Expression Patterns, Activities and Carbohydrate-Metabolizing Regulation of Sucrose Phosphate Synthase, Sucrose Synthase and Neutral Invertase in Pineapple Fruit during Development and Ripening
Abstract
:1. Introduction
2. Results and Discussion
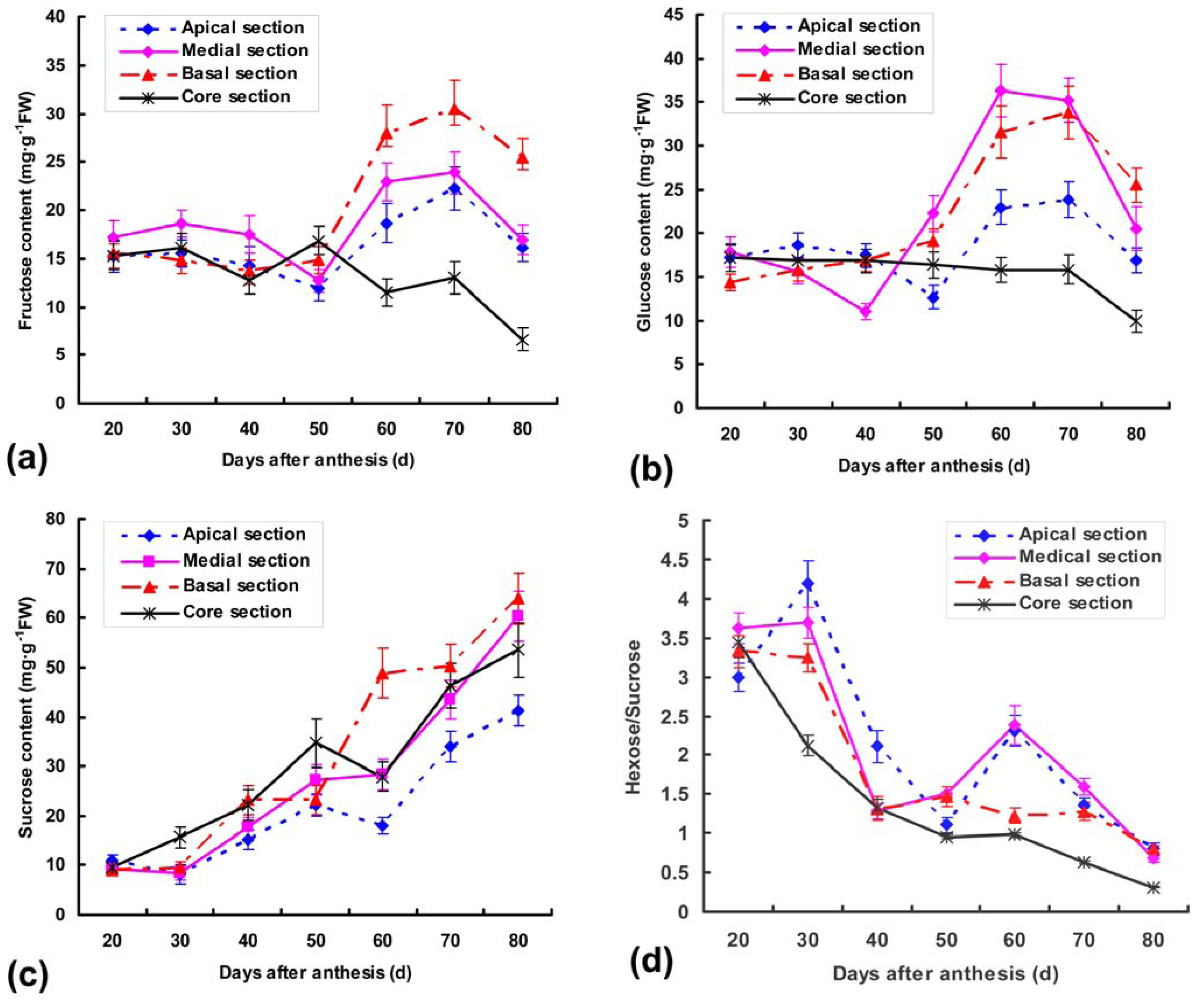
2.1. Soluble sugar Contents
2.2. Isolation of Ac-Sps, Ac-Ni and Ac-Susy Genes and Sequence Analysis
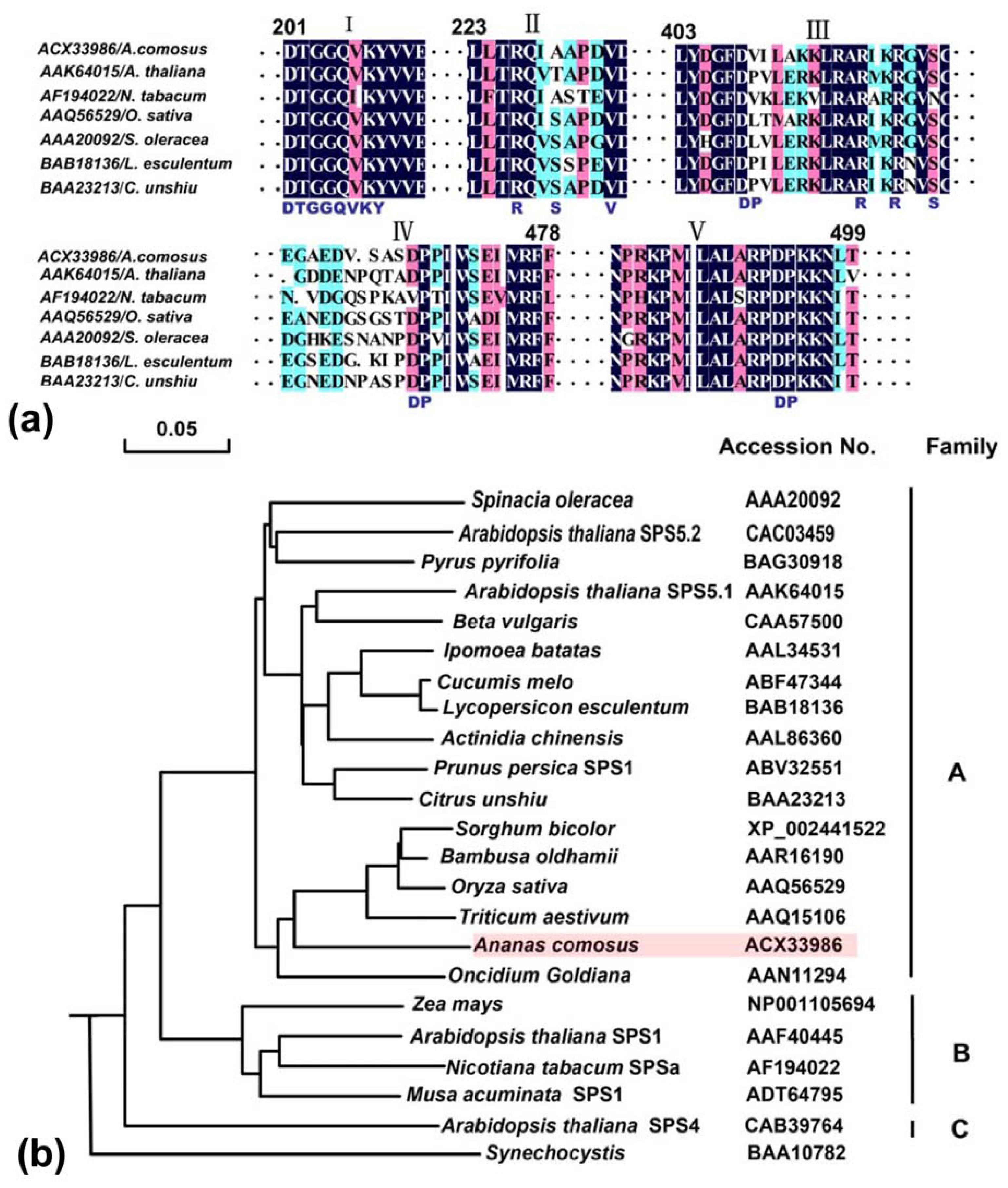
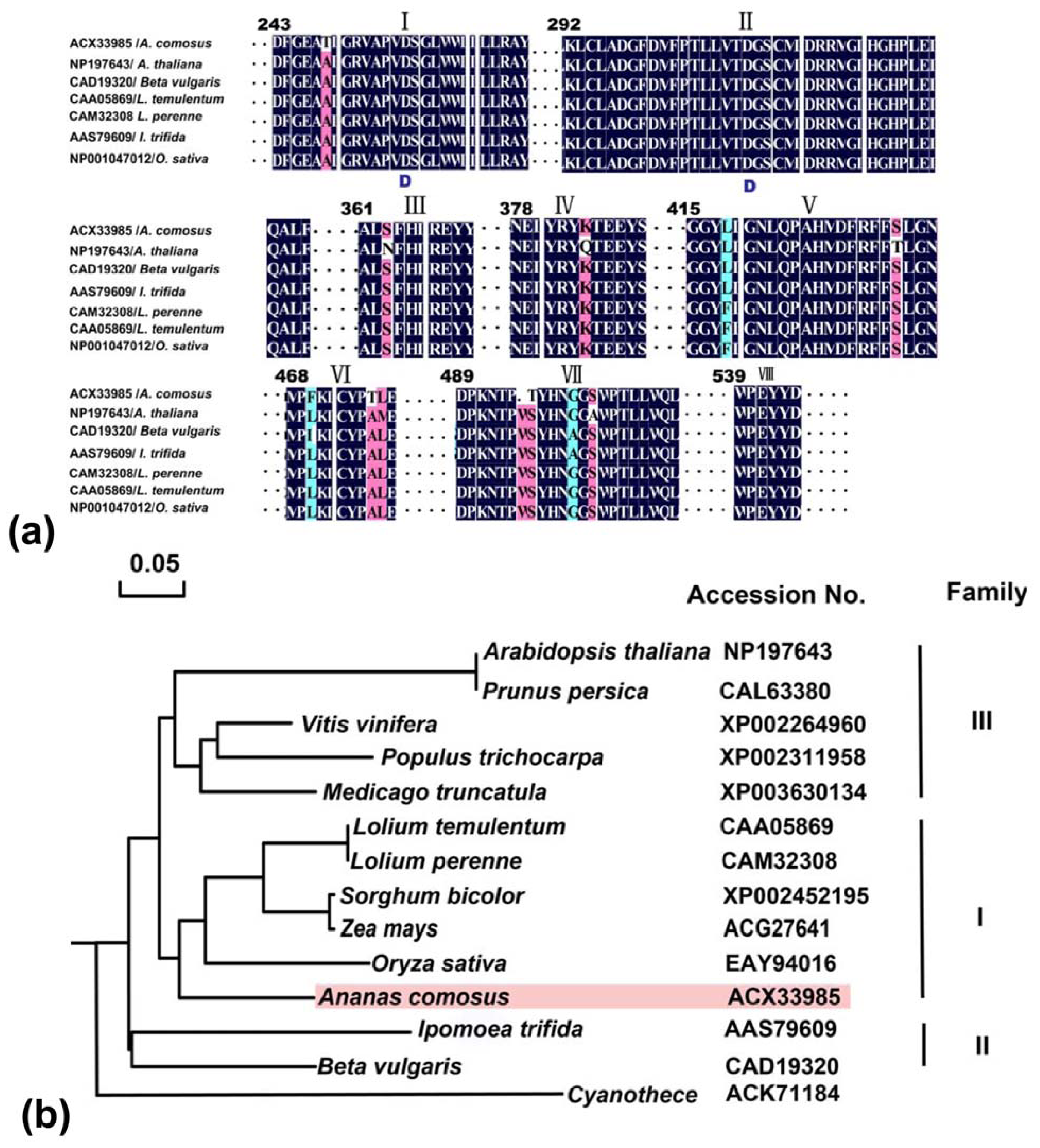
2.2.1. Multiple Alignments of Three Genes
2.2.2. Domain Characteristics and Phylogenetic Analysis of Ac-SPS
2.2.3. Domain Characteristics and Phylogenetic Analysis of Ac-NI
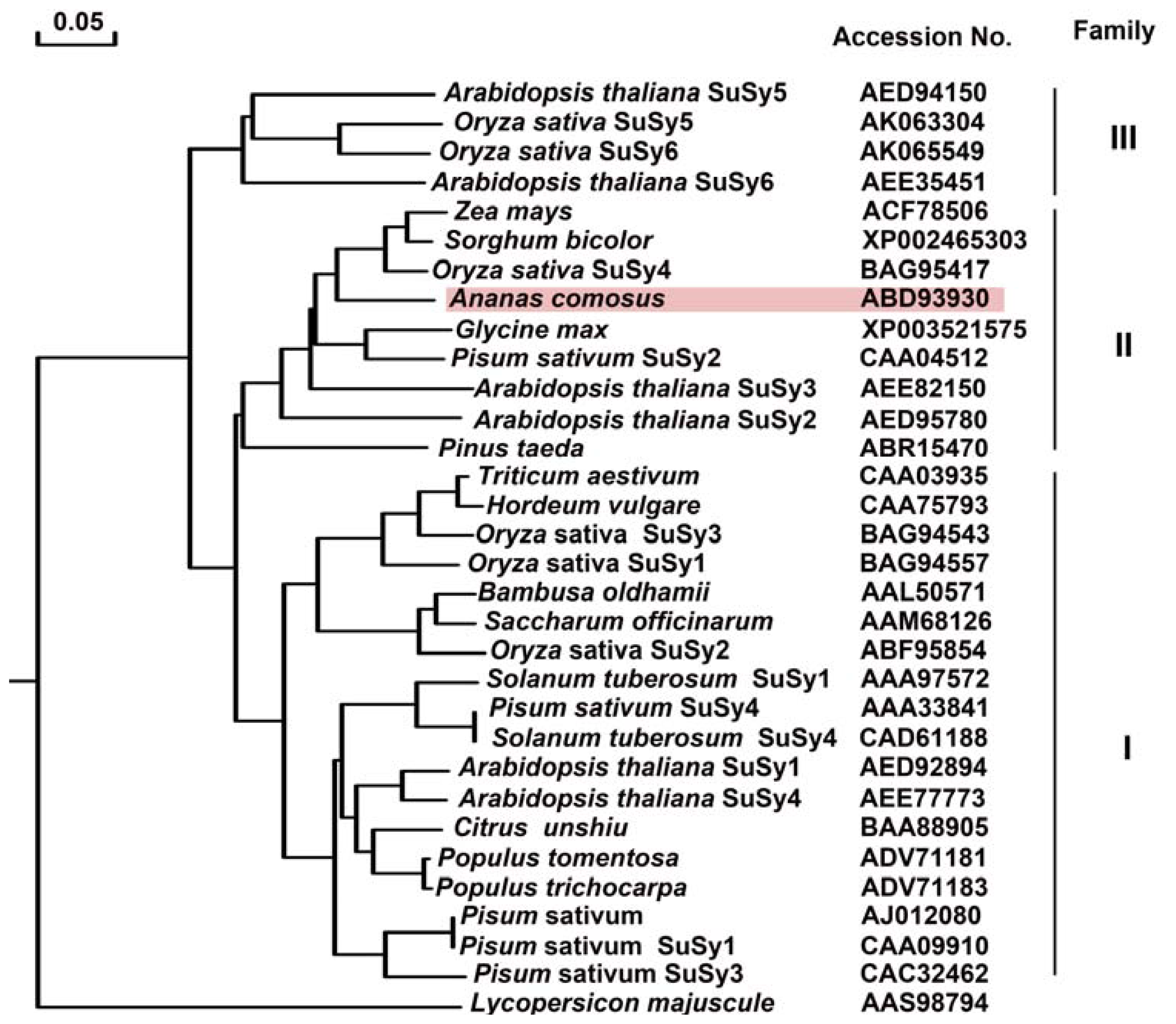
2.2.4. Phylogenetic Tree Analysis of Ac-SuSy
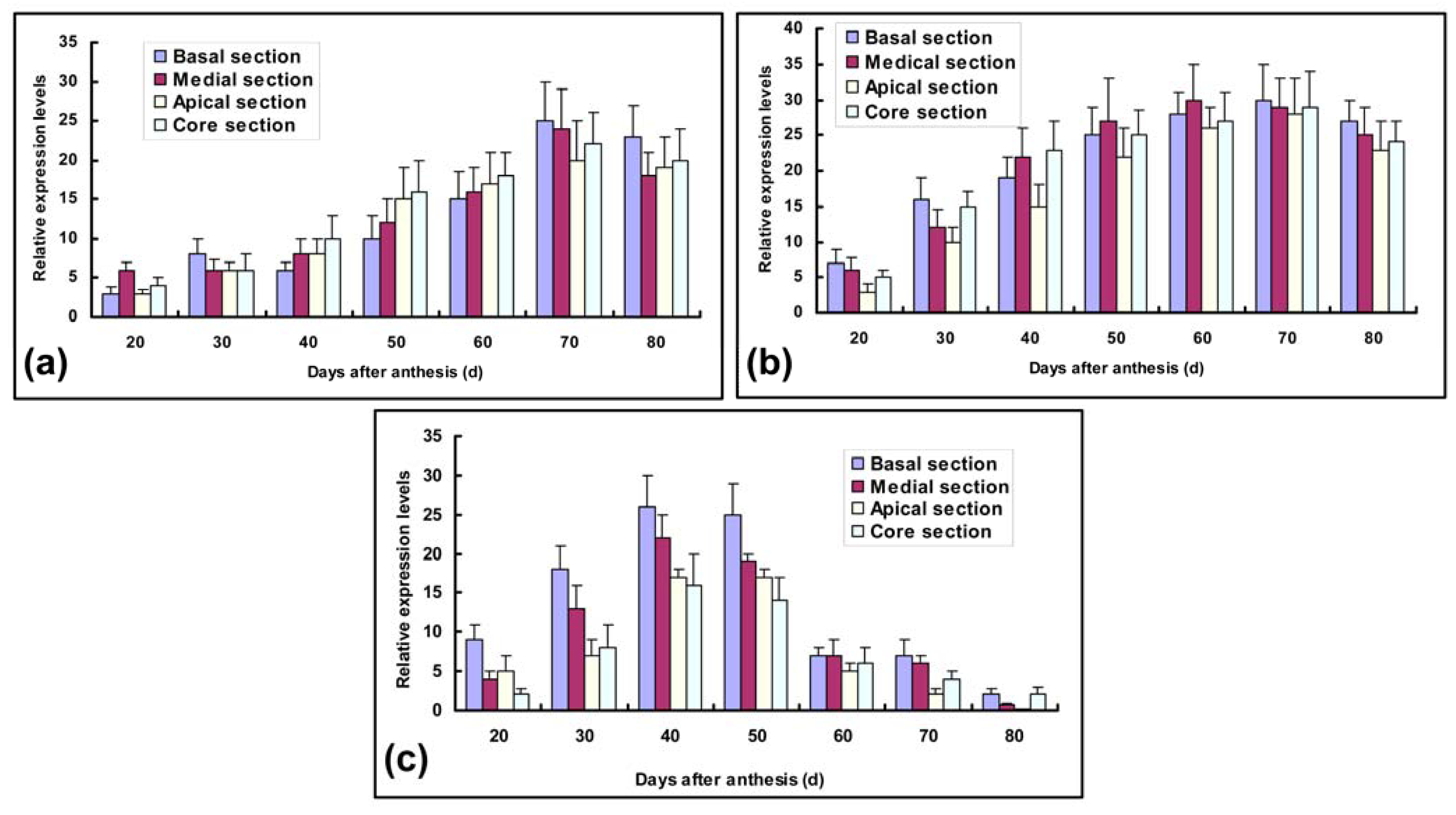
2.3. Transcript Expression Analysis of Ac-sps, Ac-susy and Ac-ni genes
2.4. Detection of Ac-SPS, Ac-SuSy and Ac-NI Activities
3. Experimental Section
3.1. Plant Materials
3.2. Plant Materials Determination of Sugar Content and Composition
3.3. Cloning and Expression Analysis of Ac-sps, Ac-susy and Ac-ni
3.3.1. Isolation of Total RNA and Sequence Amplification
3.3.2. Sequence Comparisons
3.3.3. Real-Time PCR Analysis of Gene Expressions
3.4. The Extraction and Activity Analysis of NI, SPS and SuSy
3.5. Statistical Analysis
4. Conclusions
Acknowledgments
- Conflict of InterestThe authors declare no conflict of interest.
References
- Itai, A.; Tanahashi, T. Inhibition of sucrose loss during cold storage in Japanese pear (Pyrus pyrifolia Nakai) by 1-MCP. Postharvest Biol. Technol 2008, 48, 355–363. [Google Scholar]
- Basson, C.E.; Groenewald, J.H.; Kossmann, J.; Cronjé, C.; Bauer, R. Sugar and acid-related quality attributes and enzyme activities in strawberry fruits: Invertase is the main sucrose hydrolysing enzyme. Food Chem 2010, 121, 1156–1162. [Google Scholar]
- Yativ, M.; Harary, I.; Wolf, S. Sucrose accumulation in watermelon fruits: genetic variation and biochemical analysis. J. Plant Physiol 2009, 167, 589–596. [Google Scholar]
- Winter, H.; Huber, S.C. Regulation of sucrose metabolism in higher plants: localization and regulation of activity of key enzymes. Crit. Rev. Biochem. Mol. Biol 2000, 35, 253–289. [Google Scholar]
- Li, M.J.; Feng, F.J.; Cheng, L.L. Expression patterns of genes involved in sugar metabolism and accumulation during apple fruit development. PLoS One 2012, 7, e33055. [Google Scholar]
- Trevanion, S.J.; Castleden, C.K.; Foyer, C.H.; Furbank, R.T.; Quick, W.P.; Lunn, J.E. Regulation of sucrose-phosphate synthase in wheat (Triticum aestivum) leaves. Funct. Plant Biol 2004, 31, 685–695. [Google Scholar]
- Zhang, X.M.; Du, L.Q.; Xie, J.H.; Dou, M.A.; Wang, W.; Mo, Y.W.; Sun, G.M. Cloning and expression of pineapple sucrose phosphate synthase gene during fruit development. Afr. J. Biotechnol. 2010, 9, 8296–8303. [Google Scholar]
- Koch, K.E. Sucrose metabolism: Regulatory mechanisms and pivotal roles in sugar sensing and plant development. Curr. Opin. Plant Biol 2004, 7, 235–246. [Google Scholar]
- Verma, A.K.; Upadhyay, S.K.; Verma, P.C.; Solomon, S.; Singh, S.B. Functional analysis of sucrose phosphate synthase (SPS) and sucrose synthase (SS) in sugarcane (Saccharum) cultivars. Plant Biol 2011, 13, 325–332. [Google Scholar]
- Carlson, S.J.; Chourey, P.S.; Helentjaris, T.; Datta, R. Gene-expression studies on developing kernels of maize sucrose synthase (SuSy) mutants show evidence for a third SuSy gene. Plant Mol. Biol 2002, 49, 15–29. [Google Scholar]
- Roitsch, T.; González, M.C. Function and regulation of plant invertases: sweet sensations. Trends Plant Sci 2004, 9, 606–613. [Google Scholar]
- Keutgen, A.J.; Pawelzik, E. Quality and nutritional value of strawberry fruit under long term salt stress. Food Chem 2008, 107, 1413–1420. [Google Scholar]
- Zhang, M.F.; Li, Z.L. A comparison of sugar-accumulating patterns and relative compositions in developing fruits of two oriental melon varieties as determined by HPLC. Food Chem 2005, 90, 785–790. [Google Scholar]
- Qi, X.P.; Wu, Z.C.; Li, J.H.; Mo, X.R.; Wu, S.H.; Chu, J.; Wu, P. AtCYT-INV1, a neutral invertase, is involved in osmotic stress-induced inhibition on lateral root growth in Arabidopsis. Plant Mol. Biol 2007, 64, 575–587. [Google Scholar]
- Zhang, X.M.; Dou, M.A.; Yao, Y.L.; Du, L.Q.; Li, J.G.; Sun, G.M. Dynamic analysis of sugar metabolism in different harvest seasons of pineapple (Ananas comosus L. (Merr.)). Afr. J. Biotechnol 2010, 10, 2716–2723. [Google Scholar]
- Pressman, E.; Shaked, R.; Shen, S.; Altahan, L.; Firon, N. Variations in carbohydrate content and sucrose-metabolizing enzymes in tomato (Solanum lycopersicum L.) stamen parts during pollen maturation. Am. J. Plant. Sci 2012, 3, 252–260. [Google Scholar]
- Nguyen-Quoc, B.; Foyer, C.H. A role for “futile cycles” involving invertase and sucrose synthase in sucrose metabolism of tomato fruit. J. Exp. Bot 2001, 52, 881–889. [Google Scholar]
- Castleden, C.K.; Aoki, N.; Gillespie, V.J.; MacRae, E.A.; Quick, W.P.; Buchner, P.; Foyer, C.H.; Furbank, R.T.; Lunn, J.E. Evolution and function of the sucrose phosphate synthase gene families in wheat and other grasses. Plant Physiol 2004, 135, 1753–1764. [Google Scholar]
- Langenkämper, G.; Fung, R.W.M.; Newcomb, R.D.; Atkinson, R.G.; Gardner, R.C.; MacRae, E.A. Sucrose phosphate synthase genes in plants belong to three different families. J. Mol. Evol 2002, 54, 322–332. [Google Scholar]
- Mitchell-Olds, T.; Clauss, M.J. Plant evolutionary genomics. Curr. Opin. Plant Biol 2002, 5, 74–79. [Google Scholar]
- Lunn, J.E. Evolution of sucrose synthesis. Plant Physiol 2002, 128, 1490–1500. [Google Scholar]
- Murayama, S.; Handa, H. Genes for alkaline/neutral invertase in rice: Alkaline/neutral invertases are located in plant mitochondria and also in plastids. Planta 2007, 225, 1193–1203. [Google Scholar]
- Vargas, W.; Andrea, C.; Salerno, G.L. Cyanobacterial alkaline/neutral invertases. Origin of sucrose hydrolysis in the plant cytosol? Planta 2003, 216, 951–960. [Google Scholar]
- Botha, F.C.; Black, K.G. Sucrose phosphate synthase and sucrose synthase activity during maturation of internodal tissue in sugarcane. Funct. Plant Biol 2000, 27, 81–85. [Google Scholar]
- Choudhury, S.R.; Roy, S.; Das, R.; Sengupta, D.N. Differential transcriptional regulation of banana sucrose phosphate synthase gene in response to ethylene, auxin, wounding, low temperature and different photoperiods during fruit ripening and functional analysis of banana SPS gene promoter. Planta 2008, 229, 207–223. [Google Scholar]
- Strand, A.; Zrenner, R.; Trevanion, S.; Stitt, M; Gustafsson, P.; Gardeström1, P. Decreased expression of two key enzymes in the sucrose biosynthesis pathway, cytosolic fructose-1,6-bisphosphatase and sucrose phosphate synthase, has remarkably different consequences for photosynthetic carbon metabolism in transgenic Arabidopsis thaliana. Plant J 2000, 23, 759–770. [Google Scholar]
- Itai, A.; Tanahashi, T. Inhibition of sucrose loss during cold storage in Japanese pear (Pyrus pyrifolia Nakai) by 1-MCP. Postharvest Biol. Technol 2008, 48, 355–363. [Google Scholar]
- Ciereszko, I.; Kleczkowski, L.A. Glucose and mannose regulate the expression of a major sucrose synthase gene in Arabidopsis via hexokinase-dependent mechanisms. Plant Physiol. Biochem 2002, 40, 907–911. [Google Scholar]
- Nonis, A.; Ruperti, B.; Falchi, R.; Casatta, E.; Enferadi, S.T.; Vizzotto, G. Differential expression and regulation of a neutral invertase encoding gene from peach (Prunus persica): Evidence for a role in fruit development. Physiol. Plantarum 2007, 129, 436–446. [Google Scholar]
- Yu, X.Y.; Wang, X.F.; Zhang, W.Q.; Qian, T.T.; Tang, G.M.; Guo, Y.K; Zheng, C.C. Antisense suppression of an acid invertase gene (MAI1) in muskmelon alters plant growth and fruit development. J. Exp. Bot. 2008, 59, 2969–2977. [Google Scholar]
- Basson, C.E.; Groenewald, J.H.; Kossmann, J.; Cronjé, C.; Bauer, R. Sugar and acid-related quality attributes and enzyme activities in strawberry fruits: Invertase is the main sucrose hydrolysing enzyme. Food Chem 2009, 121, 1156–1162. [Google Scholar]
- Kleczkowski, L.A.; Kunz, S.; Wilczynska, M. Mechanisms of UDP-Glucose synthesis in plants. Crit. Rev. Plant Sci 2010, 29, 191–203. [Google Scholar]
- Blast. Available online: http://blast.ncbi.nlm.nih.gov/Blast.cgi accessed on 12 February 2012.
- EMBL-EBI. Available online: http://www.ebi.ac.uk/Tools/msa/clustalw2/ accessed on 12 February 2012.


| Primer name | Forward and reverse primes (5′-3′) | Tm (°C) |
|---|---|---|
| Ac-sps degenerate primers | GGA(G/A)CTTGG(T/A/G/C)CG(T/G)GATTCTGATA CAGGAAC(A/T/C)TC(A/T)GA(C/T)TG(C/T)TTGTG | 52 |
| Ac-sps specific primers | GCTACGGCGAGCCAACCGAGAT GTGGCCAAGAGCGCCGTCAAC | 60 |
| Ac-susy specific primers 1 | GGATGACCGGTCTAAGCCCATCAT CCATTTCACGCAGTTTGGCGCAT | 60 |
| Ac-ni degenerate primers | CAG(T/C)T(T/G/A)CAGAG(C/T)TGGGAGA GTGTC(A/G)TA(A/C)TA(T/C)TCAGGCCA | 55 |
| Ac-ni specific primers | GCTCTGGGGATTTGTCAGTTCAG CGATCAATCATGCACGAGCCGT | 60 |
| 18S rRNA specific primers 2 | GCCGGCGACGCATCATTCAA CGCGCCTGCTGCCTTCCTTG | 60 |
© 2012 by the authors; licensee Molecular Diversity Preservation International, Basel, Switzerland. This article is an open-access article distributed under the terms and conditions of the Creative Commons Attribution license (http://creativecommons.org/licenses/by/3.0/).
Share and Cite
Zhang, X.-M.; Wang, W.; Du, L.-Q.; Xie, J.-H.; Yao, Y.-L.; Sun, G.-M. Expression Patterns, Activities and Carbohydrate-Metabolizing Regulation of Sucrose Phosphate Synthase, Sucrose Synthase and Neutral Invertase in Pineapple Fruit during Development and Ripening. Int. J. Mol. Sci. 2012, 13, 9460-9477. https://doi.org/10.3390/ijms13089460
Zhang X-M, Wang W, Du L-Q, Xie J-H, Yao Y-L, Sun G-M. Expression Patterns, Activities and Carbohydrate-Metabolizing Regulation of Sucrose Phosphate Synthase, Sucrose Synthase and Neutral Invertase in Pineapple Fruit during Development and Ripening. International Journal of Molecular Sciences. 2012; 13(8):9460-9477. https://doi.org/10.3390/ijms13089460
Chicago/Turabian StyleZhang, Xiu-Mei, Wei Wang, Li-Qing Du, Jiang-Hui Xie, Yan-Li Yao, and Guang-Ming Sun. 2012. "Expression Patterns, Activities and Carbohydrate-Metabolizing Regulation of Sucrose Phosphate Synthase, Sucrose Synthase and Neutral Invertase in Pineapple Fruit during Development and Ripening" International Journal of Molecular Sciences 13, no. 8: 9460-9477. https://doi.org/10.3390/ijms13089460
APA StyleZhang, X.-M., Wang, W., Du, L.-Q., Xie, J.-H., Yao, Y.-L., & Sun, G.-M. (2012). Expression Patterns, Activities and Carbohydrate-Metabolizing Regulation of Sucrose Phosphate Synthase, Sucrose Synthase and Neutral Invertase in Pineapple Fruit during Development and Ripening. International Journal of Molecular Sciences, 13(8), 9460-9477. https://doi.org/10.3390/ijms13089460




