C.; Han, L.; Zhang, A.; Fu, Y.; Yue, X.; Wang, G.; Jia, Z.; Pu, P.; Zhang, Q.; Kang, C. MicroRNA-221 and microRNA-222 regulate gastric carcinoma cell proliferation and radioresistance by targeting PTEN. BMC Cancer 2010, 10, 367. [Google Scholar]
- Zhang, C.Z.; Zhang, J.X.; Zhang, A.L.; Shi, Z.D.; Han, L.; Jia, Z.F.; Yang, W.D.; Wang, G.X.; Jiang, T.; You, Y.P.; et al. MiR-221 and miR-222 target PUMA to induce cell survival in glioblastoma. Mol. Cancer 2010, 9, 229. [Google Scholar]
- Zhang, Y.; Liu, D.; Chen, X.; Li, J.; Li, L.; Bian, Z.; Sun, F.; Lu, J.; Yin, Y.; Cai, X.; et al. Secreted monocytic miR-150 enhances targeted endothelial cell migration. Mol. Cell 2010, 39, 133–144. [Google Scholar]
- Harris, T.A.; Yamakuchi, M.; Ferlito, M.; Mendell, J.T.; Lowenstein, C.J. MicroRNA-126 regulates endothelial expression of vascular cell adhesion molecule 1. Proc. Natl. Acad. Sci. USA 2008, 105, 1516–1521. [Google Scholar]
- Yang, K.; He, Y.S.; Wang, X.Q.; Lu, L.; Chen, Q.J.; Liu, J.; Sun, Z.; Shen, W.F. MiR-146a inhibits oxidized low-density lipoprotein-induced lipid accumulation and inflammatory response via targeting toll-like receptor 4. FEBS Lett 2011, 585, 854–860. [Google Scholar]
- Brea, D.; Blanco, M.; Ramos-Cabrer, P.; Moldes, O.; Arias, S.; Pérez-Mato, M.; Leira, R.; Sobrino, T.; Castillo, J. Toll-like receptors 2 and 4 in ischemic stroke: outcome and therapeutic values. J. Cereb. Blood Flow Metab 2011, 31, 1424–1431. [Google Scholar]
- Rodríguez-Yáñez, M.; Castellanos, M.; Blanco, M.; García, M.M.; Nombela, F.; Serena, J.; Leira, R.; Lizasoain, I.; Dávalos, A.; Castillo, J. New-onset hypertension and inflammatory response/poor outcome in acute ischemic stroke. Neurology 2006, 67, 1973–1978. [Google Scholar]
- Tan, K.S.; Tan, C.T.; Churilov, L.; MacKay, M.T.; Donnan, G.A. Ischaemic stroke in young adults: A comparative study between Malaysia and Australia. Neurol. Asia 2010, 15, 1–9. [Google Scholar]
- Adams, H.P., Jr; Bendixen, B.H.; Kappelle, L.J.; Biller, J.; Love, B.B.; Gordon, D.L.; Marsh, E.E., III. Classification of subtype of acute ischemic stroke: Definitions for use in a multicenter clinical trial. TOAST. Trial of Org 10172 in Acute Stroke Treatment. Stroke 1993, 24, 35–41. [Google Scholar]
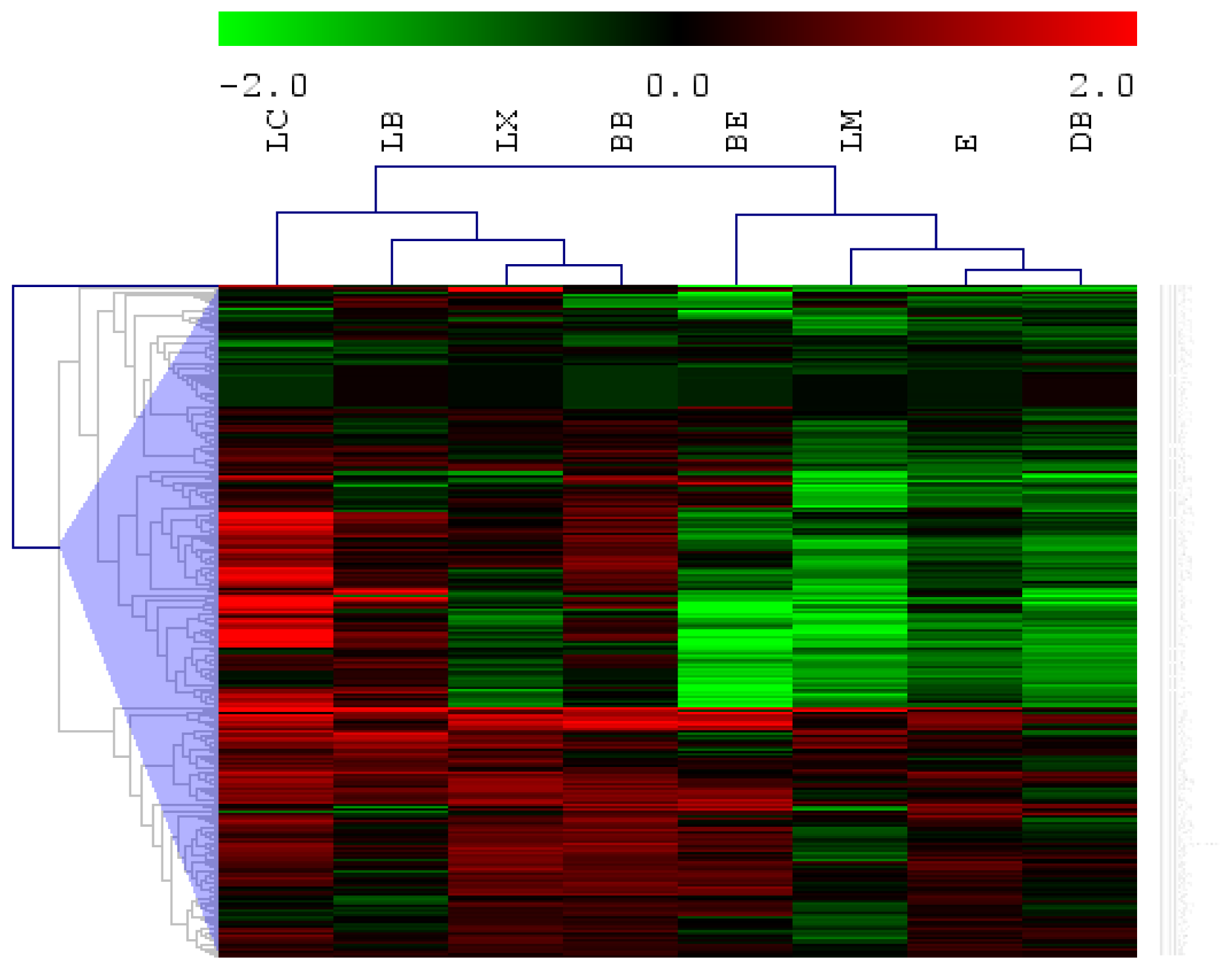

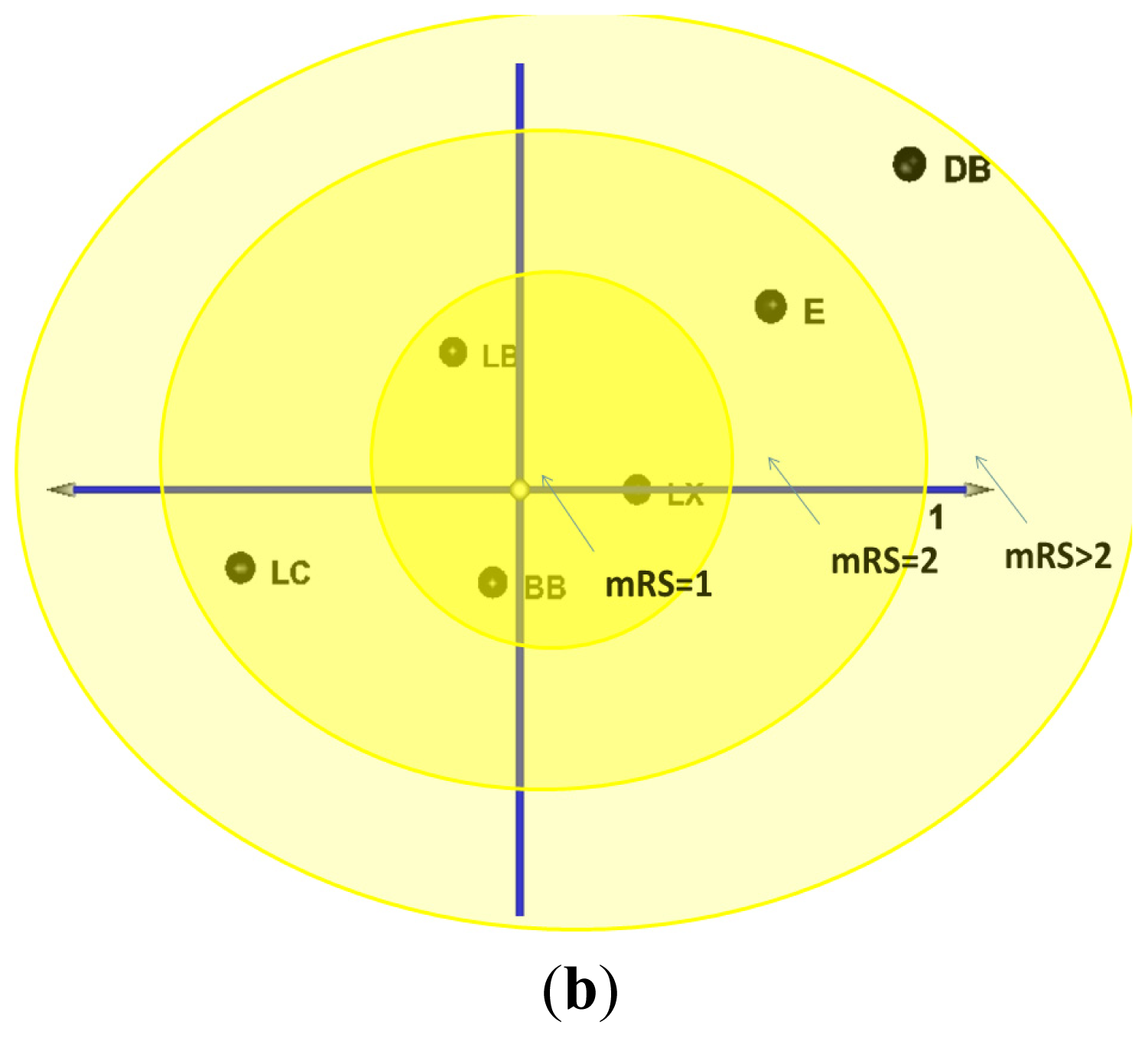
| hsa-miRNA | BB | DB | E | LB | LC | LX | BE | LM |
|---|---|---|---|---|---|---|---|---|
| miR-1258 | 2.33 | 1.57 | 1.94 | 1.87 | 4.90 | 3.35 | 2.35 | 1.56 |
| miR-125a-5p | 1.52 | 1.06 | 1.45 | 1.40 | 1.97 | 2.18 | 1.37 | 1.39 |
| miR-1260 | 1.78 | 1.60 | 1.28 | 1.48 | 2.35 | 2.27 | 1.42 | 1.10 |
| miR-1273 | 5.57 | 1.30 | 2.80 | 4.43 | 4.91 | 4.52 | 4.23 | 2.73 |
| miR-149 | 1.75 | 1.67 | 1.79 | 1.43 | 2.40 | 1.59 | 1.08 | 1.15 |
| miR-220b | 2.93 | 1.81 | 1.99 | 1.92 | 2.95 | 3.08 | 2.49 | 1.27 |
| miR-23a* | 1.57 | 1.01 | 1.79 | 1.16 | 1.50 | 1.85 | 1.69 | 1.39 |
| miR-25* | −1.57 | −1.32 | −1.14 | −1.36 | −1.71 | −1.20 | −1.17 | −1.09 |
| miR-26b* | 2.80 | 1.61 | 2.04 | 1.39 | 3.08 | 2.30 | 2.50 | 1.09 |
| miR-29b-1* | 1.44 | 1.47 | 1.57 | 1.17 | 1.45 | 1.63 | 1.55 | 1.13 |
| miR-302e | 1.60 | 1.06 | 1.27 | 1.89 | 1.75 | 1.70 | 1.06 | 1.43 |
| miR-34b | −1.28 | −1.37 | −1.18 | −1.18 | −1.26 | −1.70 | −1.20 | −1.86 |
| miR-483-5p | −1.10 | −1.45 | −1.23 | −1.09 | −1.66 | −1.04 | −1.07 | −1.49 |
| miR-488 | 1.41 | 1.07 | 1.28 | 2.23 | 1.82 | 2.32 | 1.20 | 1.69 |
| miR-490-3p | 4.04 | 1.65 | 2.13 | 1.05 | 2.22 | 3.21 | 3.47 | 1.05 |
| miR-498 | −1.23 | −1.24 | −1.23 | −1.53 | −1.40 | −1.03 | −1.07 | −1.20 |
| miR-506 | 1.40 | 1.55 | 2.09 | 2.49 | 2.86 | 2.35 | 1.01 | 1.35 |
| miR-659 | 1.53 | 1.02 | 1.16 | 1.01 | 1.51 | 1.44 | 1.47 | 1.08 |
| miR-890 | 2.09 | 1.03 | 1.30 | 1.20 | 2.08 | 2.28 | 1.43 | 1.35 |
| miR-920 | 2.55 | 1.22 | 1.93 | 1.31 | 1.59 | 2.34 | 2.67 | 1.42 |
| miR-934 | 1.99 | 1.02 | 1.26 | 1.03 | 1.23 | 1.98 | 1.72 | 1.12 |
| hsa-miRNA | Large Artery | Cardioembolic | Lacunar | |||||
|---|---|---|---|---|---|---|---|---|
| BB | DB | E | LB | LC | LX | BE | LM | |
| miR-208a | 1.10 | 1.38 | 1.12 | 1.14 | 1.26 | 1.17 | −1.23 | −1.38 |
| miR-519d | 1.38 | 1.31 | 1.07 | 1.08 | 1.54 | 1.61 | −1.09 | −1.86 |
| miR-605 | 1.71 | 1.15 | 1.34 | 1.11 | 1.75 | 2.11 | −1.04 | −1.79 |
| miR-634 | 1.33 | 1.08 | 1.16 | 1.30 | 1.36 | 1.56 | −1.09 | −1.35 |
| miR-99b* | 1.17 | 1.17 | 1.22 | 1.17 | 1.27 | 1.16 | −1.03 | −1.07 |
| miR-1246 | −1.04 | −1.23 | −1.18 | −1.06 | −1.40 | −1.20 | 1.03 | −1.26 |
| miR-377 | 2.76 | 1.07 | 1.14 | 3.59 | 4.51 | 1.61 | −1.34 | 3.75 |
| miR-513a-5p | −1.65 | −1.29 | −1.37 | −1.30 | −2.00 | −1.24 | 1.15 | −1.07 |
| miR-767-5p | 1.19 | 1.16 | 1.08 | 2.46 | 2.36 | 1.60 | −1.10 | 1.84 |
| miR-875-3p | 1.75 | 1.05 | 1.19 | 2.36 | 1.92 | 1.94 | −1.24 | 2.19 |
| miR-1274a | 1.19 | 1.42 | 1.26 | 1.09 | 1.89 | 1.43 | 1.04 | −1.48 |
| miR-1280 | 1.59 | 1.31 | 1.21 | 1.23 | 1.77 | 1.76 | 1.29 | −1.46 |
| miR-200c* | 1.65 | 1.34 | 1.71 | 1.61 | 1.84 | 1.87 | 1.10 | −1.02 |
| miR-375 | 1.17 | 1.08 | 1.35 | 1.13 | 1.42 | 1.55 | 1.09 | −1.33 |
| miR-494 | 1.25 | 1.42 | 1.40 | 1.07 | 1.43 | 1.33 | 1.22 | −1.09 |
| miR-520d-5p | 2.03 | 1.27 | 1.33 | 1.38 | 2.37 | 2.13 | 1.41 | −1.16 |
| miR-551a | 1.32 | 1.15 | 1.18 | 1.30 | 1.26 | 1.11 | 1.05 | −1.02 |
| miR-629* | 1.51 | 1.50 | 1.59 | 1.10 | 1.62 | 1.88 | 1.12 | −1.37 |
| miR-656 | 1.58 | 1.28 | 1.64 | 1.03 | 1.45 | 1.89 | 1.42 | −1.25 |
| miR-657 | 1.57 | 1.13 | 1.67 | 1.10 | 1.26 | 2.25 | 1.35 | −1.05 |
| miR-664 | 1.35 | 1.12 | 1.17 | 1.21 | 1.13 | 1.37 | 1.12 | −1.02 |
| miR-766 | 2.05 | 1.25 | 1.18 | 1.34 | 1.89 | 1.85 | 1.71 | −1.32 |
| hsa-miRNA | p-value (LA mRs ≤ 2 vs. LA mRs > 2) | Fold-Change (LA mRs ≤ 2 vs. LA mRs > 2) |
|---|---|---|
| hsa-miR-1201 | 0.0018 | 9.3616 |
| hsa-miR-125b | 0.0044 | 8.7958 |
| hsa-miR-125b-2* | 0.0039 | 6.4179 |
| hsa-miR-1275 | 0.0000 | 6.5432 |
| hsa-miR-1304 | 0.0258 | 9.3241 |
| hsa-miR-138-2* | 0.0213 | 8.2105 |
| hsa-miR-150 | 0.0023 | 11.3980 |
| hsa-miR-181a-2* | 0.0106 | 11.4422 |
| hsa-miR-195 | 0.0205 | 15.1663 |
| hsa-miR-200b* | 0.0004 | 6.4327 |
| hsa-miR-200c* | 0.0371 | 1.3122 |
| hsa-miR-208a | 0.0289 | −1.1674 |
| hsa-miR-214 | 0.0011 | 8.6163 |
| hsa-miR-221 | 0.0363 | 8.0340 |
| hsa-miR-361-5p | 0.0029 | 6.3937 |
| hsa-miR-509-5p | 0.0018 | 9.0491 |
| hsa-miR-519e* | 0.0081 | 6.7593 |
| hsa-miR-550 | 0.0053 | 7.0590 |
| hsa-miR-551b* | 0.0012 | 6.0334 |
| hsa-miR-574-3p | 0.0021 | 5.7031 |
| hsa-miR-636 | 0.0001 | −5.3117 |
| hsa-miR-664* | 0.0146 | 13.2992 |
| hsa-miR-768-5p | 0.0006 | 6.2516 |
| hsa-miR-874 | 0.0017 | 7.7018 |
| hsa-miR-937 | 0.0019 | 6.2523 |
| hsa-miR-938 | 0.0029 | 7.4285 |
| hsa-miR-99b | 0.0048 | 7.5901 |
© 2013 by the authors; licensee Molecular Diversity Preservation International, Basel, Switzerland. This article is an open-access article distributed under the terms and conditions of the Creative Commons Attribution license (http://creativecommons.org/licenses/by/3.0/).
Share and Cite
Tan, J.R.; Tan, K.S.; Koo, Y.X.; Yong, F.L.; Wang, C.W.; Armugam, A.; Jeyaseelan, K. Blood microRNAs in Low or No Risk Ischemic Stroke Patients. Int. J. Mol. Sci. 2013, 14, 2072-2084. https://doi.org/10.3390/ijms14012072
Tan JR, Tan KS, Koo YX, Yong FL, Wang CW, Armugam A, Jeyaseelan K. Blood microRNAs in Low or No Risk Ischemic Stroke Patients. International Journal of Molecular Sciences. 2013; 14(1):2072-2084. https://doi.org/10.3390/ijms14012072
Chicago/Turabian StyleTan, Jun Rong, Kay Sin Tan, Yu Xuan Koo, Fung Lin Yong, Chee Woon Wang, Arunmozhiarasi Armugam, and Kandiah Jeyaseelan. 2013. "Blood microRNAs in Low or No Risk Ischemic Stroke Patients" International Journal of Molecular Sciences 14, no. 1: 2072-2084. https://doi.org/10.3390/ijms14012072




