XRCC3 Gene Polymorphism Is Associated with Survival in Japanese Lung Cancer Patients
Abstract
:1. Introduction
2. Results
3. Discussion
4. Experimental Section
4.1. Study Subjects
4.2. Genotyping
4.3. Statistical Analysis
5. Conclusions
- Conflict of InterestThe authors declare no conflict of interest.
Abbreviations
| NER | nucleotide excision repair |
| BER | base excision repair |
| DSBR | double-strand break repair |
| OGG1 | 8-oxoguanine DNA glycosylase |
| MUTYH/MYH | Mut Y homolog |
| APEX1/APE1 | apurinic/apyrimidinic endonuclease-1 |
| XRCC1 | X-ray repair cross-complementing group 1 |
| XRCC3 | X-ray repair cross-complementing group 3 |
| CI | confidence interval |
| NSCLC | non-small-cell lung cancer |
| SD | standard deviation |
| HR | hazard ratio |
| MST | median survival time |
References
- Pisani, P.; Parkin, D.M.; Ferlay, J. Estimates of the worldwide mortality from eighteen major cancers in 1985. Implications for prevention and projections of future burden. Int. J. Cancer 1993, 55, 891–903. [Google Scholar]
- Parkin, D.M.; Bray, F.; Ferlay, J.; Pisani, P. Global cancer statistics, 2002. CA Cancer J. Clin 2005, 55, 74108. [Google Scholar]
- Yu, Z.; Chen, J.; Ford, B.N.; Brackley, M.E.; Glickman, B.W. Human DNA repair systems: An overview. Environ. Mol. Mutagen 1999, 33, 3–20. [Google Scholar]
- Wood, R.D.; Mitchell, M.; Sgouros, J.; Lindahl, T. Human DNA repair genes. Science 2001, 291, 1284–1289. [Google Scholar]
- Le Marchand, L.; Donlon, T.; Lum-Jones, A.; Seifried, A.; Wilkens, L.R. Association of the hOGG1 Ser326Cys polymorphism with lung cancer risk. Cancer Epidemiol. Biomark. Prev 2002, 11, 409–412. [Google Scholar]
- Kasahara, M.; Osawa, K.; Yoshida, K.; Miyaishi, A.; Osawa, Y.; Inoue, N.; Tsutou, A.; Tabuchi, Y.; Tanaka, K.; Yamamoto, M.; et al. Association of MUTYH Gln324His and APEX1 Asp148Glu with colorectal cancer and smoking in a Japanese population. J. Exp. Clin. Cancer Res 2008, 27, 49. [Google Scholar]
- Miyaishi, A.; Osawa, K.; Osawa, Y.; Inoue, N.; Yoshida, K.; Kasahara, M.; Tsutou, A.; Tabuchi, Y.; Sakamoto, K.; Tsubota, N.; et al. MUTYH Gln324His gene polymorphism and genetic susceptibility for lung cancer in a Japanese population. J. Exp. Clin. Cancer Res 2009, 28, 10. [Google Scholar]
- Bennett, R.A.; Wilson, D.M., III; Wong, D.; Demple, B. Interaction of human apurinic endonuclease and DNA polymerase beta in the base excision repair pathway. Proc. Natl. Acad. Sci. USA 1997, 94, 7166–7169. [Google Scholar]
- Osawa, K.; Miyaishi, A.; Uchino, K.; Osawa, Y.; Inoue, N.; Nakarai, C.; Tsutou, A.; Kido, Y.; Yoshimura, M.; Tsubota, N.; et al. APEX1 Asp148Glu gene polymorphism is a risk factor for lung cancer in relation to smoking in Japanese. Asian Pac. J. Cancer Prev 2010, 11, 1181–1186. [Google Scholar]
- Matullo, G.; Palli, D.; Peluso, M.; Guarrera, S.; Carturan, S.; Celentano, E.; Krogh, V.; Munnia, A.; Tumino, R.; Polidoro, S.; et al. XRCC1, XRCC3, XPD gene polymorphisms, smoking and (32)P-DNA adducts in a sample of healthy subjects. Carcinogenesis 2001, 22, 1437–1445. [Google Scholar]
- Gurubhagavatula, S.; Liu, G.; Park, S.; Zhou, W.; Su, L.; Wain, J.C.; Lynch, T.J.; Neuberg, D.S.; Christiani, D.C. XPD and XRCC1 genetic polymorphisms are prognostic factors in advanced non-small-cell lung cancer patients treated with platinum chemotherapy. J. Clin. Oncol 2004, 22, 2594–2601. [Google Scholar]
- Ryu, J.S.; Hong, Y.C.; Han, H.S.; Lee, J.E.; Kim, S.; Park, Y.M.; Kim, Y.C.; Hwang, T.S. Association between polymorphisms of ERCC1 and XPD and survival in non-small-celllung cancer patients treated with cisplatin combination chemotherapy. Lung Cancer 2004, 44, 311–316. [Google Scholar]
- Wei, Q.; Spitz, M.R. The role of DNA repair capacity in susceptibility to lung cancer: A review. Cancer Metastasis Rev 1997, 16, 295–307. [Google Scholar]
- Osawa, K. SNPs in ERCC1 and drug response to cisplatin in non-small-cell lung cancer patients. Pharmacogenomics 2011, 12, 445–447. [Google Scholar]
- Osawa, K. Gene Polymorphisms and Chemotherapy in Non-small Cell Lung Cancer. Zhongguo Fei Ai Za Zhi 2009, 12, 837–840. [Google Scholar]
- Takenaka, T.; Yano, T.; Kiyohara, C.; Miura, N.; Kouso, H.; Ohba, T.; Kometani, T.; Shoji, F.; Yoshino, I.; Maehara, Y. Effects of excision repair cross-complementation group 1 (ERCC1) single nucleotide polymorphisms on the prognosis of non-small cell lung cancer patients. Lung Cancer 2010, 67, 101–107. [Google Scholar]
- Wu, X.; Shell, S.M.; Yang, Z.; Zou, Y. Phosphorylation of nucleotide excision repair factor xeroderma pigmentosum group A by ataxia telangiectasia mutated and Rad3-related-dependent checkpoint pathway promotes cell survival in response to UV irradiation. Cancer Res 2006, 66, 2997–3005. [Google Scholar]
- Sreeja, L.; Syamala, V.S.; Syamala, V.; Hariharan, S.; Raveendran, P.B.; Vijayalekshmi, R.V.; Madhavan, J.; Ankathil, R. Prognostic importance of DNA repair gene polymorphisms of XRCC1 Arg399Gln and XPD Lys751Gln in lung cancer patients from India. J. Cancer Res. Clin. Oncol 2008, 134, 645–652. [Google Scholar]
- Hsieh, W.C.; Cheng, Y.W.; Lin, C.J.; Chou, M.C.; Chen, C.Y.; Lee, H. Prognostic significance of X-ray cross-complementing group 1 T-77C polymorphism in resected non-small cell lung cancer. Jpn. J. Clin. Oncol 2009, 39, 81–85. [Google Scholar]
- Liao, W.Y.; Shih, J.Y.; Chang, G.C.; Cheng, Y.K.; Yang, J.C.; Chen, Y.M.; Yu, C.J. Genetic polymorphism of XRCC1 Arg399Gln is associated with survival in non-small-cell lung cancer patients treated with gemcitabine/platinum. J. Thorac. Oncol 2012, 7, 973–981. [Google Scholar]
- Yin, Z.; Zhou, B.; He, Q.; Li, M.; Guan, P.; Li, X.; Cui, Z.; Xue, X.; Su, M.; Ma, R.; et al. Association between polymorphisms in DNA repair genes and survival of non-smoking female patients with lung adenocarcinoma. BMC Cancer 2009, 9, 439. [Google Scholar]
- De las Peñas, R.; Sanchez-Ronco, M.; Alberola, V.; Taron, M.; Camps, C.; Garcia-Carbonero, R.; Massuti, B.; Queralt, C.; Botia, M.; Garcia-Gomez, R.; et al. Spanish Lung Cancer Group. Polymorphisms in DNA repair genes modulate survival in cisplatin/gemcitabine-treated non-small-cell lung cancer patients. Ann. Oncol 2006, 17, 668–675. [Google Scholar]
- Butkiewicz, D.; Rusin, M.; Sikora, B.; Lach, A.; Chorąży, M. An association between DNA repair gene polymorphisms and survival in patients with resected non-small cell lung cancer. Mol. Biol. Rep 2011, 38, 5231–5241. [Google Scholar]
- Ahrendt, S.A.; Hu, Y.; Buta, M.; McDermott, M.P.; Benoit, N.; Yang, S.C.; Wu, L.; Sidransky, D. p53 mutations and survival in stage I non-small-cell lung cancer: Results of a prospective study. J. Natl. Cancer Inst 2003, 95, 961–970. [Google Scholar]
- Osawa, Y.; Osawa, K.; Miyaishi, A.; Higuchi, M.; Tsutou, A.; Matsumura, S.; Tabuchi, Y.; Tsubota, N.; Takahashi, J. NAT2 and CYP1A2 polymorphisms and lung cancer risk in relation to smoking status. Asian Pac. J. Cancer Prev 2007, 8, 103–108. [Google Scholar]
| Variable | Patients n | Median Survival (Months) | Log-rank p value | Adjusted HR (95% CI) | p value |
|---|---|---|---|---|---|
| Gender | |||||
| Male | 65 | 52 | - | 1.00 | - |
| Female | 34 | 81 | 0.001 | 0.32 (0.16–0.65) | 0.001 |
| Age (years) | |||||
| <65 | 36 | 78 | 1.00 | ||
| ≥65 | 63 | 52 | 0.002 | 2.75 (1.41–5.36) | 0.003 |
| Histological subtype | |||||
| adenocarcinoma | 65 | 67 | - | 1.00 | - |
| squamous cell carcinoma | 29 | 49 | 0.020 | 1.93 (1.10–3.40) | 0.023 |
| others | 5 | 41 | - | - | - |
| Smoking status | |||||
| Non-smokers (Pack-years = 0) | 31 | 80 | - | 1.00 | - |
| Smokers (Pack-years > 0) | 67 | 53 | 0.002 | 3.02 (1.47–6.22) | 0.003 |
| No information | 1 | - | - | - | - |
| Stage | |||||
| I & II | 74 | 69 | - | 1.00 | - |
| III & IV | 20 | 41 | 0.001 | 2.60 (1.42–4.75) | 0.002 |
| No information | 5 | 15 | - | - | - |
| T stage | |||||
| T1 | 39 | 82 | - | 1.00 | - |
| T2&T3&T4 | 55 | 49 | <0.0001 | 3.68 (1.88–7.23) | <0.0001 |
| No information | 5 | 15 | - | - | - |
| Lymph node metastasis | |||||
| N0 | 66 | 68 | - | 1.00 | - |
| N1&N2 | 28 | 49 | 0.030 | 1.87 (1.05–3.33) | 0.034 |
| No information | 5 | 15 | - | - | - |
| Distant metastasis | |||||
| M0 | 89 | 66 | - | 1.00 | - |
| M1 | 5 | 50 | 0.783 | 1.18 (0.37–3.79) | 0.784 |
| No information | 5 | 14 | - | - | - |
| Recurrence | |||||
| No | 50 | 83 | 1.00 | ||
| Yes | 44 | 43 | <0.0001 | 6.15 (3.09–12.24) | <0.0001 |
| No information | 5 | 4 | - | - | - |
| Genotype | Patients n | MST (mon) | Log-rank p value | HR (95% CI) | Adjusted HR | ||
|---|---|---|---|---|---|---|---|
| p value | (95% CI) a | p value | |||||
| OGG1 | |||||||
| Ser/Ser | 25 | 58 | 1.00 | - | 1.00 | - | |
| Ser/Cys | 50 | 70 | 0.910 | 0.88 (0.45–1.72) | 0.712 | 0.96 (0.44–2.10) | 0.927 |
| Cys/Cys | 24 | 63 | 0.99 (0.46–2.10) | 0.972 | 0.90 (0.40–2.06) | 0.808 | |
| Ser/Cys, Cys/Cys | 74 | 66 | 0.784 | 0.92 (0.49–1.72) | 0.786 | 0.94 (0.47–1.89) | 0.854 |
| MUTYH | |||||||
| Gln/Gln | 20 | 58 | 1.00 | - | 1.00 | - | |
| Gln/His | 53 | 63 | 0.914 | 1.13 (0.55–2.33) | 0.731 | 0.96 (0.42–2.17) | 0.919 |
| His/His | 26 | 70 | 1.02 (0.45–2.32) | 0.972 | 0.85 (0.33–2.18) | 0.851 | |
| Gln/His, His/His | 79 | 63 | 0.797 | 1.09 (0.55–2.18) | 0.798 | 0.93 (0.42–2.03) | 0.845 |
| APEX | |||||||
| Asp/Asp | 40 | 58 | 1.00 | - | 1.00 | - | |
| Asp/Glu | 48 | 62 | 0.649 | 0.88 (0.50–1.55) | 0.652 | 1.02 (0.55–1.88) | 0.963 |
| Glu/Glu | 11 | 71 | 0.61 (0.21–1.78) | 0.37 | 0.51 (0.15–1.76) | 0.289 | |
| Asp/Glu, Glu/Glu | 59 | 64 | 0.505 | 0.83 (0.48–1.44) | 0.508 | 0.91 (0.50–1.66) | 0.766 |
| XRCC1 | |||||||
| Arg/Arg | 44 | 62 | - | 1.00 | - | 1.00 | - |
| Arg/Gln | 49 | 56 | 0.162 | 1.20 (0.69–2.08) | 0.524 | 0.85 (0.46–1.55) | 0.588 |
| Gln/Gln | 6 | 91 | 0.22 (0.03–1.62) | 0.136 | 0.37 (0.05–2.87) | 0.342 | |
| Arg/Gln, Gln/Gln | 55 | 60 | 0.897 | 1.04 (0.60–1.79) | 0.898 | 0.80 (0.44–1.46) | 0.470 |
| XRCC3 | |||||||
| Thr/Thr | 88 | 66 | - | 1.00 | - | 1.00 | |
| Thr/Met | 11 | 28 | 0.202 | 1.67 (0.75–3.72) | 0.209 | 1.94 (0.83–4.53) | 0.128 |
| Met/Met | 0 | - | - | - | - | - | - |
| Thr/Met, Met/Met | 11 | 28 | 0.202 | 1.67 (0.75–3.72) | 0.209 | 1.94 (0.83–4.53) | 0.128 |
| Genotype | Patients n | MST (mon) | Log-rank p value | HR (95% CI) | Adjusted HR | ||
|---|---|---|---|---|---|---|---|
| p value | (95% CI) a | p value | |||||
| Adenocarcinoma | |||||||
| OGG1 | |||||||
| Ser/Ser | 16 | 62 | - | 1.00 | - | 1.00 | - |
| Ser/Cys, Cys/Cys | 49 | 68 | 0.784 | 0.89 (0.38–2.08) | 0.785 | 0.80 (0.29–2.26) | 0.803 |
| MUTYH | |||||||
| Gln/Gln | 13 | 70 | 1.00 | 1.00 | |||
| Gln/His, His/His | 52 | 66 | 0.65 | 1.25 (0.48–3.27) | 0.653 | 1.54 (0.44–5.36) | 0.498 |
| APEX | |||||||
| Asp/Asp | 28 | 63 | - | 1.00 | - | 1.00 | - |
| Asp/Glu, Glu/Glu | 37 | 70 | 0.549 | 0.80 (0.39–1.67) | 0.553 | 1.14 (0.48–2.73) | 0.763 |
| XRCC1 | |||||||
| Arg/Arg | 26 | 71 | 1.00 | - | 1.00 | - | |
| Arg/Gln, Gln/Gln | 39 | 64 | 0.522 | 1.28 (0.60–2.76) | 0.526 | 0.87 (0.35–2.20) | 0.775 |
| XRCC3 | |||||||
| Thr/Thr | 58 | 67 | - | 1.00 | - | 1.00 | - |
| Thr/Met, Met/Met | 7 | 64 | 0.995 | 1.00 (0.30–3.32) | 0.995 | 1.32 (0.38–4.64) | 0.661 |
| Squamous Cell Carcinoma | |||||||
| OGG1 | |||||||
| Ser/Ser | 8 | 52 | - | 1.00 | - | 1.00 | - |
| Ser/Cys, Cys/Cys | 21 | 48 | 0.824 | 1.11 (0.43–2.88) | 0.825 | 1.25 (0.44–3.55) | 0.670 |
| MUTYH | |||||||
| Gln/Gln | 5 | 64 | - | 1.00 | - | 1.00 | - |
| Gln/His, His/His | 24 | 45 | 0.377 | 1.72 (0.51–5.85) | 0.385 | 4.73 (0.51–5.85) | 0.153 |
| APEX | |||||||
| Asp/Asp | 11 | 40 | 1.00 | - | 1.00 | - | |
| Asp/Glu, Glu/Glu | 18 | 54 | 0.328 | 0.65 (0.27–1.56) | 0.334 | 0.43 (0.14–1.32) | 0.139 |
| XRCC1 | |||||||
| Arg/Arg | 16 | 50 | 1.00 | - | 1.00 | - | |
| Arg/Gln, Gln/Gln | 13 | 46 | 0.991 | 1.00 (0.42–2.37) | 0.991 | 1.17 (0.46–2.94) | 0.743 |
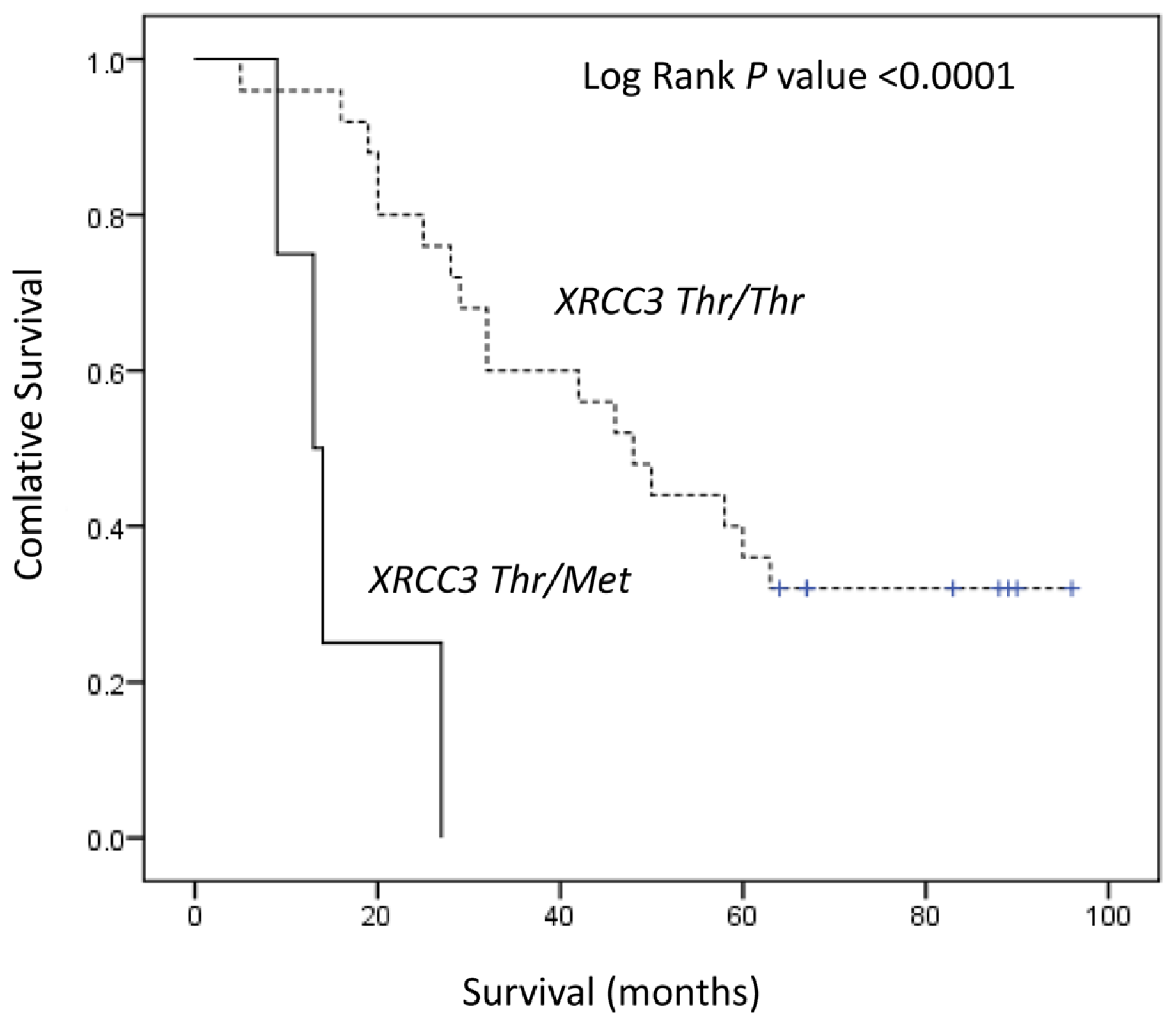
| XRCC3 | |||||||
| Thr/Thr | 25 | 48 | - | 1.00 | - | 1.00 | - |
| Thr/Met, Met/Met | 4 | 13 | <0.0001 | 9.35 (2.52–34.68) | 0.001 | 9.05 (1.89–44.39) | 0.006 |
© 2012 by the authors; licensee Molecular Diversity Preservation International, Basel, Switzerland. This article is an open-access article distributed under the terms and conditions of the Creative Commons Attribution license (http://creativecommons.org/licenses/by/3.0/).
Share and Cite
Osawa, K.; Nakarai, C.; Uchino, K.; Yoshimura, M.; Tsubota, N.; Takahashi, J.; Kido, Y. XRCC3 Gene Polymorphism Is Associated with Survival in Japanese Lung Cancer Patients. Int. J. Mol. Sci. 2012, 13, 16658-16667. https://doi.org/10.3390/ijms131216658
Osawa K, Nakarai C, Uchino K, Yoshimura M, Tsubota N, Takahashi J, Kido Y. XRCC3 Gene Polymorphism Is Associated with Survival in Japanese Lung Cancer Patients. International Journal of Molecular Sciences. 2012; 13(12):16658-16667. https://doi.org/10.3390/ijms131216658
Chicago/Turabian StyleOsawa, Kayo, Chiaki Nakarai, Kazuya Uchino, Masahiro Yoshimura, Noriaki Tsubota, Juro Takahashi, and Yoshiaki Kido. 2012. "XRCC3 Gene Polymorphism Is Associated with Survival in Japanese Lung Cancer Patients" International Journal of Molecular Sciences 13, no. 12: 16658-16667. https://doi.org/10.3390/ijms131216658




