Individual Variations in Inorganic Arsenic Metabolism Associated with AS3MT Genetic Polymorphisms
Abstract
:1. Introduction
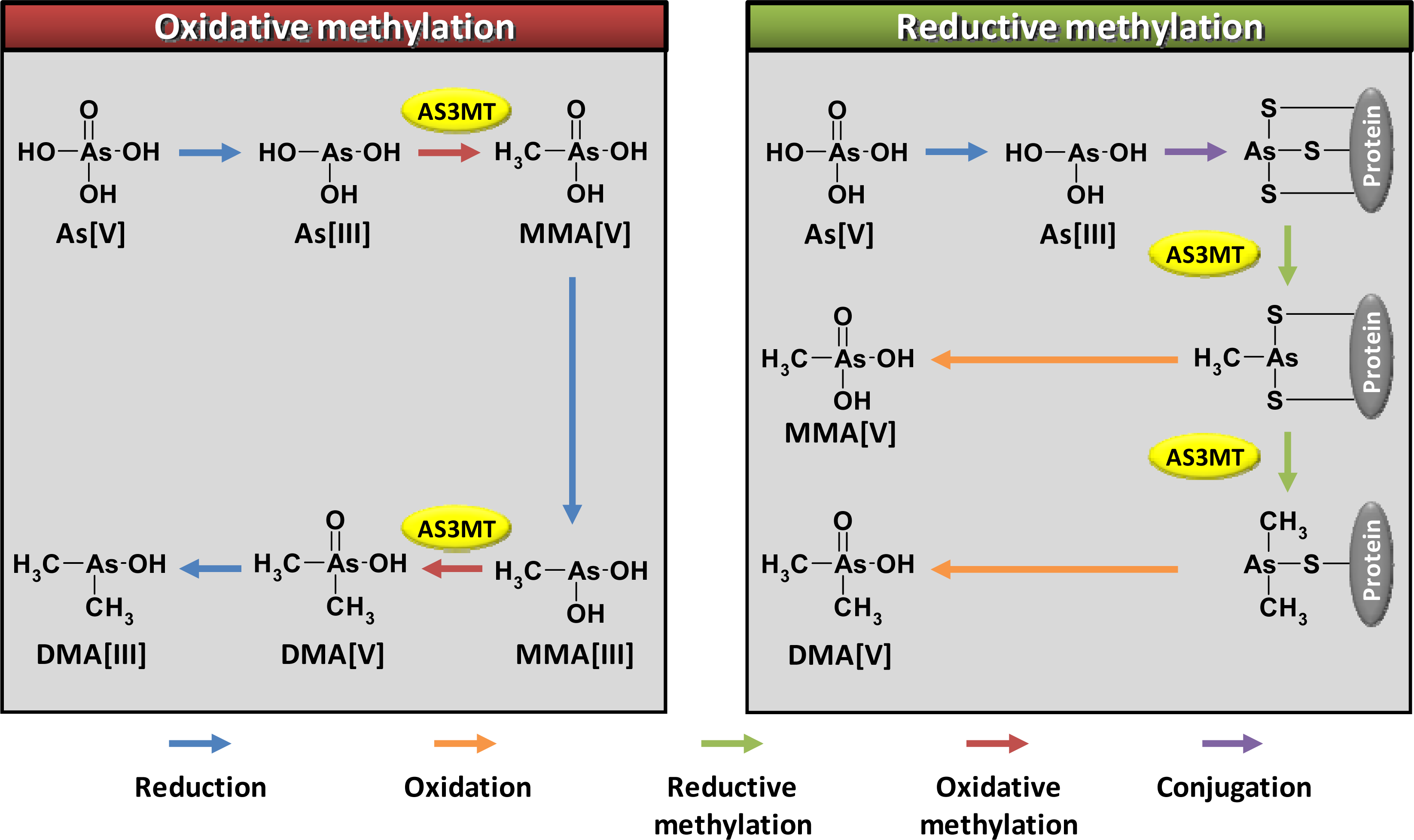
2. Arsenic Methylation by Human AS3MT
3. Association of Arsenic Methylation with Genotypes in Human AS3MT
3.1. In Vitro Studies
3.2. Human Case Studies
3.2.1. Mexico
3.2.2. Argentina
3.2.3. Chile
3.2.4. Central Europe
3.2.5. India
3.2.6. Taiwan
3.2.7. Vietnam
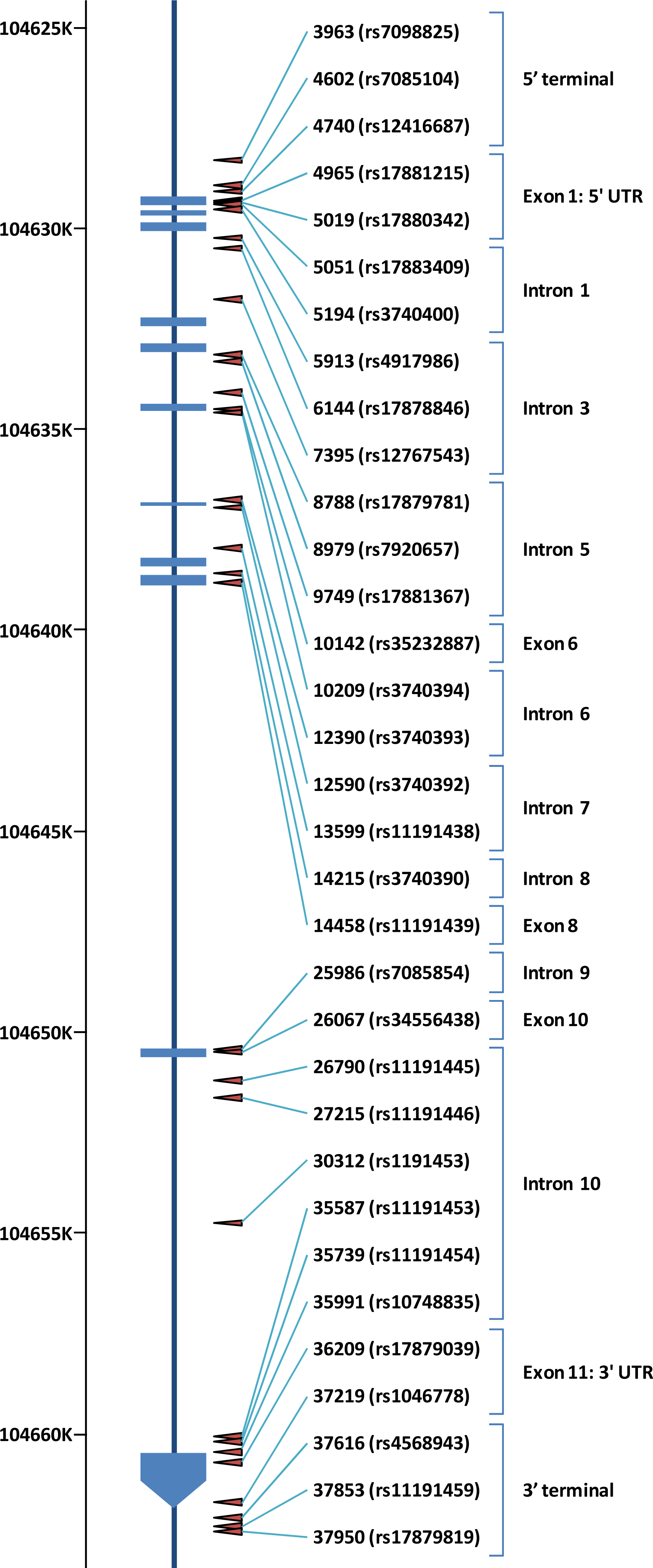
4. Allele Frequency of SNPs in AS3MT
5. Conclusions and Future Perspectives
Acknowledgments
References
- Guidelines for Drinking Water Quality, 3rd ed; World Health Organization: Geneva, Switzerland, 2004.
- Mandal, BK; Suzuki, KT. Arsenic round the world: A review. Talanta 2002, 58, 201–235. [Google Scholar]
- Nordstrom, DK. Public health. Worldwide occurrences of arsenic in ground water. Science 2002, 296, 2143–2145. [Google Scholar]
- Smedley, PL; Kinniburgh, DG. A review of the source, behavior and distribution of arsenic in natural waters. Appl. Geochem 2002, 17, 517–568. [Google Scholar]
- Wu, MM; Kuo, TL; Hwang, YH; Chen, CJ. Dose-response relation between arsenic concentration in well water and mortality from cancers and vascular diseases. Am. J. Epidemiol 1989, 130, 1123–1132. [Google Scholar]
- Tondel, M; Rahman, M; Magnuson, A; Chowdhury, IA; Faruquee, MH; Ahmad, SA. The relationship of arsenic levels in drinking water and the prevalence rate of skin lesions in Bangladesh. Environ. Health Perspect 1999, 107, 727–729. [Google Scholar]
- Vahter, M. Variation in human metabolism of arsenic. In Arsenic Exposure and Health Effects; Chappell, WR, Abernathy, CO, Calderon, RL, Eds.; Elsevier Science, Ltd: Oxford, UK, 1999; pp. 267–279. [Google Scholar]
- Concha, G; Nermell, B; Vahter, MV. Metabolism of inorganic arsenic in children with chronic high arsenic exposure in northern Argentina. Environ. Health Perspect 1998, 106, 355–359. [Google Scholar]
- Del Razo, LM; Garcia-Vargas, GG; Vargas, H; Albores, A; Gonsebatt, ME; Montero, R; Ostrosky-Wegman, P; Kelsh, M; Cebrian, ME. Altered profile of urinary arsenic metabolites in adults with chronic arsenicism. A pilot study. Arch. Toxicol 1997, 71, 211–217. [Google Scholar]
- Valenzuela, OL; Borja-Aburto, VH; Garcia-Vargas, GG; Cruz-Gonzalez, MB; Garcia-Montalvo, EA; Calderon-Aranda, ES; Del Razo, LM. Urinary trivalent methylated arsenic species in a population chronically exposed to inorganic arsenic. Environ. Health Perspect 2005, 113, 250–254. [Google Scholar]
- Steinmaus, C; Bates, MN; Yuan, Y; Kalman, D; Atallah, R; Rey, OA; Biggs, ML; Hopenhayn, C; Moore, LE; Hoang, BK; Smith, AH. Arsenic methylation and bladder cancer risk in case-control studies in Argentina and the United States. J. Occup. Environ. Med 2006, 48, 478–488. [Google Scholar]
- Tseng, CH. Arsenic methylation, urinary arsenic metabolites and human diseases: Current perspective. J. Environ. Sci. Health C 2007, 25, 1–22. [Google Scholar]
- Vahter, M. Methylation of inorganic arsenic in different mammalian species and population groups. Sci. Progr. London 1999, 82, 69–88. [Google Scholar]
- Tseng, CH. A review on environmental factors regulating arsenic methylation in humans. Toxicol. Appl. Pharmacol 2009, 235, 338–350. [Google Scholar]
- Vahter, M; Concha, G; Nermell, B; Nilsson, R; Dulout, F; Natarajan, AT. A unique metabolism of inorganic arsenic in native Andean women. Eur. J. Pharmacol 1995, 293, 455–462. [Google Scholar]
- Chiou, HY; Hsueh, YM; Hsieh, LL; Hsu, LI; Yi-Hsiang, H; Hsieh, FI; Wei, ML; Chen, HC; Yang, HT; Leu, LC; et al. Arsenic methylation capacity, body retention, and null genotypes of glutathione S-transferase M1 and T1 among current arsenic-exposed residents in Taiwan. Mutat. Res 1997, 386, 197–207. [Google Scholar]
- Agusa, T; Fujihara, J; Takeshita, H; Iwata, H; Minh, TB; Trang, PTK; Viet, PH; Tanabe, S. Relationship between genetic polymorphism of arsenic (+3 oxidation state) methyltransferase (AS3MT) and profiles of urinary arsenic compounds in Vietnamese. Biomed. Res. Trace Elem 2009, 19, 265–267. [Google Scholar]
- Agusa, T; Iwata, H; Fujihara, J; Kunito, T; Takeshita, H; Minh, TB; Trang, PTK; Viet, PH; Tanabe, S. Genetic polymorphisms in AS3MT and arsenic metabolism in residents of the Red River Delta, Vietnam. Toxicol. Appl. Pharmacol 2009, 236, 131–141. [Google Scholar]
- Agusa, T; Iwata, H; Fujihara, J; Kunito, T; Takeshita, H; Tu, BM; Pham, TKT; Pham, HV; Tanabe, S. Genetic polymorphisms in glutathione S-transferase (GST) superfamily and arsenic metabolism in residents of the Red River Delta, Vietnam. Toxicol. Appl. Pharmacol 2010, 242, 352–362. [Google Scholar]
- Agusa, T; Iwata, H; Fujihara, J; Kunito, T; Takeshita, H; Minh, TB; Trang, PTK; Viet, PH; Tanabe, S. Interindividual variation in arsenic metabolism in a Vietnamese population: Association with 17 single nucleotide polymorphisms in AS3MT. In Interdisciplinary Studies on Environmental Chemistry Vol 3 Biological Responses to Chemical Contaminants: From Molecular to Community Level; Hamamura, N, Suzuki, S, Mendo, S, Barroso, CM, Iwata, H, Tanabe, S, Eds.; TERRAPUB: Tokyo, Japan, 2010; pp. 113–119. [Google Scholar]
- Agusa, T; Kunito, T; Kubota, R; Inoue, S; Fujihara, J; Minh, TB; Ha, NN; Tu, NPC; Trang, PTK; Chamnan, C; et al. Exposure, metabolism, and health effects of arsenic in residents from arsenic-contaminated groundwater areas of Vietnam and Cambodia: A review. Rev. Environ. Health 2010, 25, 193–220. [Google Scholar]
- Hernández, A; Marcos, R. Genetic variations associated with interindividual sensitivity in the response to arsenic exposure. Pharmacogenomics 2008, 9, 1113–1132. [Google Scholar]
- Challenger, F. Biological methylation. Chem. Rev 1945, 36, 315–361. [Google Scholar]
- Cullen, WR; Reimer, KJ. Arsenic speciation in the environment. Chem. Rev 1989, 89, 713–764. [Google Scholar]
- Petrick, JS; Ayala-Fierro, F; Cullen, WR; Carter, DE; Vasken Aposhian, H. Monomethylarsonous acid (MMAIII) is more toxic than arsenite in Chang human hepatocytes. Toxicol. Appl. Pharmacol 2000, 163, 203–207. [Google Scholar]
- Styblo, M; Del Razo, LM; Vega, L; Germolec, DR; LeCluyse, EL; Hamilton, GA; Reed, W; Wang, C; Cullen, WR; Thomas, DJ. Comparative toxicity of trivalent and pentavalent inorganic and methylated arsenicals in rat and human cells. Arch. Toxicol 2000, 74, 289–299. [Google Scholar]
- Hayakawa, T; Kobayashi, Y; Cui, X; Hirano, S. A new metabolic pathway of arsenite: Arsenic-glutathione complexes are substrates for human arsenic methyltransferase Cyt19. Arch. Toxicol 2005, 79, 183–191. [Google Scholar]
- Naranmandura, H; Suzuki, N; Suzuki, KT. Trivalent arsenicals are bound to proteins during reductive methylation. Chem. Res. Toxicol 2006, 19, 1010–1018. [Google Scholar]
- Lin, S; Shi, Q; Nix, FB; Styblo, M; Beck, MA; Herbin-Davist, KM; Hall, LL; Simeonsson, JB; Thomas, DJ. A novel S-adenosyl-L-methionine: Arsenic(III) methyltransferase from rat liver cytosol. J. Bio. Chem 2002, 277, 10795–10803. [Google Scholar]
- Wood, TC; Salavagionne, OE; Mukherjee, B; Wang, LW; Klumpp, AF; Thomae, BA; Eckloff, BW; Schaid, DJ; Wieben, ED; Weinshilboum, RM. Human arsenic methyltransferase (AS3MT) pharmacogenetics—Gene resequencing and functional genomics studies. J. Bio. Chem 2006, 281, 7364–7373. [Google Scholar]
- HapMap Public Release #28; National Center for Biotechnology Information: Bethesda, MD, USA, 2010. Available online: http://hapmap.ncbi.nlm.nih.gov/cgi-perl/gbrowse/hapmap28_B36/ (accessed on 10 March 2011).
- Kojima, C; Ramirez, DC; Tokar, EJ; Himeno, S; Drobná, Z; Stýblo, M; Mason, RP; Waalkes, MP. Requirement of arsenic biomethylation for oxidative DNA damage. J. Nat. Cancer Inst 2009, 101, 1670–1681. [Google Scholar]
- NCBI Reference Assembly; National Center for Biotechnology Information: Bethesda, MD, USA, 2010. Available online: http://www.ncbi.nlm.nih.gov/projects/SNP/ (accessed on 10 March 2011).
- Drobna, Z; Waters, SB; Walton, FS; LeCluyse, EL; Thomas, DJ; Styblo, M. Interindividual variation in the metabolism of arsenic in cultured primary human hepatocytes. Toxicol. Appl. Pharmacol 2004, 201, 166–177. [Google Scholar]
- Meza, MM; Yu, LZ; Rodriguez, YY; Guild, M; Thompson, D; Gandolfi, AJ; Klimecki, WT. Developmentally restricted genetic determinants of human arsenic metabolism: Association between urinary methylated arsenic and CYT19 polymorphisms in children. Environ. Health Perspect 2005, 113, 775–781. [Google Scholar]
- Meza, M; Gandolfi, AJ; Klimecki, WT. Developmental and genetic modulation of arsenic biotransformation: A gene by environment interaction? Toxicol. Appl. Pharmacol 2007, 222, 381–387. [Google Scholar]
- Valenzuela, OL; Drobna, Z; Hernandez-Castellanos, E; Sanchez-Pena, LC; Garcia-Vargas, GG; Borja-Aburto, VH; Styblo, M; Del Razo, LM. Association of AS3MT polymorphisms and the risk of premalignant arsenic skin lesions. Toxicol. Appl. Pharmacol 2009, 239, 200–207. [Google Scholar]
- Gomez-Rubio, P; Meza-Montenegro, MM; Cantu-Soto, E; Klimecki, WT. Genetic association between intronic variants in AS3MT and arsenic methylation efficiency is focused on a large linkage disequilibrium cluster in chromosome 10. J. Appl. Toxicol 2010, 30, 260–270. [Google Scholar]
- Sampayo-Reyes, A; Hernández, A; El-Yamani, N; López-Campos, C; Mayet-Machado, E; Rincón-Castañeda, CB; Limones-Aguilar, ML; López-Campos, JE; de León, MB; González-Hernández, S; Hinojosa-Garza, D; Marcos, R. Arsenic induces DNA damage in environmentally exposed Mexican children and adults. Influence of GSTO1 and AS3MT Polymorphisms. Toxicol. Sci 2010, 117, 63–71. [Google Scholar]
- Schläwicke Engström, K; Broberg, K; Concha, G; Nermell, B; Warholm, M; Vahter, M. Genetic polymorphisms influencing arsenic metabolism: Evidence from Argentina. Environ. Health Perspect 2007, 115, 599–605. [Google Scholar]
- Núñez, ALN; Hill, HG; Martinez, AL. Guía Museo Arqueológico. In Universidad Católica del Norte Chile San Pedro de Atacama (II Regíon): Instituto de Investigaciones; Arqueológicas y Museo, RP, Gustavo Le Paige, SJ, Eds.; Universidad Católica del Norte Chile: Antofagasta, Chile, 1991. [Google Scholar]
- Figueroa, LT; Razmilic, BB; González, MU. Corporal distribution of arsenic in mummied bodies owned to an arsenical habitat. In International Seminar Proceedings: Arsenic in the Environment and Its Incidence on Health; Sancha, AM, Ed.; Universidad de Chile, Facultad de Ciencias Fisicas y Matematicas: Santiago, Chile, 1992; pp. 77–82. [Google Scholar]
- Schläwicke Engström, K; Nermell, B; Concha, G; Stromberg, U; Vahter, M; Broberg, K. Arsenic metabolism is influenced by polymorphisms in genes involved in one-carbon metabolism and reduction reactions. Mutat. Res 2009, 667, 4–14. [Google Scholar]
- Hernandez, A; Xamena, N; Surralles, J; Sekaran, C; Tokunaga, H; Quinteros, D; Creus, A; Marcos, R. Role of the Met(287)Thr polymorphism in the AS3MT gene on the metabolic arsenic profile. Mutat. Res 2008, 637, 80–92. [Google Scholar]
- Hernandez, A; Xamena, N; Sekaran, C; Tokunaga, H; Sampayo-Reyes, A; Quinteros, D; Creus, A; Marcos, R. High arsenic metabolic efficiency in AS3MT(287)Thr allele carriers. Pharmacogenet. Genomics 2008, 18, 349–355. [Google Scholar]
- Lindberg, AL; Kumar, R; Goessler, W; Thirumaran, R; Gurzau, E; Koppova, K; Rudnai, P; Leonardi, G; Fletcher, T; Vahter, M. Metabolism of low-dose inorganic arsenic in a Central Europena population: Influence of sex and genetic polymorphisms. Environ. Health Perspect 2007, 115, 1081–1086. [Google Scholar]
- Chakraborti, D; Rahman, MM; Paul, K; Chowdhury, UK; Sengupta, MK; Lodh, D; Chanda, CR; Saha, KC; Mukherjee, SC. Arsenic calamity in the Indian subcontinent: What lessons have been learned? Talanta 2002, 58, 3–22. [Google Scholar]
- De Chaudhuri, S; Ghosh, P; Sarma, N; Majumdar, P; Sau, TJ; Basu, S; Roychoudhury, S; Ray, K; Giri, AK. Genetic variants associated with arsenic susceptibility: Study of purine nucleoside phosphorylase, arsenic (+3) methyltransferase, and glutathione S-transferase omega genes. Environ. Health Perspect 2008, 116, 501–505. [Google Scholar]
- Chung, CJ; Hsueh, YM; Bai, CH; Huang, YK; Huang, YL; Yang, MH; Chen, CJ. Polymorphisms in arsenic metabolism genes, urinary arsenic methylation profile and cancer. Canc. Causes Contr 2009, 20, 1653–1661. [Google Scholar]
- Berg, M; Tran, HC; Nguyen, TC; Pham, HV; Schertenleib, R; Giger, W. Arsenic contamination of groundwater and drinking water in Vietnam: A human health threat. Environ. Sci. Technol 2001, 35, 2621–2626. [Google Scholar]
- Nga, TTV; Inoue, M; Khatiwada, NR; Takizawa, S. Heavy metal tracers for the analysis of groundwater contamination: Case study in Hanoi City, Vietnam. Water Sci. Tech. Water Sup 2003, 3, 343–350. [Google Scholar]
- Trang, PTK; Berg, M; Viet, PH; Van Mui, N; Van der Meer, JR. Bacterial bioassay for rapid and accurate analysis of arsenic in highly variable groundwater samples. Environ. Sci. Tech 2005, 39, 7625–7630. [Google Scholar]
- Berg, M; Luzi, S; Trang, PTK; Viet, PH; Giger, W; Stuben, D. Arsenic removal from groundwater by household sand filters: Comparative field study, model calculations, and health benefits. Environ. Sci. Tech 2006, 40, 5567–5573. [Google Scholar]
- Shinkai, Y; Van Truc, D; Sumi, D; Canh, D; Kumagai, Y. Arsenic and other metal contamination of groundwater in the Mekong River Delta, Vietnam. J. Health Sci 2007, 53, 344–346. [Google Scholar]
- Buschmann, J; Berg, M; Stengel, C; Winkel, L; Sampson, ML; Trang, PTK; Viet, PH. Contamination of drinking water resources in the Mekong delta floodplains: Arsenic and other trace metals pose serious health risks to population. Environ. Int 2008, 34, 756–764. [Google Scholar]
- Norrman, J; Sparrenbom, CJ; Berg, M; Nhan, DD; Nhan, PQ; Rosqvist, H; Jacks, G; Sigvardsson, E; Baric, D; Moreskog, J; Harms-Ringdahl, P; Van Hoan, N. Arsenic mobilisation in a new well field for drinking water production along the Red River, Nam Du, Hanoi. Appl. Geochem 2008, 23, 3127–3142. [Google Scholar]
- Agusa, T; Kunito, T; Fujihara, J; Kubota, R; Minh, TB; Trang, PTK; Iwata, H; Subramanian, A; Viet, PH; Tanabe, S. Contamination by arsenic and other trace elements in tube-well water and its risk assessment to humans in Hanoi, Vietnam. Environ. Pollut 2006, 139, 95–106. [Google Scholar]
- Agusa, T; Inoue, S; Kunito, T; Minh, TB; Ha, NN; Tu, NPC; Trang, PTK; Iwata, H; Viet, PH; Tuyen, BC; Tanabe, S. Human exposure to arsenic from groundwater in the Red River and the Mekong River Deltas in Vietnam. Int. J. Environ. Stud 2009, 66, 49–57. [Google Scholar]
- Agusa, T; Kunito, T; Minh, TB; Trang, PTK; Iwata, H; Viet, PH; Tanabe, S. Relationship of urinary arsenic metabolites to intake estimates in residents of the Red River Delta, Vietnam. Environ. Pollut 2009, 157, 396–403. [Google Scholar]
- Agusa, T; Kunito, T; Kubota, R; Inoue, S; Fujihara, J; Minh, TB; Ha, NN; Tu, NPC; Trang, PTK; Chamnan, C; et al. Exposure, metabolism, and health effects of arsenic in residents from arsenic-contaminated groundwater areas of Vietnam and Cambodia: A review. Rev. Environ. Health 2010, 25, 193–220. [Google Scholar]
- Nguyen, VA; Bang, S; Viet, PH; Kim, KW. Contamination of groundwater and risk assessment for arsenic exposure in Ha Nam province, Vietnam. Environ. Int 2009, 35, 466–472. [Google Scholar]
- Fujihara, J; Kunito, T; Agusa, T; Yasuda, T; Iida, R; Fujii, Y; Takeshita, H. Population differences in the human arsenic (+3 oxidation state) methyltransferase (AS3MT) gene polymorphism detected by using genotyping method. Toxicol. Appl. Pharmacol 2007, 225, 251–254. [Google Scholar]
- Fujihara, J; Soejima, M; Koda, Y; Kunito, T; Takeshita, H. Asian specific low mutation frequencies of the M287T polymorphism in the human arsenic (+3 oxidation state) methyltransferase (AS3MT) gene. Mutat. Res.-Genet. Toxicol. E. M 2008, 654, 158–161. [Google Scholar]
- Fujihara, J; Fujii, Y; Agusa, T; Kunito, T; Yasuda, T; Moritani, T; Takeshita, H. Ethnic differences in five intronic polymorphisms associated with arsenic metabolism within human arsenic (+3 oxidation state) methyltransferase (AS3MT) gene. Toxicol. Appl. Pharmacol 2009, 234, 41–46. [Google Scholar]
- Fujihara, J; Soejima, M; Yasuda, T; Koda, Y; Agusa, T; Kunito, T; Tongu, M; Yamada, T; Takeshita, H. Global analysis of genetic variation in human arsenic (+3 oxidation state) methyltransferase (AS3MT). Toxicol. Appl. Pharmacol 2010, 243, 292–299. [Google Scholar]
- Zhu, J; Chen, Z; Lallemand-Breitenbach, V; De Thé, H. How acute promyclocytic leukaemia revived arsenic. Nat. Rev. Cancer 2002, 2, 705–713. [Google Scholar]


| SNP ID | rs # | Location | Nucleotide change | Ancestral | Allele | Japanese (n = 141) | South Koreans (n = 230) | Chinese (n = 54) | Mongolians (n = 58) | Tibetans (n = 65) | Nepalese (n = 31) |
|---|---|---|---|---|---|---|---|---|---|---|---|
| 3963 | rs7098825 | 5′ terminal | C/T | T | C | 0.266 | 0.298 | 0.241 | 0.009 | 0.331 | 0.355 |
| T | 0.734 | 0.702 | 0.759 | 0.991 | 0.669 | 0.645 | |||||
| 4602 | rs7085104 | 5′ terminal | A/G | A | A | 0.567 | 0.565 | 0.436 | 0.672* | 0.454 | 0.694 |
| G | 0.433 | 0.435 | 0.565 | 0.327* | 0.546 | 0.307 | |||||
| 4740 | rs12416687 | 5′ terminal | C/T | T | C | 0.060 | 0.067 | 0.130 | 0.061 | 0.054 | 0.000 |
| T | 0.940 | 0.933 | 0.871 | 0.940 | 0.946 | 1.000 | |||||
| 5913 | rs4917986 | Intron 3 | C/T | T | C | NA | NA | NA | NA | NA | NA |
| T | NA | NA | NA | NA | NA | NA | |||||
| 6144 | rs17878846 | Intron 3 | A/T | A | A | 0.734 | 0.730 | 0.695* | 0.931 | 0.708* | 0.678 |
| T | 0.266 | 0.270 | 0.306* | 0.069 | 0.292* | 0.323 | |||||
| 7395 | rs12767543 | Intron 3 | A/G | G | A | 0.287 | 0.274* | 0.426 | 0.069 | 0.185 | 0.275 |
| G | 0.713 | 0.727* | 0.574 | 0.931 | 0.815 | 0.726 | |||||
| 8979 | rs7920657 | Intron 5 | A/T | T | A | 0.057 | 0.076 | 0 | 0.077 | 0.069 | 0.000 |
| T | 0.944 | 0.924 | 1.000 | 0.922 | 0.931 | 1.000 | |||||
| 9749 | rs17881367 | Intron 5 | A/G | A | A | 0.944 | 0.924 | 1.000 | 0.922* | 0.931 | 1.000 |
| G | 0.057 | 0.076 | 0 | 0.077* | 0.069 | 0.000 | |||||
| 10209 | rs3740394 | Intron 6 | A/G | T | A | 0.990 | 0.985 | 1.000 | 0.983 | 0.985 | 0.984 |
| G | 0.011 | 0.015 | 0 | 0.017 | 0.016 | 0.016 | |||||
| 12390 | rs3740393 | Intron 6 | C/G | C | C | 0.373* | 0.279 | 0.278 | 0.431 | 0.338 | 0.339 |
| G | 0.628* | 0.722 | 0.722 | 0.569 | 0.662 | 0.661 | |||||
| 12590 | rs3740392 | Intron 7 | C/T | T | C | 0.178* | 0.172 | 0.251 | 0.215 | 0.185 | 0.162 |
| T | 0.823* | 0.829 | 0.751 | 0.784 | 0.816 | 0.839 | |||||
| 14215 | rs3740390 | Intron 8 | C/T | C | C | 0.483* | 0.587* | 0.343 | 0.587* | 0.262 | 0.645 |
| T | 0.518* | 0.413* | 0.658 | 0.414* | 0.739 | 0.355 | |||||
| 144581 | rs11191439 | Exon 9 | C/T | T | C | 0.004 | 0.009 | 0 | 0.026 | 0.023 | 0.000 |
| T | 0.997 | 0.992 | 1.000 | 0.974 | 0.977 | 1.000 | |||||
| 25986 | rs7085854 | Intron 8 | C/T | T | C | 0.057 | 0.122 | 0.259 | 0.052 | 0.162 | 0.145 |
| T | 0.944 | 0.879 | 0.741 | 0.949 | 0.839 | 0.855 | |||||
| 26790 | rs11191445 | Intron 10 | C/T | C | C | 0.990* | 0.985 | 1.000 | 0.983 | 0.977 | 0.968 |
| T | 0.011* | 0.015 | 0 | 0.017 | 0.023 | 0.033 | |||||
| 27215 | rs11191446 | Intron 10 | A/G | A | A | 0.926 | 0.918 | 0.972 | 0.949 | 0.900 | 0.952 |
| G | 0.075 | 0.083 | 0.028 | 0.052 | 0.100 | 0.049 | |||||
| 35587 | rs11191453 | Intron 10 | C/T | T | C | 0.344* | 0.370* | 0.333 | 0.155* | 0.516* | 0.404 |
| T | 0.656* | 0.631* | 0.666 | 0.845* | 0.485* | 0.597 | |||||
| 35991 | rs10748835 | Intron 10 | A/G | A | A | 0.543 | 0.570 | 0.602 | 0.561 | 0.654 | 0.597 |
| G | 0.457 | 0.431 | 0.398 | 0.440 | 0.346 | 0.404 | |||||
| 37616 | rs4568943 | 3′ terminal | A/C | A | A | 0.656* | 0.735 | 0.519* | 0.906* | 0.639 | 0.678 |
| C | 0.344* | 0.266 | 0.482* | 0.095* | 0.362 | 0.323 | |||||
| 37853 | rs11191459 | 3′ terminal | A/G | G | A | NA | NA | NA | NA | NA | NA |
| G | NA | NA | NA | NA | NA | NA | |||||
| 37950 | rs17879819 | 3′ terminal | C/T | C | C | 0.710 | 0.713 | 0.676 | 0.923 | 0.638 | 0.629 |
| T | 0.291 | 0.287 | 0.324 | 0.078 | 0.361 | 0.371 | |||||
| References | [65] | [65] | [65] | [65] | [65] | [65] |
| SNP ID | rs # | Location | Nucleotide change | Ancestral | Allele | Vietnamese (n = 100) | Sri Lanka-Tamils (n = 29) | Sri Lanka-Sinhalese (n = 30) | Ovambos (n = 102) | Ghanaians (n = 87) |
|---|---|---|---|---|---|---|---|---|---|---|
| 3963 | rs7098825 | 5′ terminal | C/T | T | C | NA | 0.086 | 0.117 | 0.054 | 0.121 |
| T | NA | 0.914 | 0.883 | 0.946 | 0.879 | |||||
| 4602 | rs7085104 | 5′ terminal | A/G | A | A | 0.420 | 0.845 | 0.817 | 0.525 | 0.598 |
| G | 0.580 | 0.155 | 0.184 | 0.475 | 0.403 | |||||
| 4740 | rs12416687 | 5′ terminal | C/T | T | C | 0 | 0.017 | 0.017 | 0.133 | 0.104 |
| T | 0.720 | 0.983 | 0.984 | 0.868 | 0.897 | |||||
| 5913 | rs4917986 | Intron 3 | C/T | T | C | 0.080 | NA | NA | NA | NA |
| T | 0.920 | NA | NA | NA | NA | |||||
| 6144 | rs17878846 | Intron 3 | A/T | A | A | 0.750 | 0.845 | 0.867 | 1.000 | 1.000 |
| T | 0.250 | 0.155 | 0.134 | 0 | 0 | |||||
| 7395 | rs12767543 | Intron 3 | A/G | G | A | 0.335* | 0.121 | 0.167 | 0.113 | 0.304 |
| G | 0.665* | 0.880 | 0.834 | 0.888 | 0.695 | |||||
| 8979 | rs7920657 | Intron 5 | A/T | T | A | 0 | 0.035 | 0.017 | 0.142 | 0.075 |
| T | 0.590 | 0.966 | 0.984 | 0.858 | 0.926 | |||||
| 9749 | rs17881367 | Intron 5 | A/G | A | A | NA | 0.966 | 0.984 | 0.858 | 0.926 |
| G | NA | 0.035 | 0.017 | 0.142 | 0.075 | |||||
| 10209 | rs3740394 | Intron 6 | A/G | T | A | NA | 0.983 | 0.984 | 0.868 | 0.920 |
| G | NA | 0.017 | 0.017 | 0.133 | 0.081 | |||||
| 12390 | rs3740393 | Intron 6 | C/G | C | C | 0.225 | 0.189 | 0.251 | 0.540 | 0.299 |
| G | 0.775 | 0.810 | 0.751 | 0.461 | 0.702 | |||||
| 12590 | rs3740392 | Intron 7 | C/T | T | C | 0.360 | 0.155 | 0.084 | 0.221 | 0.189 |
| T | 0.640 | 0.845 | 0.917 | 0.780 | 0.810 | |||||
| 14215 | rs3740390 | Intron 8 | C/T | C | C | 0.755 | 0.810 | 0.867 | 0.554 | 0.414* |
| T | 0.245 | 0.189 | 0.134 | 0.446 | 0.587* | |||||
| 144581 | rs11191439 | Exon 9 | C/T | T | C | 0 | 0 | 0.017 | 0.093 | 0.075 |
| T | 0.980 | 1.000 | 0.984 | 0.907 | 0.926 | |||||
| 25986 | rs7085854 | Intron 8 | C/T | T | C | NA | 0.172 | 0.250 | 0.299 | 0.316 |
| T | NA | 0.828 | 0.750 | 0.701 | 0.684 | |||||
| 26790 | rs11191445 | Intron 10 | C/T | C | C | NA | 0.983 | 0.984 | 0.814 | 0.966 |
| T | NA | 0.017 | 0.017 | 0.186 | 0.035 | |||||
| 27215 | rs11191446 | Intron 10 | A/G | A | A | NA | 0.966 | 0.984 | 0.873 | 0.920 |
| G | NA | 0.035 | 0.017 | 0.128 | 0.081 | |||||
| 35587 | rs11191453 | Intron 10 | C/T | T | C | 0.270* | 0.207 | 0.134 | 0.045 | 0.063 |
| T | 0.730* | 0.793 | 0.867 | 0.957 | 0.937 | |||||
| 35991 | rs10748835 | Intron 10 | A/G | A | A | 0.585 | 0.535 | 0.367 | 0.706 | 0.402 |
| G | 0.415 | 0.466 | 0.633 | 0.294 | 0.598 | |||||
| 37616 | rs4568943 | 3′ terminal | A/C | A | A | NA | 0.794 | 0.867 | 0.941* | 0.931 |
| C | NA | 0.207 | 0.134 | 0.059* | 0.069 | |||||
| 37853 | rs11191459 | 3′ terminal | A/G | G | A | 0.495 | NA | NA | NA | NA |
| G | 0.505 | NA | NA | NA | NA | |||||
| 37950 | rs17879819 | 3′ terminal | C/T | C | C | NA | 0.793 | 0.867 | 0.961* | 0.937 |
| T | NA | 0.207 | 0.134 | 0.040* | 0.063 | |||||
| References | [20] | [65] | [65] | [65] | [65] |
| SNP ID | rs # | Location | Nucleotide change | Ancestral | Mexico (n = 144) | Mexico (n = 122) | Mexico (n = 405) | Argentina (n = 147) |
|---|---|---|---|---|---|---|---|---|
| 3963 | rs7098825 | 5′ terminal | C/T | T | ||||
| 4602 | rs7085104 | 5′ terminal | A/G | A | NS | %MMA[III + V]: AA + AG > GG %DMA[III + V]: GG > AA + AG DMA[III + V]/MMA[III + V]: GG > AA + AG | ||
| 4740 | rs12416687 | 5′ terminal | C/T | T | NS | |||
| 4965 | rs17881215 | Exon 1: 5′ UTR | C/G | G | ||||
| 5019 | rs17880342 | Exon 1: 5′ UTR | C/T | C | NA | |||
| 5051 | rs17883409 | Intron 1 | * | NAV | NS | NA | ||
| 5194 | rs3740400 | Intron 1 | A/C | C | NA | |||
| 5913 | rs4917986 | Intron 3 | C/T | T | ||||
| 6144 | rs17878846 | Intron 3 | A/T | A | ||||
| 7395 | rs12767543 | Intron 3 | A/G | G | DMA[V]/MMA[V]: AA + AG > GG2 | S | ||
| 8788 | rs17881367 | Intron 5 | -/TTT | NAV | NA | |||
| 8979 | rs7920657 | Intron 5 | A/T | T | ||||
| 9749 | rs17881367 | Intron 5 | A/G | A | ||||
| 10209 | rs3740394 | Intron 6 | C/T | T | NS | |||
| 12390 | rs3740393 | Intron 6 | C/G | C | DMA[V]/MMA[V]: CC + CG > GG2 | S | %MMA[V]: GG > CC, CG %DMA[V]: CC, CG > GG DMA[V]/MMA[V]: CC, CG > GG | |
| 12590 | rs3740392 | Intron 7 | C/T | T | NS | NA | ||
| 13599 | rs10883797 | Intron 7 | C/G | C | NS | |||
| 14215 | rs3740390 | Intron 8 | C/T | C | S | %MMA[V]: CC > TT, CT %DMA[V]: TT, CT > CC DMA[V]/MMA[V]: TT, CT > CC | ||
| 144581 | rs11191439 | Exon 9 | C/T | T | %MMA[III + V]: CT + CC > TT %As[V]: CT + CC > TT | NS | NA | |
| 25986 | rs7085854 | Intron 9 | C/T | T | NS | |||
| 27215 | rs11191446 | Intron 10 | A/G | A | ||||
| 30312 | rs1191453 | Intron 10 | C/T | T | DMA[V]/MMA[V]: CC > CT > TT | |||
| 35587 | rs11191453 | Intron 10 | C/T | T | DMA[V]/MMA[V]: CC + CT > TT2 MMA[V]/As[III]: TT > CC + CT2 | NS | S | |
| 35739 | rs11191454 | Intron 10 | A/G | A | S | |||
| 35991 | rs10748835 | Intron 10 | A/G | A | %MMA[V]: GG > AA, AG %DMA[V]: AA, AG > GG DMA[V]/MMA[V]: AA, AG > GG | |||
| 36209 | rs17879039 | Exon 11: 3′ UTR | A/G | G | ||||
| 37219 | rs1046778 | Exon 11: 3′ UTR | C/T | T | S | |||
| 37616 | rs4568943 | 3′ terminal | A/C | A | ||||
| 37853 | rs11191459 | 3′ terminal | A/G | G | ||||
| 37950 | rs17879819 | 3′ terminal | C/T | C | ||||
| LD cluster | ||||||||
| LD1 | 7395-12390-35587 | 7395-12390-14215-30312 -35587-35739-37219 | 12390-14215-35991 | |||||
| LD2 | ||||||||
| LD3 | ||||||||
| LD4 | ||||||||
| References | [35, 36] | [37] | [38] | [40] |
| SNP ID | rs # | Location | Nucleotide change | Ancestral | Argentina (n = 104) | Chile (n = 50) | Chile (n = 207) |
|---|---|---|---|---|---|---|---|
| 3963 | rs7098825 | 5′ terminal | C/T | T | |||
| 4602 | rs7085104 | 5′ terminal | A/G | A | %MMA[V]: AA + AG > GG %DMA[V]: GG > AG + AA | ||
| 4740 | rs12416687 | 5′ terminal | C/T | T | |||
| 4965 | rs17881215 | Exon 1: 5′ UTR | C/G | G | %MMA: CG > GG | ||
| 5019 | rs17880342 | Exon 1: 5′ UTR | C/T | C | |||
| 5051 | rs17883409 | Intron 1 | * | NAV | |||
| 5194 | rs3740400 | Intron 1 | A/C | C | %MMA[V]: AA + AC > CC %DMA[V]: CC > AC + AA | NS | |
| 5913 | rs4917986 | Intron 3 | C/T | T | |||
| 6144 | rs17878846 | Intron 3 | A/T | A | |||
| 7395 | rs12767543 | Intron 3 | A/G | G | |||
| 8788 | rs17881367 | Intron 5 | -/TTT | NAV | |||
| 8979 | rs7920657 | Intron 5 | A/T | T | |||
| 9749 | rs17881367 | Intron 5 | A/G | A | |||
| 10209 | rs3740394 | Intron 6 | C/T | T | |||
| 12390 | rs3740393 | Intron 6 | C/G | C | |||
| 12590 | rs3740392 | Intron 7 | C/T | T | |||
| 13599 | rs10883797 | Intron 7 | C/G | C | |||
| 14215 | rs3740390 | Intron 8 | C/T | C | |||
| 144581 | rs11191439 | Exon 9 | C/T | T | %MMA: CT + CC > TT | %MMA: CT + CC > TT | |
| 25986 | rs7085854 | Intron 9 | C/T | T | |||
| 27215 | rs11191446 | Intron 10 | A/G | A | |||
| 30312 | rs1191453 | Intron 10 | C/T | T | |||
| 35587 | rs11191453 | Intron 10 | C/T | T | |||
| 35739 | rs11191454 | Intron 10 | A/G | A | |||
| 35991 | rs10748835 | Intron 10 | A/G | A | |||
| 36209 | rs17879039 | Exon 11: 3′ UTR | A/G | G | NS | ||
| 37219 | rs1046778 | Exon 11: 3′ UTR | C/T | T | |||
| 37616 | rs4568943 | 3′ terminal | A/C | A | |||
| 37853 | rs11191459 | 3′ terminal | A/G | G | |||
| 37950 | rs17879819 | 3′ terminal | C/T | C | |||
| LD cluster | |||||||
| LD1 | 4602-5194-12390-14215-359913 | 4965-14458 | |||||
| LD2 | |||||||
| LD3 | |||||||
| LD4 | |||||||
| References | [43] | [44] | [45] |
| SNP ID | rs # | Location | Nucleotide change | Ancestral | Central Europe4 (n = 415) | Taiwan (n = 208) | Vietnam (n = 100) |
|---|---|---|---|---|---|---|---|
| 3963 | rs7098825 | 5′ terminal | C/T | T | NS | ||
| 4602 | rs7085104 | 5′ terminal | A/G | A | %DMA[V]: AA, AG > GG | ||
| 4740 | rs12416687 | 5′ terminal | C/T | T | %DMA[V]: TT > CT, CC | ||
| 4965 | rs17881215 | Exon 1: 5′ UTR | C/G | G | |||
| 5019 | rs17880342 | Exon 1: 5′ UTR | C/T | C | |||
| 5051 | rs17883409 | Intron 1 | * | NAV | |||
| 5194 | rs3740400 | Intron 1 | A/C | C | |||
| 5913 | rs4917986 | Intron 3 | C/T | T | %MMA[V]: CT > TT | ||
| 6144 | rs17878846 | Intron 3 | A/T | A | NS | ||
| 7395 | rs12767543 | Intron 3 | A/G | G | NS | ||
| 8788 | rs17881367 | Intron 5 | -/TTT | NAV | |||
| 8979 | rs7920657 | Intron 5 | A/T | T | %DMA[V]: AT, TT > AA | ||
| 9749 | rs17881367 | Intron 5 | A/G | A | %MMA[V]: AG > AA | ||
| 10209 | rs3740394 | Intron 6 | C/T | T | |||
| 12390 | rs3740393 | Intron 6 | C/G | C | %MMA[V]: GG > CG %DMA[V]: CG > GG DMA[V]/MMA[V]: CG > GG | %MMA[V]: GG > CC DMA[V]/MMA[V]: CG > GG | |
| 12590 | rs3740392 | Intron 7 | C/T | T | %DMA[V]: TT > CC DMA[V]/MMA[V]: TT > CC | ||
| 13599 | rs10883797 | Intron 7 | C/G | C | |||
| 14215 | rs3740390 | Intron 8 | C/T | C | NS | ||
| 144581 | rs11191439 | Exon 9 | C/T | T | %DMA[V]: TT > CT + CC %MMA[V]: CT + CC > TT | MMA[V]/IA: CT > TT | |
| 25986 | rs7085854 | Intron 9 | C/T | T | |||
| 27215 | rs11191446 | Intron 10 | A/G | A | %MMA[V]: AG > AA DMA[V]/MMA[V]: AA > AG MMA[V]/IA: AG > AA | ||
| 30312 | rs1191453 | Intron 10 | C/T | T | |||
| 35587 | rs11191453 | Intron 10 | C/T | T | DMA[V]/MMA[V]: CT > TT | ||
| 35739 | rs11191454 | Intron 10 | A/G | A | |||
| 35991 | rs10748835 | Intron 10 | A/G | A | %DMA[V]: AG > AA | ||
| 36209 | rs17879039 | Exon 11: 3′ UTR | A/G | G | |||
| 37219 | rs1046778 | Exon 11: 3′ UTR | C/T | T | |||
| 37616 | rs4568943 | 3′ terminal | A/C | A | |||
| 37853 | rs11191459 | 3′ terminal | A/G | G | %DMA[V]: GG, AG > AA %MMA[V]: GG > AA | ||
| 37950 | rs17879819 | 3′ terminal | C/T | C | NS | ||
| LD cluster | |||||||
| LD1 | 3963-6144-12390-14215-35587-37950 | ||||||
| LD2 | 4602-35991-37853 | ||||||
| LD3 | 4740-12590 | ||||||
| LD4 | 5913-9749-27215 | ||||||
| References | [46] | [49] | [18, 20] |
© 2011 by the authors; licensee MDPI, Basel, Switzerland. This article is an open-access article distributed under the terms and conditions of the Creative Commons Attribution license (http://creativecommons.org/licenses/by/3.0/).
Share and Cite
Agusa, T.; Fujihara, J.; Takeshita, H.; Iwata, H. Individual Variations in Inorganic Arsenic Metabolism Associated with AS3MT Genetic Polymorphisms. Int. J. Mol. Sci. 2011, 12, 2351-2382. https://doi.org/10.3390/ijms12042351
Agusa T, Fujihara J, Takeshita H, Iwata H. Individual Variations in Inorganic Arsenic Metabolism Associated with AS3MT Genetic Polymorphisms. International Journal of Molecular Sciences. 2011; 12(4):2351-2382. https://doi.org/10.3390/ijms12042351
Chicago/Turabian StyleAgusa, Tetsuro, Junko Fujihara, Haruo Takeshita, and Hisato Iwata. 2011. "Individual Variations in Inorganic Arsenic Metabolism Associated with AS3MT Genetic Polymorphisms" International Journal of Molecular Sciences 12, no. 4: 2351-2382. https://doi.org/10.3390/ijms12042351




