Optimization of Xylanase Production from Penicillium sp.WX-Z1 by a Two-Step Statistical Strategy: Plackett-Burman and Box-Behnken Experimental Design
Abstract
:1. Introduction
2. Results
2.1. Screening of Important Nitrogen Sources for Xylanase Production by Penicillium sp.WX-Z1
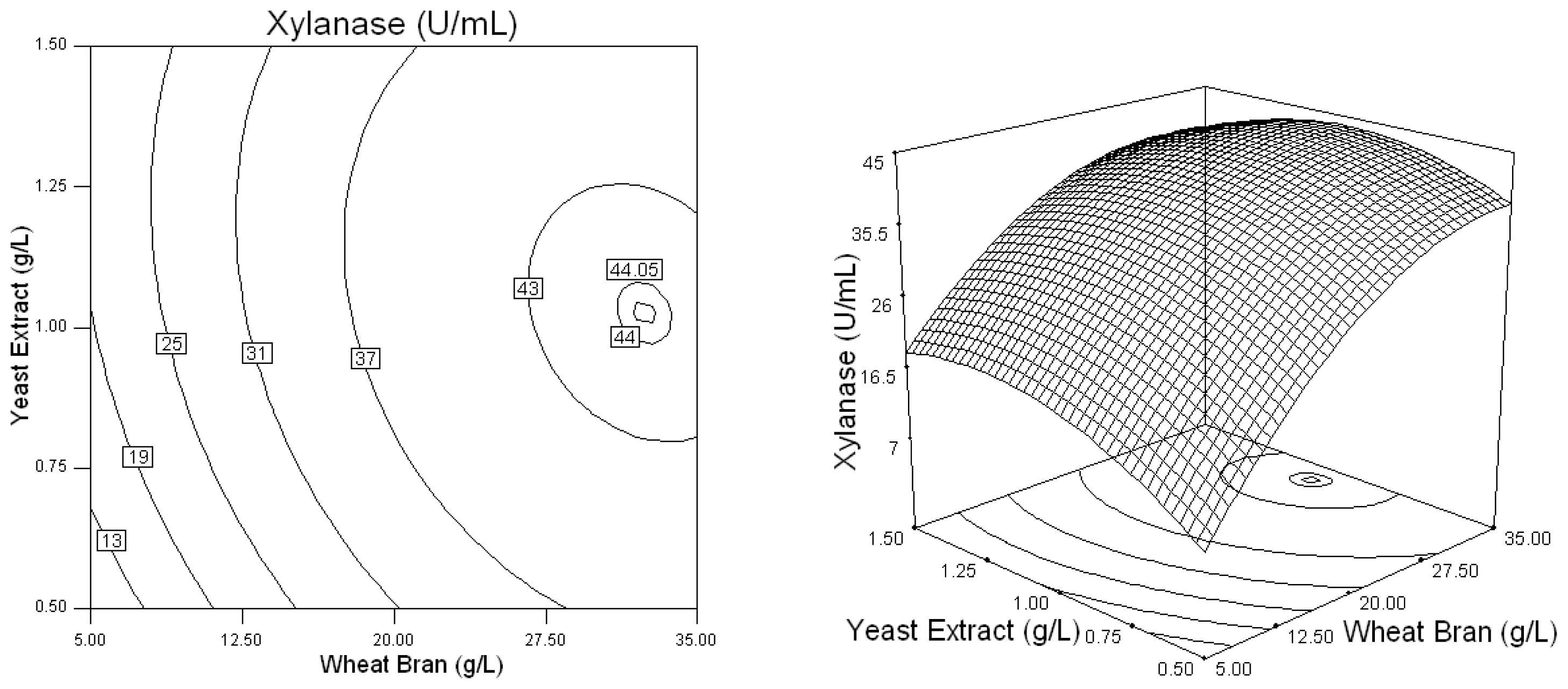
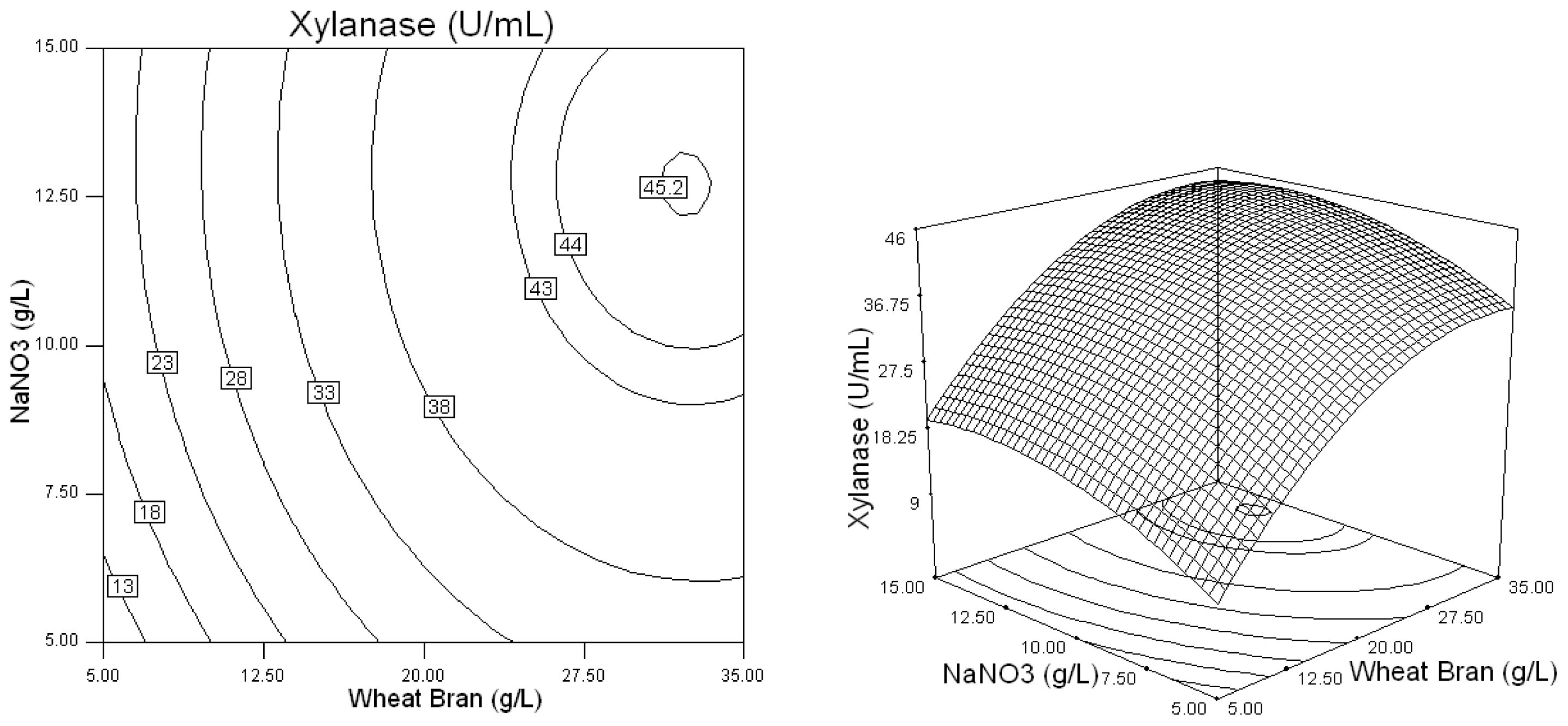
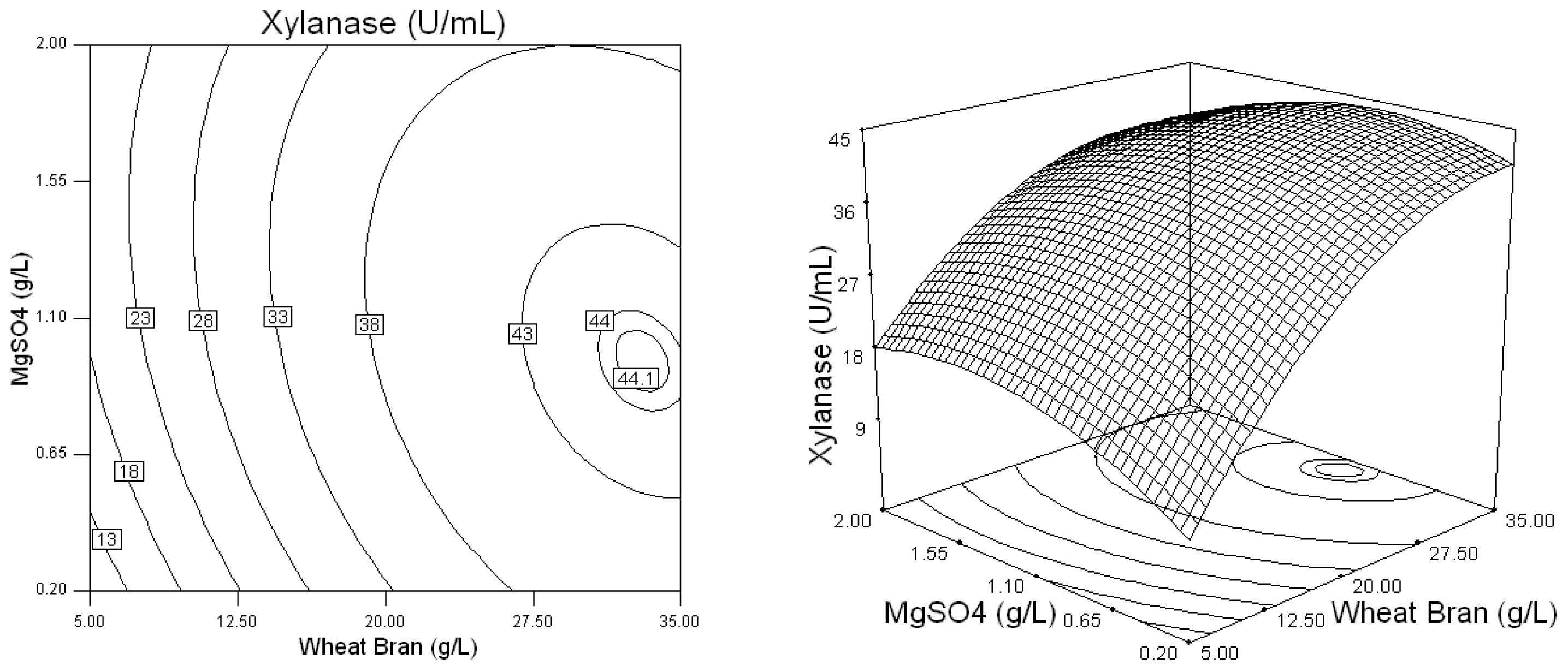
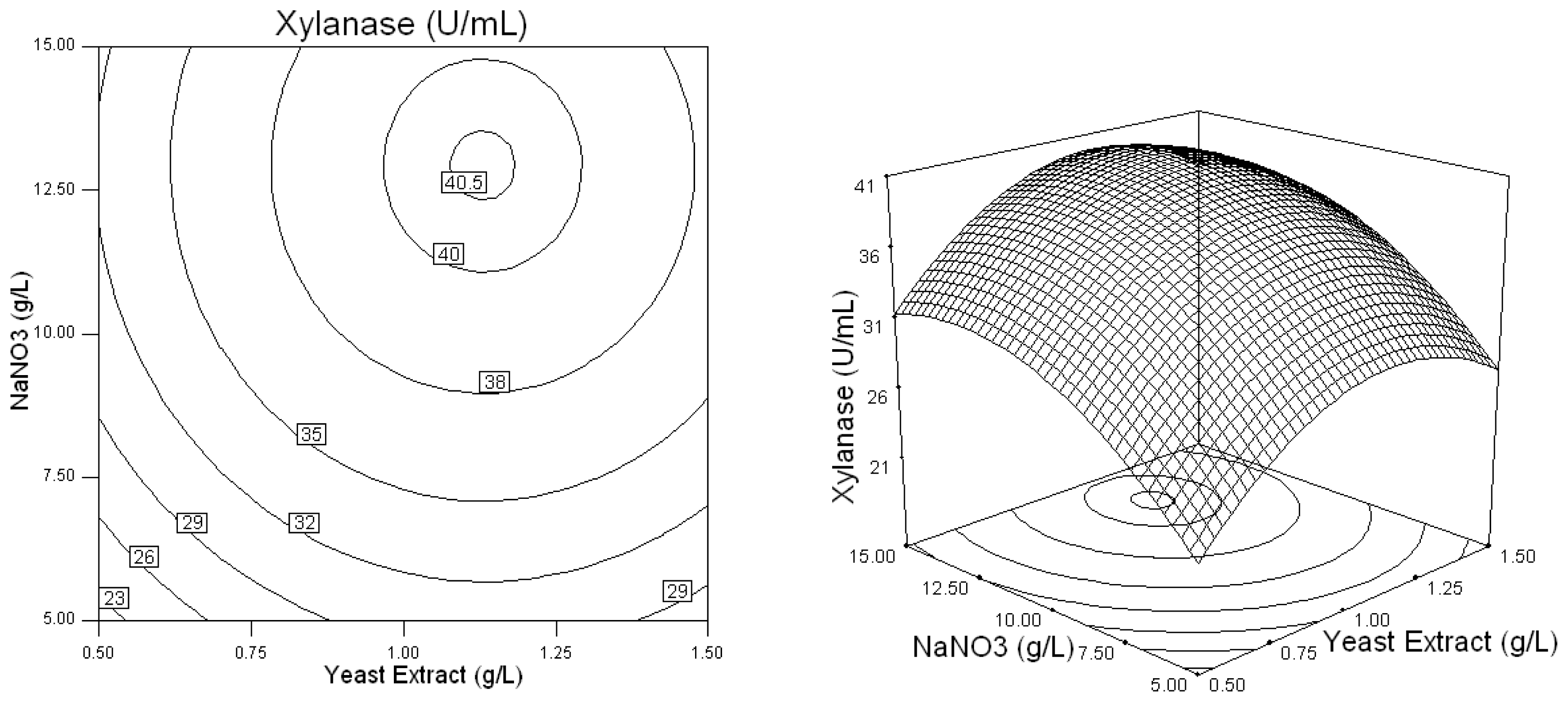
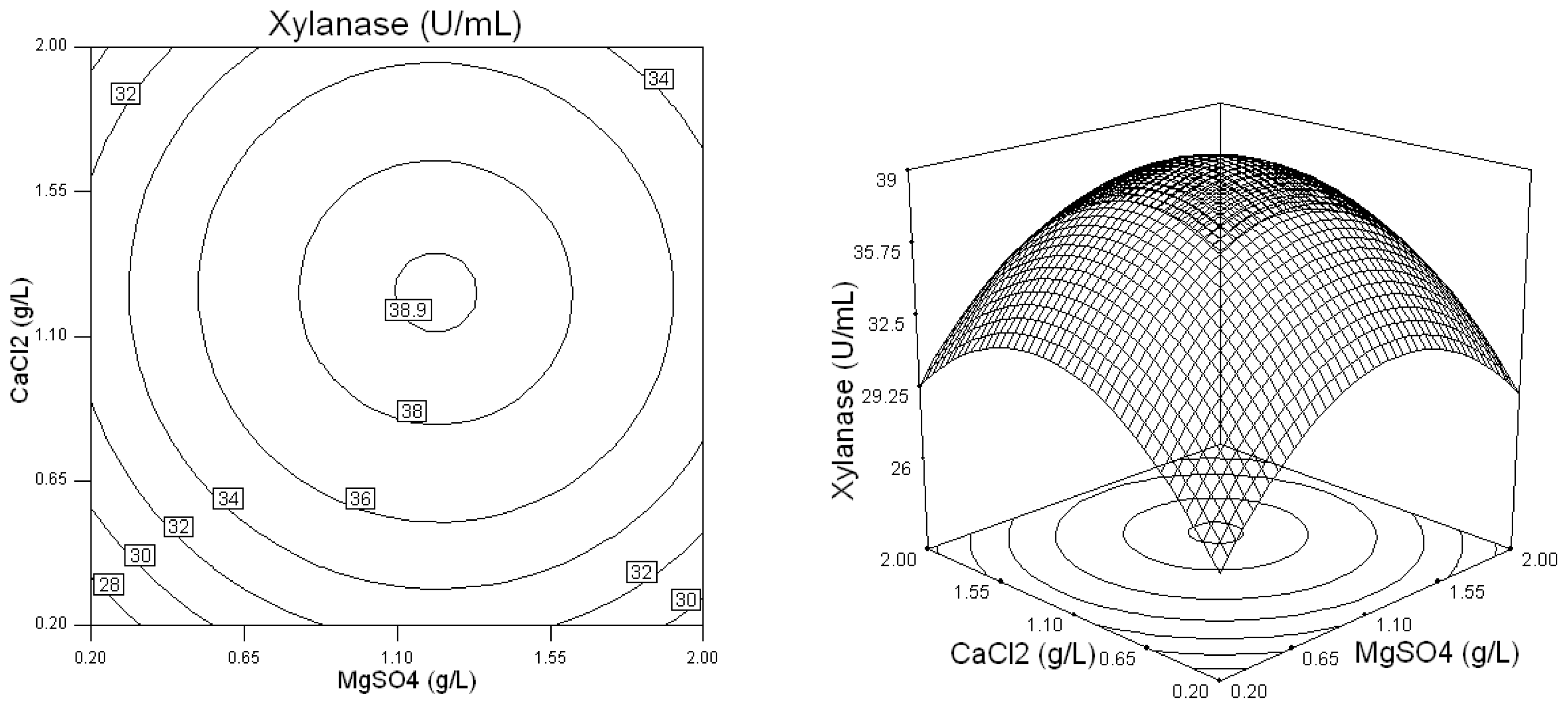
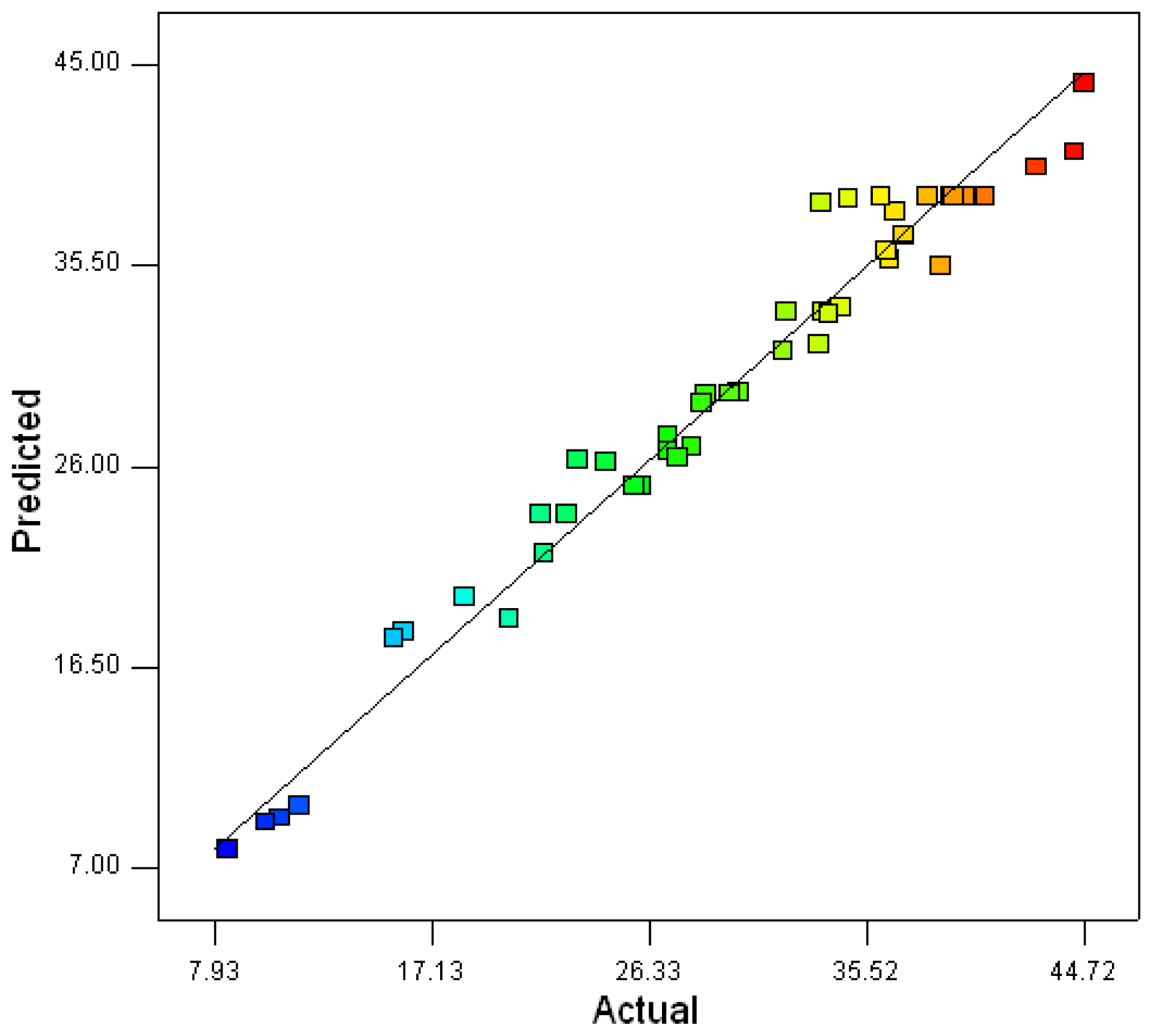
2.2. Optimization of Screened Nutrient Sources for Xylanase Production by Penicillium sp.WX-Z1 Using Response Surface Methodology
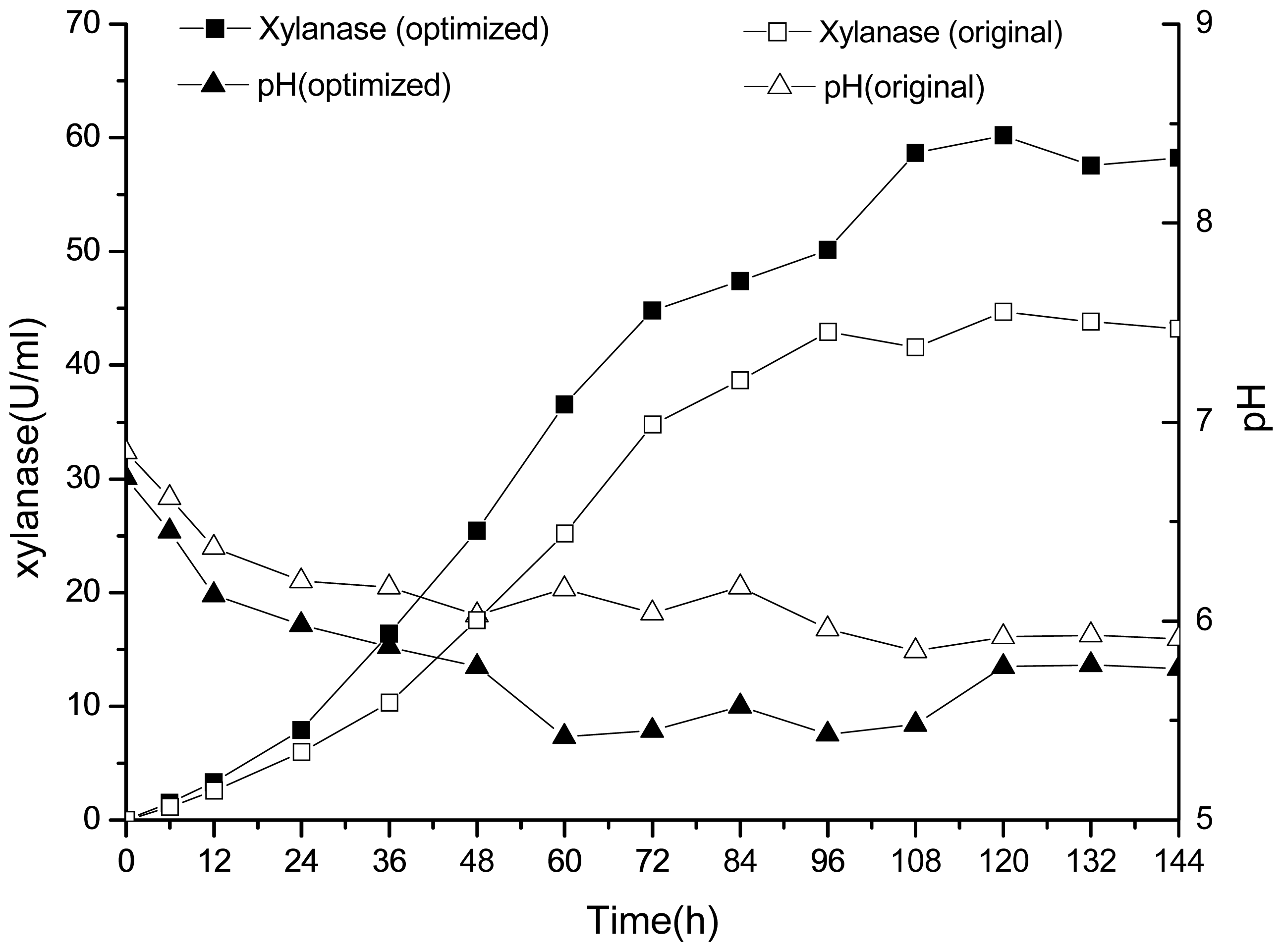
2.3. 30-L Bioreactor Fermentation Results
3. Discussion
4. Experimental Section
4.1. Microorganism
4.2. Medium and Cultivation
4.3. Optimization Procedure
4.3.1. Plackett-Burman Design for Identification of Important Nutrients
4.3.2. Box-Behnken Design
4.4. Bioreactor Cultivation Conditions
4.5. Analytical Method
5. Conclusions
Acknowledgments
- Conflict of Interest The authors declare that they have no conflicts of interest. Fengjie Cui and Liming Zhao conceived of the study, participated in its design and coordination, and drafted the manuscript.
References
- Diaz, A.B.; Bolivar, J.; de Ory, I.; Caro, I.; Blandino, A. Applicability of enzymatic extracts obtained by solid state fermentation on grape pomace and orange peels mixtures in must clarification. Lwt-Food Sci. Technol 2011, 44, 840–846. [Google Scholar]
- Hang, Y.D.; Woodams, E.E. Xylanolytic activity of commercial juice-processing enzyme preparations. Lett. Appl. Microbiol 1997, 24, 389–392. [Google Scholar]
- Bhushan, S.; Kalia, K.; Sharma, M.; Singh, B.; Ahuja, P.S. Processing of apple pomace for bioactive molecules. Crit. Rev. Biotechnol 2008, 28, 285–296. [Google Scholar]
- Das, M.; Gupta, S.; Kapoor, V.; Banerjee, R.; Bal, S. Enzymatic polishing of rice—A new processing technology. Lwt-Food Sci. Technol 2008, 41, 2079–2084. [Google Scholar]
- Biely, P. Microbial xylanolytic systems. Trends Biotechnol 1985, 3, 286–290. [Google Scholar]
- Viikari, L.; Kantelinen, A.; Sundquist, J.; Linko, M. Xylanases in bleaching: From an idea to the industry. FEMS Microbiol. Rev 1994, 13, 335–350. [Google Scholar]
- Daneault, C.; Leduc, C.; Valade, J.L. The use of xylanases in kraft pulp bleaching—A review. Tappi J 1994, 77, 125–131. [Google Scholar]
- Polizeli, M.L.T.M.; Rizzatti, A.C.S.; Monti, R.; Terenzi, H.F.; Jorge, J.A.; Amorim, D.S. Xylanases from fungi: Properties and industrial applications. Appl. Microbiol. Biotechnol 2005, 67, 577–591. [Google Scholar]
- Verma, D.; Kanagaraj, A.; Jin, S.X.; Singh, N.D.; Kolattukudy, P.E.; Daniell, H. Chloroplast-Derived enzyme cocktails hydrolyse lignocellulosic biomass and release fermentable sugars. Plant Biotechnol. J 2010, 8, 332–350. [Google Scholar]
- Wang, H.Y. Effects of rhamnolipid on the cellulase and xylanase in hydrolysis of wheat straw. Bioresour. Technol 2011, 102, 6515–6521. [Google Scholar]
- Sapag, A.; Wouters, J.; Lambert, C.; de Ioannes, P.; Eyzaguirre, J.; Depiereux, E. The endoxylanases from family 11: Computer analysis of protein sequences reveals important structural and phylogenetic relationships. J. Biotechnol 2002, 95, 109–131. [Google Scholar]
- Lemos, J.L.S.; Fontes, M.C.D.; Pereira, N. Xylanase production by Aspergillus awamori in solid-state fermentation and influence of different nitrogen sources. Appl. Biochem. Biotechnol 2001, 91, 681–689. [Google Scholar]
- Gupta, S.; Bhushan, B.; Hoondal, G.S. Enhanced production of xylanase from Staphylococcus sp SG-13 using amino acids. World J. Microb. Biotechnol 1999, 15, 511–512. [Google Scholar]
- Beg, Q.K.; Bhushan, B.; Kapoor, M.; Hoondal, G.S. Production and characterization of thermostable xylanase and pectinase from Streptomyces sp. QG-11-3. J. Ind. Microbiol. Biotechnol 2000, 24, 396–402. [Google Scholar]
- Plackett, R.L.; Burman, J.P. The design of optimum multifactorial experiments. Biometrika 1946, 37, 305–325. [Google Scholar]
- Salihu, A.; Alam, M.Z.; AbdulKarim, M.I.; Salleh, H.M. Optimization of lipase production by Candida cylindracea in palm oil mill effluent based medium using statistical experimental design. J. Mol. Catal. Enzym 2011, 69, 66–73. [Google Scholar]
- Cui, F.J.; Liu, Z.Q.; Li, Y.; Ping, L.F.; Ping, L.Y.; Zhang, Z.C.; Lin, L.; Dong, Y.; Huang, D.M. Production of mycelial biomass and exo-polymer by Hericium erinaceus CZ-2: Optimization of nutrients levels using response surface methodology. Biotechnol. Bioproc 2010, 15, 299–307. [Google Scholar]
- Periyasamy, A.K.; Muthu, M.; Kannan, V. Optimization of nutrients for the production of RNase by Bacillus firmus VKPACU1 using response surface methodology. Biotechnol. Bioprocess 2010, 14, 202–206. [Google Scholar]
- Chen, X.S.; Tang, L.; Li, S.; Liao, L.J.; Zhang, J.H.; Mao, Z.G. Optimization of medium for enhancement of epsilon-Poly-l-Lysine production by Streptomyces sp. M-Z18 with glycerol as carbon source. Bioresour. Technol 2011, 102, 1727–1732. [Google Scholar]
- Li, Y.; Cui, F.J.; Liu, Z.Q.; Xu, Y.Y.; Zhao, H. Improvement of xylanase production by Penicillium oxalicum ZH-30 using response surface methodology. Enzym. Microb. Technol 2007, 40, 1381–1388. [Google Scholar]
- Bocchini, D.A.; Alves-Prado, H.F.; Baida, L.C.; Roberto, I.C.; Gomes, E.; da Silva, R. Optimization of xylanase production by Bacillus circulans D1 in submerged fermentation using response surface methodology. Process Biochem 2002, 38, 727–731. [Google Scholar]
- Li, Y.; Lin, J.; Meng, D.J.; Lu, J.; Gu, G.X.; Mao, Z.G. Effect of pH, cultivation time and substrate concentration on the endoxylanase production by Aspergillus awamori ZH-26 under submerged fermentation using central composite rotary design. Food Technol. Biotechnol 2006, 44, 473–477. [Google Scholar]
- Liu, W.; Lu, Y.L.; Ma, G.R. Induction and glucose repression of endo-beta-xylanase in the yeast Trichosporon cutaneum SL409. Process Biochem 1999, 34, 67–72. [Google Scholar]
- Fernandezespinar, M.; Pinaga, F.; Degraaff, L.; Visser, J.; Ramon, D.; Valles, S. Purification, characterization and regulation of the synthesis of an Aspergillus-nidulans acidic xylanase. Appl. Microbiol. Biotechnol 1994, 42, 555–562. [Google Scholar]
- Kurzatkowski, W.; Torronen, A.; Filipek, J.; Mach, R.L.; Herzog, P.; Sowka, S.; Kubicek, C.P. Glucose-Induced secretion of Trichoderma reesei xylanases. Appl. Environ. Microb 1996, 62, 2859–2865. [Google Scholar]
- Kimura, T.; Ito, J.; Kawano, A.; Makino, T.; Kondo, H.; Karita, S.; Sakka, K.; Ohmiya, K. Purification, characterization, and molecular cloning of acidophilic xylanase from Penicillium sp. 40. Biosci. Biotechnol. Biochem 2000, 64, 1230–1237. [Google Scholar]
- Soundari, S.G.; Sashi, V.; Malathy, N.S. Effect of nitrogen sources, bivalent cations and agitation rate on-xylanase production. Plant Arch 2007, 7, 615–616. [Google Scholar]
- Senthilkumar, S.R.; Ashokkumar, B.; Raj, K.C.; Gunasekaran, P. Optimization of medium composition for alkali-stable xylanase production by Aspergillus fischeri Fxn 1 in solid-state fermentation using central composite rotary design. Bioresour. Technol 2005, 96, 1380–1386. [Google Scholar]
- Ding, C.H.; Jiang, Z.Q.; Li, X.T.; Li, L.T.; Kusakabe, I. High activity xylanase production by Streptomyces olivaceoviridis E-86. World J. Microb. Biotechnol 2004, 20, 7–10. [Google Scholar]
- Techapun, C.; Sinsuwongwat, S.; Watanabe, M.; Sasaki, K.; Poosaran, N. Production of cellulase-free xylanase by a thermotolerant Streptomyces sp. grown on agricultural waste and media optimization using mixture design and Plackett-Burman experimental design methods. Biotechnol. Lett 2002, 24, 1437–1442. [Google Scholar]
- Box, G.E.; Behnken, D.W. Some new three level designs for the study of quantitative variable. Technometrics 1960, 2, 456–475. [Google Scholar]
- Cui, F.J.; Li, Y.; Liu, Z.Q.; Zhao, H.; Ping, L.F.; Ping, L.Y. Optimization of fermentation conditions for production of xylanase by a newly isolated strain, Penicillium thiersii ZH-19. World J. Microb. Biotechnol 2009, 25, 721–725. [Google Scholar]
| Variable code | Variable | Value | |
|---|---|---|---|
| −1 | +1 | ||
| X1 | Wheat bran (g/L) | 5 | 35 |
| X2 | Glucose (g/L) | 5 | 15 |
| X3 | Tryptone (g/L) | 0.5 | 1.5 |
| X4 | Yeast extract (g/L) | 0.5 | 1.5 |
| X5 | Urea (g/L) | 0.5 | 1.5 |
| X6 | NH4Cl (g/L) | 5 | 15 |
| X7 | (NH4)2SO4 (g/L) | 5 | 15 |
| X8 | NaNO3 (g/L) | 5 | 15 |
| X9 | K2HPO4 (g/L) | 5 | 15 |
| X10 | KH2PO4 (g/L) | 5 | 15 |
| X11 | Tween (g/L) | 0.5 | 1.5 |
| X12 | EDTA (g/L) | 0.1 | 1 |
| X13 | MgSO4 (g/L) | 0.2 | 2 |
| X14 | CaCl2 (g/L) | 0.2 | 2 |
| X15 | FeSO4 (g/L) | 0.01 | 0.05 |
| X16 | MnSO4 (g/L) | 0.1 | 0.5 |
| X17 | ZnCl2 (g/L) | 0.01 | 0.05 |
| X18 | pH | 6 | 7 |
| X19 | Inoculum size | 1 × 106 | 1 × 108 |
| No. | X1 | X2 | X3 | X4 | X5 | X6 | X7 | X8 | X9 | X10 | X11 | X12 | X13 | X14 | X15 | X16 | X17 | X18 | X19 | Y |
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| 1 | 1 | −1 | 1 | 1 | −1 | −1 | −1 | −1 | 1 | −1 | 1 | −1 | 1 | 1 | 1 | 1 | −1 | −1 | −1 | 24.55 |
| 2 | −1 | 1 | −1 | −1 | 1 | 1 | 1 | 1 | −1 | 1 | −1 | 1 | −1 | −1 | −1 | −1 | 1 | 1 | 1 | 4.30 |
| 3 | 1 | 1 | −1 | 1 | 1 | −1 | −1 | −1 | −1 | 1 | −1 | 1 | −1 | 1 | 1 | 1 | 1 | −1 | −1 | 20.68 |
| 4 | −1 | −1 | 1 | −1 | −1 | 1 | 1 | 1 | 1 | −1 | 1 | −1 | 1 | −1 | −1 | −1 | −1 | 1 | 1 | 8.20 |
| 5 | −1 | 1 | 1 | −1 | 1 | 1 | −1 | −1 | −1 | −1 | 1 | −1 | 1 | −1 | 1 | 1 | 1 | 1 | −1 | 3.10 |
| 6 | 1 | −1 | −1 | 1 | −1 | −1 | 1 | 1 | 1 | 1 | −1 | 1 | −1 | 1 | −1 | −1 | −1 | −1 | 1 | 26.10 |
| 7 | −1 | −1 | 1 | 1 | −1 | 1 | 1 | −1 | −1 | −1 | −1 | 1 | −1 | 1 | −1 | 1 | 1 | 1 | −1 | 6.09 |
| 8 | 1 | 1 | −1 | −1 | 1 | −1 | −1 | 1 | 1 | 1 | 1 | −1 | 1 | −1 | 1 | −1 | −1 | −1 | 1 | 22.81 |
| 9 | 1 | −1 | −1 | 1 | 1 | −1 | 1 | 1 | −1 | −1 | −1 | −1 | 1 | −1 | 1 | −1 | 1 | 1 | −1 | 28.50 |
| 10 | −1 | 1 | 1 | −1 | −1 | 1 | −1 | −1 | 1 | 1 | 1 | 1 | −1 | 1 | −1 | 1 | −1 | −1 | 1 | 1.30 |
| 11 | 1 | 1 | −1 | −1 | 1 | 1 | −1 | 1 | 1 | −1 | −1 | −1 | −1 | 1 | −1 | 1 | −1 | 1 | −1 | 22.20 |
| 12 | −1 | −1 | 1 | 1 | −1 | −1 | 1 | −1 | −1 | 1 | 1 | 1 | 1 | −1 | 1 | −1 | 1 | −1 | 1 | 6.63 |
| 13 | 1 | 1 | 1 | −1 | −1 | 1 | 1 | −1 | 1 | 1 | −1 | −1 | −1 | −1 | 1 | −1 | 1 | −1 | −1 | 13.71 |
| 14 | −1 | −1 | −1 | 1 | 1 | −1 | −1 | 1 | −1 | −1 | 1 | 1 | 1 | 1 | −1 | 1 | −1 | 1 | 1 | 15.11 |
| 15 | 1 | 1 | 1 | 1 | −1 | −1 | 1 | 1 | −1 | 1 | 1 | −1 | −1 | −1 | −1 | 1 | −1 | 1 | −1 | 25.29 |
| 16 | −1 | −1 | −1 | −1 | 1 | 1 | −1 | −1 | 1 | −1 | −1 | 1 | 1 | 1 | 1 | −1 | 1 | −1 | 1 | 3.62 |
| 17 | −1 | 1 | 1 | 1 | 1 | −1 | −1 | 1 | 1 | −1 | 1 | 1 | −1 | −1 | −1 | −1 | 1 | −1 | −1 | 6.78 |
| 18 | 1 | −1 | −1 | −1 | −1 | 1 | 1 | −1 | −1 | 1 | −1 | −1 | 1 | 1 | 1 | 1 | −1 | 1 | 1 | 22.13 |
| 19 | 1 | −1 | 1 | 1 | 1 | 1 | −1 | −1 | 1 | 1 | −1 | 1 | 1 | −1 | −1 | −1 | −1 | 1 | −1 | 22.71 |
| 20 | −1 | 1 | −1 | −1 | −1 | −1 | 1 | 1 | −1 | −1 | 1 | −1 | −1 | 1 | 1 | 1 | 1 | −1 | 1 | 6.18 |
| 21 | −1 | 1 | −1 | 1 | 1 | 1 | 1 | −1 | −1 | 1 | 1 | −1 | 1 | 1 | −1 | −1 | −1 | −1 | −1 | 5.98 |
| 22 | 1 | −1 | 1 | −1 | −1 | −1 | −1 | 1 | 1 | −1 | −1 | 1 | −1 | −1 | 1 | 1 | 1 | 1 | 1 | 22.91 |
| 23 | 1 | −1 | 1 | −1 | 1 | 1 | 1 | 1 | −1 | −1 | 1 | 1 | −1 | 1 | 1 | −1 | −1 | −1 | −1 | 23.68 |
| 24 | −1 | 1 | −1 | 1 | −1 | −1 | −1 | −1 | 1 | 1 | −1 | −1 | 1 | −1 | −1 | 1 | 1 | 1 | 1 | 5.21 |
| 25 | −1 | 1 | −1 | 1 | −1 | 1 | 1 | 1 | 1 | −1 | −1 | 1 | 1 | −1 | 1 | 1 | −1 | −1 | −1 | 9.18 |
| 26 | 1 | −1 | 1 | −1 | 1 | −1 | −1 | −1 | −1 | 1 | 1 | −1 | −1 | 1 | −1 | −1 | 1 | 1 | 1 | 19.71 |
| 27 | −1 | −1 | 1 | −1 | 1 | −1 | 1 | 1 | 1 | 1 | −1 | −1 | 1 | 1 | −1 | 1 | 1 | −1 | −1 | 8.33 |
| 28 | 1 | 1 | −1 | 1 | −1 | 1 | −1 | −1 | −1 | −1 | 1 | 1 | −1 | −1 | 1 | −1 | −1 | 1 | 1 | 20.55 |
| 29 | −1 | −1 | −1 | 1 | −1 | 1 | −1 | 1 | 1 | 1 | 1 | −1 | −1 | 1 | 1 | −1 | 1 | 1 | −1 | 12.87 |
| 30 | 1 | 1 | 1 | −1 | 1 | −1 | 1 | −1 | −1 | −1 | −1 | 1 | 1 | −1 | −1 | 1 | −1 | −1 | 1 | 16.01 |
| 31 | −1 | −1 | −1 | −1 | 1 | −1 | 1 | −1 | 1 | 1 | 1 | 1 | −1 | −1 | 1 | 1 | −1 | 1 | −1 | 2.89 |
| 32 | 1 | 1 | 1 | 1 | −1 | 1 | −1 | 1 | −1 | −1 | −1 | −1 | 1 | 1 | −1 | −1 | 1 | −1 | 1 | 26.11 |
| 33 | 1 | −1 | −1 | −1 | −1 | 1 | −1 | 1 | −1 | 1 | 1 | 1 | 1 | −1 | −1 | 1 | 1 | −1 | −1 | 23.65 |
| 34 | −1 | 1 | 1 | 1 | 1 | −1 | 1 | −1 | 1 | −1 | −1 | −1 | −1 | 1 | 1 | −1 | −1 | 1 | 1 | 5.32 |
| 35 | 1 | 1 | −1 | −1 | −1 | −1 | 1 | −1 | 1 | −1 | 1 | 1 | 1 | 1 | −1 | −1 | 1 | 1 | −1 | 21.09 |
| 36 | −1 | −1 | 1 | 1 | 1 | 1 | −1 | 1 | −1 | 1 | −1 | −1 | −1 | −1 | 1 | 1 | −1 | −1 | 1 | 7.84 |
| 37 | −1 | 1 | 1 | −1 | −1 | −1 | −1 | 1 | −1 | 1 | −1 | 1 | 1 | 1 | 1 | −1 | −1 | 1 | −1 | 12.23 |
| 38 | 1 | −1 | −1 | 1 | 1 | 1 | 1 | −1 | 1 | −1 | 1 | −1 | −1 | −1 | −1 | 1 | 1 | −1 | 1 | 16.65 |
| 39 | −1 | −1 | −1 | −1 | −1 | −1 | −1 | −1 | −1 | −1 | −1 | −1 | −1 | −1 | −1 | −1 | −1 | −1 | −1 | 2.82 |
| 40 | 1 | 1 | 1 | 1 | 1 | 1 | 1 | 1 | 1 | 1 | 1 | 1 | 1 | 1 | 1 | 1 | 1 | 1 | 1 | 29.72 |
| Factors | Medium components | Effect | Standard error | t-Ratio | p-Value | Confidence level (%) |
|---|---|---|---|---|---|---|
| X1 | Wheat bran | 15.74 | 0.165 | 95.39 | 0.0067 | 99.33 |
| X2 | Glucose | −1.36 | 0.165 | −8.25 | 0.0767 | 92.33 |
| X3 | Trypton | −0.12 | 0.165 | −0.70 | 0.6125 | 38.75 |
| X4 | Yeast extract | 3.05 | 0.165 | 18.49 | 0.0344 | 96.56 |
| X5 | Urea | −0.54 | 0.165 | −3.29 | 0.1878 | 81.22 |
| X6 | NH4Cl | −0.78 | 0.165 | −4.72 | 0.1330 | 86.7 |
| X7 | (NH4)2SO4 | −0.54 | 0.165 | −3.27 | 0.1891 | 81.09 |
| X8 | NaNO3 | 5.06 | 0.165 | 30.68 | 0.0207 | 97.93 |
| X9 | K2HPO4 | −0.52 | 0.165 | −3.16 | 0.1949 | 80.51 |
| X10 | KH2PO4 | 0.27 | 0.165 | 1.65 | 0.3471 | 65.29 |
| X11 | Tween | 0.54 | 0.165 | 3.25 | 0.1898 | 81.02 |
| X12 | EDTA | 0.39 | 0.165 | 2.34 | 0.2572 | 74.28 |
| X13 | MgSO4 | 2.35 | 0.165 | 14.24 | 0.0446 | 95.54 |
| X14 | CaCl2 | 2.16 | 0.165 | 13.11 | 0.0485 | 95.15 |
| X15 | FeSO4 | 0.77 | 0.165 | 4.68 | 0.1339 | 86.61 |
| X16 | MnSO4 | −0.24 | 0.165 | −1.42 | 0.3897 | 61.03 |
| X17 | ZnCl2 | −0.55 | 0.165 | −3.35 | 0.1846 | 81.54 |
| X18 | pH | 1.88 | 0.165 | 11.37 | 0.0558 | 94.42 |
| X19 | Inoculum size | 0.50 | 0.165 | −3.00 | 0.2044 | 79.56 |
| Runs | Wheat bran (X1) (g/L) | Yeast extract (X2) (g/L) | NaNO3 (X3) (g/L) | MgSO4 (X4) (g/L)4 | CaCl2 (X5) (g/L) | Xylanase (U/mL) |
|---|---|---|---|---|---|---|
| 1 | 5 | 0.5 | 10 | 1.1 | 1.1 | 8.52 |
| 2 | 5 | 1.5 | 10 | 1.1 | 1.1 | 20.42 |
| 3 | 35 | 0.5 | 10 | 1.1 | 1.1 | 33.60 |
| 4 | 35 | 1.5 | 10 | 1.1 | 1.1 | 34.76 |
| 5 | 20 | 1 | 5 | 0.2 | 1.1 | 21.72 |
| 6 | 20 | 1 | 5 | 2 | 1.1 | 24.52 |
| 7 | 20 | 1 | 15 | 0.2 | 1.1 | 33.72 |
| 8 | 20 | 1 | 15 | 2 | 1.1 | 36.52 |
| 9 | 20 | 0.5 | 10 | 1.1 | 0.2 | 26.02 |
| 10 | 20 | 0.5 | 10 | 1.1 | 2 | 28.12 |
| 11 | 20 | 1.5 | 10 | 1.1 | 0.2 | 30.12 |
| 12 | 20 | 1.5 | 10 | 1.1 | 2 | 34.42 |
| 13 | 5 | 1 | 5 | 1.1 | 1.1 | 10.72 |
| 14 | 5 | 1 | 15 | 1.1 | 1.1 | 18.52 |
| 15 | 35 | 1 | 5 | 1.1 | 1.1 | 38.67 |
| 16 | 35 | 1 | 15 | 1.1 | 1.1 | 44.72 |
| 17 | 20 | 1 | 10 | 0.2 | 0.2 | 23.33 |
| 18 | 20 | 1 | 10 | 0.2 | 2 | 28.72 |
| 19 | 20 | 1 | 10 | 2 | 0.2 | 28.52 |
| 20 | 20 | 1 | 10 | 2 | 2 | 33.52 |
| 21 | 5 | 1 | 10 | 0.2 | 1.1 | 10.12 |
| 22 | 5 | 1 | 10 | 2 | 1.1 | 15.92 |
| 23 | 35 | 1 | 10 | 0.2 | 1.1 | 44.32 |
| 24 | 35 | 1 | 10 | 2 | 1.1 | 37.12 |
| 25 | 20 | 0.5 | 5 | 1.1 | 1.1 | 21.89 |
| 26 | 20 | 0.5 | 15 | 1.1 | 1.1 | 31.99 |
| 27 | 20 | 1.5 | 5 | 1.1 | 1.1 | 27.12 |
| 28 | 20 | 1.5 | 15 | 1.1 | 1.1 | 37.12 |
| 29 | 20 | 1 | 5 | 1.1 | 0.2 | 22.87 |
| 30 | 20 | 1 | 5 | 1.1 | 2 | 27.11 |
| 31 | 20 | 1 | 15 | 1.1 | 0.2 | 32.12 |
| 32 | 20 | 1 | 15 | 1.1 | 2 | 36.34 |
| 33 | 5 | 1 | 10 | 1.1 | 0.2 | 11.52 |
| 34 | 5 | 1 | 10 | 1.1 | 2 | 15.52 |
| 35 | 35 | 1 | 10 | 1.1 | 0.2 | 42.72 |
| 36 | 35 | 1 | 10 | 1.1 | 2 | 36.72 |
| 37 | 20 | 0.5 | 10 | 0.2 | 1.1 | 25.72 |
| 38 | 20 | 0.5 | 10 | 2 | 1.1 | 27.52 |
| 39 | 20 | 1.5 | 10 | 0.2 | 1.1 | 29.72 |
| 40 | 20 | 1.5 | 10 | 2 | 1.1 | 33.92 |
| 41 | 20 | 1 | 10 | 1.1 | 1.1 | 36.12 |
| 42 | 20 | 1 | 10 | 1.1 | 1.1 | 38.12 |
| 43 | 20 | 1 | 10 | 1.1 | 1.1 | 39.12 |
| 44 | 20 | 1 | 10 | 1.1 | 1.1 | 39.67 |
| 45 | 20 | 1 | 10 | 1.1 | 1.1 | 40.55 |
| 46 | 20 | 1 | 10 | 1.1 | 1.1 | 39.23 |
| Model term | Degree of freedom | Estimate | Standard error | t Value | p > |t| |
|---|---|---|---|---|---|
| Intercept | 1 | 38.80 | 0.93 | 41.53 | <0.0001 * |
| X1 | 1 | 12.59 | 0.57 | 22.00 | <0.0001 * |
| X2 | 1 | 2.76 | 0.57 | 4.83 | <0.0001 * |
| X3 | 1 | 4.78 | 0.57 | 8.35 | <0.0001 * |
| X4 | 1 | 1.26 | 0.57 | 2.21 | 0.0368 * |
| X5 | 1 | 1.45 | 0.57 | 2.54 | 0.0177 * |
| X1X1 | 1 | −7.55 | 0.77 | −9.74 | <0.0001 * |
| X2X1 | 1 | −2.69 | 1.14 | −2.35 | 0.0272 * |
| X2X2 | 1 | −5.29 | 0.77 | −6.83 | <0.0001 * |
| X3X1 | 1 | −0.44 | 1.14 | −0.38 | 0.7055 |
| X3X2 | 1 | −0.03 | 1.14 | −0.02 | 0.9827 |
| X3X3 | 1 | −4.06 | 0.77 | −5.25 | <0.0001 * |
| X4X1 | 1 | −3.25 | 1.14 | −2.84 | 0.0088* |
| X4X2 | 1 | 0.60 | 1.14 | 0.52 | 0.6047 |
| X4X3 | 1 | 0 | 1.14 | 0.00 | 1.0000 |
| X4X4 | 1 | −4.96 | 0.77 | −6.40 | <0.0001 * |
| X5X1 | 1 | −2.50 | 1.14 | −2.18 | 0.0385 * |
| X5X2 | 1 | 0.55 | 1.14 | 0.48 | 0.6350 |
| X5X3 | 1 | −0.005 | 1.14 | −0.00 | 0.9965 |
| X5X4 | 1 | −0.09 | 1.14 | −0.09 | 0.9328 |
| X5X5 | 1 | −4.73 | 0.77 | −6.11 | <0.0001 * |
© 2012 by the authors; licensee Molecular Diversity Preservation International, Basel, Switzerland. This article is an open-access article distributed under the terms and conditions of the Creative Commons Attribution license (http://creativecommons.org/licenses/by/3.0/).
Share and Cite
Cui, F.; Zhao, L. Optimization of Xylanase Production from Penicillium sp.WX-Z1 by a Two-Step Statistical Strategy: Plackett-Burman and Box-Behnken Experimental Design. Int. J. Mol. Sci. 2012, 13, 10630-10646. https://doi.org/10.3390/ijms130810630
Cui F, Zhao L. Optimization of Xylanase Production from Penicillium sp.WX-Z1 by a Two-Step Statistical Strategy: Plackett-Burman and Box-Behnken Experimental Design. International Journal of Molecular Sciences. 2012; 13(8):10630-10646. https://doi.org/10.3390/ijms130810630
Chicago/Turabian StyleCui, Fengjie, and Liming Zhao. 2012. "Optimization of Xylanase Production from Penicillium sp.WX-Z1 by a Two-Step Statistical Strategy: Plackett-Burman and Box-Behnken Experimental Design" International Journal of Molecular Sciences 13, no. 8: 10630-10646. https://doi.org/10.3390/ijms130810630





