gef Gene Expression in MCF-7 Breast Cancer Cells is Associated with a Better Prognosis and Induction of Apoptosis by p53-Mediated Signaling Pathway
Abstract
:1. Introduction
2. Results and Discussion
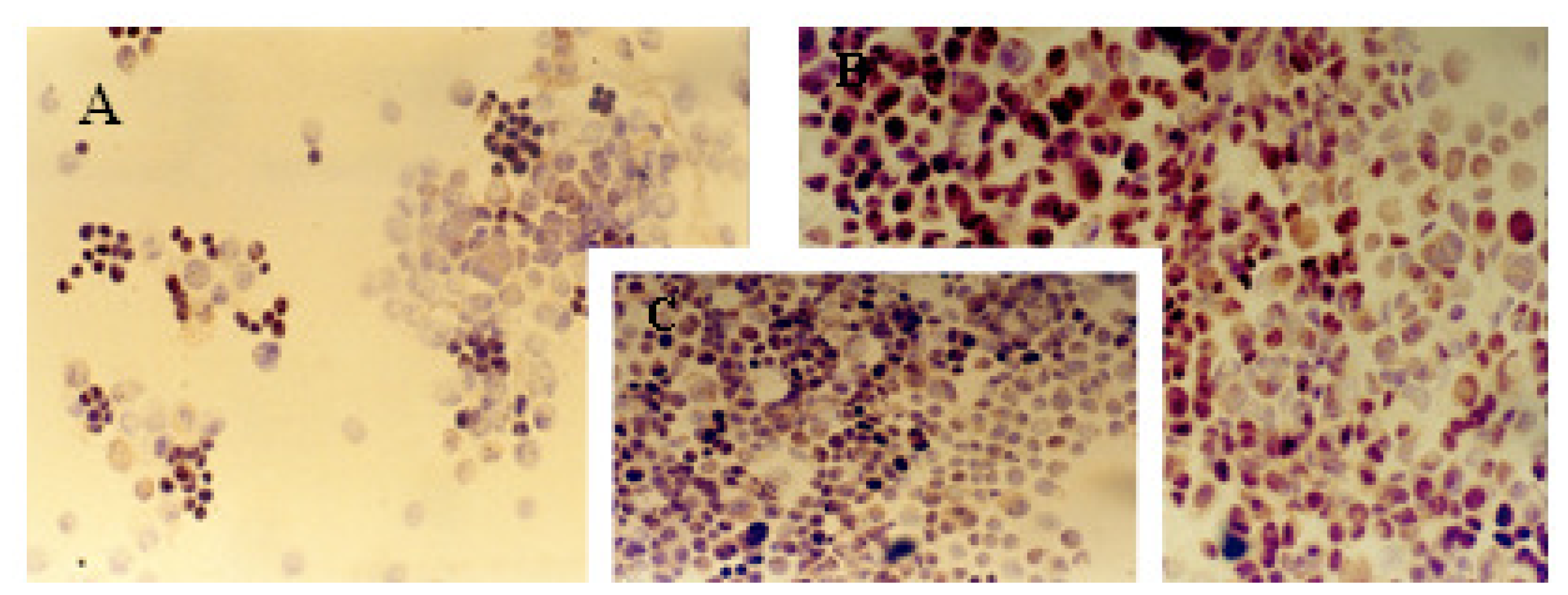
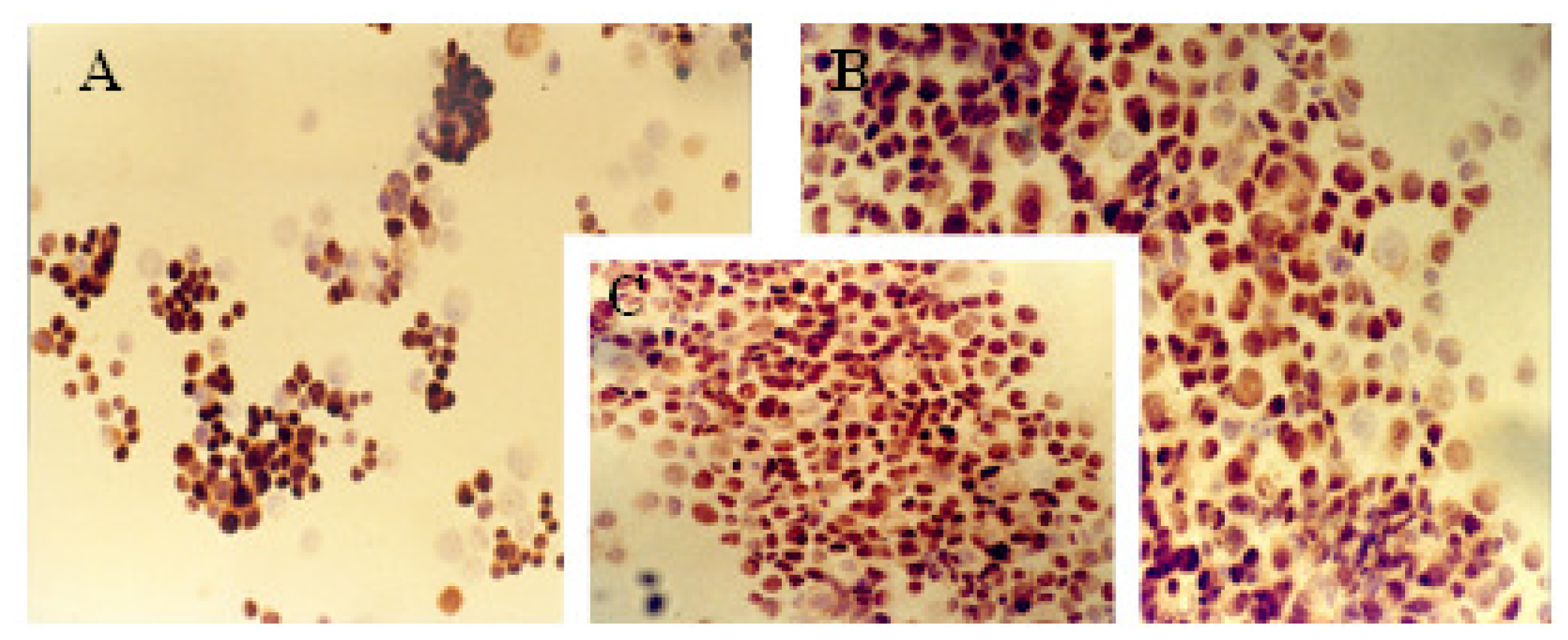
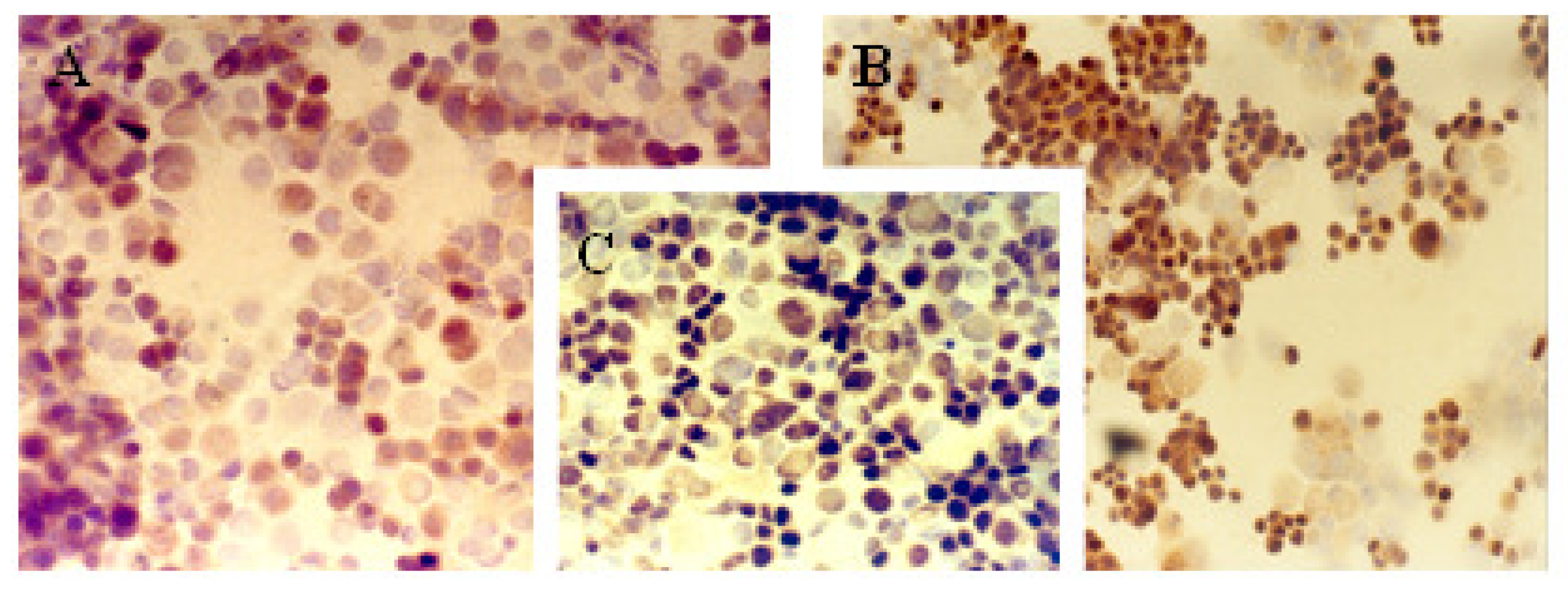
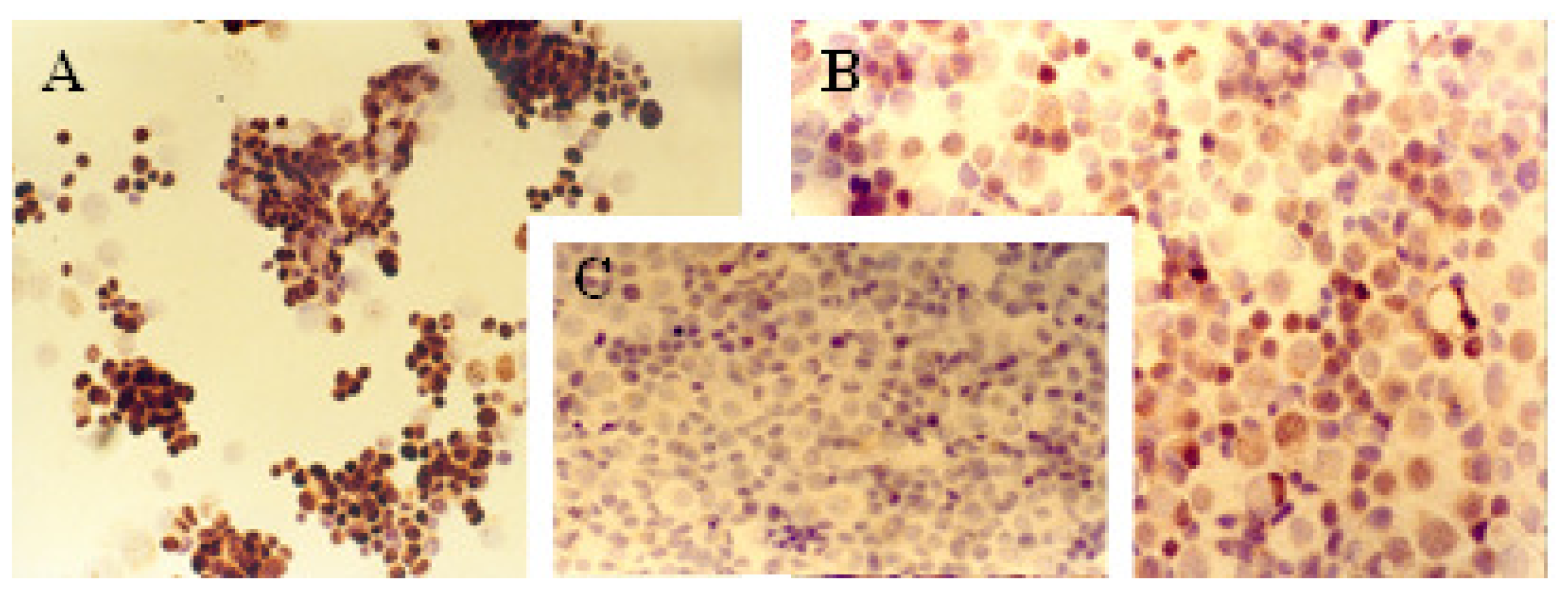
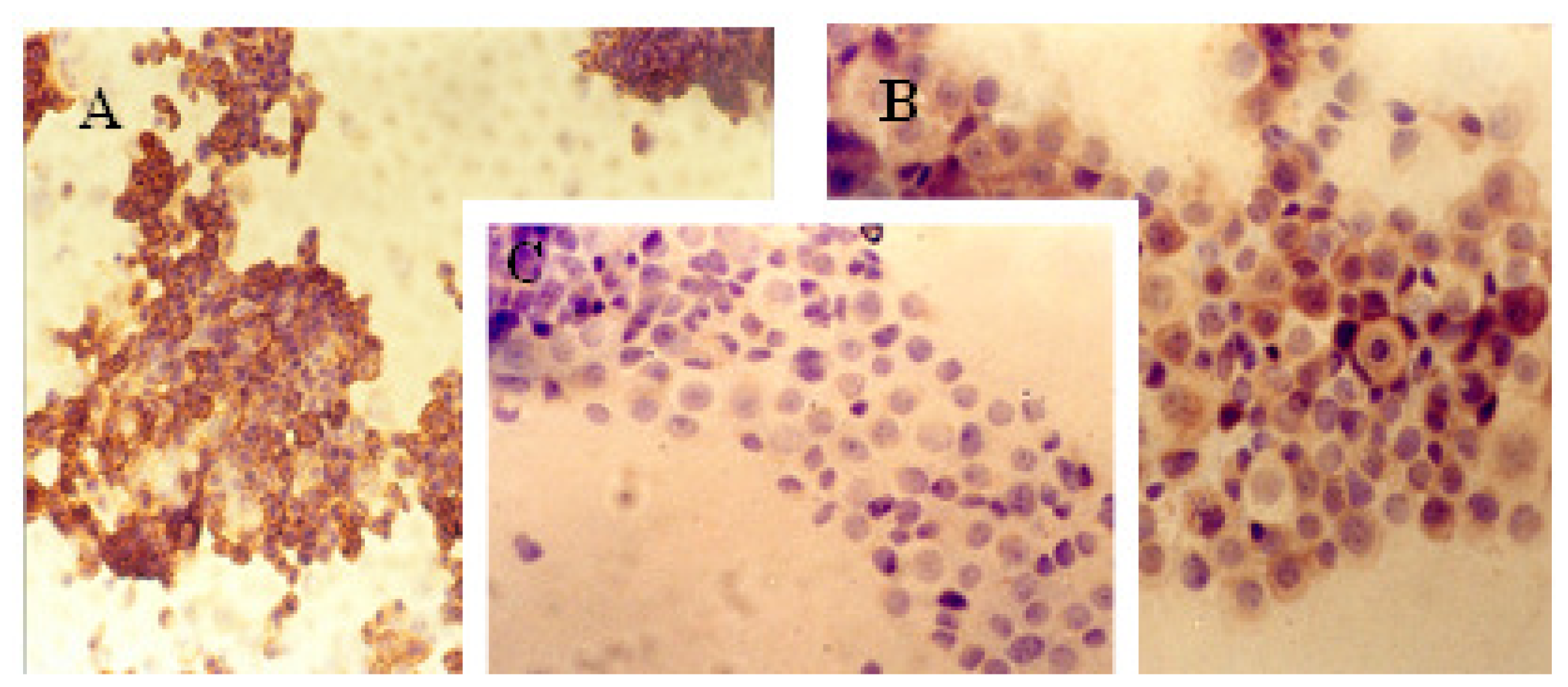
2.1. Immunocytochemical Analysis
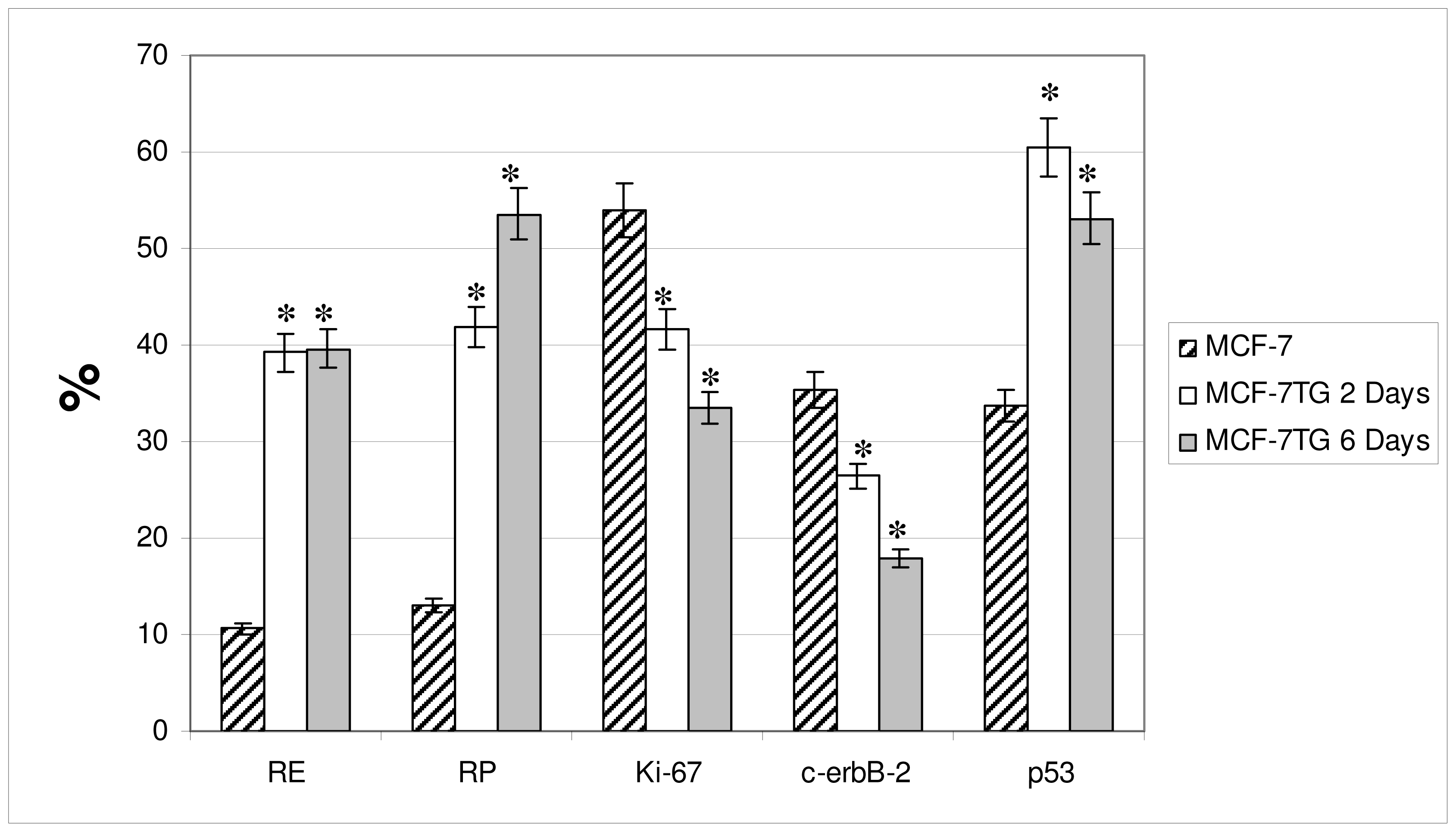
2.2. Indirect Immunofluorescence Analysis
Progesterone Receptor (PR)
Ki-67 Antigen
p53 Protein
c-erbB-2 Oncogene
3. Experimental Section
3.1. Cell Lines and Culture Conditions
3.2. Immunocytochemical Analysis
3.3. Indirect Immunofluorescence Analysis
3.4. Statistical Analysis
4. Conclusions
Acknowledgements
References
- Marasà, S.; Sciancalepore, G.; Marasà, L. Breast cancer less than 1 cm: Bio-morphologic characterization with ER, PgR, Ki67, Her-2/Neu, MDV, MAGS, p53, EGF-R. Pathologica 2008, 100, 156–161. [Google Scholar]
- Jovanovic, J.; Rønneberg, J.A.; Tost, J.; Kristensen, V. The epigenetics of breast cancer. Mol. Oncol 2010, 4, 242–254. [Google Scholar]
- Kurdistani, S.K. Histone modifications in cancer biology and prognosis. Prog. Drug Res 2011, 67, 91–106. [Google Scholar]
- Lee, E.Y.; Muller, W.J. Oncogenes and tumor suppressor genes. Cold Spring Harb. Perspect. Biol 2010, 2. [Google Scholar] [CrossRef]
- Green, D.R.; Kroemer, G. Cytoplasmic functions of the tumor suppressor p53. Nature 2009, 458, 1127–1130. [Google Scholar]
- Fisher, E.R.; Constantino, J.; Fisher, B.; Redmond, C. Pathologic findings from the national surgical adjuvant breast project (Protocol 4). Cancer 1993, 71, 2141–2150. [Google Scholar]
- Kröger, N.; Milde-Langosch, K.; Riethdorf, S.; Schmoor, C.; Schumacher, M.; Zander, A.R.; Löning, T. Prognostic and predictive effects of immunohistochemical factors in high-risk primary breast cancer patients. Clin. Cancer Res 2006, 12, 159–168. [Google Scholar]
- Bostrom, P.; Soderstrom, M.; Vahlberg, T.; Soderstrom, K.O.; Roberts, P.J.; Carpen, O.; Hirsimaki, P. MMP-1 expression has an independent prognostic value in breast cancer. BMC Cancer 2011, 11. [Google Scholar] [CrossRef]
- Ross, J.S.; Christos hatzis, W.; Symmans, F.; Lajos, G. Commercialized multigene predictors of clinical outcome for breast cancer. Oncologist 2008, 13, 477–493. [Google Scholar]
- Malamou-Mitsi, V.; Gogas, H.; Dafni, U.; Bourli, A.; Fillipides, T.; Sotiropoulou, M.; Vlachodimitropoulos, D.; Papadopoulos, S.; Tzaida, O.; Kafiri, G.; et al. Evaluation of the prognostic and predictive value of p53 and Bcl-2 in breast cancer patients participating in a randomized study with dose-dense sequential adjuvant chemotherapy. Ann. Oncol 2006, 17, 1504–1511. [Google Scholar]
- Ross, J.S.; Symmans, W.F; Pusztai, L.; Hortobagyi, G.N. Standardizing slide-based assays in breast cancer: Hormone receptors, HER2, and sentinel lymph nodes. Clin. Cancer Res 2007, 13, 2831–2835. [Google Scholar]
- Ring, B.Z.; Seitz, R.S.; Beck, R.; Shasteen, W.J.; Tarr, S.M.; Cheang, M.C.; Yoder, B.J.; Budd, G.T.; Nielsen, T.O.; Hicks, D.G.; et al. Novel prognostic immunohistochemical biomarker panel for estrogen receptor-positive breast cancer. J. Clin. Oncol 2006, 24, 3039–3047. [Google Scholar]
- Service, R.F. New role for estrogen in cancer? Science 1998, 279, 1631–1633. [Google Scholar]
- Osborne, C.K. Tamoxifen in the treatment of breast cancer. N. Engl. J. Med 1998, 339, 1609–1618. [Google Scholar]
- Rastelli, F.; Crispino, S. Factors predictive of response to hormone therapy in breast cancer. Tumori 2008, 94, 370–383. [Google Scholar]
- Mendelson, J.; Baselga, J. Status of epidermal growth factor receptor antagonist in biology and treatment of cancer. J. Clin. Oncol 2003, 21, 2787–2799. [Google Scholar]
- Jeziorski, A.; Blonski, J.Z.; Niewiadomska, H. The expression of products of oncogens c-erbB-2 and EGFR and proliferating antigens Ki67 and PCNA in primary invasive ductal cancer of female breast. J. Exp. Clin. Cancer Res 2000, 19, 61–67. [Google Scholar]
- Dowsett, M.; Smith, I.E.; Ebbs, S.R.; Dixon, J.M.; Skene, A.; A’Hern, R.; Salter, J.; Detre, S.; Hills, M.; Walsh, G. Prognostic value of Ki67 expression after short-term presurgical endocrine therapy for primary breast cancer. J. Natl. Cancer Inst 2007, 99, 167–170. [Google Scholar]
- Kerlikowske, K.; Molinaro, A.M.; Gauthier, M.L.; Berman, H.K.; Waldman, F.; Bennington, J.; Sanchez, H.; Jimenez, C.; Stewart, K.; Chew, K.; et al. Biomarker expression and risk of subsequent tumors after initial ductal carcinoma in situ diagnosis. J. Natl. Cancer Inst 2010, 102, 627–637. [Google Scholar]
- Cheang, M.C.; Chia, S.K.; Voduc, D.; Gao, D.; Leung, S.; Snider, J.; Watson, M.; Davies, S.; Bernard, P.S.; Parker, J.S.; et al. Ki67 index, HER2 status, and prognosis of patients with luminal B breast cancer. J. Natl. Cancer Int 2009, 101, 736–750. [Google Scholar]
- Riley, T.; Sontag, E.; Chen, P.; Levine, A. Transcriptional control of human p53-regulated genes. Nat. Rev. Mol. Cell Biol 2008, 9, 402–412. [Google Scholar]
- Diaz, R.M.; Bateman, A.; Emiliusen, L.; Fielding, A.; Trono, D.; Russell, S.J.; Vile, R.G. A lentiviral vector expressing a fusogenic glycoprotein for cancer gene therapy. Gene Ther. 2000, 7, 1656–1663. [Google Scholar]
- Kirkham, L.A.; Bateman, A.R.; Melcher, A.A.; Vile, R.G.; Fielding, A.K. Lack of specificity of cell-surface protease targeting of a cytotoxic hyperfusogenic gibbon ape leukaemia virus envelope glycoprotein. J. Gene Med 2002, 4, 592–600. [Google Scholar]
- Bateman, A.R.; Harrington, K.J.; Kottke, T.; Ahmed, A.; Melcher, A.A.; Gough, M.J.; Linardakis, E.; Riddle, D.; Dietz, A.; Lohse, C.M.; et al. Viral fusogenic membrane glycoproteins kill solid tumor cells by nonapoptotic mechanisms that promote cross presentation of tumor antigens by dendritic cells. Cancer Res 2002, 62, 6566–6578. [Google Scholar]
- Gerdes, K.; Poulsen, L.K.; Thisted, T.; Nielsen, A.; Anderssen, P.H. The hok killer gene family in Gram negative bacteria. New Biol 1990, 2, 964–956. [Google Scholar]
- Boulaiz, H.; Prados, J.; Melguizo, C.; Garcia, A.; Marchal, J.A.; Carrillo, E.; Vélez, C.; Aranega, A. Inhibition of cell proliferation and apoptosis induction in human melanoma MCF7 cell line by gef gene. Br. J. Cancer 2003, 89, 192–198. [Google Scholar]
- Boulaiz, H.; Prados, J.; Marchal, J.A.; Concha, A.; Melguizo, C.; Carrillo, E.; Vélez, C.; Martínez, A.; Aránega, A. Modification of Ki-67 antigen and Cell surface changes during gef gene induced apoptosis in human breast cancer MCF-7 cells. Cell Mol. Biol 2005, 51, 87–92. [Google Scholar]
- Schepelmann, S.; Springer, C.J. Viral vectors for gene-directed enzyme prodrug therapy. Curr. Gene Ther 2006, 6, 647–670. [Google Scholar]
- Ladd, B.; O’Konek, J.J.; Ostruszka, L.J.; Shewach, D.S. Unrepairable DNA double-strand breaks initiate cytotoxicity with HSV-TK/ganciclovir. Cancer Gene Ther 2011, 51, 751–759. [Google Scholar]
- Zarovni, N.; Vago, R.; Solda, T.; Monaco, L.; Fabbrini, M.S. Saporin as a novel suicide gene in anticancer gene therapy. Cancer Gene Ther 2006, 1, 1–9. [Google Scholar]
- Rama, A.R.; Prados, J.; Melguizo, C.; Burgos, M.; Alvarez, P.J.; Rodriguez-Serrano, F.; Ramos, J.L.; Aranega, A. Synergistic antitumoral effect of combination E gene therapy and Doxorubicin in MCF-7 breast cancer cells. Biomed. Pharmacother 2011, 65, 260–270. [Google Scholar]
- Rosen, P.P.; Lesser, M.L.; Arroyo, C.D.; Cranor, M.; Borgen, P.; Norton, L. P53 in node negative breast carcinoma: An immunohistochemical study of epidemiological risk factors, histological features and prognosis. J. Clin. Oncol 1995, 13, 821–830. [Google Scholar]
- Arpino, G.; Weiss, H.; Lee, A.V.; Schiff, R.; de Placido, S.; Osborne, C.K. Estrogen receptor-positive, progesterone receptor-negative breast cancer: Association with growth factor receptor expression and tamoxifen resistance. J. Natl. Cancer Inst 2005, 9, 7, 1254–1261. [Google Scholar]
- Ciccarese, M.; Vito, L.; de Laurentis, M. Controversies in adjuvant endocrine therapy for pre- and post-menopausal women with breast cancer. Eur. J. Cancer 2008, 6, 4–9. [Google Scholar]
- Mansour, E.G.; Ravdin, Y.; Dressler, L. Prognostic factors in early breast carcinoma. Cancer 1994, 74, 381–400. [Google Scholar]
- Agrup, M.; Stal, O.; Olsen, K.; Wingren, S. C-erbB-2 overexpression and survival in early onset breast cancer. Breast Cancer Res. Treat 2000, 63, 23–29. [Google Scholar]
- Leong, A.S.-Y. Immunohistological markers for tumor prognostication. Curr. Diag. Pathol 2001, 7, 176–186. [Google Scholar]
- Sahin, A.A. Biologic and clinical significance of HER-2/neu (c- erbB-2) in breast cancer. Adv. Anat. Pathol 2000, 7, 158–166. [Google Scholar]
- Ross, J.; Linette, G.; Stec, J.; Clark, E.; Ayers, M.; Leschly, N.; Symmans, W.F.; Hortobagyi, G.N.; Pusztai, L. Breast cancer biomarkers molecular medicine. Expert. Rev. Mol. Diagn 2003, 3, 573–585. [Google Scholar]
- Van Diest, P.J.; van de Wall, E.; Baak, J.P. Prognostic value of proliferation in invasive breast cancer: A review. J. Clin. Pathol 2004, 57, 675–681. [Google Scholar]
- Bubb, R.S.; Komaki, R.; Hachiya, T.; Milas, I.; Ro, J.Y.; Langford, L.; Sawaya, R.; Putnam, J.B.; Allen, P.; Cox, J.D.; et al. Association of Ki-67, p53, and bcl-2 expression of the primary non-small-cell lung cancer lesion with brain metastatic lesion. Int. J. Rad. Oncol. Biol. Phys 2002, 53, 1216–1224. [Google Scholar]
- Barbareschi, M. Prognostic value of the immunohistochemical expression of p53 in breast carcinomas. Apll. Imunohistochem 1996, 4, 106–116. [Google Scholar]
- Allred, D.C.; Harvey, J.M.; Berardo, M.; Gary, M.; Clark, G.M. Prognostic and predictive factors in breast cancer by immunohistochemical analysis. Mod. Pathol 1998, 11, 155–168. [Google Scholar]
- Bankfalvi, A.; Tory, K.; Kemper, M.; Breukelmann, D.; Cubick, C.; Poremba, C.; Füzesi, L.; Lellè, R.J.; Böcker, W. Clinical relevance of immunohistochemical expression of p53-targeted gene products mdm-2, p21 and Bcl-2 in breast carcinoma. Pathol. Res. Pract 2000, 196, 489–501. [Google Scholar]
- Bottini, A.; Berruti, A.; Bersiga, A.; Brizzi, M.P.; Brunelli, A.; Gorzegno, G.; DiMarco, B.; Aguggini, S.; Bolsi, G.; Cirillo, F.; et al. p53 but not Bcl-2 immunostaining is predictive of poor clinical complete response to primary chemotherapy in breast cancer patients. Clin. Cancer Res 2000, 6, 2751–2758. [Google Scholar]
- Clahsen, P.C.; van de Velde, C.J.; Duval, C.; Pallud, C.; Mandard, A.M.; Delobelle-Deroide, A.; van den Broek, L.; Sahmoud, T.M.; van de Vijver, M.J. p53 protein accumulation and response to adjuvant chemotherapy in premenopausal women with node-negative early breast cancer. J. Clin. Oncol 1998, 16, 470–479. [Google Scholar]
- Rozan, S.; Vincent-Salomon, A.; Zafrani, B.; Validire, P.; de Cremoux, P.; Bernoux, A.; Nieruchalski, M.; Fourquet, A.; Clough, K.; Dieras, V.; et al. No significant predictive value of c-erbB-2 or p53 expression regarding sensitivy to primary chemotherapy or radiotherapy in breast cancer. Int. J. Cancer 1998, 79, 27–33. [Google Scholar]
- Sjöström, J.; Blomqvist, C.; Heikkilävon, P.; von Boguslawski, P.K.; Räisänen-Sokolowski, A.; Bengtsson, N.O.; Mjaaland, I.; Malmström, P.; Ostenstadt, B.; Bergh, J.; et al. Predictive value of p53, mdm-2, p21, and mib-1 for chemotherapy response. Clin. Cancer Res 2000, 6, 3103–3110. [Google Scholar]
- Falette, M.P.; Paperin, I.; Treilleux, A.C.; Gratadour, N.; Peloux, H.; Mignotte, N.; Tooke, E.; Löfman, M.; Inganäs, M.; Bremond, A.; et al. Prognostic value of p53 gene mutations in a large series of node-negative breast cancer patients. Cancer Res 1998, 58, 1451–1455. [Google Scholar]
- Dai, C.; Gu, W. p53 post-translational modification: Deregulated in tumorigenesis. Trends Mol. Med 2010, 16, 528–536. [Google Scholar]
- Gentile, M.; Bergman Jungeström, M.; Olsen, K.E.; Söderkvist, P.; Wingren, S. p53 and survival in early onset breast cancer: Analysis of gene mutations, loss of heterozygosity protein accumulation. Eur. J. Cancer 1999, 35, 1202–1207. [Google Scholar]
- Blagosklonny, M.V. Loss of function and p53 protein stabilization. Oncogene 1997, 15, 1889–1893. [Google Scholar]
- Vagunda, V.; Smardová, J.; Vagundová, M.; Jandáková, E.; Zaloudík, J.; Koukalová, H. Correlations of breast carcinoma biomarkers and p53 tested by FASAY and immunohistochemistry. Pathol. Res. Pract 2003, 199, 795–801. [Google Scholar]
- Seo, J.H.; Moon, H.S.; Kim, I.Y.; Guo, D.D.; Lee, H.G.; Choi, Y.J.; Cho, C.S. PEGylated conjugated linoleic acid stimulation of apoptosis via a p53-mediated signaling pathway in MCF-7 breast cancer cells. Eur. J. Pharm. Biopharm 2008, 70, 621–626. [Google Scholar]
- SPSS, version 7; SPSS: Chicago, IL, USA, 1997.
| Cell line | Protein markers | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| ER | PR | c-erbB-2 | Ki-67 | p53 | ||||||
| A | B | A | B | A | B | A | B | A | B | |
| MCF-7 | 13% | ++ | 20% | ++ | 62% | +++ | 72% | +++ | 33% | ++ |
| MCF-7TG dex 2 days | 50% | ++ | 57% | ++ | 27% | + | 18% | ++ | 59% | +++ |
| MCF-7TG dex 6 days | 55% | ++ | 61% | + | 17% | + | 8% | +/− | 55% | +++ |
| Antibody | Source | Dilution | Staining pattern in cancer cells |
|---|---|---|---|
| ER | Dako (Barcelona, Spain) | 1:200 | Nuclear staining |
| PR | Dako (Barcelona, Spain) | 1:250 | Nuclear staining |
| c-erbB-2 | Dako (Barcelona, Spain) | 1:250 | Membrane staining |
| Ki-67 | Dako (Barcelona, Spain) | 1:100 | Nuclear staining |
| p53 | Dako (Barcelona, Spain) | 1:100 | Nuclear and Cytoplasm staining |
© 2011 by the authors; licensee MDPI, Basel, Switzerland. This article is an open-access article distributed under the terms and conditions of the Creative Commons Attribution license (http://creativecommons.org/licenses/by/3.0/).
Share and Cite
Boulaiz, H.; Álvarez, P.J.; Prados, J.; Marchal, J.; Melguizo, C.; Carrillo, E.; Peran, M.; Rodríguez, F.; Ramírez, A.; Ortíz, R.; et al. gef Gene Expression in MCF-7 Breast Cancer Cells is Associated with a Better Prognosis and Induction of Apoptosis by p53-Mediated Signaling Pathway. Int. J. Mol. Sci. 2011, 12, 7445-7458. https://doi.org/10.3390/ijms12117445
Boulaiz H, Álvarez PJ, Prados J, Marchal J, Melguizo C, Carrillo E, Peran M, Rodríguez F, Ramírez A, Ortíz R, et al. gef Gene Expression in MCF-7 Breast Cancer Cells is Associated with a Better Prognosis and Induction of Apoptosis by p53-Mediated Signaling Pathway. International Journal of Molecular Sciences. 2011; 12(11):7445-7458. https://doi.org/10.3390/ijms12117445
Chicago/Turabian StyleBoulaiz, Houria, Pablo J. Álvarez, Jose Prados, Juan Marchal, Consolación Melguizo, Esmeralda Carrillo, Macarena Peran, Fernando Rodríguez, Alberto Ramírez, Raúl Ortíz, and et al. 2011. "gef Gene Expression in MCF-7 Breast Cancer Cells is Associated with a Better Prognosis and Induction of Apoptosis by p53-Mediated Signaling Pathway" International Journal of Molecular Sciences 12, no. 11: 7445-7458. https://doi.org/10.3390/ijms12117445






