Personalized Targeted Therapy for Lung Cancer
Abstract
: Lung cancer has long been recognized as an extremely heterogeneous disease, since its development is unique in every patient in terms of clinical characterizations, prognosis, response and tolerance to treatment. Personalized medicine refers to the use of markers to predict which patient will most likely benefit from a treatment. In lung cancer, the well-developed epidermal growth factor receptor (EGFR) and the newly emerging EML4-anaplastic lymphoma kinase (ALK) are important therapeutic targets. This review covers the basic mechanism of EGFR and EML4-ALK activation, the predictive biomarkers, the mechanism of resistance, and the current targeted tyrosine kinase inhibitors. The efficacy of EGFR and ALK targeted therapies will be discussed in this review by summarizing the prospective clinical trials, which were performed in biomarker-based selected patients. In addition, the revolutionary sequencing and systems strategies will also be included in this review since these technologies will provide a comprehensive understanding in the molecular characterization of cancer, allow better stratification of patients for the most appropriate targeted therapies, eventually resulting in a more promising personalized treatment. The relatively low incidence of EGFR and ALK in non-Asian patients and the lack of response in mutant patients limit the application of the therapies targeting EGFR or ALK. Nevertheless, it is foreseeable that the sequencing and systems strategies may offer a solution for those patients.1. Introduction
Lung cancer has the highest mortality rate of all cancers, and is the second most diagnosed cancer in both men and women, behind prostate and breast cancer, respectively [1–3]. It has been reported that more than 1.6 million cases are diagnosed each year along with 1.3 million deaths [3]. Recently, the United States has released cancer statistics showing an occurrence of 221,130 new cases of lung cancer in 2011, accounting for ~14% of all cancer cases expected to be diagnosed. Approximately 85%–90% of lung cancer cases are caused by voluntary or involuntary (second hand) cigarette smoking [4]. Lung cancer is mainly divided into two classes, which are non-small cell lung cancer (NSCLC, ~85%) and small cell lung cancer (SCLC, ~15%), according to biology therapy and prognosis [1]. NSCLC could be further divided into squamous cell carcinoma (SCC), adenocarcinoma and large cell lung carcinoma (LCLC). Genetic susceptibility plays a crucial role in the occurrence of lung cancer, especially in young patients [2].
Lung cancer is aggressive and its treatment remains one of the most challenging tasks in the medical world. Conventional treatment modalities include surgery, radiation therapy and chemotherapy. The selection of therapy is dependent upon the cancer type (small cell or non-small cell), development stage, and genetic characterization. Patients diagnosed with lung cancer often receive more than one type of treatment. The discovery and development of molecular inhibitors have had a major impact in the treatment of NSCLC [5]. In the last decade, four molecular targeted agents were approved for treatment of lung cancer: gefitinib (2002), erlotinib (2003), bevacizumab (2006), and crizotinib (2011) [6]. The 1-year survival rate for lung cancer was 43% in 2003–2006. However, the overall 5-year survival rate for all stages is still as low as 16%–17% for NSCLC and even lower in SCLC (6%). Despite the 5-year survival rate reaching 53% for patients diagnosed at an early stage, only 15% of such cases were determined in a timely manner when the tumor was still localized [2].
Cancer has long been recognized as an extremely heterogeneous disease, since its development is unique in every patient in terms of clinical characterizations, prognosis, response and tolerance to treatment [7,8]. The inception of the human genome project gave birth to “personalized medicine” in cancer care treatment. This revolutionary method promises to help patients improve outcomes, avoid unnecessary treatments and reduce health care cost [8]. In oncology, the term “personalized medicine” refers to moving away from a “one size fits all” strategy in cancer therapy by the use of identifying markers that reliably predict which patient will most likely benefit from treatment [9]. Over the past 20 years, research in targeted treatment has focused on novel treatment programs. Personalized targeted therapy has already been applied towards the treatment of various cancers such as: NSCLC, squamous-cell carcinoma of the head and neck [10], colorectal cancer [11], pancreatic cancer [12], and breast cancer [13]. In the future, cancer care targeted at genes and proteins of tumor cells may assist in early detection and help physicians design the most appropriate treatment package tailored to each individual patient.
In this review, we will summarize the pathways, mechanisms, and the current corresponding inhibitors for epidermal growth factor receptor (EGFR), as well as the newly emerging EML4-anaplastic lymphoma kinase (ALK) targets. The efficacy of tyrosine kinase inhibitors (TKIs) were evaluated by summarizing prospective validating biomarker-based clinical trials. Revolutionary sequencing and systems strategies will also be reviewed since we expect these methodologies to make personalized medicine a reality by looking at whole genome sequencing and the possible aberrations, rather than just one target (such as EGFR, KRAS or EML4-ALK).
2. Molecular Targets
2.1. EGFR
EGFR is a member of the ErbB family of cell surface receptor tyrosine kinase (RTK). The EGFR family consists of four members: EGFR (or ErbB-1), HER-2 (or ErbB-2), HER-3 (or ErbB-3), and HER-4 (or ErbB4) [14]. It has been demonstrated that RTKs play a crucial role in tumorigenesis by controlling signal transduction pathways that regulate proliferation and apoptosis [15]. The RTKs (with the exception of HER-2) are activated by the binding of specific activating soluble ligands, which occur in the extracellular portion of the RTKs [16]. This interaction between ligands and receptors promotes the formation of functional active homodimers (EGFR dimer) or heterodimers (HER3 or HER4 dimer). It also activates the intracellular kinase domain and a subsequent ATP-dependent cross-autophosphorylation of C-terminal tail of the receptor [14,17]. Eventually, the phosphorylated residues recruit a diverse set of cytoplasmic signaling molecules as a docking site, which triggers downstream intracellular signaling pathways including the PI3K/AKT prosurvival, STAT transcription, and RAS/RAF/MEK proliferation pathways (Figure 1) [18,19]. It is virtually sufficient to deregulate the signaling pathways by stimulating apoptosis and growth cessation [19,20]. In NSCLC patients, 50%–80% have shown an over-expression of EGFR [21], which is associated with angiogenesis and poor prognosis [22]. The association between EGFR alteration and pathogenesis makes it a prime candidate for targeted treatments.
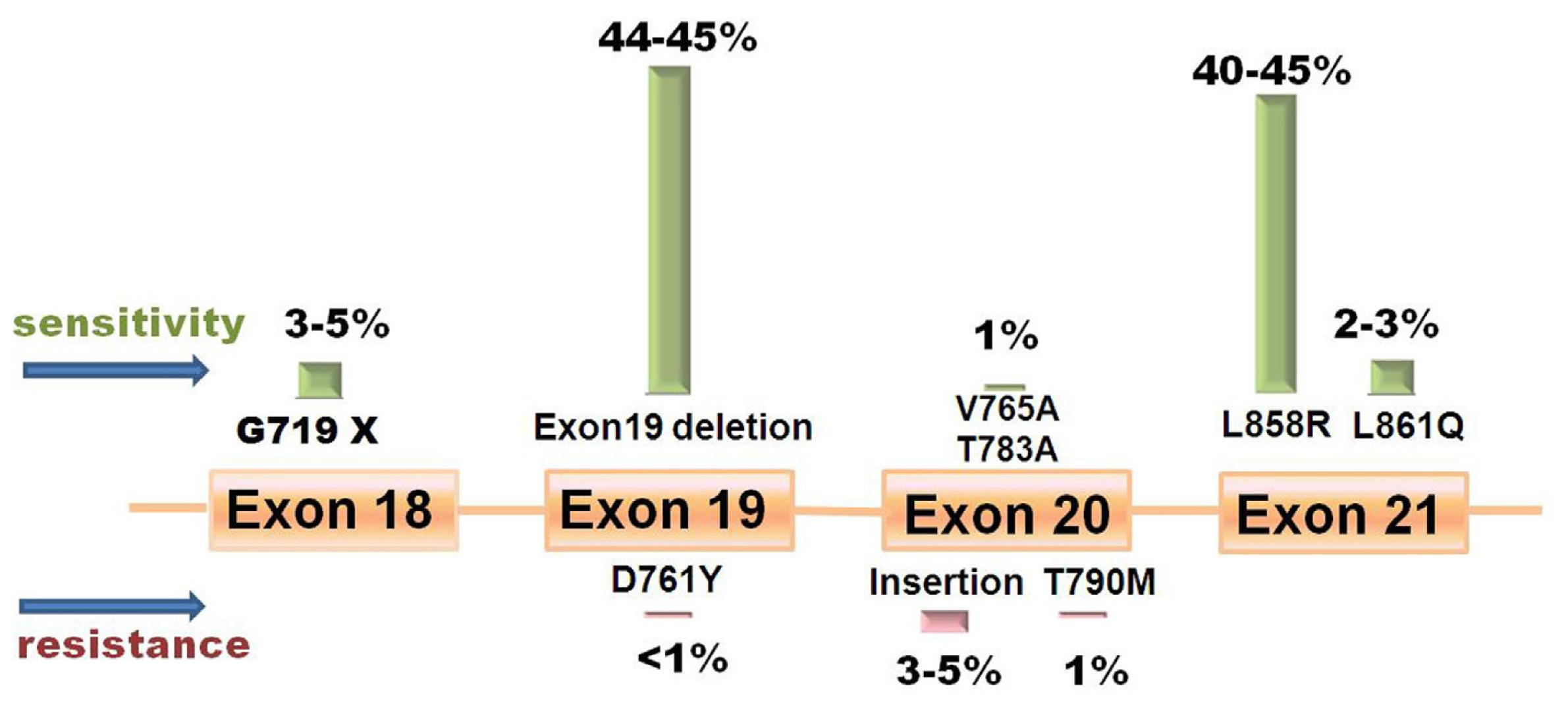
Potent response predictors are necessary to help doctors predict which patients will most likely to respond to EGFR TKIs. These predictors are ultimately used to obtain an optimal treatment while avoiding resistance. In 2004, three influential studies discovered a series of somatic mutations in the kinase domain of EGFR which are associated with the response to EGFR TKI therapy [23–25]. To date, EGFR mutations are presumed to be the strongest predictive biomarker for the efficacy of EGFR TKI therapy [26] due to much higher response rate (RR, 37.5%–100% vs. 2.9%–23% [27]; 70% vs. 33.2% as a first-line treatment; 47.4% vs. 28.5% as a second-line treatment [28]) and longer overall survival (OS, 13–23 months vs. 5–17 months [27]) in mutant patients. Mok [29] summarized six clinical trials to compare the response to EGFR TKIs and chemotherapy in patients carrying positive mutations. Patients have responded better to EGFR TKIs than to chemotherapy demonstrated by a higher RR (62.1%–84.6% vs. 10.5%–47.3%) and longer progression-free survival (PFS) (8.4–13.1 months vs. 4.6–6.7 months). In April 2011, the American Society of Clinical Oncology (ASCO) has issued a provisional clinical opinion, which suggested that initiating first-line therapy with an EGFR TKI should be based on positive EGFR mutation tests in patients with newly diagnosed advanced NSCLC [30]. EGFR mutations are more common in non-smoking East Asian females and those with adenocarcinoma histology (95% were found in adenocarcinomas) [31–36]. There are several reviews summarizing the frequency and distribution of EGFR mutations (Figure 2) [14,15,29,33,37–39].
EGFR gene copy number is also considered to be a good predictor for response to EGFR TKI therapy. It has been demonstrated in several studies that an increased copy number is associated with a higher overall RR, a longer PFS, and an OS benefit during treatment with erlotinib or gefitinib [40–42]. In fact, EGFR mutation was validated to be more selective than EGFR gene number [43].
2.2. EML4-ALK
The ALK tyrosine kinase receptor has gained much attention recently as a newly emerging relevant biomarker and therapeutic target in NSCLC. ALK is one of the members of the insulin receptor family located at chromosome 2 and encodes a trans-membrane receptor tyrosine kinase [44,45]. The activation of ALK is primarily through the formation of fusion genes (Figure 1) [46]. EML4-ALK translocation is the most common ALK gene rearrangement [47]. The intracellular kinase domain of ALK fuses with the N-terminal of EML4, and then encodes a cytoplasmic chimeric protein with kinase activity, which subsequently drives tumor growth [47]. EML4-ALK rearrangements in NSCLC patients are mainly found in younger non-smoking patients with adenocarcinoma [48,49]. EML4-ALK rearrangements are mutually exclusive with EGFR or KRAS mutations [47,50]. It has been reported that approximately 2%–11% of tumors carrying positive EML4-ALK, which is rarely found in SCC [33,36,51,52].
2.3. KRAS
KRAS mutations are a negative predictor of response to EGFR TKIs, mainly accounting for primary resistance [53]. Most KRAS mutations in lung adenocarcinoma are associated with smoking. KRAS positive mutations are limited to NSCLC (predominantly adenocarcinomas) and are mutually exclusive to mutations in EGFR and ALK [54]. In several countries, patients harboring a KRAS mutation have been excluded from EGFR TKI therapy [55].
2.4. The Potential Targets under Development
Mammalian target of rapamycin (mTOR), with serine/threonine kinase activity, appears to trigger the activation of PI3K pathway through ligand binding and eventually regulates the cell cycle. The development of mTOR inhibitors provides a great opportunity for helping patients with solid tumors. To date, studies utilizing mTOR inhibitors in NSCLC patients have reached phase I/II clinical trials as a monotherapy or in combination. These mTOR inhibitors are known as sirolimus, temsirolimus, everolimus, and others [56].
The amplification of fibroblast growth factor receptor 1 (FGFR1), predominantly in SCC (up to ~20%), is considered to be a potential target for treatment with anti-FGFR1 agents [57]. Dy et al. reported dose-dependent tumor cell death (6 out of 9 lung cancer cell lines) due to the treatment of Y15 (1,2,4,5-benzentetraamine tetra hydrochloride). Y15 is a small molecular inhibitor of focal adhesion kinase (FAK), which is a non-receptor tyrosine kinase [58]. Mutations in the DDR2 kinase gene were indicated to drive SCC. The cell lines harboring DDR2 mutations were sensitive to dasatinib, which blocked cellular transformation. DDR2 mutations are found in 4% of SCC patients [59].
3. Resistance
Some of the patients who have benefited from EGFR TKI therapy eventually generated resistance to the drug. This acquired resistance to TKI therapy is mostly due to a specifically acquired EGFR mutation known as T790M (at exon 20) [60]. T790M mutations account for more than 50% of acquired resistance in adenocarcinomas [33]. Besides the T790M mutation, the amplification of MET also activates similar downstream pathways, which accounts for 20% of acquired resistance [61].
The underlying mechanism for the remaining 30%–40% of resistance to EGFR TKIs is unknown. Ogawa et al. [62] found that the death-associated protein kinase (DAPK) is hypermethylated in resistant cells. The generation of a population of cancer cells with stem cell properties might be another possible reason of resistance to EGFR TKIs [63].
Patients treated with ALK inhibitors may acquire a resistance similar to patients taking an EGFR TKI. Choi et al. [64] have found secondary mutations within the kinase domain of EML4-ALK in tumor cells along with acquired resistance. The tumor cells harboring either C1156Y or L1196M mutations had a very low response to ALK inhibitors. L1196M mutant cells were more resistant to crizotinib than C1156Y mutant cells. The mutations in the PIK3CA gene and histologic change from NSCLC to SCLC were also found to be potential resistance mechanisms [65].
4. Targeted Agents
The main approach to block the EGFR pathway is by competing with ATP for binding to the tyrosine kinase domain. The EGFR TKIs are summarized in Table 1. Gefitinib and erlotinib are reversible inhibitors of the EGFR kinase and are also called “first-generation” small molecular inhibitors. Gefitinib was the first targeted agent entered into clinical trials currently approved by the FDA. Gefitinib “should be used only in cancer patients who have already taken the medicine and whose doctor believes it is helping them” [66]. New patients should not be given this drug due to a lack of OS benefit as shown in the ISEL trial [67]. Gefitinib is now widely prescribed in Asia. Erlotinib has received global approval as the treatment in second-line and third-line therapy. The first-generation of reversible EGFR TKIs usually generated resistance within one-year of treatment [68] prompting the development of a second-generation (Table 1). The second-generation TKIs may overcome resistance to the treatment of erlotinib or gefitinib via the T790M gatekeeper mutation. However, this activity needs to be further validated since it has also been reported that afatinib, a second-generation TKI, was not qualitatively superior in preventing the acquired resistance [69]. Several irreversible EGFR inhibitors blocked multiple EGFR family members, interrupting the cooperative signal pathway among EGFR members and resulted in a more complete blockage. It is not surprising that dacomitinib (PF299804) has a significantly longer PFS than erlotinib (p = 0.017) in patients carrying the wild type EGFR, since it‘s a potent irreversible inhibitor of EGFR, HER2, and HER4 [70]. The second-generation EGFR TKIs may have better efficacy as well as a delayed resistance, and may work in patients resistant to reversible inhibitors. There are also multiple pathways inhibitors at various clinical stages, which are shown in Table 1.
Crizotinib had received accelerated approval by the FDA, in August 2011 to treat patients diagnosed with late-stage (locally advanced or metastatic) NSCLC carrying positive ALK rearrangement. The corresponding diagnostic test method, Vysis ALK Break Apart FISH Probe Kit, was approved in tandem. AP26113 is a potent dual small-molecule inhibitor of ALK and EGFR (including T790M). A phase I dose escalation trial was initiated on Sep 2011 (NCT01449461), and phase II clinical trials with selected patients are expected to start in late 2012 according to the ARIAD website [71]. LDK378 is a selective small molecule ALK inhibitor. Preliminary responses were observed in its first-in-human trial [72]. Additional ALK inhibitors are summarized in Table 2.
5. Personalized Clinical Practices in Biomarker-Based Selected Patients
The last few decades have seen remarkable progress in target biomarker discovery, validation, biomarker measurement, research and development of targeted agents, and preclinical/clinical studies evaluating the efficacy of target inhibitors. Eventually, the prospective clinical trials in selected patients stratified by a target will bring personalized treatment from promise to reality.
5.1. Selecting Patients Based on Histology
The response to EGFR TKIs was initially thought to be related to the clinical characterization of patients, such as Asian female non-smokers with adenocarcinomas. There were studies focused on the evaluation of efficacy in these patients. Rizvi et al. [73] completed a clinical trial in patients who were enriched with the EGFR mutation (less than 15 packs per year for their cigarette smoking history and/or a component of bronchioloalveolar carcinoma (BAC)). The overall RR in all eligible patients was 42% (21 out of 50), while the RR in the mutation patients was 81% (17 out of 21). A phase II trial of erlotinib was completed recently. Forty-nine patients, who had stage III B/IV pulmonary adenocarcinoma or BAC and were non-smokers or former light smokers, were enrolled in this study. The overall RR was 25.5% in all patients, 66.7% in mutant patients, and 14.8% in wild type patients. Milella et al. reported a prospective phase II study in which patients were divided into four groups (EGFR mutation; highly polysomic/amplified EGFR; EGFR and/or pAKT positive; adenocarcinoma/BAC with a non-smoking history) and were given EGFR TKIs as second or subsequent line treatment. The 1st and 4th groups attained the best and second best overall RR (25% and 20%, respectively), disease control was highest in group 1 and group 4 (>50%), PFS and OS (p = 0.02 and 0.01, respectively) [74]. The selection of patients should be based on EGFR mutation. Clinical characterization can be a secondary criterion if EGFR mutation is not available.
5.2. Targeted Treatment in EGFR Mutant Patients
EGFR is the most developed target in lung cancer. Many preclinical and clinical studies have demonstrated that EGFR mutations are potent and selective prediction biomarkers for treating patients with EGFR TKIs. The discovery and validation of targets, including EGFR mutations, enables us move toward the eventual goal of personalized treatment based on the uniqueness of each patient. We have summarized the clinical trials in biomarker-based selection of patients.
The Biomarker-integrated Approaches of Targeted Therapy for Lung Cancer Elimination (BATTLE) trial in NSCLC was a pioneering phase II clinical trial to demonstrate the use of biomarkers to guide the treatment of patients with advanced NSCLC refractory given prior chemotherapy [75]. The BATTLE trial was more focused on the patient and utilized real-time biopsies revealing the uniqueness of each tumor and provided physicians a powerful tool for giving targeted therapies which are most likely to show efficacy. A total of 244 patients were eligible for this study and the first 97 patients were equally randomly assigned into 4 groups which were given erlotinib, vandetanib, erlotinib plus bexarotene and sorafenib, respectively. Eleven potential biomarkers were identified in the eligible patients. Once a satisfactory number of baseline results had been collected, the remaining 158 patients were assigned to treatment arms which were most likely going to give the best response based on their tumor type. The overall 8-week disease control rate (DCR) was 46% (34% for erlotinib; 33% for vandetanib; 50% for erlotinib/bexarotene and 58% for sorafenib); median PFS was 1.9 months; median OS was 35%. The 8-week landmark analysis indicated that median survival with 8-week disease control was 9.6 months, while it is 7.5 months without 8-week disease control. The major findings from the BATTLE trial reported that patients given treatment based on their tumor biomarkers showed more benefit compared to patients with unselected therapy. An effective treatment-marker-group pairing (defined as 0.8 posterior probability of exceeding a DCR of 30%) revealed that combinations of erlotinib in the VEGF/VEGFR-2 group; vandetanib in the EGFR group; erlotinib/bexarotene in EGFR, retinoid X receptor, cyclin D1, and non-marker groups; sorafenib in KRAS/BRAF, VEGF/VEGFR-2 and no marker group. An analysis of individual markers for treatment efficacy has demonstrated that EGFR mutation rates can (1) predict the response to erlotinib (p = 0.04), (2) predict a high VEGFR-2 expression for vandetanib (p = 0.05), and (3) predict a high cyclin D1 expression for erlotinib/bexarotene (p = 0.01). These predictions confirmed the authors’ hypotheses that biomarkers could better predict 8-week disease control. “The BATTLE trial represents an important model for future genotype-driven studies of targeted therapies in lung cancer,” says William Pao. Integrating the results of real-time biomarker detection, the BATTLE trial is moving towards tailoring treatment in specific patient populations for desirable individualized treatment. “Ultimately, we would like to be able to screen patients for tumor characteristics and give them appropriate therapies up front” (Dr. Edward S Kim). The following BATTLE-2 trial is ongoing.
We summarized recent clinical trials with selected patients that carry positive EGFR mutations in Table 3. The efficacy and toxicity of EGFR TKIs as first-line treatment was compared with chemotherapy in several phase III clinical trials (Table 3). Prospective trials comparing the efficacy of chemotherapy and EGFR TKIs in EGFR-mutant patients can provide further insight on the most appropriate treatment in this population. NSCLC Patients from Europe [76], China [77], and Japan [78] carrying positive EGFR mutations were enrolled in the studies. Erlotinib or gefitinib was compared to cisplatin/docetaxel or gemcitabine/carboplatin. Two [76,78] of the three studies found significant longer PFS (9.7 vs. 5.2 months, HR 0.37, p < 0.0001; 9.2 months vs. 6.3 months, HR 0.489, p < 0.0001) and higher RR (64% vs. 18%) in EGFR TKIs treatment group (Table 3). The another study [77] compared the two groups with PFS, which is longer in the erlotinib group compared to the chemotherapy group (13.1 vs. 4.6 months, HR 0.16, p < 0.0001). Chemotherapy resulted in more grade 3 or 4 toxicity compared to erlotinib [76–78]. The efficacy of EGFR TKIs’ therapy is consistent among the three populations, indicating the absence of ethnic differences. The EURTAC trial was among the first prospective head-to-head phase III study along with the other two studies. It has validated the ASCO proposal considering routine pretreatment assessment of EGFR mutations in patients with NSCLC [76]. LUX-lung 3, the largest prospective trial in EGFR mutation positive lung cancer using pemetrexed/cisplatin as a comparison, indicating that afatinib might be a potent first-line treatment in EGFR positive patients due to the improved PFS (11.1 months vs. 6.9 months) [79]. A better response can be expected when EGFR mutant patients are treated with EGFR TKIs rather than chemotherapy. The OS is usually not available in two arm controlled studies due to a common crossover or carryover effect [29,80].
The efficacy and safety of EGFR TKIs as a first-line treatment were prospectively evaluated in biomarker-based selected patients (Table 3). The RR ranged from 53.3% (Korean patients) [81] to 75% (Japanese patients) [82]. The PFS was 7.1 months to 398 days. The incidence of grade 3 toxicity is ~13% [82]. DCR was from 86.7% [81] to 96% [82]. The OS was also observed in two of these studies (819 days, 17.5 months) [81,83], approximately two-fold higher than chemotherapy in unselected NSCLC patients [83]. Median survival time is 17.8 and 20 months [84,85]. Moreover, Han et al. [86] demonstrated that erlotinib as neoadjuvant treatment in patients with stage IIIA-N2 NSCLC and with an activating EGFR mutation is feasible. Inoue et al. [84] observed the response to gefitinib treatment in EGFR mutation-positive patients with extremely poor performance status. This was the first reported observation showing patients receiving benefit from gefitinib in this population. This favorable response further validates biomarker-based treatment selection is feasible and proves that EGFR mutant patients can benefit from this selection strategy. The PFS of erlotinib and gefitinib monotherapy in EGFR mutant patients (Table 3) is comparable (9.7–13.1 months vs. 7.1–13.3 months). The RR ranged from 53.3% to 75% and the DCR ranged from 86.7% to 96% [81–88].
There are two biomarker-guided clinical studies, in which patients were assigned to either an EGFR TKI group or a chemotherapy group based on their EGFR mutation status (Table 3) [89,90]. Patients were given erlotinib or gefitinib if they were carriers of the EGFR mutation, while those with wild type EGFR received chemotherapy with/without cisplatin (depending on the BRCA1 mRNA levels). EGFR mutant patients responded to erlotinib/gefitinib better than patients with the wild type EGFR.
Pietanza et al. [91] and Sequist et al. [92] reported on the efficacy of XL647 and neratinib, respectively, in patients who either had resistance or generated resistance from prior treatment, but neither of these studies reported positive results (Table 3). There are also many ongoing biomarkerbased clinical trials, such as the PROSE trial [93] and UMIN 000005086 [94].
5.3. Clinical Trials in Patients with Wild Type EGFR
A limited EGFR mutation incidence prompted researchers to conduct studies to evaluate the efficacy of EGFR TKIs in patients with wild type EGFR (Table 4). Kobayashi et al. [95] concluded that “EGFR-TKI using erlotinib may be an alternative option for patients resistant to cytotoxic chemotherapy, even in those with EGFR wild-type NSCLC” based on the results from a phase II clinical trial in patients carrying wild type EGFR (Table 4). A similar result could be found in another clinical trial, which was performed by Matsuura et al. [96] in 2011. Erlotinib was given to patients carrying wild type EGFR as a third-line treatment. The acceptable RR (15%) and OS (6.7 months) indicated that erlotinib could be a potential third-line treatment option for patients without an EGFR mutation. Yoshioka et al. [97] also reported a phase II clinical trial in which Japanese patients carrying the wild type EGFR previously received one to three chemotherapy regimens. The objective RR was lower than the authors had initially expected. The efficacy of erlotinib in wild type EGFR patients is limited to this study. Garassino et al. [98] compared erlotinib with docetaxel and found that docetaxel is superior to erlotinib as a second line treatment. The treatment of EGFR TKIs in patients with EGFR heterogeneity was controversial according to the above prospective clinical studies, which is consistent with the conclusions from previous retrospective studies. The IPASS study has demonstrated that non-mutant patients receiving gefitinib have inferior outcomes (HR 2.85, p < 0.001) [100]. On the other hand, the BR.21 trial [101] discovered that patients carrying the wild type mutation seemed to benefit from the administration of erlotinib compared to the placebo arm (not significant). Sharma et al. [15] found that approximately 10%–20% of lung cancer patients without activating EGFR mutations had a partial response to gefitinib. Overall, the usage of EGFR TKIs in patients without a mutation needs to be studied more to confirm the efficacy or inefficacy.
Metro et al. [99] finished a clinical trial in the patients carrying wild type EGFR. The relationship between KRAS mutation status and the response to EGFR-TKI was observed in this trial. The median PFS was 1.6 months in KRAS mutants group, while it was 3.0 months in KRAS wild type group, indicating wild type EGFR plus mutant KRAS were associated with an increased resistance. Moreover, the patients with KRAS codon 13 mutants experienced worse responses than the patients harboring KRAS codon 12 mutants, which revealed that specific KRAS oncogene substitutions may lead to differential sensitivity and should be considered when predict the resistance/sensitivity.
5.4. Selecting Patients Based on KRAS Mutation
Janne et al. [102] presented their clinical study on KRAS positive patients at the 2012 ASCO meeting. It was the first prospective study to evaluate the clinical benefit of a targeted therapy for patients with KRAS mutant cancer. There were 422 patients enrolled in this study with stage IIIB-IV KRAS mutant NSCLC, who had received prior chemotherapy. Two treatment combinations compared selumetinib (AZD6244, ARRY-142866) + docetaxel vs. docetaxel + placebo. OS was longer in SEL/DOC group (9.4 months vs. 5.2 months; without significant difference). The patients in SEL/DOC group had significant better RR (DOC 0%, SEL/DOC 37%, p < 0.0001) and PFS (DOC 2.1 months, SEL/DOC 5.3 months, p = 0.0138) than those were given DOC alone, indicating that a KRAS mutant patient can benefit from targeted therapy (SEL + DOC). The BATTLE trial demonstrated that KRAS mutation tumors benefited from the therapy of sorafenib [75]. Riely et al. found that ridaforolimus, an inhibitor of mTOR, was associated with prolonged PFS in KRAS mutant patients compare with placebo (4 months vs. 2 months, p = 0.013, HR 0.36). [103]
5.5. Selecting Patients Based on ALK Rearrangement
The development of ALK inhibitors has been more rapid than EGFR TKIs. There are two reports considering the PROFILE 1005 trial during the ASCO meeting for two consecutive years [104,105]. The ongoing latter global clinical study in NSCLC patients with ALK-rearranged indicates that crizotinib gives a high RR, good PFS, modest toxicity, and improvement in patient-reported symptoms [105]. A phase I trial performed by Shaw et al. [106] was also a biomarker guided study. Patients with positive ALK were divided into a crizotinib treatment group and a control group. In the latter group, patients were treated with any second-line therapy. The authors show that crizotinib-treated ALK-positive patients had a higher OS for 1–2 years compared to the ALK-positive control group. The survival in crizotinib-treated ALK-positive patients was similar to the EGFR mutant group who were treated by EGFR TKIs indicating that ALK positive patients like EGFR mutant patients would benefit from a strategy looking at these biomarkers.
6. Sequencing and Systems Strategies
After the completion of the Human Genome Project in 2003, great improvement in genome-wide mapping technologies, such as next-generation sequencing (NGS), lead to decreased time and financial cost. This technological improvement makes it affordable and practical, leading to ‘sequencing and systems strategy’ [107]. It eases clinical decision making to match patients with the appropriate targeted treatment based on their genome information. The specific pathway targeted therapy as described above, relies on one (or a few) favorite genes. The sequencing and systems strategy is treating cancer differently by sequencing the whole genome using both tumor and normal tissue, surveying the global landscape of cancer, and then matching therapies to targets based on all the possible aberration [107]. Whole-genome sequencing (WGS) of primary tumors and matched metastatic biopsies provides a novel systematic discovery of mutational spectra underlying tumors, resulting in very rich genomic data. This informative genome data set could offer a more systematic consideration of treatment since cancer is usually associated with a variety of genetic alterations (such as structural abnormality, copy number gain, somatic single nucleotide variants, etc.). The thorough understanding of this diverse heterogeneity in the cancer genome sequence may potentially help to build a mutation-based taxonomy, shape personalized therapy [107], and predict the risk of developing a cancer [108].
Several WGS studies have been completed in lung cancer. Over 30 sites of significant somatic copy number alteration were identified in the Cancer Genome Atlas, a large-scale collaborative study, among 178 patients with SCC histology. Exome sequencing revealed 13 significantly mutated genes (false discovery rate < 0.01). The high expression was found in TP53, CDKN2A, PTEN, KEAP1, and NFE2L2. Four distinct expression subtypes, NFE2L2 and KEAP1 mutations, FGFR kinase alterations, increased global methylation and the highest rate of tobacco use, were identified by mRNA expression profiling. Whole genome shotgun sequencing detected the rearrangements involving several known tumor suppressors in 20 tumor/normal sample pairs, which were further confirmed by RNA sequencing including PTEN, RB1, NOTCH1, NF1, and CDKN2A. It was found that 75% patients (127 out of 178) had potential therapeutic targets [109]. Lee et al. [110] performed the NGS of a primary lung tumor and adjacent normal specimens from a 51-year-old male Caucasian patient, who had NSCLC and smoked for 15 years. More than 50,000 single nucleotide variants were identified in non-expressed genes and promoter regions, accounting for 17.7% of WGS data. The authors validated 530 variants, in which one lied in KRAS proto-oncogene, other 391 variants were found in coding regions, and 43 variations were structural alterations. Ju et al. [111] reported an integrated analysis of massively parallel whole-genome and transcriptome sequencing for a paired cancer/normal tissue from a young non-smoking patient with adenocarcinoma. This study discovered that the fusion of KIF5B and RET might result in a subset of NSCLC, indicating that a chimeric oncogene would make a potentially promising molecular target for diagnosis and personalized treatment. A WGS approach could also explore the prognostic indicator. Belvedera et al. [112] generated a computational index (GH index), which is derived from whole-genome copy number analysis, to evaluate the overall genomic damage. GH index showed a potential value to predict prognostic in SCC patients and may stratify patients better.
Genome-wide mapping provides a good opportunity to design clinical trials more thoughtfully and tailor individual treatments more evidently. While it also presents several logistical challenges, including the selection of exome or whole-genome sequencing; distinction between real drivers and disturbing passengers; bioinformatic support for the interpretation of genomic data and the eventual clinical implementation [113]. Currently, the studies utilizing this technology are usually performed as international collaborative for many common cancers, due to the time and cost-effective concerns [108].
7. Perspective
Overall, the landscape in lung cancer is rapidly developing with the discovery and identification of new molecular targets as well as new therapies that specifically inhibit cancerous activity. The integrated pipeline from discovery of a target to the biomarker-based treatment selection brings personalized medicine into reality. There is abundant evidence showing that patients who carry these targets can benefit from the treatment of corresponding inhibitors such as EGFR mutation and ALK arrangements. In this review, we summarized the currently completed clinical practices based on selected patients. These clinical trials have shown promising RR, ideal PFS and modest adverse effects, indicating that the biomarker-directed selection strategy is feasible in patients with a positive mutation. Most of the clinical trials had investigated erlotinib and gefitinib. The second-generation EGFR TKIs and the newly emerging ALK inhibitors have not been fully validated yet.
On the other hand, the limitation of EGFR and EML4-ALK inhibitors should not be ignored. The treatment of EGFR TKIs in patients harboring positive mutations led to a RR of 50%–75%. The intrinsic reason for a lack of response in the target population is unknown, which may improve the efficacy of targeted therapy greatly. EGFR and EML4-ALK mutations attribute to ~28% and ~11% [36] in American non-smokers and the incidence is even lower in smokers. Patients without these mutations (~87%) will not benefit from these particular inhibitors. This phenomenon has limited the overall progress in lung cancer therapy. Moreover, the activating targets are mainly found in adenocarcinoma, a NSCLC. The targeted therapy for SCC, LCLC and SCLC will need to be developed to make a broader impact on NSCLC. The whole-genome technology and the corresponding “sequencing and systems strategy” may be the key to this situation. This technology will very likely provide us with a comprehensive understanding in molecular characterization of cancer, allow us stratify patients more intelligently and match targeted therapy to the right patients, eventually resulting in a fulfilled and fully promising personalized treatment.
Since there are numerous ongoing research and clinical trials, we will expect more potent targets and biomarkers to be discovered and validated giving optimism that more targeted agents will be developed in the near future.
- Conflicts of InterestThe authors declare no conflict of interest.
References
- Herbst, R.S.; Heymach, J.V.; Lippman, S.M. Lung cancer. N. Engl. J. Med 2008, 359, 1367–1380.
- American cancer society. Cancer facts & figures 2011, Available online: http://www.cancer.org/acs/groups/content/@epidemiologysurveilance/documents/document/acspc-029771.pdf , accessed on 24 June 2012.
- Jemal, A.; Bray, F.; Center, M.M.; Ferlay, J.; Ward, E.; Forman, D. Global cancer statistics, 2011. CA. Cancer J. Clin 2011, 61, 69–90.
- National Comprehensive Cancer Network. Available online: http://www.nccn.org/professionals/physician_gls/pdf/nscl.pdf , accessed on 10 September 2012.
- Sequist, L.V.; Lynch, T.J. EGFR tyrosine kinase inhibitors in lung cancer: An evolving story. Annu. Rev. Med 2008, 59, 429–442.
- Cagle, P.T.; Chirieac, L.R. Advances in treatment of lung cancer with targeted therapy. Arch. Pathol. Lab. Med 2012, 136, 504–509.
- Burgess, D.J. Cancer genetics: Initially complex, always heterogeneous. Nat. Rev. Cancer 2011, 11, 153.
- Bunnell, C.A.; Shulman, L.N. Will we be able to care for cancer patients in the future? Oncology (Williston Park) 2010, 14, 1343–1348.
- Hoelder, S.; Clarke, P.A.; Workman, P. Discovery of small molecule cancer drugs: Successes, challenges and opportunities. Mol. Oncol 2012, 6, 155–176.
- Prince, A.; Aguirre-Ghizo, J.; Genden, E.; Posner, M.; Sikora, A. Head and neck squamous cell carcinoma: New translational therapies. Mt. Sinai. J. Med 2010, 77, 684–699.
- Ballestrero, A.; Garuti, A.; Cirmena, G.; Rocco, I.; Palermo, C.; Nencioni, A.; Scabini, S.; Zoppoli, G.; Parodi, S.; Patrone, F. Patient-tailored treatments with anti-EGFR monoclonal antibodies in advanced colorectal cancer: KRAS and beyond. Curr. Cancer Drug Targets 2012, 12, 316–328.
- Ko, A.H.; Tempero, M.A. Personalized medicine for pancreatic cancer: A step in the right direction. Gastroenterology 2009, 136, 43–45.
- Olopade, O.I.; Grushko, T.A.; Nanda, R.; Huo, D. Advances in breast cancer: Pathways to personalized medicine. Clin. Cancer Res 2008, 14, 7988–7999.
- Mitsudomi, T.; Yatabe, Y. Mutations of the epidermal growth factor receptor gene and related genes as determinants of epidermal growth factor receptor tyrosine kinase inhibitors sensitivity in lung cancer. Cancer Sci 2007, 98, 1817–1824.
- Sharma, S.V.; Bell, D.W.; Settleman, J.; Haber, D.A. Epidermal growth factor receptor mutations in lung cancer. Nat. Rev. Cancer 2007, 7, 169–181.
- Ciardiello, F.; Tortora, G. EGFR antagonists in cancer treatment. N. Engl. J. Med 2008, 358, 1160–1174.
- Scaltriti, M.; Baselga, J. The epidermal growth factor receptor pathway: A model for targeted therapy. Clin. Cancer Res 2006, 12, 5268–5272.
- Citri, A.; Yarden, Y. EGF-ERBB signalling: Towards the systems level. Nat. Rev. Mol. Cell Biol 2006, 7, 505–516.
- Gazdar, A.F.; Minna, J.D. Deregulated EGFR signaling during lung cancer progression: Mutations, amplicons, and autocrine loops. Cancer Prev. Res. (Phila) 2008, 1, 156–160.
- Sharma, S.V.; Settleman, J. Oncogenic shock: Turning an activated kinase against the tumor cell. Cell Cycle 2006, 5, 2878–2880.
- Martine, E.N.; Hahn, M.S.; McKenna, G.W. Molecular Biology and Genetics of Lung Cancer. In Advances in Radiation Oncology in Lung Cancer; Jeremić, B., Ed.; Springer-Verlag Berlin: Heidelberg, Germany, 2005; p. 6.
- Belani, C.P.; Goss, G.; Blumenschein, G., Jr. Recent clinical developments and rationale for combining targeted agents in non-small cell lung cancer (NSCLC). Cancer Treat. Rev. 2012, 38, 173–184.
- Lynch, T.J.; Bell, D.W.; Sordella, R.; Gurubhagavatula, S.; Okimoto, R.A.; Brannigan, B.W.; Harris, P.L.; Haserlat, S.M.; Supko, J.G.; Haluska, F.G.; et al. Activating mutations in the epidermal growth factor receptor underlying responsiveness of non-small-cell lung cancer to gefitinib. N. Engl. J. Med 2004, 350, 2129–2139.
- Paez, J.G.; Jänne, P.A.; Lee, J.C.; Tracy, S.; Greulich, H.; Gabriel, S.; Herman, P.; Kaye, F.J.; Lindeman, N.; Boggon, T.J.; et al. EGFR mutations in lung cancer: Correlation with clinical response to gefitinib therapy. Science 2004, 304, 1497–1500.
- Pao, W.; Miller, V.; Zakowski, M.; Doherty, J.; Politi, K.; Sarkaria, I.; Singh, B.; Heelan, R.; Rusch, V.; Fulton, L.; et al. EGF receptor gene mutations are common in lung cancers from “never smokers” and are associated with sensitivity of tumors to gefitinib and erlotinib. Proc. Natl. Acad. Sci. USA 2004, 101, 13306–13311.
- Fukuoka, M.; Wu, Y.L.; Thongprasert, S.; Sunpaweravong, P.; Leong, S.S.; Sriuranpong, V.; Chao, T.Y.; Nakagawa, K.; Chu, D.T.; Saijo, N.; et al. Biomarker analyses and final overall survival results from a phase III, randomized, open-label, first-line study of gefitinib versus carboplatin/paclitaxel in clinically selected patients with advanced non-small-cell lung cancer in Asia (IPASS). J. Clin. Oncol 2011, 29, 2866–2874.
- Ulivi, P.; Calistri, D.; Zoli, W.; Amadori, D. Predictive molecular markers for EGFR-TKI in non-small cell lung cancer patients: New insights and critical aspects. J. Nucleic Acids Investig 2010, 1, 47–54.
- Petrelli, F.; Borgonovo, K.; Cabiddu, M.; Barni, S. Efficacy of EGFR tyrosine kinase inhibitors in patients with EGFR-mutated non-small-cell lung cancer: A meta-analysis of 13 randomized trials. Clin. Lung Cancer 2012, 13, 107–114.
- Mok, T.S. Personalized medicine in lung cancer: What we need to know. Nat. Rev. Clin. Oncol 2011, 8, 661–668.
- Keedy, V.L.; Temin, S.; Somerfield, M.R.; Beasley, M.B.; Johnson, D.H.; McShane, L.M.; Milton, D.T.; Strawn, J.R.; Wakelee, H.A.; Giaccone, G. American Society of Clinical Oncology provisional clinical opinion: Epidermal growth factor receptor (EGFR) mutation testing for patients with advanced non-small-cell lung cancer considering first-line EGFR tyrosine kinase inhibitor therapy. J. Clin. Oncol 2011, 29, 2121–2127.
- Stella, G.M.; Luisetti, M.; Inghilleri, S.; Cemmi, F.; Scabini, R.; Zorzetto, M.; Pozzi, E. Targeting EGFR in non-small-cell lung cancer: Lessons, experiences, strategies. Respir. Med 2012, 106, 173–183.
- Sequist, L.V.; Bell, D.W.; Lynch, T.J.; Haber, D.A. Molecular predictors of response to epidermal growth factor receptor antagonists in non-small-cell lung cancer. J. Clin. Oncol 2007, 25, 587–595.
- Cheng, L.; Alexander, R.E.; Maclennan, G.T.; Cummings, O.W.; Montironi, R.; Lopez-Beltran, A.; Cramer, H.M.; Davidson, D.D.; Zhang, S. Molecular pathology of lung cancer: Key to personalized medicine. Mod. Pathol 2012, 25, 347–369.
- Shigematsu, H.; Lin, L.; Takahashi, T.; Nomura, M.; Suzuki, M.; Wistuba, I.I.; Fong, M.K.; Lee, H.; Toyooka, S.; Shimizu, N.; et al. Clinical and biological features associated with epidermal growth factor receptor gene mutations in lung cancers. J. Natl. Cancer Inst 2005, 97, 339–346.
- Riely, G.J.; Politi, K.A.; Miller, V.A.; Pao, W. Update on epidermal growth factor receptor mutations in non-small cell lung cancer. Clin. Cancer Res 2006, 12, 7232–7241.
- Couraud, S.; Zalcman, G.; Milleron, B.; Morin, F.; Souquet, P.J. Lung cancer in never smokers—A review. Eur. J. Cancer 2012, 48, 1299–1311.
- Yasuda, H.; Kobayashi, S.; Costa, D.B. EGFR exon 20 insertion mutations in non-small-cell lung cancer: Preclinical data and clinical implications. Lancet Oncol 2012, 13, e23–e31.
- Ladanyi, M.; Pao, W. Lung adenocarcinoma: Guiding EGFR-targeted therapy and beyond. Mod. Pathol 2008, 21, S16–S22.
- Shigematsu, H.; Gazdar, A.F. Somatic mutations of epidermal growth factor receptor signaling pathway in lung cancers. Int. J. Cancer 2006, 118, 257–262.
- Lee, Y.; Shim, H.S.; Park, M.S.; Kim, J.H.; Ha, S.J.; Kim, S.H.; Cho, B.C. High EGFR gene copy number and skin rash as predictive markers for EGFR tyrosine kinase inhibitors in patients with advanced squamous cell lung carcinoma. Clin. Cancer Res 2012, 18, 1760–1768.
- Hirsch, F.R.; Varella-Garcia, M.; McCoy, J.; West, H.; Xavier, A.C.; Gumerlock, P.; Bunn, P.A., Jr; Franklin, W.A.; Crowley, J.; Gandara, D.R.; et al. Increased epidermal growth factor receptor gene copy number detected by fluorescence in situ hybridization associates with increased sensitivity to gefitinib in patients with bronchioloalveolar carcinoma subtypes: A Southwest Oncology Group study. J. Clin. Oncol. 2005, 23, 6838–6845.
- Cappuzzo, F.; Hirsch, F.R.; Rossi, E.; Bartolini, S.; Ceresoli, G.L.; Bemis, L.; Haney, J.; Witta, S.; Danenberg, K.; Domenichini, I.; et al. Epidermal growth factor receptor gene and protein and gefitinib sensitivity in non-small-cell lung cancer. J. Natl. Cancer Inst 2005, 97, 643–655.
- Sholl, L.M.; Xiao, Y.; Joshi, V.; Yeap, B.Y.; Cioffredi, L.A.; Jackman, D.M.; Lee, C.; Jänne, P.A.; Lindeman, N.I. EGFR mutation is a better predictor of response to tyrosine kinase inhibitors in non-small cell lung carcinoma than FISH, CISH, and immunohistochemistry. Am. J. Clin. Pathol 2010, 133, 922–934.
- Lee, C.C.; Jia, Y.; Li, N.; Sun, X.; Ng, K.; Ambing, E.; Gao, M.Y.; Hua, S.; Chen, C.; Kim, S.; et al. Crystal structure of the ALK (anaplastic lymphoma kinase) catalytic domain. Biochem. J 2010, 430, 425–437.
- Morris, S.W.; Naeve, C.; Mathew, P.; James, P.L.; Kirstein, M.N.; Cui, X.; Witte, D.P. ALK, the chromosome 2 gene locus altered by the t(2;5) in non-Hodgkin’s lymphoma, encodes a novel neural receptor tyrosine kinase that is highly related to leukocyte tyrosine kinase (LTK). Oncogene 1997, 14, 2175–2188.
- Scagliotti, G.; Stahel, R.A.; Rosell, R.; Thatcher, N.; Soria, J.C. ALK translocation and crizotinib in non-small cell lung cancer: An evolving paradigm in oncology drug development. Eur. J. Cancer 2012, 48, 961–973.
- Soda, M.; Choi, Y.L.; Enomoto, M.; Takada, S.; Yamashita, Y.; Ishikawa, S.; Fujiwara, S.; Watanabe, H.; Kurashina, K.; Hatanaka, H.; et al. Identification of the transforming EML4-ALK fusion gene in non-small-cell lung cancer. Nature 2007, 448, 561–566.
- Kwak, E.L.; Bang, Y.J.; Camidge, D.R.; Shaw, A.T.; Solomon, B.; Maki, R.G.; Ou, S.H.; Dezube, B.J.; Jänne, P.A.; Costa, D.B.; et al. Anaplastic lymphoma kinase inhibition in non-small-cell lung cancer. N. Engl. J. Med 2010, 363, 1693–1703.
- Chen, J.; Liu, H.; Yang, T.; Wei, S.; Zhou, Q. Clinical significance of the EML4-ALK fusion gene and association with EGFR and KRAS gene mutation in 208 Chinese patients with non-small cell lung cancer. J. Clin. Oncol 2012, 30, e17514.
- Sun, Y.; Ren, Y.; Fang, Z.; Li, C.; Fang, R.; Gao, B.; Han, X.; Tian, W.; Pao, W.; Chen, H.; et al. Lung adenocarcinoma from East Asian never-smokers is a disease largely defined by targetable oncogenic mutant kinases. J. Clin. Oncol 2010, 28, 4616–4620.
- Janku, F.; Stewart, D.J.; Kurzrock, R. Targeted therapy in non-small-cell lung cancer—Is it becoming a reality? Nat. Rev. Clin. Oncol 2010, 7, 401–414.
- Pao, W.; Girard, N. New driver mutations in non-small-cell lung cancer. Lancet Oncol 2011, 12, 175–180.
- Linardou, H.; Dahabreh, I.J.; Kanaloupiti, D.; Siannis, F.; Bafaloukos, D.; Kosmidis, P.; Papadimitriou, C.A.; Murray, S. Assessment of somatic K-RAS mutations as a mechanism associated with resistance to EGFR-targeted agents: A systematic review and meta-analysis of studies in advanced non-small-cell lung cancer and metastatic colorectal cancer. Lancet Oncol 2008, 9, 962–972.
- Pao, W.; Wang, T.Y.; Riely, G.J.; Miller, V.A.; Pan, Q.; Ladanyi, M.; Zakowski, M.F.; Heelan, R.T.; Kris, M.G.; Varmus, H.E. KRAS mutations and primary resistance of lung adenocarcinomas to gefitinib or erlotinib. PLoS Med 2005, 2, e17.
- Shepherd, F.A.; Tsao, M.S. Epidermal growth factor receptor biomarkers in non-small-cell lung cancer: A riddle, wrapped in a mystery, inside an enigma. J. Clin. Oncol 2010, 28, 903–905.
- Ekman, S.; Wynes, M.W.; Hirsch, F.R. The mTOR pathway in lung cancer and implications for therapy and biomarker analysis. J. Thorac. Oncol 2012, 7, 947–953.
- Marti, M.A.; Martinez, P.; Aura, C.; Cedres, S.; Sullivan, I.; Jimenez, J.; Prudkin, L.; Montero, A.M.; Murtra-Garrell, N.; Zamora, E.; et al. Amplification of fibroblast growth factor receptor type 1 gene (FGFR1) in samples from 101 NSCLC patients (pts) with squamous cell carcinoma (SCC) histology. J. Clin. Oncol 2012, 30. abstr7041.
- Dy, K.G.; Shao, H.; Ho, B.; Yemma, M.; Adjei, A.; Golubovskaya, V.; Cance, V. Preclinical investigation of the antitumor efficacy in lung cancer of Y15, a novel focal adhesion kinase inhibitor. Mol. Cancer Ther 2011, 10, S1.
- Hammerman, P.S.; Sos, M.L.; Ramos, A.H.; Xu, C.; Dutt, A.; Zhou, W.; Brace, L.E.; Woods, B.A.; Lin, W.; Zhang, J.; et al. Mutations in the DDR2 kinase gene identify a novel therapeutic target in squamous cell lung cancer. Cancer Discov 2011, 1, 78–89.
- Pao, W.; Miller, V.A.; Politi, K.A.; Riely, G.J.; Somwar, R.; Zakowski, M.F.; Kris, M.G.; Varmus, H. Acquired resistance of lung adenocarcinomas to gefitinib or erlotinib is associated with a second mutation in the EGFR kinase domain. PLoS Med 2005, 2, e73.
- Ma, C.; Wei, S.; Song, Y. T790M and acquired resistance of EGFR TKI: A literature review of clinical reports. J. Thorac. Dis 2011, 3, 10–18.
- Ogawa, T.; Liggett, T.E.; Melnikov, A.A.; Monittom, C.L.; Kusuke, D.; Shiga, K.; Kobayashi, T.; Horii, A.; Chatterjee, A.; Levenson, V.V.; et al. Methylation of death-associated protein kinase is associated with cetuximab and erlotinib resistance. Cell Cycle 2012, 11, 1656–1663.
- Ghosh, G.; Lian, X.; Kron, S.J.; Palecek, S.P. Properties of resistant cells generated from lung cancer cell lines treated with EGFR inhibitors. BMC Cancer 2012, 12, 95.
- Choi, Y.L.; Soda, M.; Yamashita, Y.; Ueno, T.; Takashima, J.; Nakajima, T.; Yatabe, Y.; Takeuchi, K.; Hamada, T.; Haruta, H.; et al. ALK Lung Cancer Study Group. EML4-ALK mutations in lung cancer that confer resistance to ALK inhibitors. N. Engl. J. Med 2010, 363, 1734–1739.
- Sequist, L.V.; Waltman, B.A.; Dias-Santagata, D.; Digumarthy, S.; Turke, A.B.; Fidias, P.; Bergethon, K.; Shaw, A.T.; Gettinger, S.; Cosper, A.K.; et al. Genotypic and histological evolution of lung cancers acquiring resistance to EGFR inhibitors. Sci. Transl. Med 2011, 3, doi:10.1126/scitranslmed.3002003.
- Gefitinib (marketed as Iressa) Information, Available online: http://www.fda.gov/Drugs/DrugSafety/PostmarketDrugSafetyInformationforPatientsandProviders/ucm110473.htm , accessed on 07 July 2012.
- Thatcher, N.; Chang, A.; Parikh, P.; Rodrigues Pereira, J.; Ciuleanu, T.; von Pawel, J.; Thongprasert, S.; Tan, E.H.; Pemberton, K.; Archer, V.; et al. Gefitinib plus best supportive care in previously treated patients with refractory advanced non-small-cell lung cancer: Results from a randomized, placebo-controlled, multicentre study (Iressa Survival Evaluation in Lung Cancer). Lancet 2005, 366, 1527–1537.
- Metro, G.; Crinò, L. Advances on EGFR mutation for lung cancer. Transl. Lung Cancer Res 2012, 1, 5–13.
- Kim, Y.; Ko, J.; Cui, Z.; Abolhoda, A.; Ahn, J.S.; Ou, S.H.; Ahn, M.J.; Park, K. The EGFR T790M mutation in acquired resistance to an irreversible second-generation EGFR inhibitor. Mol. Cancer Ther 2012, 11, 784–791.
- Boyer, J.M.; Blackhall, H.F.; Park, K.; Barrios, H.C.; Krzakowski, J.M.; Taylor, I.; Liang, Q.J.; Denis, J.L.; O’Connell, P.J.; Ramalingam, S.S. Efficacy and safety of PF299804 versus erlotinib (E): A global, randomized phase II trial in patients (pts) with advanced non-small cell lung cancer (NSCLC) after failure of chemotherapy (CT). J. Clin. Oncol 2010, 28, 18s.
- Available online: http://www.ariad.com/wt/tertiarypage/alk_inhibitor , accessed on 24 June 2012.
- Mehra, R.; Camidge, D.R.; Sharma, S.; Felip, E.; Tan, D.S.; Vansteenkiste, J.F.; de Pas, T.M.; Kim, D.W.; Santoro, A.; Liu, G.; et al. First-in-human phase I study of the ALK inhibitor LDK378 in advanced solid tumors. J. Clin. Oncol 2012, 30. abstr 3007.
- Rizvi, A.N.; Pao, W.; Kris, G.M.; Rusch, V.; Ladanyi, M.; Ginex, K.P.; Zakowski, F.M.; Tyson, B.L.; Heelan, T.R.; Varmus, H. A prospective study to correlate EGFR mutations with gefitinib response. J. Clin. Oncol 2005, 23. abstr7091.
- Milella, M.; Nuzzo, C.; Bria, E.; Sperduti, I.; Visca, P.; Buttitta, F.; Antoniani, B.; Merola, R.; Gelibter, A.; Cuppone, F.; et al. EGFR molecular profiling in advanced NSCLC: A prospective phase II study in molecularly/clinically selected patients pretreated with chemotherapy. J. Thorac. Oncol 2012, 7, 672–680.
- Kim, E.S.; Herbst, R.S.; Wistuba, I.I.; Lee, J.J.; Blumenschein, G.R., Jr; Tsao, A.; Stewart, D.J.; Hicks, M.E.; Erasmus, J., Jr; Gupta, S.; et al. The BATTLE trial: Personalizing therapy for lung cancer. Cancer Discov. 2011, 1, 44–53.
- Rosell, R.; Carcereny, E.; Gervais, R.; Vergnenegre, A.; Massuti, B.; Felip, E.; Palmero, R.; Garcia-Gomez, R.; Pallares, C.; Sanchez, J.M.; et al. Erlotinib versus standard chemotherapy as first-line treatment for European patients with advanced EGFR mutation-positive non-small-cell lung cancer (EURTAC): A multicentre, open-label, randomised phase 3 trial. Lancet Oncol 2012, 13, 239–246.
- Zhou, C.; Wu, Y.L.; Chen, G.; Feng, J.; Liu, X.Q.; Wang, C.; Zhang, S.; Wang, J.; Zhou, S.; Ren, S.; et al. Erlotinib versus chemotherapy as first-line treatment for patients with advanced EGFR mutation-positive non-small-cell lung cancer (OPTIMAL, CTONG-0802): A multicentre, open-label, randomised, phase 3 study. Lancet Oncol 2011, 12, 735–742.
- Mitsudomi, T.; Morita, S.; Yatabe, Y.; Negoro, S.; Okamoto, I.; Tsurutani, J.; Seto, T.; Satouchi, M.; Tada, H.; Hirashima, T.; et al. Gefitinib versus cisplatin plus docetaxel in patients with non-small-cell lung cancer harbouring mutations of the epidermal growth factor receptor (WJTOG3405): An open label, randomised phase 3 trial. Lancet Oncol 2010, 11, 121–128.
- Yang, C.J.; Schuler, H.M.; Yamamoto, N.; O’Byrne, J.K.; Hirsh, V.; Mok, T.; Geater, L.S.; Orlov, V.S.; Tsai, C.; Boyer, J.M.; et al. LUX-Lung 3: A randomized, open-label, phase III study of afatinib versus pemetrexed and cisplatin as first-line treatment for patients with advanced adenocarcinoma of the lung harboring EGFR-activating mutations. 2012, 30, 7500.
- Inoue, A.; Kobayashi, K.; Maemondo, M.; Sugawara, S.; Oizumi, S.; Isobe, H.; Gemma, A.; Saijo, Y.; Yoshizawa, H.; Morita, S.; et al. Final overall survival results of NEJ002, a phase III trial comparing gefitinib to carboplatin (CBDCA) plus paclitaxel (TXL) as the first-line treatment for advanced non-small cell lung cancer (NSCLC) with EGFR mutations. J. Clin. Oncol 2011, 29. abstr 7519.
- Kim, D.W.; Lee, S.H.; Lee, J.S.; Lee, M.A.; Kang, J.H.; Kim, S.Y.; Shin, S.W.; Kim, H.K.; Heo, D.S. A multicenter phase II study to evaluate the efficacy and safety of gefitinib as first-line treatment for Korean patients with advanced pulmonary adenocarcinoma harboring EGFR mutations. Lung Cancer 2011, 71, 65–69.
- Tamura, K.; Okamoto, I.; Kashii, T.; Negoro, S.; Hirashima, T.; Kudoh, S.; Ichinose, Y.; Ebi, N.; Shibata, K.; Nishimura, T.; et al. Multicentre prospective phase II trial of gefitinib for advanced non-small cell lung cancer with epidermal growth factor receptor mutations: Results of the West Japan Thoracic Oncology Group trial (WJTOG0403). Br. J. Cancer 2008, 98, 907–914.
- Sequist, L.V.; Martins, R.G.; Spigel, D.; Grunberg, S.M.; Spira, A.; Jänne, P.A.; Joshi, V.A.; McCollum, D.; Evans, T.L.; Muzikansky, A.; et al. First-line gefitinib in patients with advanced non-small-cell lung cancer harboring somatic EGFR mutations. J. Clin. Oncol 2008, 26, 2442–2449.
- Inoue, A.; Kobayashi, K.; Usui, K.; Maemondo, M.; Okinaga, S.; Mikami, I.; Ando, M.; Yamazaki, K.; Saijo, Y.; Gemma, A.; et al. First-line gefitinib for patients with advanced non-small-cell lung cancer harboring epidermal growth factor receptor mutations without indication for chemotherapy. J. Clin. Oncol 2009, 27, 1394–1400.
- Sugio, K.; Uramoto, H.; Onitsuka, T.; Mizukami, M.; Ichiki, Y.; Sugaya, M.; Yasuda, M.; Takenoyama, M.; Oyama, T.; Hanagiri, T.; et al. Prospective phase II study of gefitinib in non-small cell lung cancer with epidermal growth factor receptor gene mutations. Lung Cancer 2009, 64, 314–318.
- Han, B.; Xiong, L.; Sun, J.; Li, R.; Lou, Y.; Zhang, Y. Erlotinib as neoadjuvant treatment in patients with stage IIIA-N2 non-small cell lung cancer (NSCLC) with activating epidermal growth factor receptor (EGFR) mutation (NCT01217619, ESTERN). J. Clin. Oncol 2012, 30. abstr e17551.
- Yang, J.C.; Shih, J.Y.; Su, W.C.; Hsia, T.C.; Tsai, C.M.; Ou, S.H.; Yu, C.J.; Chang, G.C.; Ho, C.L.; Sequist, L.V.; et al. Afatinib for patients with lung adenocarcinoma and epidermal growth factor receptor mutations (LUX-Lung 2): A phase II trial. Lancet Oncol 2012, 13, 539–548.
- Kris, G.M.; Mok, T.; Ou, S.H.; Martins, R.; Kim, W.D.; Goldberg, Z.; Zhang, H.; Taylor, I.; Letrent, P.S.; Janne, A.P. First-line dacomitinib (PF-00299804), an irreversible pan-HER tyrosine kinase inhibitor, for patients with EGFR-mutant lung cancers. J. Clin. Oncol 2012, 30. abstr 7530.
- Zhong, W.; Yang, X.; Liao, R.; Nie, Q.; Dong, S.; Su, J.; Zhang, X.; Zhou, Q.; Yang, J.; Wu, L.Y. Induction erlotinib or gemcitabine/carboplatin factorial assignment therapy in stage IIIA-N2 non-small cell lung cancer. J. Clin. Oncol 2011, 29. abstr e17512.
- Rosell, R.; Perez-Roca, L.; Sanchez, J.J.; Cobo, M.; Moran, T.; Chaib, I.; Provencio, M.; Domine, M.; Sala, M.A.; Jimenez, U.; et al. Customized treatment in non-small-cell lung cancer based on EGFR mutations and BRCA1 mRNA expression. PLoS One 2009, 4, e5133.
- Pietanza, M.C.; Gadgeel, S.M.; Dowlati, A.; Lynch, T.J.; Salgia, R.; Rowland, K.M., Jr; Wertheim, M.S.; Price, K.A.; Riely, G.J.; Azzoli, C.G.; et al. Phase II study of the multitargeted tyrosine kinase inhibitor XL647 in patients with non-small-cell lung cancer. J. Thorac. Oncol. 2012, 7, 856–865.
- Sequist, L.V.; Besse, B.; Lynch, T.J.; Miller, V.A.; Wong, K.K.; Gitlitz, B.; Eaton, K.; Zacharchuk, C.; Freyman, A.; Powell, C.; et al. Neratinib, an irreversible pan-ErbB receptor tyrosine kinase inhibitor: Results of a phase II trial in patients with advanced non-small-cell lung cancer. J. Clin. Oncol 2010, 28, 3076–3083.
- Sorlini, C.; Barni, S.; Petrelli, F.; Novello, S.; de Marinis, F.; de Pas, M.T.; Grossi, F.; Bearz, A.; Mencoboni, M.; Aieta, M; et al. PROSE: Randomized proteomic stratified phase III study of second line erlotinib versus chemotherapy in patients with inoperable non-small cell lung cancer (NSCLC). J. Clin. Oncol 2011, 29. abstr TPS214.
- Hisamoto, A.; Sasaki, J.; Takigawa, N.; Shioyama, Y.; Kishimoto, J.; Takemoto, M.; Hotta, K.; Tanimoto, M.; Ichinose, Y.; Kiura, K; et al. A phase II trial of induction gefitinib monotherapy followed by cisplatin-docetaxel and concurrent thoracic irradiation in patients with EGFR-mutant locally advanced non-small-cell lung cancer (LA-NSCLC): LOGIK0902/OLCSG0905 intergroup trial. J. Clin. Oncol 2012, 30. abstr 7045.
- Kobayashi, T.; Koizumi, T.; Agatsuma, T.; Yasuo, M.; Tsushima, K.; Kubo, K.; Eda, S.; Kuraishi, H.; Koyama, S.; Hachiya, T.; et al. A phase II trial of erlotinib in patients with EGFR wild-type advanced non-small-cell lung cancer. Cancer Chemother. Pharmacol 2012, 69, 1241–1246.
- Matsuura, S.; Inui, N.; Ozawa, Y.; Nakamura, Y.; Toyoshima, M.; Yasuda, K.; Yamada, T.; Shirai, T.; Suganuma, H.; Yokomura, K.; et al. Phase II study of erlotinib as third-line monotherapy in patients with advanced non-small-cell lung cancer without epidermal growth factor receptor mutations. Jpn. J. Clin. Oncol 2011, 41, 959–963.
- Yoshioka, H.; Hotta, K.; Kiura, K.; Takigawa, N.; Hayashi, H.; Harita, S.; Kuyama, S.; Segawa, Y.; Kamei, H.; Umemura, S.; et al. A phase II trial of erlotinib monotherapy in pretreated patients with advanced non-small cell lung cancer who do not possess active EGFR mutations: Okayama Lung Cancer Study Group trial 0705. J. Thorac. Oncol 2010, 5, 99–104.
- Garassino, C.M.; Martelli, O.; Bettini, A.; Floriani, I.; Copreni, E.; Lauricella, C.; Ganzinelli, M.; Marabese, M.; Broggini, M.; Veronese, S.; et al. TAILOR: A phase III trial comparing erlotinib with docetaxel as the second-line treatment of NSCLC patients with wild-type (wt) EGFR. J. Clin. Oncol 2012, 30. abstr LBA7501.
- Metro, G.; Chiari, R.; Duranti, S.; Siggillino, A.; Fischer, M.J.; Giannarelli, D.; Ludovini, V.; Bennati, C.; Marcomigni, L.; Baldi, A.; et al. Impact of specific mutant KRAS on clinical outcome of EGFR-TKI-treated advanced non-small cell lung cancer patients with an EGFR wild type genotype. Lung Cancer 2012. in press.
- Mok, T.S.; Wu, Y.L.; Thongprasert, S.; Yang, C.H.; Chu, D.T.; Saijo, N.; Sunpaweravong, P.; Han, B.; Margono, B.; Ichinose, Y.; et al. Gefitinib or carboplatin-paclitaxel in pulmonary adenocarcinoma. N. Engl. J. Med 2009, 361, 947–957.
- Zhu, C.Q.; da Cunha Santos, G.; Ding, K.; Sakurada, A.; Cutz, J.C.; Liu, N.; Zhang, T.; Marrano, P.; Whitehead, M.; Squire, J.A.; et al. Role of KRAS and EGFR as biomarkers of response to erlotinib in National Cancer Institute of Canada Clinical Trials Group Study BR.21. J. Clin. Oncol 2008, 26, 4268–4275.
- Janne, A.P.; Shaw, T.A.; Pereira, J.R.; Jeannin, G.; Vansteenkiste, J.; Barrios, H.C.; Franke, A.F.; Grinsted, L.; Smith, D.P.; Zazulina, V.; et al. Phase II double-blind, randomized study of selumetinib (SEL) plus docetaxel (DOC) versus DOC plus placebo as second-line treatment for advanced KRAS mutant non-small cell lung cancer (NSCLC). J. Clin. Oncol 2012, 30. abstr 7503.
- Riely, J.G.; Brahmer, R.J.; Planchard, D.; Crinò, L.; Doebele, C.R.; Mas Lopez, L.; Gettinger, N.S.; Schumann, C.; Li, X.; Atkins, B.M.; et al. A randomized discontinuation phase II trial of ridaforolimus in non-small cell lung cancer (NSCLC) patients with KRAS mutations. J. Clin. Oncol 2012, 30. abstr 7531.
- Crinò, L.; Kim, D.; Riely, J.G.; Janne, A.P.; Blackhall, H.F.; Camidge, R.D.; Hirsh, V.; Mok, T.; Solomon, J.B.; Park, K.; et al. Initial phase II results with crizotinib in advanced ALK-positive non-small cell lung cancer (NSCLC): PROFILE 1005. J. Clin. Oncol 2011, 29. abstr 7514.
- Kim, D.W.; Ahn, M.J.; Shi, Y.; De Pas, T.M.; Yang, P.C.; Riely, J.G.; Crinò, L.; Evans, L.T.; Liu, X.; Han, J.Y.; et al. Results of a global phase II study with crizotinib in advanced ALK-positive non-small cell lung cancer (NSCLC). J. Clin. Oncol 2012, 30. abstr 7533.
- Shaw, A.T.; Yeap, B.Y.; Solomon, B.J.; Riely, G.J.; Gainor, J.; Engelman, J.A.; Shapiro, G.I.; Costa, D.B.; Ou, S.H.; Butaney, M.; et al. Effect of crizotinib on overall survival in patients with advanced non-small-cell lung cancer harbouring ALK gene rearrangement: A retrospective analysis. Lancet Oncol 2011, 12, 1004–1012.
- Cho, W.; Ziogas, D.E.; Katsios, C.; Roukos, D.H. Emerging personalized oncology: Sequencing and systems strategies. Futur. Oncol 2012, 8, 637–641.
- Daniels, M.; Goh, F.; Wright, C.M.; Sriram, K.B.; Relan, V.; Clarke, B.E.; Duhig, E.E.; Bowman, R.V.; Yang, I.A.; Fong, K.M.; et al. Whole genome sequencing for lung cancer. J. Thorac. Dis 2012, 4, 155–163.
- Govindan, R.; Hammerman, S.P.; Hayes, N.D.; Wilkerson, D.M.; Baylin, S.; Meyerson, M. Comprehensive genomic characterization of squamous cell carcinoma of the lung. J. Clin. Oncol 2012, 30. abstr 7006.
- Lee, W.; Jiang, Z.; Liu, J.; Haverty, P.M.; Guan, Y.; Stinson, J.; Yue, P.; Zhang, Y.; Pant, K.P.; Bhatt, D.; et al. The mutation spectrum revealed by paired genome sequences from a lung cancer patient. Nature 2010, 465, 473–477.
- Ju, Y.S.; Lee, W.C.; Shin, J.Y.; Lee, S.; Bleazard, M.T.; Won, J.K.; Kim, Y.T.; Kim, J.I.; Kang, J.H.; Seo, J.S. A transforming KIF5B and RET gene fusion in lung adenocarcinoma revealed from whole-genome and transcriptome sequencing. Genome Res 2012, 22, 436–445.
- Belvedere, O.; Berri, S.; Chalkley, R.; Conway, C.; Barbone, F.; Pisa, F.; MacLennan, K.; Daly, C.; Alsop, M.; Morgan, J.; et al. A computational index derived from whole-genome copy number analysis is a novel tool for prognosis in early stage lung squamous cell carcinoma. Genomics 2012, 99, 18–24.
- Tran, B.; Dancey, J.E.; Kamel-Reid, S.; McPherson, J.D.; Bedard, P.L.; Brown, A.M.; Zhang, T.; Shaw, P.; Onetto, N.; Stein, L.; et al. Cancer genomics: Technology, discovery, and translation. J. Clin. Oncol 2012, 30, 647–660.




|
Table 1. Summary of EGFR TKIs for NSCLC. |
| Agent | Molecular properties | Approved status | Company |
|---|---|---|---|
| First-generation | |||
| Iressa/gefitinib | EGFR | Marketed in over 64 countries. It is a third-line treatment of NSCLC, after platinum-and docetaxel-based chemotherapy failed. | AstraZeneca |
| Tarceva/erlotinib | EGFR | Approved by several agencies, including U.S. Food and Drug Administration and European Medicines Agency, as second- and third-line treatment of NSCLC after platinum-based chemotherapy failed. | OSI/Roche/Genentech |
| Icotinib | EGFR | State Food and Drug Administration of China approved for the treatment of patients with advanced stage NSCLC. | Zhejiang Beta Pharma |
| Second-generation | |||
| Afatinib/BIBW 2992 | EGFR/T790M | Phase III clinical trial | Boehringer-Ingelheim |
| Dacomitinib/PF299804 | Pan-EGFR/T790M | Phase III clinical trial | Pfizer |
| Neratinib/HKI-272 | EGFR, HER1, HER2 | Phase I/II clinical trial | Pfizer |
| AP26113 | EGFR/T790M, ALK | Phase I/II clinical trial | Ariad |
| Neratinib/HKI-272 | EGFR, HER2 | Phase II clinical trial | Wyeth |
| AV412 | EGFR, HER2 | Phase I clinical trial | AVEO Pharmaceuticals |
| Lapatinib | EGFR, HER2 | Phase III clinical trial | GSK |
| Multiple signal transduction pathway inhibitors | |||
| XL647 | EGFR, HER2, VEGFR | Phase II clinical trial | Exelixis |
| Vandetanib/caprelsa | EGFR, VEGFR2 | Phase III clinical trial | AstraZeneca |
| BMS-690514 | Pan-EGFR, VEGFR | Phase II clinical trial | Bristol-Myers Squibb |
EGFR: epidermal growth factor receptor; NSCLC: non-small cell lung cancer; TKIs: tyrosine kinase inhibitors.

|
Table 2. Summary of ALK inhibitors for NSCLC. |
| Agent | Molecular properties | Approved status | Company |
|---|---|---|---|
| Xalkori/crizotinib | c-MET, ALK | Approved by FDA for patients with late-stage NSCLC carrying positive ALK. | Pfizer |
| AP26113 | EGFR/T790M, ALK | Phase I/II clinical trial | Ariad |
| LDK378 | ALK | Phase I clinical trial | Novartis |
| AF802/CH5424802 | ALK | Phase I/II trial | Chugai |
| ASP3026 | ALK | Phase I clinical trial | Astrella |
| X-396 | ALK | Pre-clinic | Xcovery |
| GSK-1838705A | ALK | Pre-clinic | GSK |
| NMS-E628 | ALK | Pre-clinic | - |
ALK: anaplastic lymphoma kinase; EGFR: epidermal growth factor receptor; NSCLC: non-small cell lung cancer.

|
Table 3. The clinical trials in selected patients carrying EGFR mutation. |
| Author | Description | Drug | Patients (No. of patients) | End point | Results |
|---|---|---|---|---|---|
| Compare EGFR tyrosine kinase inhibitors (TKIs) with chemotherapy in EGFR mutant patients | |||||
| Rosell et al. [76] | Phase III (EURTAC) | Erlotinib vs. cisplatin/docetaxel (or gemcitabine) | EGFR+ (174) | PFS | PFS was 9.7 months (erlotinib) and 5.2 months (chemotherapy) (HR 0.37, p < 0.0001), the RR was 64% (erlotinib) and 18% (chemotherapy). |
| Zhou et al. [77] | Phase III (OPTIMAL) | Erlotinib vs. gemcitabine/carboplatin (GC) | EGFR+ (154) | PFS | PFS was 13.1 months (erlotinib) and 4.6 months (GC) (HR 0.16, p < 0.0001). |
| Mitsudomi et al. [78] | Phase III (WJTOG3405) | Gefitinib vs. cisplatin/docetaxel (CD) | EGFR+ (177) | PFS | PFS was 9.2 months (gefitinib) and 6.3 months (CD) (p < 0.0001). |
| Yang et al. [79] | Phase III (LUX-Lung 3) | Afatinib vs. pemetrexed/cisplatin | EGFR+ (345) | PFS | Prolonged PFS was found in afatinib group (11.1 vs. 6.9 months, HR 0.58, p = 0.0004). |
| Inoue et al. [80] | Phase III (NEJ002) | Gefitinib vs. CBDCA+PTX (CP) | EGFR+ (228) | OS | OS was not significant different between gefitinib and CP groups. The median survival time and 2-year survival rate were 27.7 months, 57.9% (gefitinib), and 26.6 months, 53.7% (CP) (HR 0.887; p = 0.483). |
| EGFR TKIs was given alone in EGFR mutant patients: single arm design | |||||
| Kim et al. [81] | Phase II | First-line gefitinib | EGFR+ (45) | Objective RR | Objective RR: 53.3%; DCR: 86.7%, the median PFS: 398 days; median OS: 819 days. |
| Tamura et al. [82] | Phase II (WJTOG0403) | First-line gefitinib | EGFR+ (28) | RR | The overall RR was 75%, the DCR was 96% and the median PFS was 11.5 months. |
| Sequist et al. [83] | Phase II | First-line gefitinib | EGFR+ (31) | RR | The RR was 55% and median PFS was 9.2 months. |
| Inoue et al. [84] | Phase II | First-line gefitinib | EGFR+ (22) | Overall RR | The overall RR was 66%, and the DCR was 90%, PS improvement rate was 79% (p < 0.00005), and 68% improved from ≥ PS 3 at baseline to ≤ PS 1. The median PFS, median survival time, and 1-year survival rate were 6.5 months, 17.8 months, and 63%. |
| Sugio et al. [85] | Phase II | Gefitinib monotherapy | EGFR+(19) | - | The overall RR, DCR, median PFS and median survival time were 63.2%, 89.5%, 7.1 months and 20 months. |
| Han et al. [86] | Phase II (ESTERN) | Erlotinib as neoadjuvant treatment | EGFR+ (5) | Radical resection rate | One male patient with stable disease after neoadjuvant treatment got right upper lobe resection. |
| Yang et al. [87] | Phase II (LUX-Lung 2) | First- or second-line afatinib | EGFR+ (129) | Objective RR | The objective RR was 66%. |
| Kris et al. [88] | Phase II | First-line dacomitinib (PF-00299804) | EGFR+ (47) or patients with adenocarcinoma, no prior systemic tx, had smoked <10 pack years. | PFS; PR | In the patients with mutant EGFR, PR rate was 74%. Preliminary PFS was 96% (4 months) and 77% (1 year). Preliminary median PFS was 17 months. |
| Biomarker-based treatment selection: EGFR TKIs in the patients with positive mutation; chemotherapy in wild type patients | |||||
| Zhong et al. [89] | Phase II (LUX-Lung 2) | EGFR+ in one arm given erlotinib, EGFR− in another arm given GC | EGFR+ (24) | RR | The RR were 58% for the erlotinib arm and 33% for the GC arm (p = 0.49), the RRs were 17% for the erlotinib arm and 25% for the GC arm (p = 0.64). |
| Rosell et al. [90] | Phase II | EGFR+ were given erlotinib, and those with wild type EGFR received chemotherapy with or without cisplatin | EGFR+ (123) | - | Median survival exceeded 28 months for 12 patients with EGFR mutations, and 9–11 months for the patients with wild type EGFR. Two-year survival was 73.3% and 0%–41.2%, respectively. |
| Second-generation EGFR TKIs in patients with resistance | |||||
| Pietanza et al. [91] | Phase II | XL647 | 41 patients with relapsed or recurrent advanced NSCLC who progressed after ≥ 12 weeks of stable disease or response to erlotinib or gefitinib and/or those patients with a documented EGFR T790M | Objective RR | The objective RR was 3%, 67% of the patients harbored T790M had progression of disease, while14% of those without this mutation, 11 patients (28%) had a dose reduction due to toxicity. |
| Sequist et al. [92] | Phase II | Neratinib | 167 patients with ≥ 12 weeks of prior TKI therapy of EGFR TKIs | Objective RR | The objective RR was 3% in EGFR mutant patients, 0% in the other patients. |
DCR: disease control rate; EGFR: epidermal growth factor receptor; OS: overall survival; PFS: progression free survival; PS: performance score; PR: partial response; RR: response rate.

|
Table 4. The clinical trials in selected patients carrying wild type EGFR. |
| Author | Prescription | Drug and study design | Selection of patients | End point | Results |
|---|---|---|---|---|---|
| Kobayashi et al. [95] | Phase II | Erlotinib monotherapy | EGFR− (31) | DCR and PFS | The RR, DCR, median PFS, and survival times were 17.2%, 44.8%, 2.1 months and 7.7 months, respectively. |
| Matsuura et al. [96] | Phase II | Erlotinib monotherapy | EGFR− (20) | - | Overall RR was 15% and a DCR was 55%, median PFS and OS were 2.1 and 6.7 months, respectively. |
| Yoshioka et al. [97] | Phase II | Erlotinib monotherapy | EGFR− (30) | Object RR | Object RR was 3.3%, and the disease became stable in 18 patients (60%), the median survival time and median PFS were 9.2 and 2.1 months, respectively. |
| Garassino et al. [98] | Phase III (TAILOR | Erlotinib vs. docetaxel | EGFR− (211) | PFS | PFS was significant higher in docetaxel therapy (HR 0.70, p = 0.016) over erlotinib regimen. |
| Metro et al. [99] | - | Erlotinib or gefitinib monotherapy | EGFR− (67) | PFS; OS | Median PFS and OS were 2.9 months and 18.0 months, respectively. KRAS mutant patients had significantly shorter PFS (1.6 months) than KRAS wild type patients (3.0 months) (p = 0.04). |
DCR: disease control rate; EGFR: epidermal growth factor receptor; HR: hazard ratio; OS: overall survival; PFS: progression free survival; PR: partial response; RR: response rate.
© 2012 by the authors; licensee Molecular Diversity Preservation International, Basel, Switzerland. This article is an open-access article distributed under the terms and conditions of the Creative Commons Attribution license (http://creativecommons.org/licenses/by/3.0/).
Share and Cite
Wu, K.; House, L.; Liu, W.; Cho, W.C.S. Personalized Targeted Therapy for Lung Cancer. Int. J. Mol. Sci. 2012, 13, 11471-11496. https://doi.org/10.3390/ijms130911471
Wu K, House L, Liu W, Cho WCS. Personalized Targeted Therapy for Lung Cancer. International Journal of Molecular Sciences. 2012; 13(9):11471-11496. https://doi.org/10.3390/ijms130911471
Chicago/Turabian StyleWu, Kehua, Larry House, Wanqing Liu, and William C.S. Cho. 2012. "Personalized Targeted Therapy for Lung Cancer" International Journal of Molecular Sciences 13, no. 9: 11471-11496. https://doi.org/10.3390/ijms130911471
APA StyleWu, K., House, L., Liu, W., & Cho, W. C. S. (2012). Personalized Targeted Therapy for Lung Cancer. International Journal of Molecular Sciences, 13(9), 11471-11496. https://doi.org/10.3390/ijms130911471





