Microsatellite Analysis in Multistage Carcinogenesis of Esophageal Squamous Cell Carcinoma from Chongqing in Southern China
Abstract
:1. Introduction
2. Results
3. Discussion
4. Experimental Section
4.1. Sample Collection
4.2. Tissue Cell Acquisition by Using Microdissection
4.3. Extraction of Genomic DNA
4.4. Loss of Heterozygosity (LOH) Analysis
4.5. Statistical Analysis
5. Conclusions
Acknowledgements
- Conflict of Interest The authors declare no conflict of interest.
References
- Maley, C.C.; Galipeau, P.C.; Li, X.; Sanchez, C.A.; Paulson, T.G.; Blount, P.L.; Reid, B.J. The combination of genetic instability and clonal expansion predicts progression to esophageal adenocarcinoma. Cancer Res 2004, 64, 7629–7633. [Google Scholar]
- Maitra, A.; Wistuba, I.I.; Washington, C.; Virmani, A.K.; Ashfaq, R.; Milchgrub, S.; Gazdar, A.F.; Minna, J.D. High-resolution chromosome 3p allelotyping of breast carcinomas and precursor lesions demonstrates frequent loss of heterozygosity and a discontinuous pattern of allele loss. Am. J. Pathol 2001, 159, 119–130. [Google Scholar]
- Wang, Z.; Lai, F.M. Analysis of loss of heterozygosity on chromosome 8 in human prostate carcinoma and high grade prostatic intraepithelial neoplasia. Zhonghua Nan Ke Xue 2004, 10, 26–31. [Google Scholar]
- Karakosta, A.; Golias, C.; Charalabopoulos, A.; Peschos, D.; Batistatou, A.; Charalabopoulos, K. Genetic models of human cancer as a multistep process. Paradigm models of colorectal cancer, breast cancer, and chronic myelogenous and acute lymphoblastic leukaemia. J. Exp. Clin. Cancer Res 2005, 24, 505–514. [Google Scholar]
- Ha, P.K.; Pilkington, T.A.; Westra, W.H.; Sciubba, J.; Sidransky, D.; Califano, J.A. Progression of microsatellite instability from premalignant lesions to tumors of the head and neck. Int. J. Cancer 2002, 102, 615–617. [Google Scholar]
- Dawsey, S.M.; Lewin, K.J.; Wang, G.Q.; Liu, F.S.; Nieberg, R.K.; Yu, Y.; Li, J.Y.; Blot, W.J.; Li, B.; Taylor, P.R. Squamous esophageal histology and subsequent risk of squamous cell carcinoma of the esophagus. A prospective follow-up study from Linxian, China. Cancer 1994, 74, 1686–1692. [Google Scholar]
- Shen, D.; Nooraie, F.; Elshimali, Y.; Lonsberry, V.; He, J.; Bose, S.; Chia, D.; Seligson, D.; Chang, H.R.; Goodglick, L. Decreased expression of annexin A1 is correlated with breast cancer development and progression as determined by a tissue microarray analysis. Hum. Pathol 2006, 37, 1583–1591. [Google Scholar]
- Zhou, G.; Li, H.; DeCamp, D.; Chen, S.; Shu, H.; Gong, Y.; Flaig, M.; Gillespie, J.W.; Hu, N.; Taylor, P.R.; et al. 2D differential in-gel electrophoresis for the identification of esophageal scans cell cancer-specific protein markers. Mol. Cell Proteomics 2002, 1, 117–124. [Google Scholar]
- Yang, L.; Leung, A.C.; Ko, J.M.; Lo, P.; Tang, J.; Srivastava, G.; Oshimura, M.; Stanbridge, E.; Daigo, Y.; Nakamura, Y.; et al. Tumor suppressive role of a 2.4 Mb 9q33-q34 critical region and DEC1 in esophageal squamous cell carcinoma. Oncogene 2005, 24, 697–705. [Google Scholar]
- Huang, X.P.; Wei, F.; Liu, X.Y.; Xu, X.; Hu, H.; Chen, B.; Xia, S.; Han, Y.; Han, Y.; Cai, Y.; et al. Allelic loss on 13q in esophageal squamous cell carcinomas from northern China. Cancer Lett 2002, 185, 87–94. [Google Scholar]
- Hu, N.; Huang, J.; Emmert-Buck, M.R.; Tang, Z.Z.; Roth, M.J.; Wang, C.; Dawsey, S.M.; Li, G.; Li, W.J.; Wang, Q.H.; et al. Frequent inactivation of the TP53 gene in esophageal squamous cell carcinoma from a high-risk population in China. Clin. Cancer Res 2001, 7, 883–891. [Google Scholar]
- Koppert, L.B.; Wijnhoven, B.P.; van Dekken, H.; Tilanus, H.W.; Dinjens, W.N. The molecular biology of esophageal adenocarcinoma. J. Surg. Oncol 2005, 92, 169–190. [Google Scholar]
- Partridge, M.; Pateromichelakis, S.; Phillips, E.; Emilion, G.; Langdon, J. Profiling clonality and progression in multiple premalignant and malignant oral lesions identifies a subgroup of cases with a distinct presentation of squamous cell carcinoma. Clin. Cancer Res 2001, 7, 1860–1866. [Google Scholar]
- Cheng, L.; Bostwick, D.G.; Li, G.; Wang, Q.; Hu, N.; Vortmeyer, A.O.; Zhuang, Z. Allelic imbalance in the clonal evolution of prostate carcinoma. Cancer 1999, 85, 2017–2022. [Google Scholar]
- Heim, S.; Teixeira, M.R.; Dietrich, C.U.; Pandis, N. Cytogenetic polyclonality in tumors of the breast. Cancer Genet. Cytogenet 1997, 95, 16–19. [Google Scholar]
- He, S.; Guo, G.M.; Liu, F.X.; Huang, X.P.; Xu, X.; Cai, Y.; Han, Y.L.; Zhan, Q.M.; Wu, M.; Dong, J.T.; et al. Molecular analysis in combination with iodine staining may contribute to the risk prediction of esophageal squamous cell carcinoma. J. Cancer Res. Clin. Oncol 2008, 134, 307–315. [Google Scholar]
- Dawsey, S.M.; Lewin, K.J.; Liu, F.S.; Wang, G.Q.; Shen, Q. Esophageal morphology from Linxian, China. Squamous histologic findings in 754 patients. Cancer 1994, 73, 2027–2037. [Google Scholar]
- The Genome Database. Available online: http://www.gdb.org/ accessed on 15 October 2009.
- The NCBI genome database. Available online: http://www.ncbi.nlm.nih.gov/ accessed on 7 November 2009.
- Zhang, X.L.; Fu, W.L.; Zhao, H.X.; Zhou, L.X.; Huang, J.F.; Wang, J.H. Molecular studies of loss of heterozygosity in Chinese sporadic retinoblastoma patients. Clin. Chim. Acta 2005, 358, 75–80. [Google Scholar]
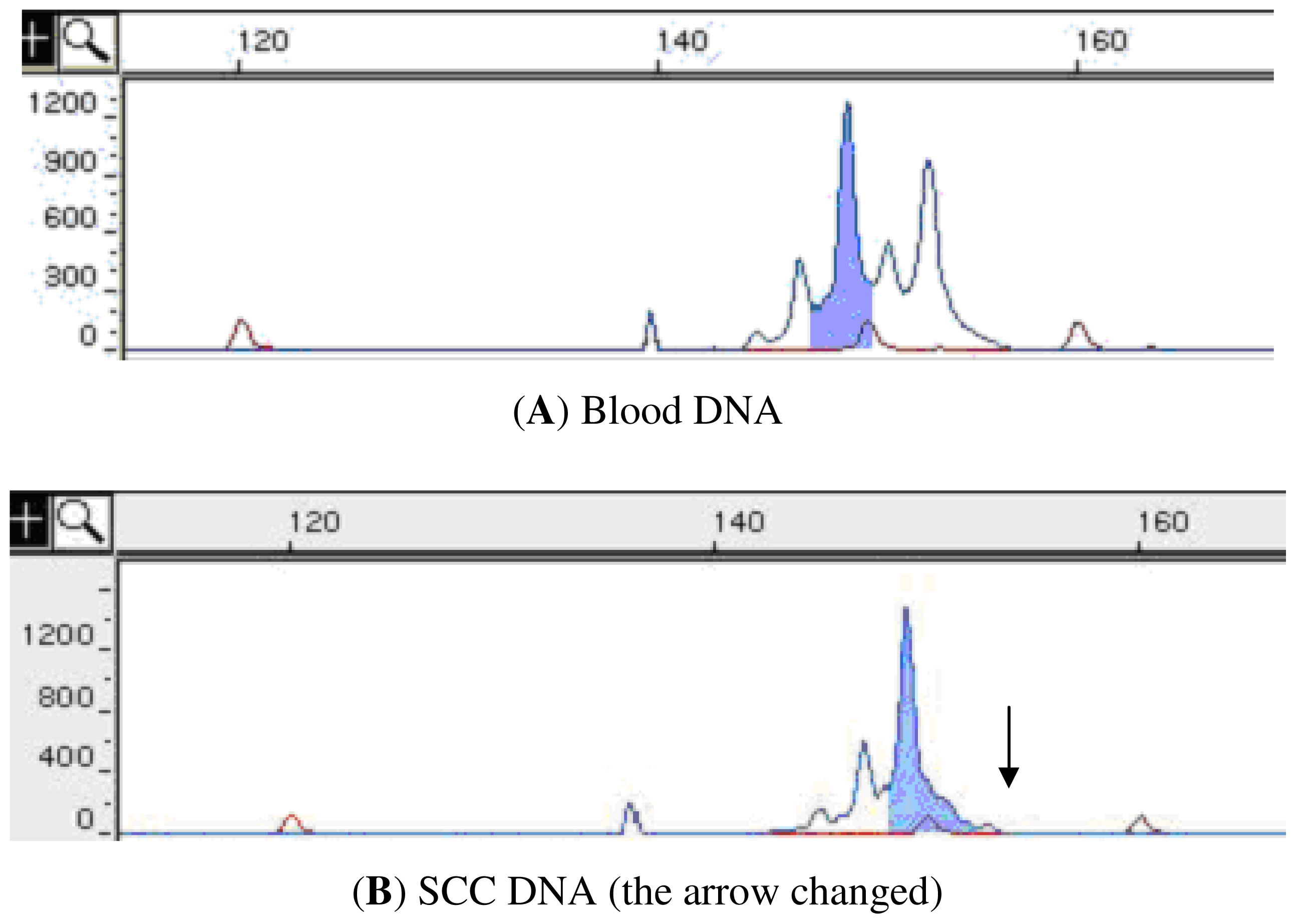
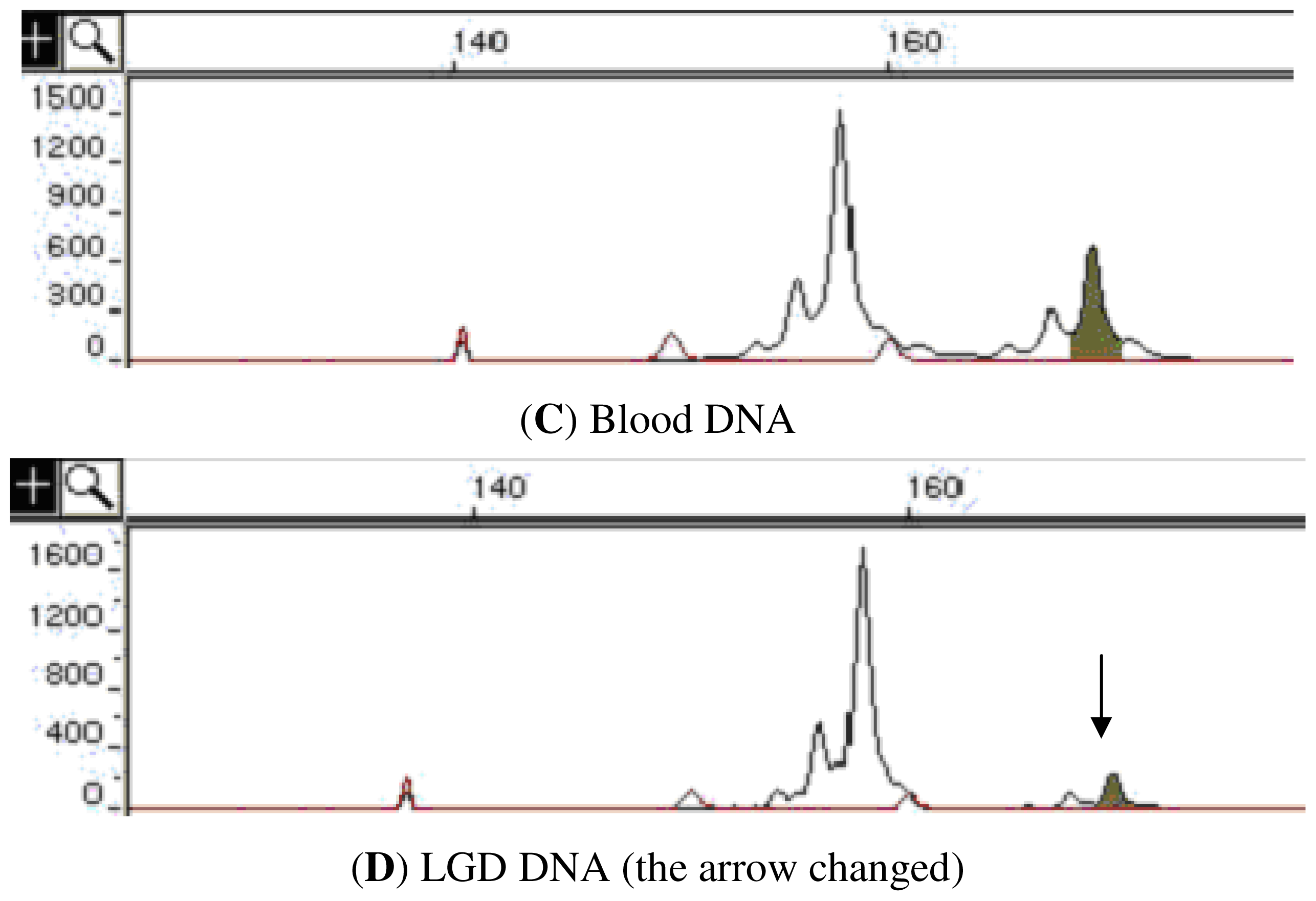
| Marker | Location | LGD a sample (%) | HGD a sample (%) | SCC a sample (%) |
|---|---|---|---|---|
| 3S1597 | 3p25 | 4/21 (19) | 12/22 (54.5) | 18/22 (81.8) |
| D3S2452 | 3p21-p14 | 7/25 (28) | 20/25 (80) | 18/25 (72) |
| D3S1285 | 3p14 | 3/19 (15.8) | 14/21 (66.7) | 16/21 (76.2) |
| D4S174 | 4p14-p13 | 2/27 (7.4) | 7/27 (25.9) | 12/27 (44.4) |
| D5S409 | 5q14-q15 | 0/16 (0) | 7/16 (43.7) | 8/16 (50) |
| D5S2501 | 5q21-q23.3 | 1/13 (7.7) | 7/13 (53.8) | 7/13 (53.8) |
| D8S261 | 8p22-p21.3 | 0/18 (0) | 5/18 (27.8) | 6/18 (33.3) |
| D9S157 | 9p23-p22 | 0/23 (0) | 16/23 (69.5) | 15/23 (65.2) |
| D9S111 | 9q12-q21.1 | 0/25 (0) | 9/25 (36) | 15/25 (60) |
| D9S125 | 9q34-q34 | 9/27 (33.3) | 21/27 (77.8) | 23/27 (85.2) |
| D11S1338 | 11p15.5 | 0/29 (0) | 10/29 (34.5) | 11/29 (37.9) |
| D13S175 | 13q13 | 0/22 (0) | 10/23 (43.5) | 12/23 (52.2) |
| D13S153 | 13q14.2 | 4/24 (16.7) | 10/24 (41.7) | 13/24 (54.2) |
| D13S173 | 13q32-q34 | 0/24 (0) | 5/24 (20.8) | 11/24 (45.8) |
| D17S786 | 17p13.1 | 5/23 (21.7) | 13/23 (56.5) | 16/23 (69.5) |
| D17S261 | 17p11.2 | 0/22 (0) | 10/22 (45.4) | 11/22 (50) |
| Total b | 35/358 (9.8) | 176/362 (48.6) | 212/362 (58.5) |
© 2011 by the authors; licensee MDPI, Basel, Switzerland. This article is an open-access article distributed under the terms and conditions of the Creative Commons Attribution license (http://creativecommons.org/licenses/by/3.0/).
Share and Cite
Liu, M.; Zhang, F.; Liu, S.; Zhao, W.; Zhu, J.; Zhang, X. Microsatellite Analysis in Multistage Carcinogenesis of Esophageal Squamous Cell Carcinoma from Chongqing in Southern China. Int. J. Mol. Sci. 2011, 12, 7401-7409. https://doi.org/10.3390/ijms12117401
Liu M, Zhang F, Liu S, Zhao W, Zhu J, Zhang X. Microsatellite Analysis in Multistage Carcinogenesis of Esophageal Squamous Cell Carcinoma from Chongqing in Southern China. International Journal of Molecular Sciences. 2011; 12(11):7401-7409. https://doi.org/10.3390/ijms12117401
Chicago/Turabian StyleLiu, Ming, Feng Zhang, Shen Liu, Wen Zhao, Jing Zhu, and Xiaoli Zhang. 2011. "Microsatellite Analysis in Multistage Carcinogenesis of Esophageal Squamous Cell Carcinoma from Chongqing in Southern China" International Journal of Molecular Sciences 12, no. 11: 7401-7409. https://doi.org/10.3390/ijms12117401




