Expression Analysis of MYC Genes from Tamarix hispida in Response to Different Abiotic Stresses
Abstract
:1. Introduction
2. Results
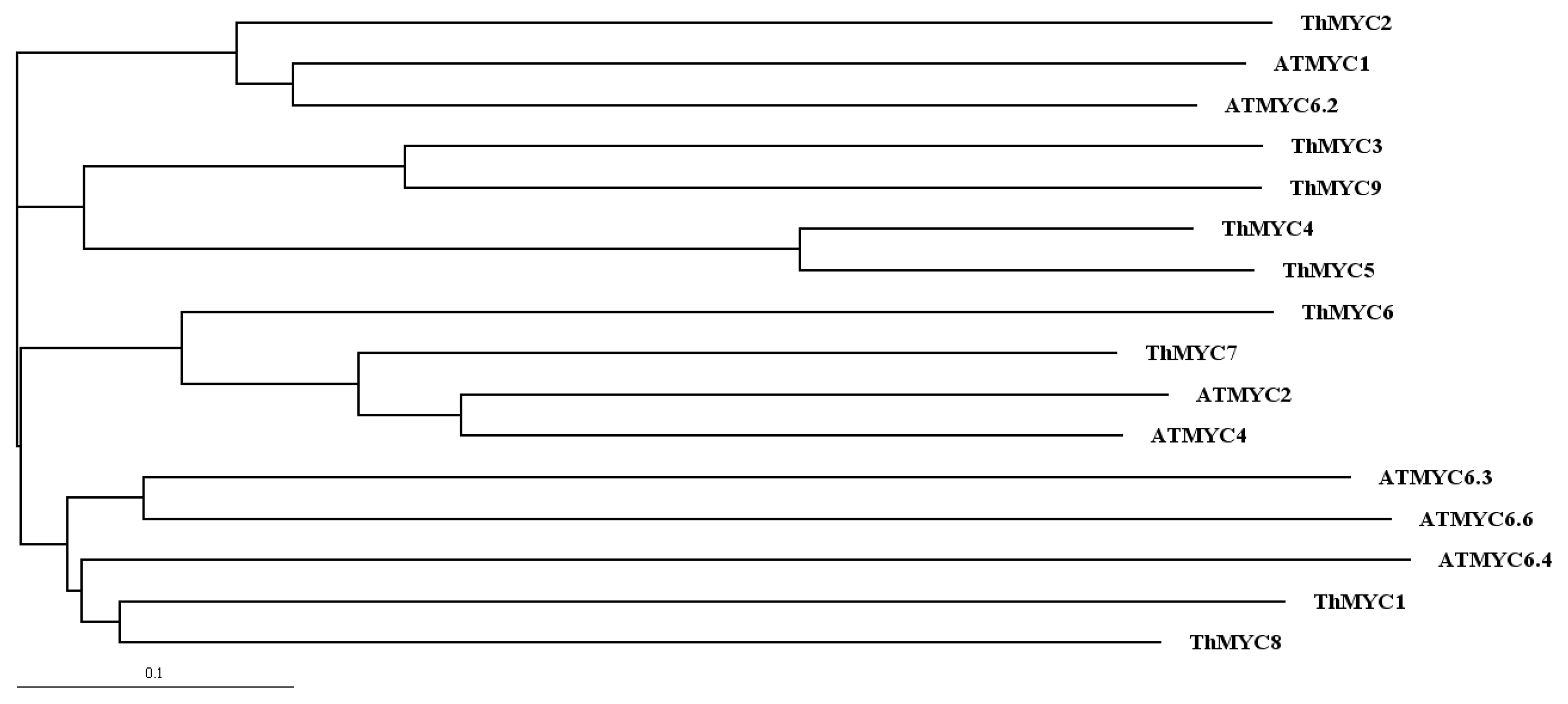
2.1. Identification and Bioinformatics Analysis of MYC Genes from T. hispida
2.2. Relative Abundances of ThMYCs in Root, Stem and Leaf tissues
2.3. Expression Patterns of ThMYCs in Response to Different Stresses
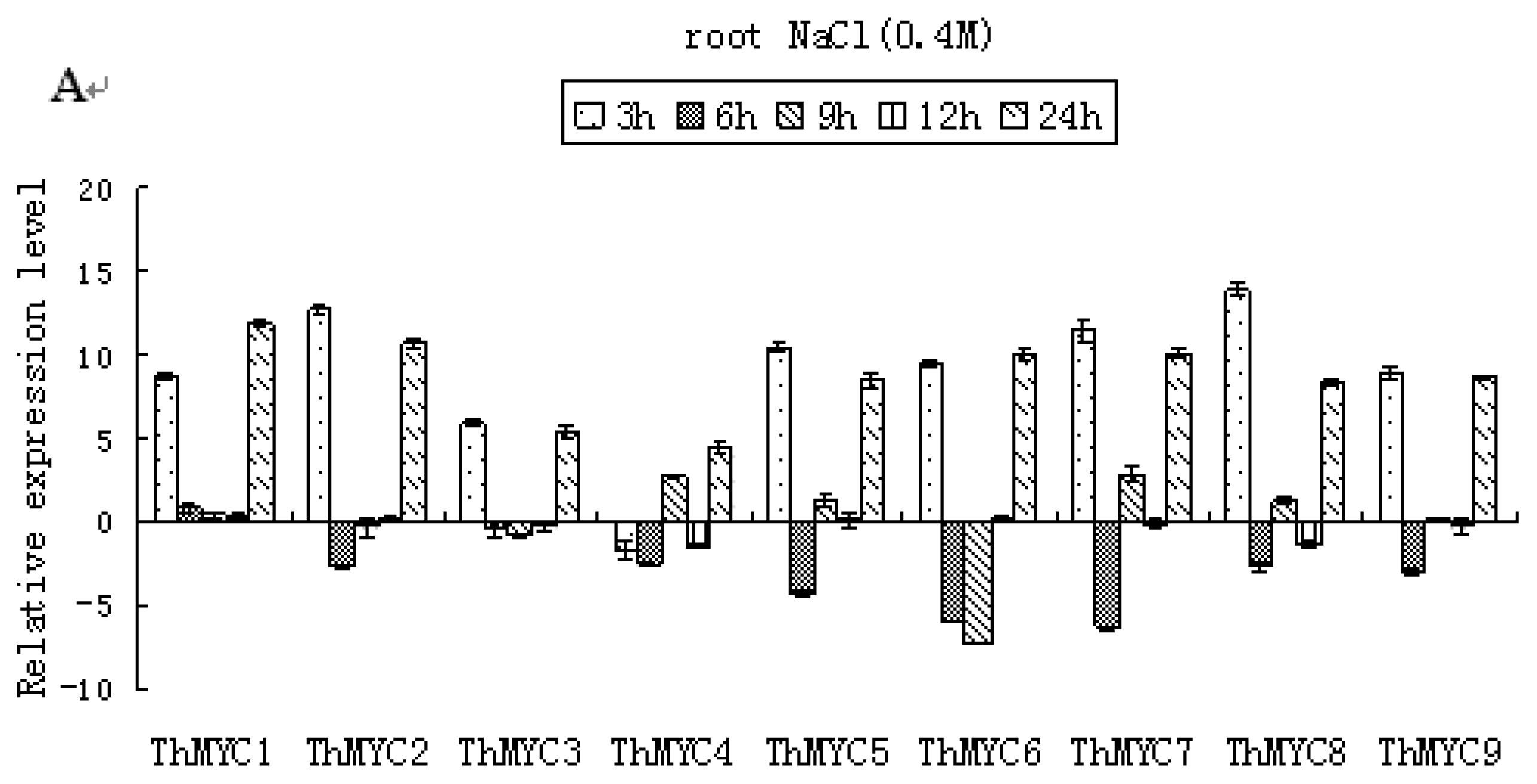
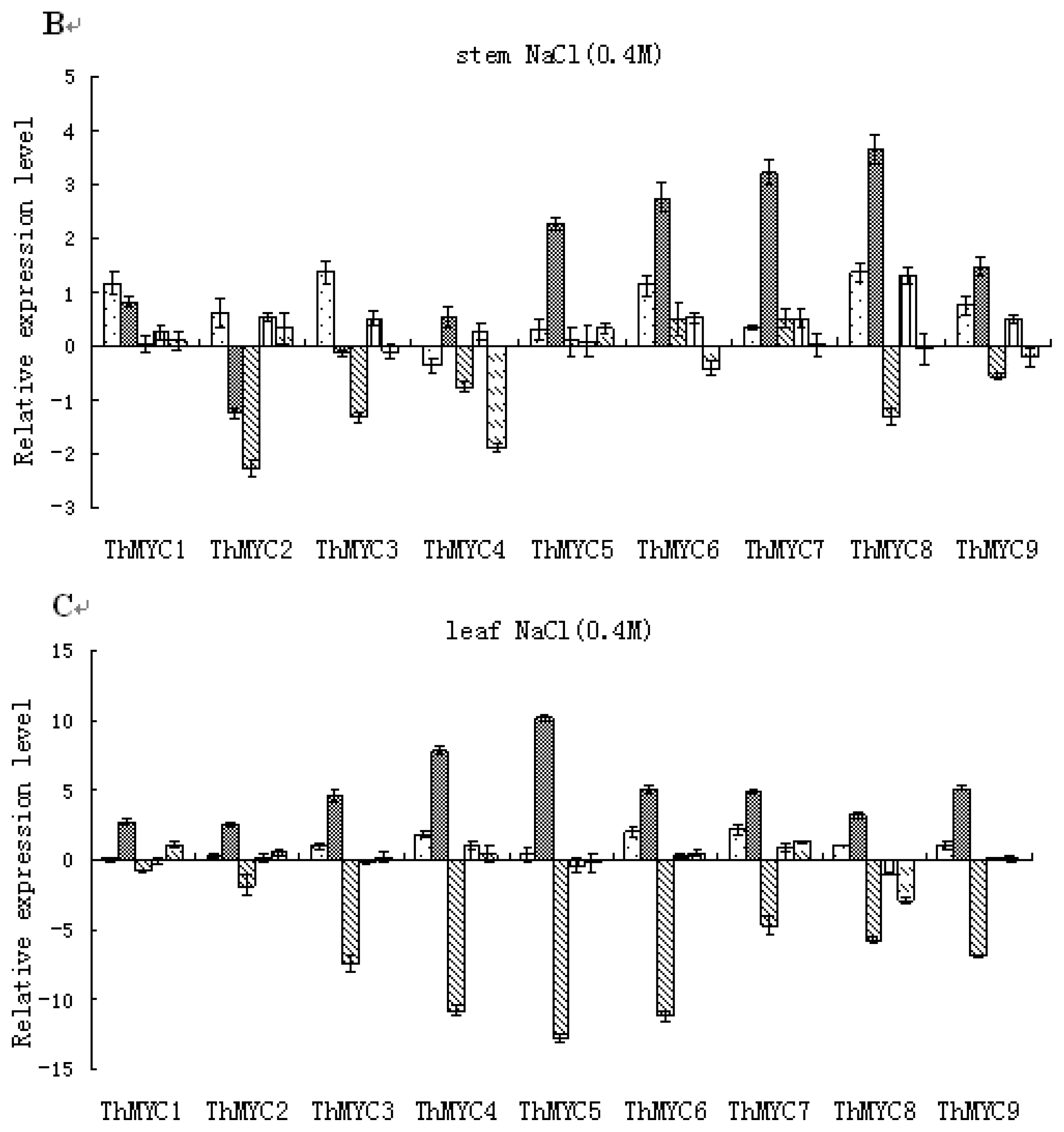
2.4. Expression Patterns of ThMYCs in Response to NaCl Stress
2.5. Expression Patterns of ThMYCs in Response to PEG Stress
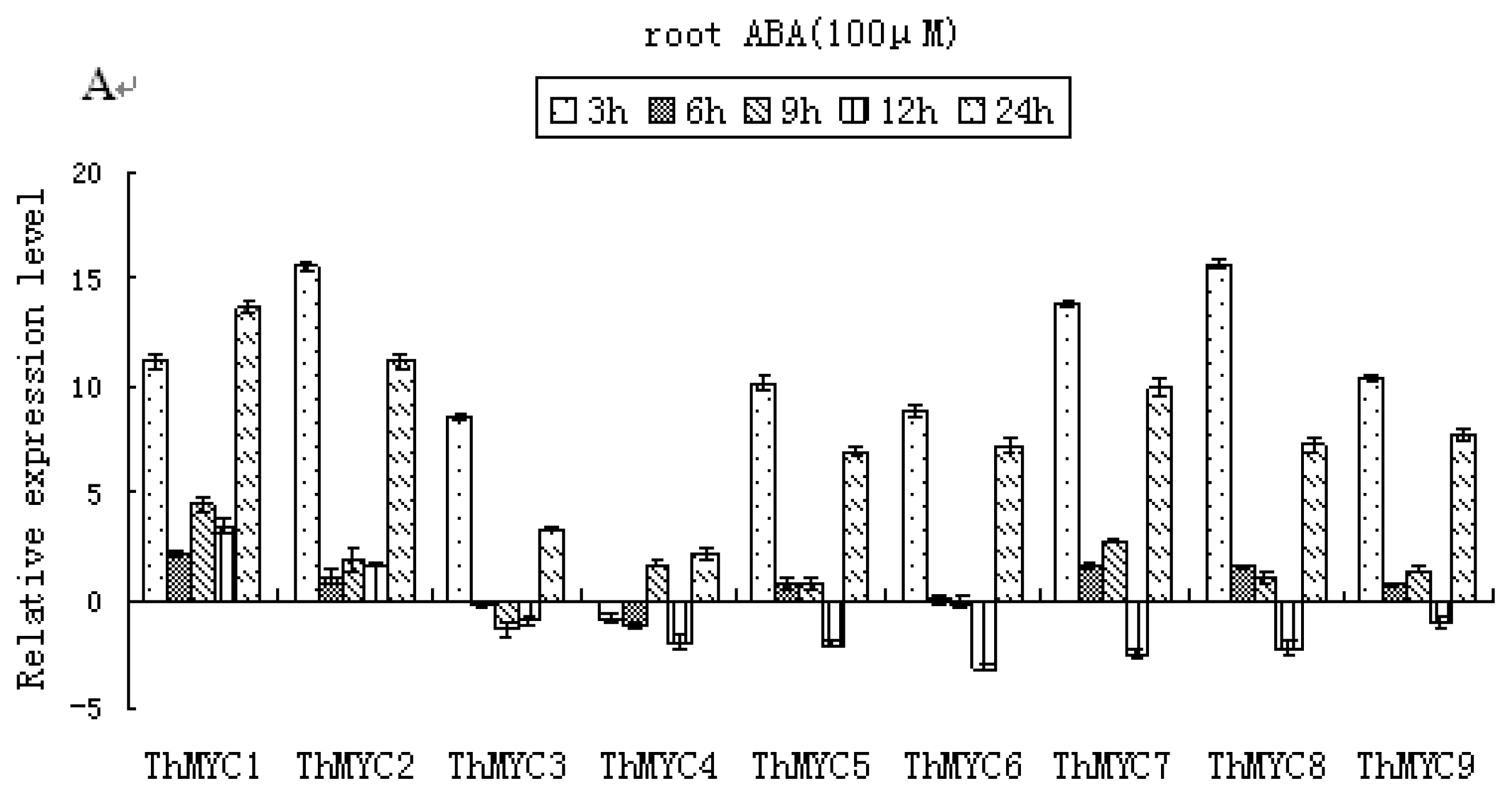
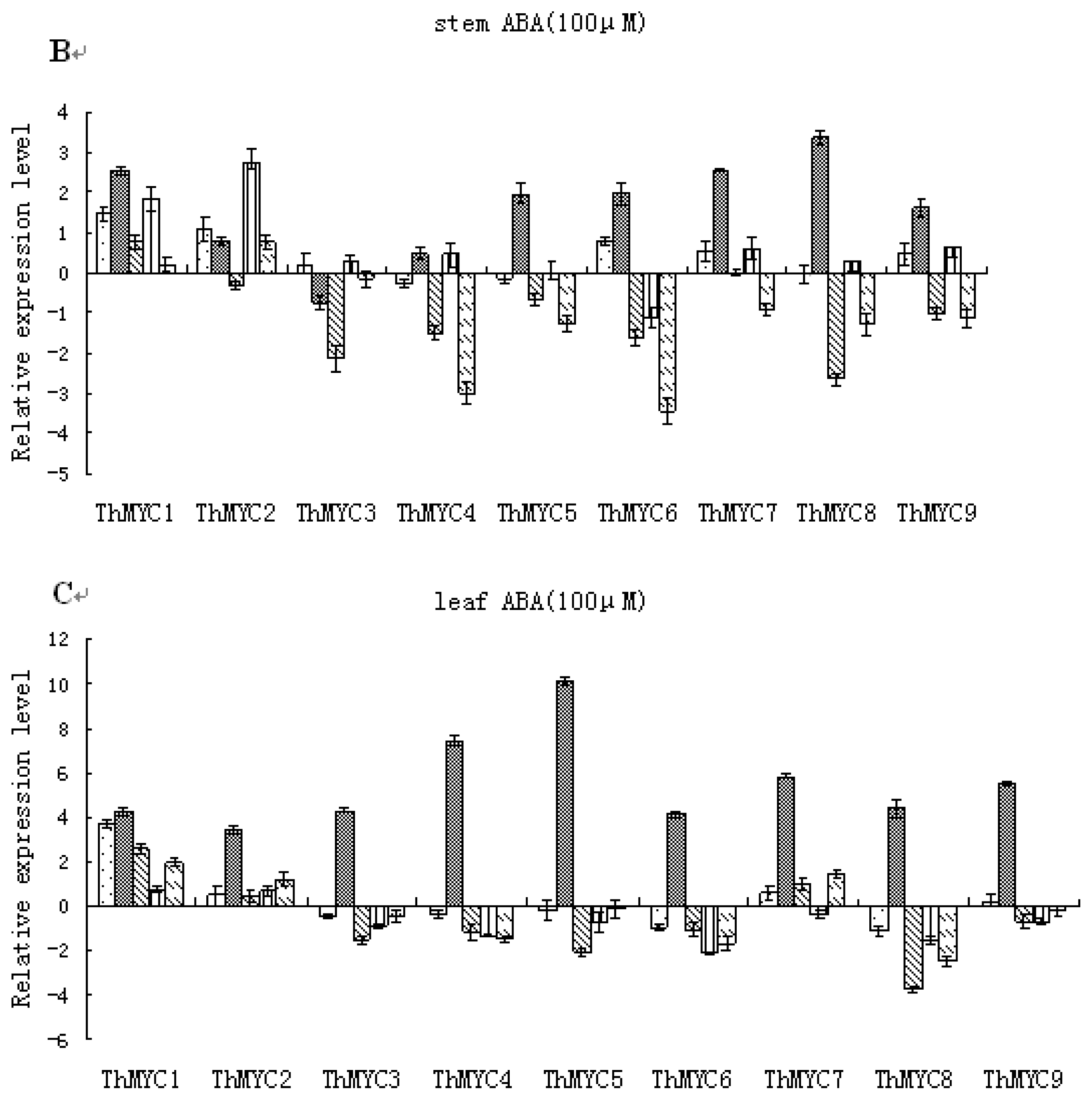
2.6. Expression Patterns of ThMYCs in Response to ABA Treatment
3. Discussion
4. Experimental Section
4.1. Plant Culture Conditions and Treatments
4.2. Cloning and Analysis of MYC Family Genes from T. hispida
4.3. Phylogenetic Analysis of MYC Sequences
4.4. MYC Gene Expression Profiles in Response to Abiotic Stresses
5. Conclusions
Acknowledgments
References
- Pires, N.; Dolan, L. Origin and diversification of basic-helix-loop-helix proteins in plants. Mol. Biol. Evol 2010, 27, 862–874. [Google Scholar]
- Berger, S.; Bell, E.; Mullet, J.E. Two methyl jasmonate-insensitive mutants show altered expression of AtVsp in response to methyl jasmonate and wounding. Plant Physiol 1996, 111, 525–531. [Google Scholar]
- Anderson, J.P.; Badruzsaufari, E.; Schenk, P.M.; Manners, J.M.; Desmond, O.J.; Ehlert, C.; Maclean, D.J.; Ebert, P.R.; Kazan, K. Antagonistic interaction between abscisic acid and jasmonate-ethylene signaling pathways modulates defense gene expression and disease resistance in Arabidopsis. The Plant Cell 2004, 16, 3460–3479. [Google Scholar]
- Boter, M.; Ruiz-Rivero, O.; Abdeen, A.; Pratt, S. Conserved MYC transcription factors play a key role in jasmonate signaling both in tomato and Arabidopsis. Genes Dev 2004, 18, 1577–1591. [Google Scholar]
- Lorenzo, O.; Chico, J.M.; Sanchez-Serrano, J.J.; Solano, R. JASMONATE-INSENSITIVE1 encodes a MYC transcription factor essential to discriminate between different jasmonate-regulated defense responses in Arabidopsis. The Plant Cell 2004, 16, 1938–1950. [Google Scholar]
- Lorenzo, O.; Solano, R. Molecular players regulating the jasmonate signalling network. Curr. Opin. Plant Biol 2005, 8, 532–540. [Google Scholar]
- Zhu, Y.; Cai, X.L.; Wang, Z.Y.; Hong, M.M. An Interaction between a MYC Protein and an EREBP Protein Is Involved in Transcriptional Regulation of the Rice Wx Gene. J. Biol. Chem 2003, 48, 47803–47811. [Google Scholar]
- Dang, C.V.; Barrett, J.; Villa-Garcia, M.; Resar, L.M.S.; Kato, G.T.; Fearon, E.R. Intracellular leucine zipper interactions suggest c-Myc hetero-oligomerization. Mol. Cell Biol 1991, 11, 954–962. [Google Scholar]
- Shimizu, T.; Toumoto, A.; Ihara, K.; Shimizu, M.; Kyogoku, Y.; Ogawa, N. Crystal structure of PHO4 bHLH domain-DNA complex: flanking base recognition. EMBO J 1997, 16, 4689–4697. [Google Scholar]
- Atchley, W.R.; Terhalle, W.; Dress, A. Positional dependence, cliques, and predictive motifs in the bHLH protein domain. J. Mol. Evol 1999, 48, 501–516. [Google Scholar]
- Martínez-García, J.F.; Huq, E.; Quail, P.H. Direct targeting of light signals to a promoter element-bound transcription factor. Science 2000, 288, 859–863. [Google Scholar]
- Toledo-Ortiz, G.; Huq, E.; Quail, P.H. The Arabidopsis basic/helix-loop-helix transcription factor family. The Plant Cell 2003, 15, 1749–1770. [Google Scholar]
- Qian, W.; Tan, G.; Liu, H.; He, S.; Gao, Y.; An, C. Identification of a bHLH-type G-box binding factor and its regulation activity with G-box and BoxI elements of the PsCHS1 promoter. Plant Cell Rep 2007, 26, 85–93. [Google Scholar]
- Abe, H.; Yamaguchi-Shinozaki, K.; Urao, T.; Iwasaki, T.; Hosokawa, D.; Shinozaki, K. Role of Arabidopsis MYC and MYB Homologs in Drought and Abscisic Acid-Regulated Gene Expression. The Plant Cell 1997, 9, 1859–1868. [Google Scholar]
- van der Fits, L.; Zhang, H.; Menke, F.L.; Deneka, M.; Memelink, J. A Catharanthus roseus BPF-1 homologue interacts with an elicitor-responsive region of the secondary metabolite biosynthetic gene Str and is induced by elicitor via a JA-independent signal transduction Pathway. Plant Mol. Biol 2000, 44, 675–685. [Google Scholar]
- Dombrecht, B.; Xue, G.P.; Sprague, S.J.; Kirkegaard, J.A.; Ross, J.J.; Reid, J.B.; Fitt, G.P.; Sewelam, N.; Schenk, P.M.; Manners, J.M.; et al. MYC2 Differentially Modulates Diverse Jasmonate-Dependent Functions in Arabidopsis. The Plant Cell 2007, 19, 2225–2245. [Google Scholar]
- Zhao, Y.; Zhou, L.M.; Chen, Y.Y.; Yang, S.G.; Tian, W.M. MYC genes with differential responses to tapping, mechanical wounding, ethrel and methyl jasmonate in laticifers of rubber tree. J. Plant Physiol 2011, 168, 1649–1658. [Google Scholar]
- Li, H.; Sun, J.; Xu, Y.; Jiang, H.; Wu, X.; Li, C. The bHLH-type transcription factor AtAIB positively regulates ABA response in Arabidopsis. Plant Mol. Biol 2007, 65, 655–665. [Google Scholar]
- Fursova, O.V.; Pogorelko, G.V.; Tarasov, V.A. Identification of ICE2, a gene involved in cold acclimation which determines freezing tolerance in Arabidopsis thaliana. Gene 2009, 429, 98–103. [Google Scholar]
- Yin, Y.; Vafeados, D.; Tao, Y.; Yoshida, S.; Asami, T.; Chory, J. A new class of transcription factors mediates brassinosteroid-regulated gene expression in Arabidopsis. Cell 2005, 120, 249–259. [Google Scholar]
- Penfield, S.; Josse, E.M.; Kannangara, R.; Gilday, A.D.; Halliday, K.J.; Graham, I.A. Cold and light control seed germination through the bHLH transcription factor SPATULA. Curr. Biol 2005, 15, 1998–2006. [Google Scholar]
- Tuteja, N. Abscisic Acid and Abiotic Stress Signaling. Plant Signal. Behav 2007, 2, 135–138. [Google Scholar]
- Muniz García, M.N.; Giammaria, V.; Grandellis, C.; Téllez-Inón, M.T.; Ulloa, R.M.; Capiati, D.A. Characterization of StABF1, a stress-responsive bZIP transcription factor from Solanum tuberosum L. that is phosphorylated by StCDPK2in vitro. Planta 2011. [Google Scholar] [CrossRef]
- Ying, S.; Zhang, D.F.; Fu, J.; Shi, Y.S.; Song, Y.C.; Wang, T.Y.; Li, Y. Cloning and characterization of a maize bZIP transcription factor, ZmbZIP72, confers drought and salt tolerance in transgenic. Arabidopsis. Planta 2011. [Google Scholar] [CrossRef]
- Gao, S.Q.; Chen, M.; Xu, Z.S.; Zhao, C.P.; Li, L.; Xu, H.J.; Tang, Y.M.; Zhao, X.; Ma, Y.Z. The soybean GmbZIP1 transcription factor enhances multiple abiotic stress tolerances in transgenic plants. Plant Mol. Biol 2011, 75, 537–553. [Google Scholar]
- Mahajan, S.; Tuteja, N. Cold, salinity and drought stresses: An overview. Arch. Biochem. Biophys 2005, 444, 139–158. [Google Scholar]
- Abe, H.; Urao, T.; Urao, T.; Ito, T.; Seki, M.; Shinozaki, K.; Yamaguchi-Shinozaki, K. Arabidopsis AtMYC2 (bHLH) and AtMYB2 (MYB) Function as Transcriptional Activators in Abscisic Acid Signaling. Plant Cell 2003, 15, 63–78. [Google Scholar]
- Pertea, G.; Huang, X.; Liang, F.; Antonescu, V.; Sultana, R.; Karamycheva, S.; Lee, Y.; White, J.; Cheung, F.; Parvizi, B.; et al. TIGR Gene Indices clustering tools (TGICL): A software system for fast clustering of large EST datasets. Bioinformatics 2003, 19, 651–652. [Google Scholar]
- ORF Finder. http://www.ncbi.nlm.nih.gov/gorf/gorf.html accessed on 12 September 2011.
- Saitou, N.; Nei, M. The neighbor-joining method: a new method for reconstructing phylogenetic trees. Mol. Biol. Evol 1987, 4, 406–425. [Google Scholar]
- Clustalw 2. http://www.ebi.ac.uk/Tools/clustalw2/ accessed on 12 September 2011.
- Boutrot, F.; Chantret, N.; Gautier, M.F. Genome-wide analysis of the rice and Arabidopsis non-specific lipid transfer protein (nsLtp) gene families and identification of wheat nsLtp genes by EST data mining. BMC Genomics 2008, 9, 86. [Google Scholar]
- ProtParam. http://www.expasy.org/tools/protparam.html accessed on 12 September 2011.
- Pfaffl, M.W.; Horgan, G.W.; Dempfle, L. Relative expression software tool (REST) for group-wise comparison and statistical analysis of relative expression results in real-time PCR. Nucleic Acids Res 2002, 30, 36. [Google Scholar]

| Gene | GenBank Accession number | Forward Primers (5′-3′) | Reverse Primers (5′-3′) |
|---|---|---|---|
| ThMYC1 | JN166785 | AGGCTTAATGACAAGTTTGTGG | AGCGTATGCGGCTGGCATTGTT |
| ThMYC2 | JN166786 | GTATCCGGATATAGTTGAGCAG | GGATGCCATCAAGAGTTGATG |
| ThMYC3 | JN166787 | AATGGTAGTGGTAGAGTCGGTG | CTCATCATTAGGTCCTGACGAT |
| ThMYC4 | JN166788 | GAAGCGATTGAGGGAAGATGAT | CTTCACATACTCCACTGCTTCT |
| ThMYC5 | JN166789 | TTGAGTGGAAGCGTTGATGGGT | TATCACTAATTCTTGTCCTTCG |
| ThMYC6 | JN166790 | TGAGTACTTGGTAGCTAGCTCT | ATCATCATCATCAGAACCACTG |
| ThMYC7 | JN166792 | TAGGAACCGAAGTCTGGATCCT | GTATAGGTATACATACCAGAGT |
| ThMYC8 | JN166793 | ATGAAAACTCTTTACTCACAGC | TCCGACCCTACGCGTATGTGTC |
| ThMYC9 | JN166794 | TCAAGCTACTGATAGCCACAGT | TCGAATGTAGTTGGAGCAAGCT |
| Actin | FJ618517 | AAACAATGGCTGATGCTG | ACAATACCGTGCTCAATAGG |
| α-tubulin | FJ618518 | CACCCACCGTTGTTCCAG | ACCGTCGTCATCTTCACC |
| β-tubulin | FJ618519 | GGAAGCCATAGAAAGACC | CAACAAATGTGGGATGCT |
| Gene | cDNA Length (bp) | Mature Protein | ||
|---|---|---|---|---|
| Amino acid length | MW (kDa) | pI | ||
| ThMYC1 | 717 | 238 | 26.42 | 7.67 |
| ThMYC2 | 1479 | 492 | 55.06 | 4.77 |
| ThMYC4 | 483 | 160 | 17.89 | 5.96 |
| ThMYC5 | 810 | 269 | 28.00 | 6.30 |
| ThMYC9 | 906 | 301 | 31.69 | 6.19 |
| Relative abundance | |||
|---|---|---|---|
| Gene | Roots | Stem | Leaves |
| ThMYC1 | 511,603.5 | 9.257782 | 12.73741 |
| ThMYC2 | 5248.417 | 2.206867 | 5.107675 |
| ThMYC3 | 1 | 1 | 1 |
| ThMYC4 | 1629.259 | 0.466624 | 0.877619 |
| ThMYC5 | 321,620.5 | 2.7007 | 2.800466 |
| ThMYC6 | 396,510.6 | 2.046748 | 3.219121 |
| ThMYC7 | 85,284.74 | 1.304352 | 1.171752 |
| ThMYC8 | 31,916.17 | 0.666803 | 1.085731 |
| ThMYC9 | 14,050.09 | 1.613051 | 2.249142 |
© 2012 by the authors; licensee Molecular Diversity Preservation International, Basel, Switzerland. This article is an open-access article distributed under the terms and conditions of the Creative Commons Attribution license (http://creativecommons.org/licenses/by/3.0/).
Share and Cite
Ji, X.; Wang, Y.; Liu, G. Expression Analysis of MYC Genes from Tamarix hispida in Response to Different Abiotic Stresses. Int. J. Mol. Sci. 2012, 13, 1300-1313. https://doi.org/10.3390/ijms13021300
Ji X, Wang Y, Liu G. Expression Analysis of MYC Genes from Tamarix hispida in Response to Different Abiotic Stresses. International Journal of Molecular Sciences. 2012; 13(2):1300-1313. https://doi.org/10.3390/ijms13021300
Chicago/Turabian StyleJi, Xiaoyu, Yucheng Wang, and Guifeng Liu. 2012. "Expression Analysis of MYC Genes from Tamarix hispida in Response to Different Abiotic Stresses" International Journal of Molecular Sciences 13, no. 2: 1300-1313. https://doi.org/10.3390/ijms13021300




