Mitochondrial Cytochrome c Oxidase and F1Fo-ATPase Dysfunction in Peppers (Capsicum annuum L.) with Cytoplasmic Male Sterility and Its Association with orf507 and Ψatp6-2 Genes
Abstract
:1. Introduction
2. Results and Discussion
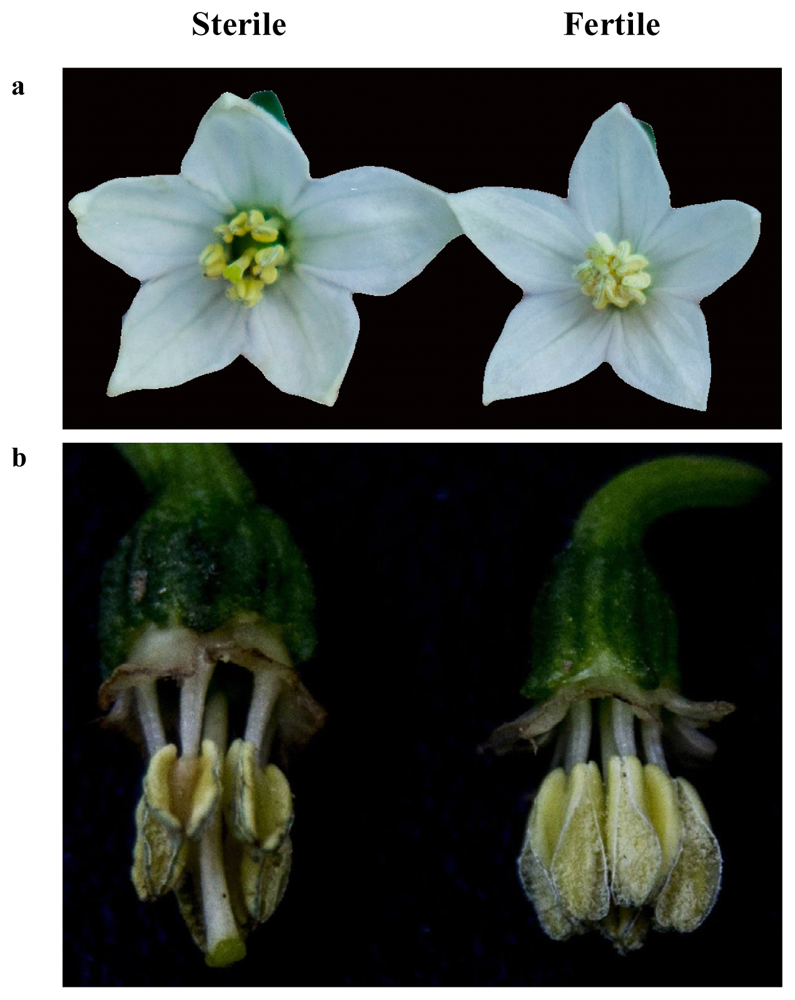
2.1. Confirmation of Pollen Phenotype in the Sterile and Maintainer Lines
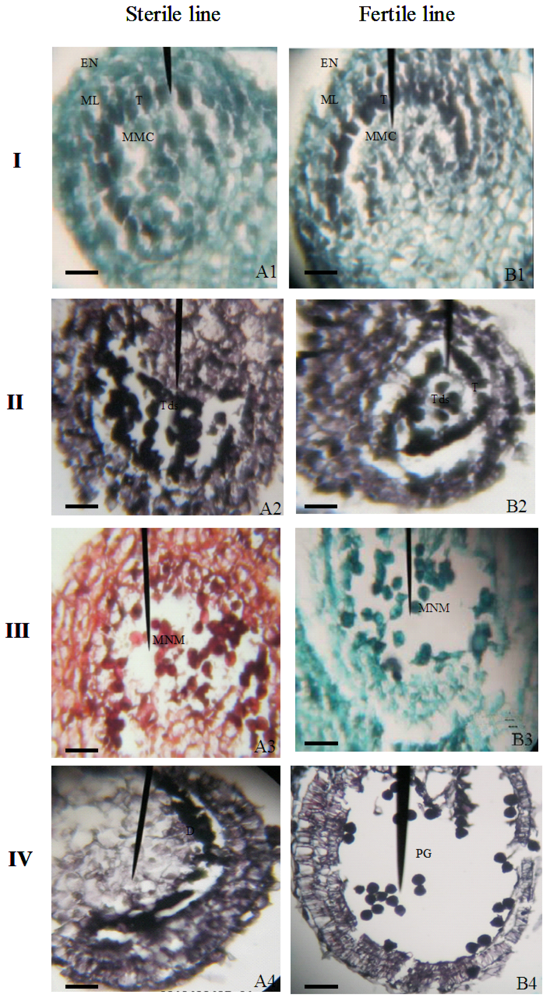
2.2. Cytological Features of Anther Development in the Sterile and Fertile Pepper Lines
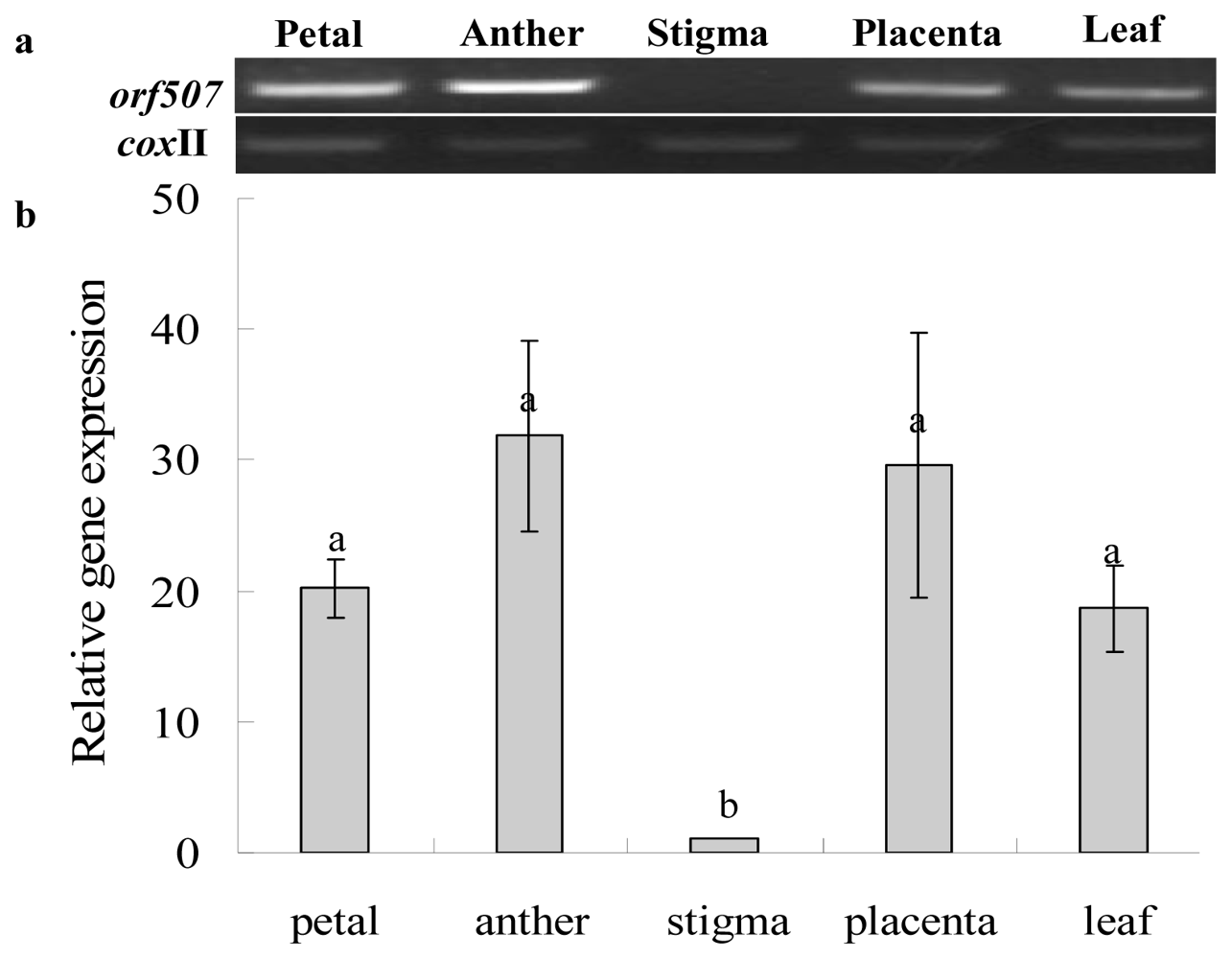
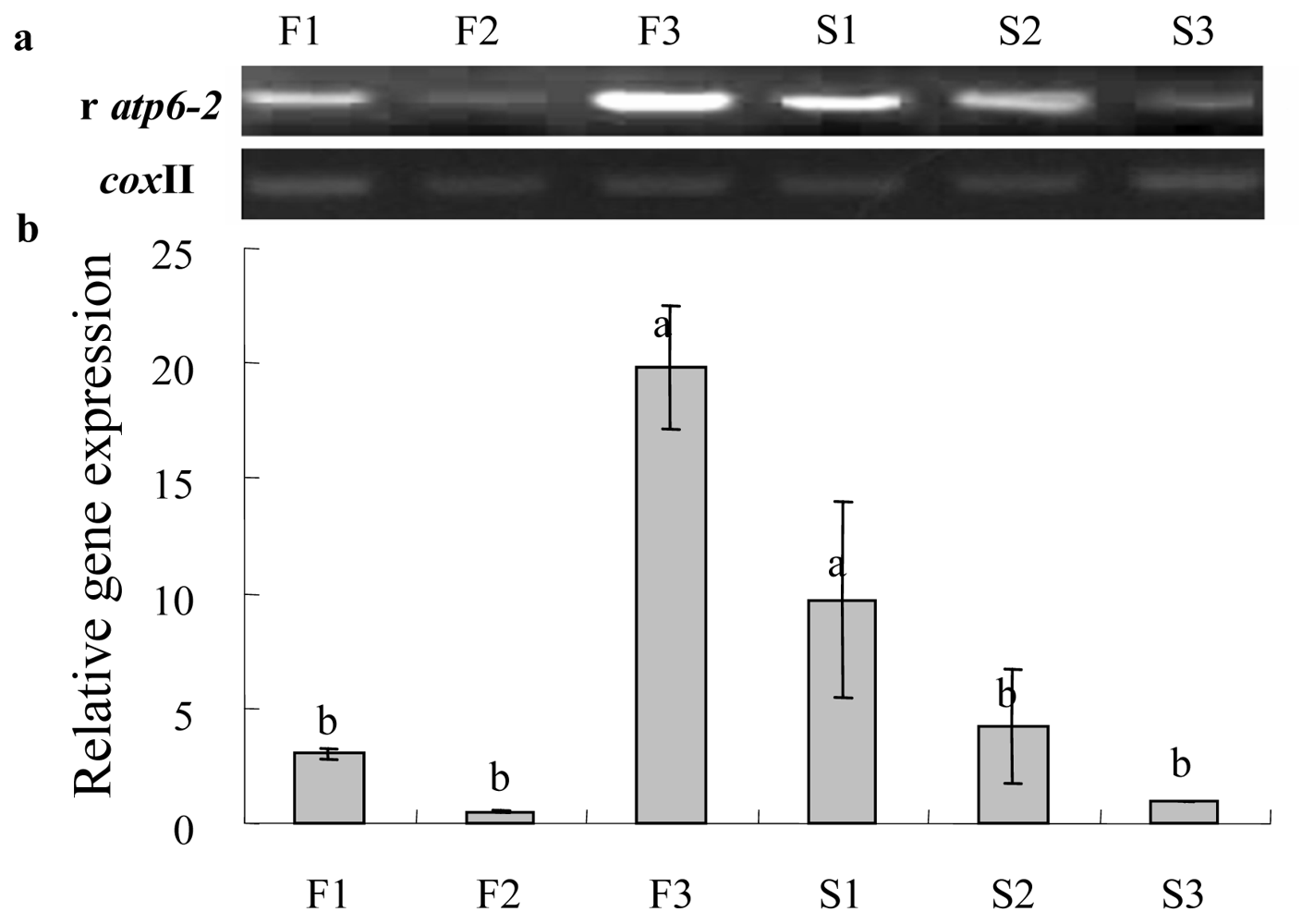
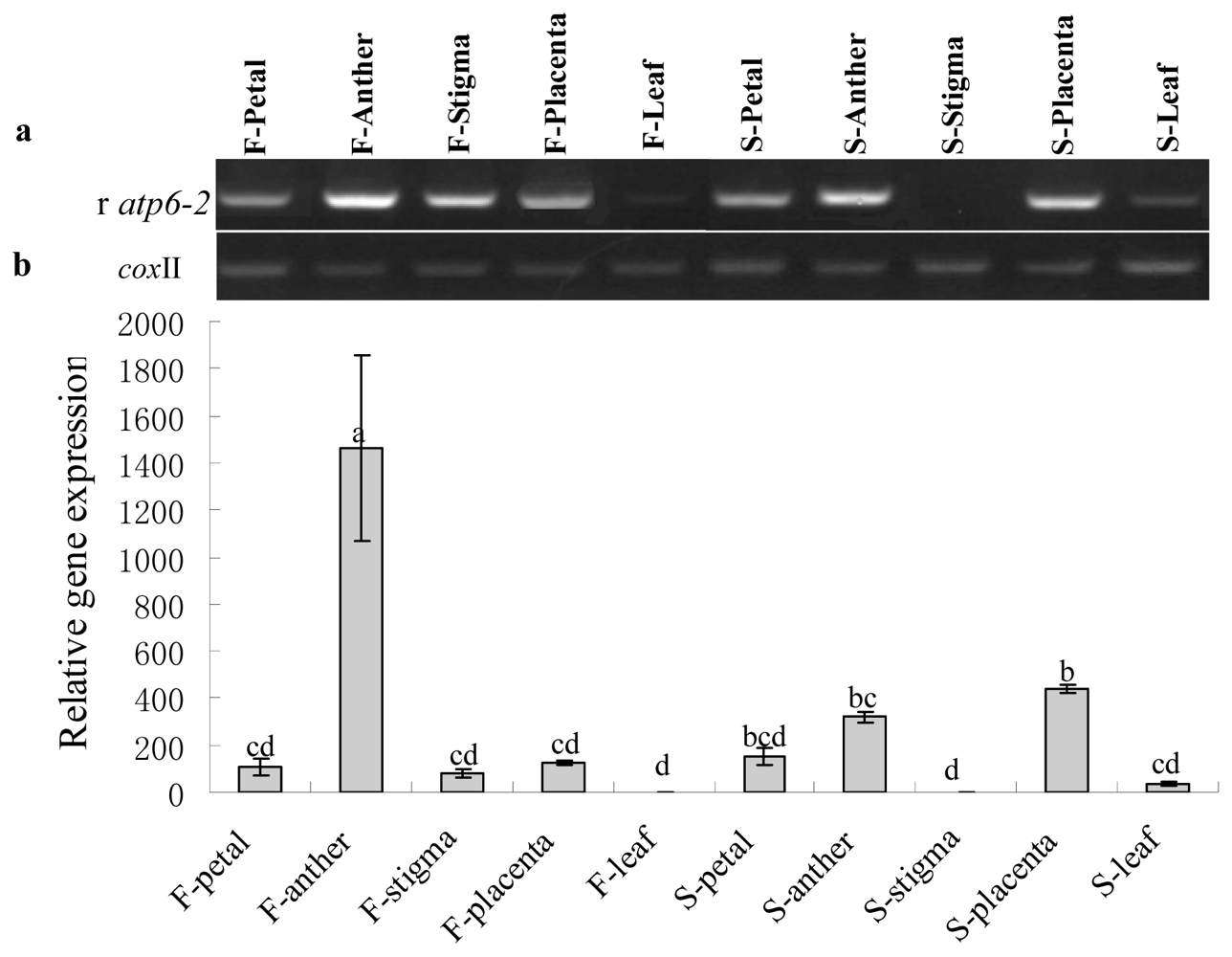
2.3. Gene Expression Analysis of Cox II-orf507 and Ψatp6-2
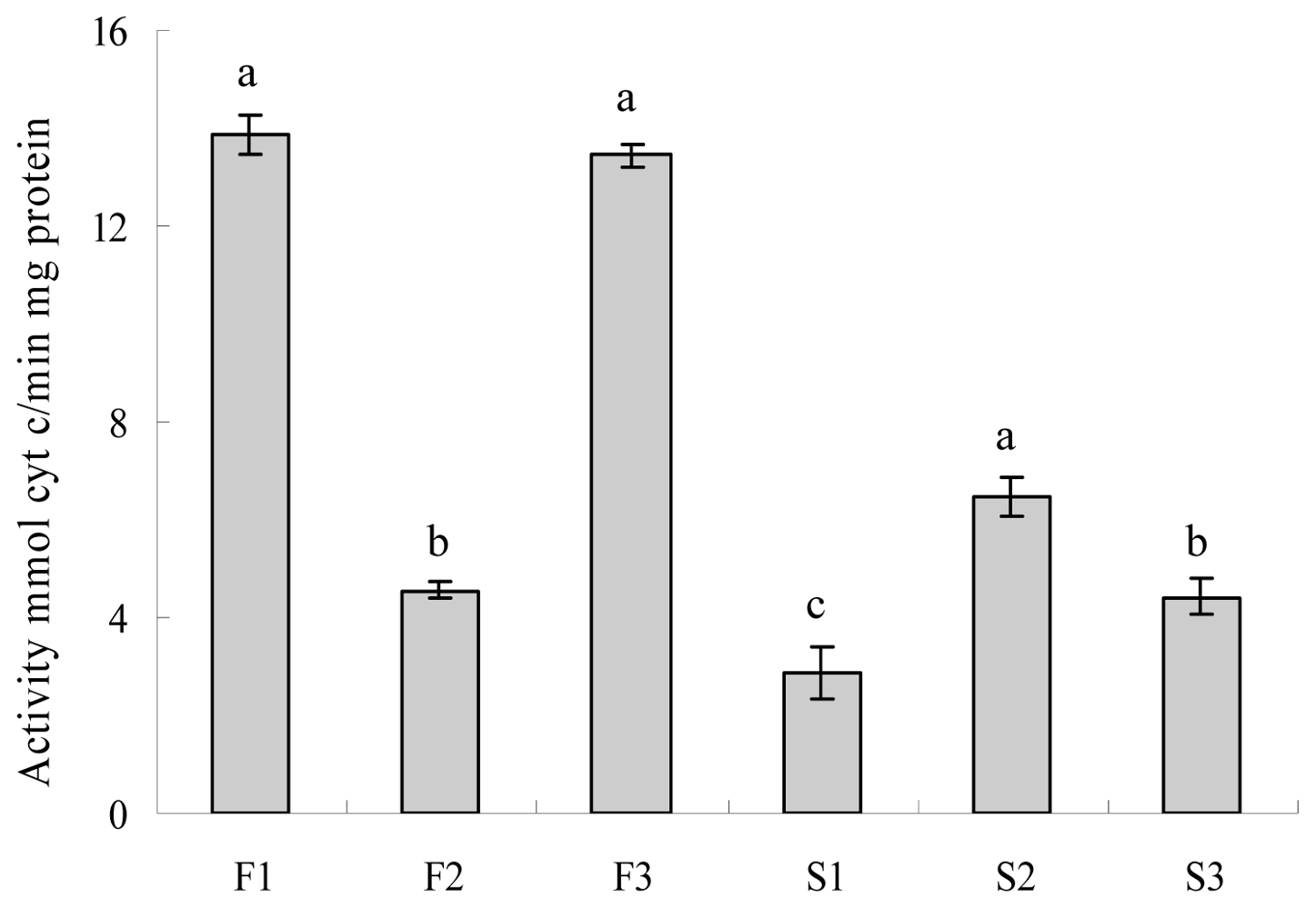
2.4. Evaluation of Cytochrome c Oxidase and ATPase Activity
3. Experimental Section
3.1. Plant Materials and Growing Conditions
3.2. Analysis of Pollen Development
3.3. Extraction of Mitochondria
3.4. Gene Expression Pattern Analysis
3.5. ATPase and Cytochrome c Oxidase Activity
4. Conclusions
Acknowledgments
- Conflict of InterestThe authors declare no conflict of interest.
References
- Schnable, P.S.; Wise, R.P. The molecular basis of cytoplasmic male sterility and fertility restoration. Trends Plant Sci 1998, 3, 175–180. [Google Scholar]
- Hanson, M.R.; Bentolila, S. Interaction of mitochondria and nuclear genes that affect male gametophyte development. Plant Cell 2004, 16, S154–S169. [Google Scholar]
- Abad, A.R.; Mehrtens, B.J.; Mackenzie, S.A. Specific expression in reproductive tissues and fate of a mitochondrial sterility-associated protein in cytoplasmic male sterile bean. Plant Cell 1995, 7, 271–285. [Google Scholar]
- Wintz, H.; Chen, H.C.; Sutton, C.A.; Conley, C.A.; Cobb, A.; Ruth, D.; Hanson, M.R. Expression of the CMS-associated urfS sequence in transgenic petunia and tobacco. Plant Mol. Biol 1995, 28, 83–92. [Google Scholar]
- Kim, S.; Lim, H.; Park, S.; Cho, K.H.; Sung, S.K.; Oh, D.G.; Kim, K.T. Identification of a novel mitochondrial genome type and development of molecular markers for cytoplasm classification in radish (Raphanus sativus L.). Theor. Appl. Genet 2007, 115, 1137–1145. [Google Scholar]
- Gulyas, G.; Shin, Y.; Kim, H.; Lee, J.S.; Hirata, Y. Altered transcript reveals an orf507 sterility-related gene in chili pepper (Capsicum annuum L.). Plant Mol. Biol. Rep 2010, 28, 605–612. [Google Scholar]
- Sabar, M.; Gagliardi, D. ORFB is a subunit of F1Fo-ATP synthase: Insight into the basis of cytoplasmic male sterility in sunflower. EMBO Rep 2003, 4, 381–386. [Google Scholar]
- Nizampatnam, N.R.; Doodhi, H.; Narasimhan, Y.K. Expression of sunflower cytoplasmic male sterility-associated open reading frame, orfH522 induces male sterility in transgenic tobacco plants. Planta 2009, 229, 987–1001. [Google Scholar]
- Zhang, H.; Li, S.Q.; Yi, P.; Wan, C.X. A honglian CMS line of rice displays aberrant Fo of F1Fo-ATPase. Plant Cell Rep 2007, 26, 1065–1071. [Google Scholar]
- Yan, Z.X.; Shao, J.Z.; Ding, Y. Catalytic assays in blue natibe gels revealed normal ATPase but deficient NADH dehydrogenase activity in ZidaoA CMS line of rice (Oryza sativa). Acta Physiol. Plant 2011, 33, 2477–2484. [Google Scholar]
- Hakansson, G.; Glimelius, K. Extensive nuclear influence on mitochondrial transcription and genome structure in male-fertile and male-sterile alloplasmic Nicotiana materials. Mol. Gen. Genet 1991, 229, 380–388. [Google Scholar]
- Hernould, M.; Suharsono, S. Male-sterility induction in transgenic tobacco plants with an unedited atp9 mitochondrial gene from wheat. Proc. Natl. Acad. Sci. USA 1993, 90, 2370–2374. [Google Scholar]
- Janska, H.; Sarria, R.; Woloszynska, M.; Arrieta-Montiel, M.; Mackenzie, S.A. Stoichiometric shifts in the common bean mitochondrial genome leading to male sterility and spontaneous reversion to fertility. Plant Cell 1998, 10, 1163–1180. [Google Scholar]
- McCormick, S. Control of male gametophyte development. Plant Cell 2004, 16, S142–S153. [Google Scholar]
- Goldberg, R.B.; Beals, T.P.; Sanders, P.M. Anther development: Basic principles and practical applications. Plant Cell 1993, 5, 1217–1229. [Google Scholar]
- Wu, H.M.; Cheung, A.Y. Programmed cell death in plant reproduction. Plant. Mol. Biol 2000, 44, 267–281. [Google Scholar]
- Piffanelli, P.; Murphy, D.J. Novel organelles and targeting mechanisms in the anther tapetum. Trends Plant. Sci 1998, 3, 250–253. [Google Scholar]
- Horner, H.T.; Rogers, M.A. A comparative light and electron microscopic study of microsporogenesis in male-fertile and male-sterile pepper (Capsicum annuum L.). Can. J. Bot 1974, 52, 435–441. [Google Scholar]
- Lee, S.L.J.; Warmke, H.E. Organelle size and number in fertile and T-cytoplasmic male-sterile corn. Am. J. Bot 1979, 66, 141–148. [Google Scholar]
- De Paepe, R.; Forchioni, A.; Chetrit, P.; Vedel, F. Specific mitochondrial proteins in pollen: Presence of an additional ATP synthase beta subunit. Proc. Natl. Acad. Sci. USA 1993, 90, 5934–5938. [Google Scholar]
- Hatefi, Y. The mitochondria electron transport and oxidative phosphorylation system. Annu. Rev. Biochem 1985, 54, 1015–1069. [Google Scholar]
- Sabar, M.; Janneke, B.; Leaver, C.J. Histochemical staining and quantification of plant mitochondrial respiratory chain complexes using blue-native polyacrylamide gel electrophoresis. Plant J 2005, 44, 893–901. [Google Scholar]
- Brunori, M.; Antonini, G. Review Cytochrome c oxidase subunit structure and proton pumping. Eur. J. Biochem 1987, 169, 1–8. [Google Scholar]
- Ostermeier, C.; Lwata, S. Cytochrome c oxidase. Curr. Opin. Struc. Biol 1996, 6, 460–466. [Google Scholar]
- Li, W.Q.; Zhang, X.Q.; Xia, C. Male gametophyte defective 1, encoding the FAd subunit of mitochondrial F1Fo-ATP synthase, is essential for pollen formation in Arabidopsis thaliana. Plant Cell Physiol 2010, 51, 923–935. [Google Scholar]
- Cabezón, E.; Butler, P.J.; Runswick, M.J.; Walker, J.E. Modulation of the oligomerization state of the bovine F1-ATPase inhibitor protein, IF1, by pH. J. Biol. Chem 2000, 275, 25460–25464. [Google Scholar]
- Cabezón, E.; Runswick, M.J.; Leslie, A.G.; Walker, J.E. The structure of bovine IF1, the regulatory subunit of mitochondrial F1-ATPase. EMBO J 2001, 20, 6990–6996. [Google Scholar]
- Hashimoto, T.; Yoshida, Y.; Tagawa, K. Simultaneous bindings of ATPase inhibitor and 9K protein to F1Fo-ATPase in the presence of 15K protein in yeast mitochondria. J. Biochem 1990, 108, 17–20. [Google Scholar]
- Ichikawa, N.; Yoshida, Y.; Hashimoto, T.; Ogasawara, N.; Yoshikawa, H.; Imamoto, F.; Taqawa, K. Activation of ATP hydrolysis by an uncoupler in mutant mitochondria lacking an intrinsic ATPase inhibitor in yeast. J. Biol. Chem 1990, 265, 6274–6278. [Google Scholar]
- Venard, R.; Brethes, D.; Giraud, M.F.; Vaillier, J.; Velours, J.; Haraux, F. Investigation of the role and mechanism of IF1 and STF1 proteins, twin inhibitory peptides which interact with the yeast mitochondrial ATP synthase. Biochemistry 2003, 42, 7626–7636. [Google Scholar]
- Warmke, H.E.; Lee, S.L. Pollen abortion in T cytoplasm malesterile corn (Zea mays): A suggested mechanism. Science 1978, 200, 561–563. [Google Scholar]
- Balk, J.; Leaver, C.J. The PET1-CMS mitochondrial mutation in sunflower is associated with premature programmed cell death and cytochrome c release. Plant Cell 2001, 13, 1803–1818. [Google Scholar]
- Wang, Z.H.; Zou, Y.J.; Li, X.Y.; Zhang, Q.Y. Cytoplasmic male sterility of rice with Boro II cytoplasm is caused by a cytotoxic peptide and is restored by two releated PPR motif genes via distinct modes of mRNA silencing. Plant Cell 2006, 18, 676–687. [Google Scholar]
- Peterson, P.A. Cytoplasmically inherited male sterility in Capsicum. Am. Nat 1958, 92, 111–119. [Google Scholar]
- Kim, D.H.; Kang, J.G.; Kim, B.D. Isolation and characterization of the cytoplasmic male sterility-associated orf456 gene of chili pepper (Capsicum annuum L.). Plant. Mol. Biol 2007, 63, 519–532. [Google Scholar]
- Kim, D.H.; Kim, B.D. The organization of mitochondrial atp6 gene region in male fertile and CMS lines of pepper (Capsicum annuum L.). Curr. Genet 2006, 49, 59–67. [Google Scholar]
- Valiyaveetil, F.I.; Fillingame, R.H. Transmembrane topography of subunit a in the Escherichia coli F1Fo ATP synthase. J. Biol. Chem 1998, 273, 16241–16247. [Google Scholar]
- Shifriss, C.; Frankel, R. A new male sterility gene in Capsicum annuum L. J. Am. Soc. Hort. Sci 1969, 94, 385–387. [Google Scholar]
- Kaul, M.L.H. Male Sterility in Higher Plants; Springer Verlag Press: New York, NY, USA, 1988. [Google Scholar]
- Bhargawa, Y.R. Microsporogenesis in pollen sterile C. annuum L. Capsicum-Newsletter 1989, 7, 29. [Google Scholar]
- Peng, J.; Gong, Z.H.; Huang, W. Studies on the abortion mechanism of microspore in chili pepper CMS line H9A. Chin. Bull. Bot 2010, 45, 44–51. [Google Scholar]
- Binh, N.T.H.; Tomkova, J.H. A case of early dissolution of the microsporocyte callose wall in male-sterile (CMS) sweet pepper (Capsicum annuum L.). Biol. Plant 1982, 24, 260–265. [Google Scholar]
- Wang, S.B.; Luo, X.D.; Dai, L.F. Meiotic observation and male gametes development in cytoplasmic male sterile of pepper (Capsicum annuum L.). Acta Hortic. Sin. 2004, 31, 807–810. [Google Scholar]
- Owen, H.A.; Makaroff, C.A. Ultrastructure of microsporogenesis and microgametogenesis in Arabidopsis thaliana (L.) Heynh. ecotype Wassilewskija (Brassicaceae). Protoplasma 1995, 185, 7–21. [Google Scholar]
- Yang, J.H.; Zhang, M.F.; Yu, J.Q. Mitochondrial nad2 gene is co-transcripted with CMS-associated orfB gene in cytoplasmic male-sterile stem mustard (Brassica juncea). Mol. Biol. Rep 2009, 36, 345–351. [Google Scholar]
- Yang, J.H.; Huai, Y.; Zhang, M.F. Mitochondrial atpA gene is altered in a new orf220-type cytoplasmic male-sterile line of stem mustard (Brassica juncea). Mol. Biol. Rep 2009, 36, 273–280. [Google Scholar]
- Vedel, F.; Lalanne, E.; Sabar, M.; Chetrit, P.; de Paepe, R. The mitochondria respiratory chain and ATP synthase complexes: Composition, structure and mutational studies. Plant Physiol. Biochem 1999, 37, 629–643. [Google Scholar]
- Boyer, P.D. The ATP synthase—A splendid molecular machine. Annu. Rev. Biochem 1997, 66, 717–749. [Google Scholar]
- Wada, T.; Long, J.C.; Zhang, D.; Vik, S.B. A novel labeling approach supports the five-transmembrane model of subunit a of the Escherichia coli ATP synthase. J. Biol. Chem 1999, 274, 17353–17357. [Google Scholar]
- Paul, M.F.; Barrientos, A.; Tzagoloff, A. A single amino acid change in subunit 6 of the yeast mitochondria ATPase suppresses a null mutation in ATP10. J. Biol. Chem 2000, 275, 29238–29243. [Google Scholar]
- García-Trejo, J.J.; Morales-Ríos, E. Regulation of the F1Fo-ATP synthase rotary nanomotor in its monomeric-bacterial and dimeric-mitochondrial forms. J. Biol. Phys 2008, 34, 197–212. [Google Scholar]
- Millar, A.H.; Liddell, A.; Leaver, C.J. Isolation and subfractionation of mitochondria from plants. Method Cell Biol 2007, 80, 65–90. [Google Scholar]
- Bisswanger, H. Practical Enzymology; Chemical Industry Press: Beijing, China, 2005. [Google Scholar]
- Prasad, T.K.; Anderson, M.D.; Stewart, C.R. Acclimation, hydrogen peroxide, and abscisic acid protect mitochondria against irreversible chilling injury in maize seedlings. Plant Physiol 1994, 105, 619–627. [Google Scholar]





| Oligonucleotide | Sequence | Reference |
|---|---|---|
| orf507 (forward) | 5′-ATGCCCAAAAGTCCCATGTA-3′ | [35] |
| orf507 (reverse) | 5′-AAAAGCGCTAAACAAATTGC-3′ | This study |
| r atp6-2 (forward) | 5′-GCTTCGCCTTCGTATAGTAGTTC-3′ | This study |
| r atp6-2 (reverse) | 5′-AGTCATAGTGCTCACCCTGTTTG-3′ | This study |
| coxII (forward) | 5′-ATGATTGTTCTAGAATGGCT-3′ | [6] |
| coxII (reverse) | 5′-TTATGGGATTAATTGATTGGATACCCG-3′ | [6] |
© 2013 by the authors; licensee Molecular Diversity Preservation International, Basel, Switzerland. This article is an open-access article distributed under the terms and conditions of the Creative Commons Attribution license (http://creativecommons.org/licenses/by/3.0/).
Share and Cite
Ji, J.; Huang, W.; Yin, C.; Gong, Z. Mitochondrial Cytochrome c Oxidase and F1Fo-ATPase Dysfunction in Peppers (Capsicum annuum L.) with Cytoplasmic Male Sterility and Its Association with orf507 and Ψatp6-2 Genes. Int. J. Mol. Sci. 2013, 14, 1050-1068. https://doi.org/10.3390/ijms14011050
Ji J, Huang W, Yin C, Gong Z. Mitochondrial Cytochrome c Oxidase and F1Fo-ATPase Dysfunction in Peppers (Capsicum annuum L.) with Cytoplasmic Male Sterility and Its Association with orf507 and Ψatp6-2 Genes. International Journal of Molecular Sciences. 2013; 14(1):1050-1068. https://doi.org/10.3390/ijms14011050
Chicago/Turabian StyleJi, Jiaojiao, Wei Huang, Chuanchuan Yin, and Zhenhui Gong. 2013. "Mitochondrial Cytochrome c Oxidase and F1Fo-ATPase Dysfunction in Peppers (Capsicum annuum L.) with Cytoplasmic Male Sterility and Its Association with orf507 and Ψatp6-2 Genes" International Journal of Molecular Sciences 14, no. 1: 1050-1068. https://doi.org/10.3390/ijms14011050




