Characterization of the Sesbania rostrata Phytochelatin Synthase Gene: Alternative Splicing and Function of Four Isoforms
Abstract
:1. Introduction
2. Results and Discussion
2.1. Cloning and Sequence Analysis of PCS cDNAs and Gene from S. rostrata
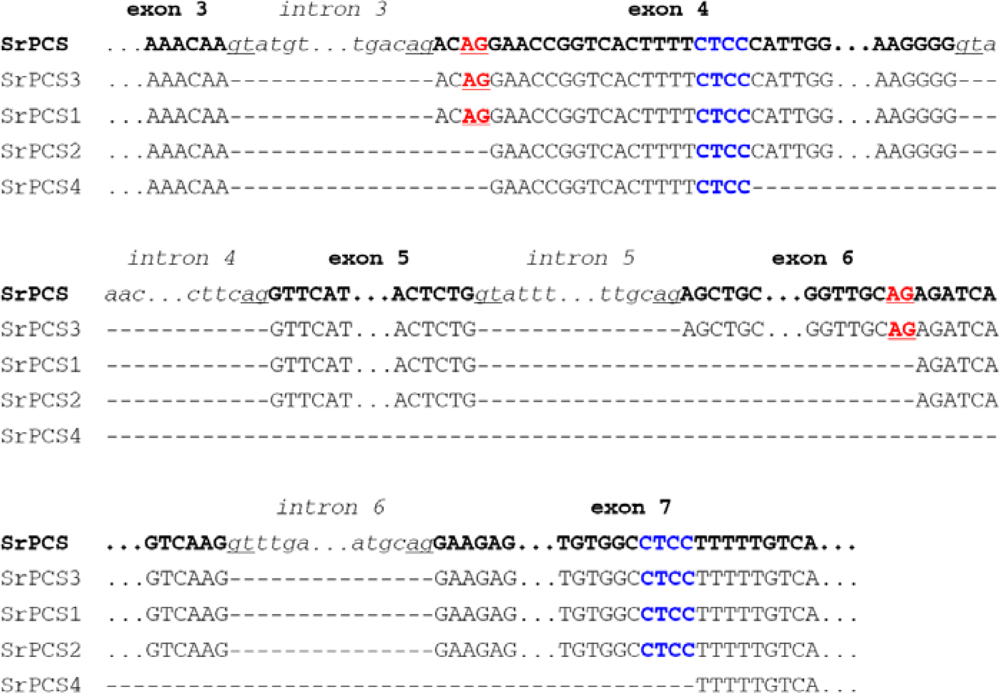
2.2. Genomic Organization and Pre-mRNA Splicing of SrPCS Gene
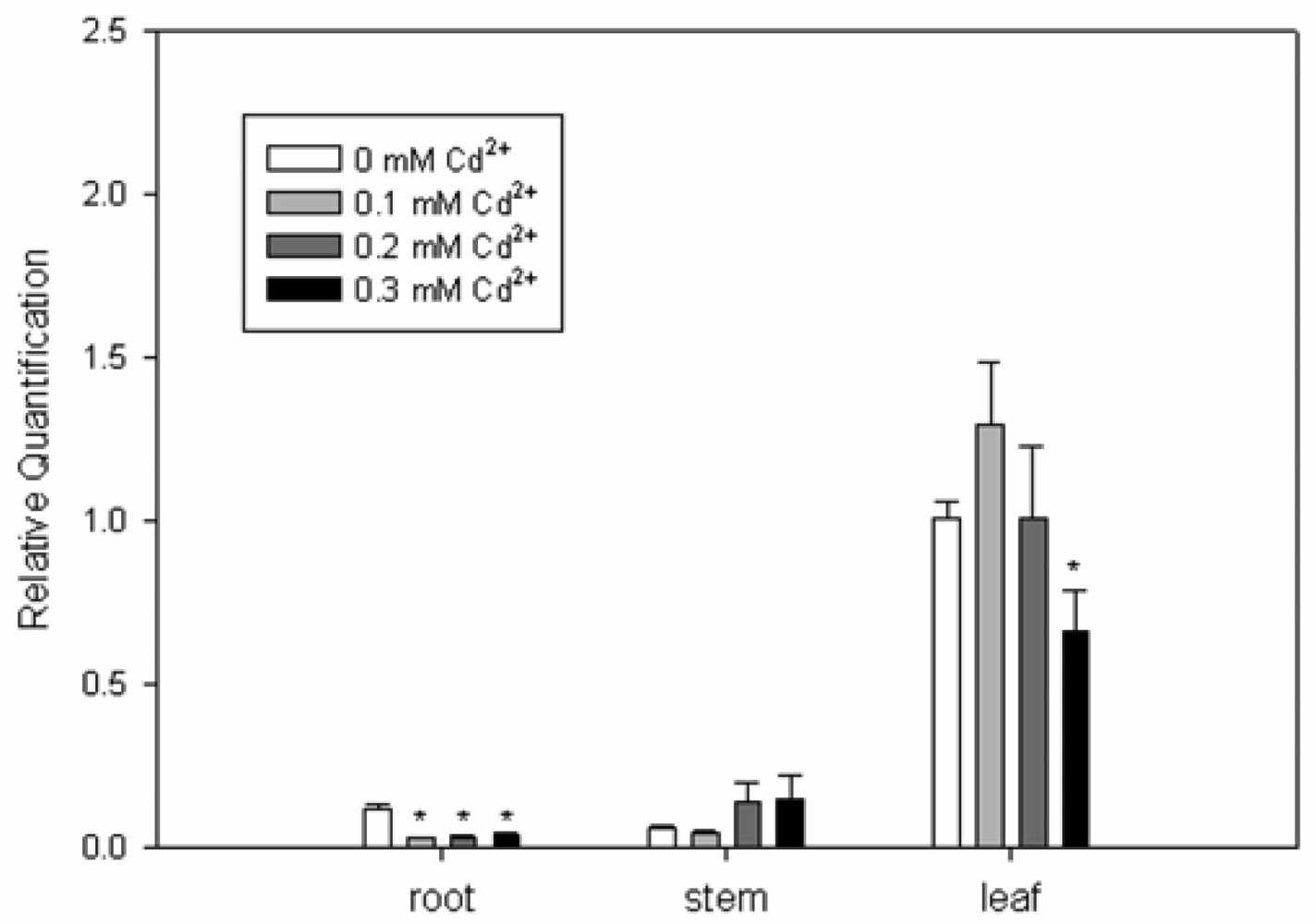
2.3. Expression Analysis of SrPCS Gene
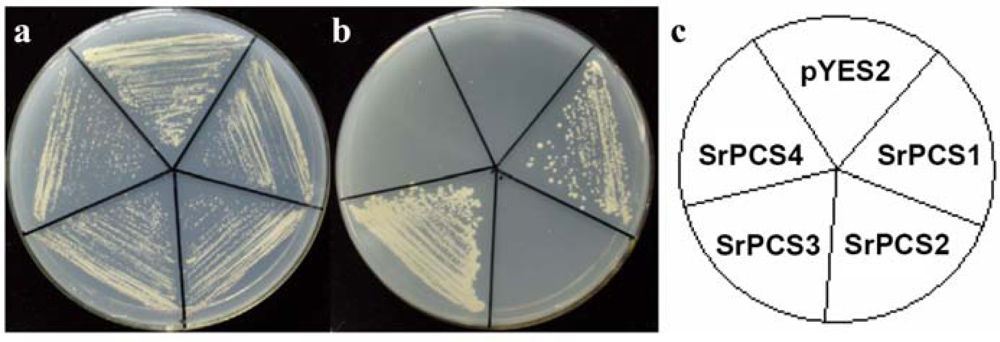
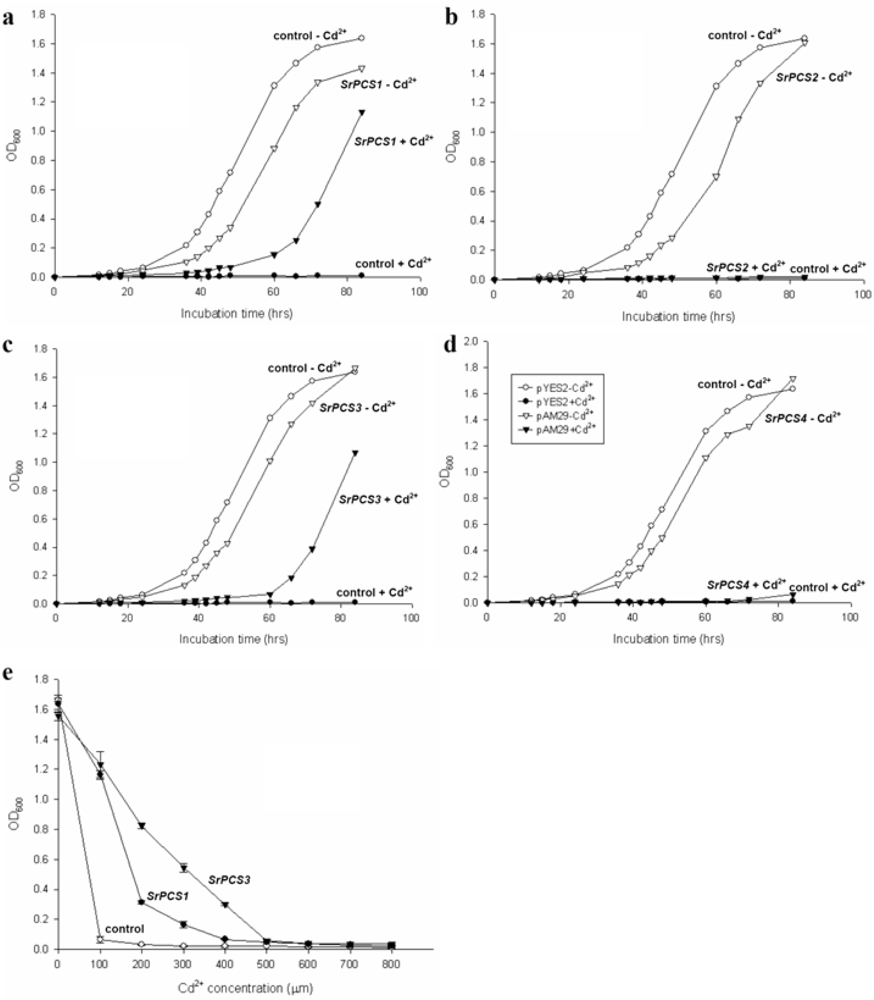
2.4. Functionality of Four SrPCS Isoforms SrPCS1–SrPCS4
3. Experimental Section
3.1. Plant Materials
3.2. Cloning of PCS cDNAs and Gene from S. rostrata
3.3. Real Time RT-PCR of SrPCS3
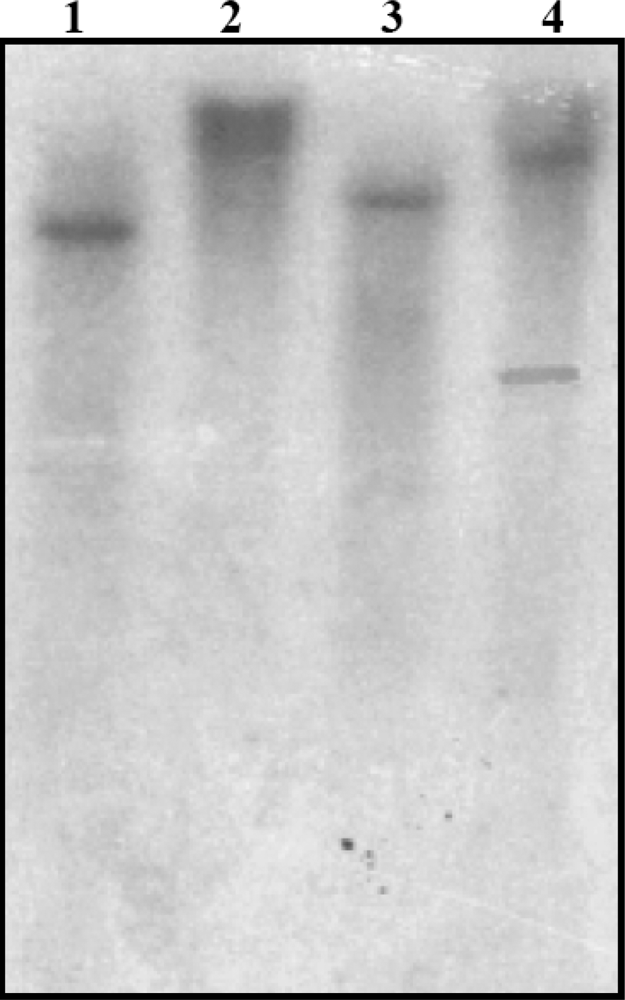
3.4. Southern Blot Analysis of SrPCS
3.5. Expression of Recombinant SrPCS1-4 in Yeast and Cd Tolerance Assay
4. Conclusions
Acknowledgments
References
- Cobbett, C; Goldsbrough, P. Phytochelatins and metallothioneins: roles in heavy metal detoxification and homeostasis. Annu. Rev. Plant. Biol 2002, 53, 159–182. [Google Scholar]
- Grill, E; Winnacker, EL; Zenk, MH. Phytochelatins: the principal heavy-metal complexing peptides of higher plants. Science 1985, 230, 674–676. [Google Scholar]
- Tennstedt, P; Peisker, D; Bottcher, C; Trampczynska, A; Clemens, S. Phytochelatin synthesis is essential for the detoxification of excess zinc and contributes significantly to the accumulation of zinc. Plant Physiol 2009, 149, 938–948. [Google Scholar]
- Cobbett, CS. Phytochelatins and their roles in heavy metal detoxification. Plant Physiol 2000, 123, 825–832. [Google Scholar]
- Grill, E; Loffler, S; Winnacker, EL; Zenk, MH. Phytochelatins, the heavy-metal-binding peptides of plants, are synthesized from glutathione by a specific gamma-glutamylcysteine dipeptidyl transpeptidase (phytochelatin synthase). Proc. Natl. Acad. Sci. USA 1989, 86, 6838–6842. [Google Scholar]
- Ha, SB; Smith, AP; Howden, R; Dietrich, WM; Bugg, S; O'Connell, MJ; Goldsbrough, PB; Cobbett, CS. Phytochelatin synthase genes from Arabidopsis and the yeast Schizosaccharomyces pombe. Plant Cell 1999, 11, 1153–1164. [Google Scholar]
- Oven, M; Page, JE; Zenk, MH; Kutchan, TM. Molecular characterization of the homophytochelatin synthase of soybean Glycine max: relation to phytochelatin synthase. J. Biol. Chem 2002, 277, 4747–4754. [Google Scholar]
- Tsuji, N; Nishikori, S; Iwabe, O; Matsumoto, S; Shiraki, K; Miyasaka, H; Takagi, M; Miyamoto, K; Hirata, K. Comparative analysis of the two-step reaction catalyzed by prokaryotic and eukaryotic phytochelatin synthase by an ion-pair liquid chromatography assay. Planta 2005, 222, 181–191. [Google Scholar]
- Vatamaniuk, OK; Mari, S; Lang, A; Chalasani, S; Demkiv, LO; Rea, PA. Phytochelatin synthase, a dipeptidyltransferase that undergoes multisite acylation with gamma-glutamylcysteine during catalysis: stoichiometric and site-directed mutagenic analysis of arabidopsis thaliana PCS1-catalyzed phytochelatin synthesis. J. Biol. Chem 2004, 279, 22449–22460. [Google Scholar]
- Vatamaniuk, OK; Mari, S; Lu, YP; Rea, PA. Mechanism of heavy metal ion activation of phytochelatin (PC) synthase: blocked thiols are sufficient for PC synthase-catalyzed transpeptidation of glutathione and related thiol peptides. J. Biol. Chem 2000, 275, 31451–31459. [Google Scholar]
- Ruotolo, R; Peracchi, A; Bolchi, A; Infusini, G; Amoresano, A; Ottonello, S. Domain organization of phytochelatin synthase: functional properties of truncated enzyme species identified by limited proteolysis. J. Biol. Chem 2004, 279, 14686–14693. [Google Scholar]
- Tsuji, N; Nishikori, S; Iwabe, O; Shiraki, K; Miyasaka, H; Takagi, M; Hirata, K; Miyamoto, K. Characterization of phytochelatin synthase-like protein encoded by alr0975 from a prokaryote, Nostoc sp. PCC 7120. Biochem. Biophys. Res. Commun 2004, 315, 751–755. [Google Scholar]
- Romanyuk, ND; Rigden, DJ; Vatamaniuk, OK; Lang, A; Cahoon, RE; Jez, JM; Rea, PA. Mutagenic definition of a papain-like catalytic triad, sufficiency of the N-terminal domain for single-site core catalytic enzyme acylation, and C-terminal domain for augmentative metal activation of a eukaryotic phytochelatin synthase. Plant Physiol 2006, 141, 858–869. [Google Scholar]
- Vivares, D; Arnoux, P; Pignol, D. A papain-like enzyme at work: Native and acyl-enzyme intermediate structures in phytochelatin synthesis. Proc. Natl. Acad. Sci. USA 2005, 102, 18848–18853. [Google Scholar]
- Rea, PA; Vatamaniuk, OK; Rigden, DJ. Weeds, worms, and more. Papain's long-lost cousin, phytochelatin synthase. Plant Physiol 2004, 136, 2463–2474. [Google Scholar]
- Vatamaniuk, OK; Mari, S; Lu, YP; Rea, PA. AtPCS1, a phytochelatin synthase from Arabidopsis: isolation and in vitro reconstitution. Proc. Natl. Acad. Sci. USA 1999, 96, 7110–7115. [Google Scholar]
- Clemens, S; Kim, EJ; Neumann, D; Schroeder, JI. Tolerance to toxic metals by a gene family of phytochelatin synthases from plants and yeast. EMBO J 1999, 18, 3325–3333. [Google Scholar]
- Ramos, J; Clemente, MR; Naya, L; Loscos, J; Perez-Rontome, C; Sato, S; Tabata, S; Becana, M. Phytochelatin synthases of the model legume Lotus japonicus. A small multigene family with differential response to cadmium and alternatively spliced variants. Plant Physiol 2007, 143, 1110–1118. [Google Scholar]
- Ramos, J; Naya, L; Gay, M; Abian, J; Becana, M. Functional characterization of an unusual phytochelatin synthase, LjPCS3, of Lotus japonicus. Plant Physiol 2008, 148, 536–545. [Google Scholar]
- Loscos, J; Naya, L; Ramos, J; Clemente, MR; Matamoros, MA; Becana, M. A reassessment of substrate specificity and activation of phytochelatin synthases from model plants by physiologically relevant metals. Plant Physiol 2006, 140, 1213–1221. [Google Scholar]
- Clemens, S; Schroeder, JI; Degenkolb, T. Caenorhabditis elegans expresses a functional phytochelatin synthase. Eur. J. Biochem 2001, 268, 3640–3643. [Google Scholar]
- Vatamaniuk, OK; Bucher, EA; Ward, JT; Rea, PA. A new pathway for heavy metal detoxification in animals. Phytochelatin synthase is required for cadmium tolerance in Caenorhabditis elegans. J. Biol. Chem 2001, 276, 20817–20820. [Google Scholar]
- Grill, E; Winnacker, EL; Zenk, MH. Phytochelatins, a class of heavy-metal-binding peptides from plants, are functionally analogous to metallothioneins. Proc. Natl. Acad. Sci. USA 1987, 84, 439–443. [Google Scholar]
- Cazale, AC; Clemens, S. Arabidopsis thaliana expresses a second functional phytochelatin synthase. FEBS Lett 2001, 507, 215–219. [Google Scholar]
- Lee, S; Korban, SS. Transcriptional regulation of Arabidopsis thaliana phytochelatin synthase (AtPCS1) by cadmium during early stages of plant development. Planta 2002, 215, 689–693. [Google Scholar]
- Somasegaran, P; Hoben, HJ. Handbook for Rhizobia: methods in legume- Rhizobia technology; Springer-Verlag: New York, NY, USA, 1994. [Google Scholar]
- Yang, ZY; Chen, FH; Yuan, JG; Zheng, ZW; Wong, MH. Responses of Sesbania rostrata and S. cannabina to Pb, Zn, Cu and Cd toxicities. J. Environ. Sci.-China 2004, 16, 670–673. [Google Scholar]
- Siebert, PD; Chenchik, A; Kellogg, DE; Lukyanov, KA; Lukyanov, SA. An improved PCR method for walking in uncloned genomic DNA. Nucleic Acids Res 1995, 23, 1087–1088. [Google Scholar]
- Nikoshkov, A; Hurd, YL. p53 splice variants generated by atypical mRNA processing confer complexity of p53 transcripts in the human brain. Biochem. Biophys. Res. Commun 2006, 351, 383–386. [Google Scholar]
- Li, L; Howe, GA. Alternative splicing of prosystemin pre-mRNA produces two isoforms that are active as signals in the wound response pathway. Plant Mol. Biol 2001, 46, 409–419. [Google Scholar]
- Kazan, K. Alternative splicing and proteome diversity in plants: the tip of the iceberg has just emerged. Trends Plant Sci 2003, 8, 468–471. [Google Scholar]
- Ali, GS; Reddy, ASN. Regulation of alternative splicing of pre-mRNAs by stresses. In Nuclear pre-mRNA Processing in Plants; Reddy, ASN, Golovkin, M, Eds.; Springer: Berlin, Heidelberg, Germany, 2008; Volume 326, p. 257. [Google Scholar]
- Heiss, S; Wachter, A; Bogs, J; Cobbett, C; Rausch, T. Phytochelatin synthase (PCS) protein is induced in Brassica juncea leaves after prolonged Cd exposure. J. Exp. Bot 2003, 54, 1833–1839. [Google Scholar]
- Stamm, S; Ben-Ari, S; Rafalska, I; Tang, Y; Zhang, Z; Toiber, D; Thanaraj, TA; Soreq, H. Function of alternative splicing. Gene 2005, 344, 1–20. [Google Scholar]
- Ben-Dov, C; Hartmann, B; Lundgren, J; Valcarcel, J. Genome-wide analysis of alternative pre-mRNA splicing. J. Biol. Chem 2008, 283, 1229. [Google Scholar]
- Barbazuk, WB; Fu, Y; McGinnis, KM. Genome-wide analyses of alternative splicing in plants: Opportunities and challenges. Genome Res 2008, 18, 1381–1392. [Google Scholar]
- Wang, BB; Brendel, V. Genomewide comparative analysis of alternative splicing in plants. Proc. Natl. Acad. Sci. USA 2006, 103, 7175–7180. [Google Scholar]
- Campbell, MA; Haas, BJ; Hamilton, JP; Mount, SM; Buell, CR. Comprehensive analysis of alternative splicing in rice and comparative analyses with Arabidopsis. BMC Genomics 2006, 7, 327. [Google Scholar]
- Wang, BB; O'Toole, M; Brendel, V; Young, ND. Cross-species EST alignments reveal novel and conserved alternative splicing events in legumes. BMC Plant Biol 2008, 8, 17. [Google Scholar]
- Raskin, I; Ensley, BD. Phytoremediation of Toxic Metals: Using Plants to Clean up the Environment; Wiley: New York, NY, USA, 2000. [Google Scholar]
- Murashige, T; Skoog, F. A revised medium for rapid growth and bioassays with tobacco tissue cultures. Physiol. Plant 1962, 15, 473–497. [Google Scholar]
- Huang, JC; Chen, F. Simultaneous amplification of 5′ and 3′ cDNA ends based on template-switching effect and inverse PCR. BioTechniques 2006, 40, 187–189. [Google Scholar]
- Sharma, AD; Gill, PK; Singh, P. DNA isolation from dry and fresh samples of polysacchariderich plants. Plant Mol. Biol. Rep 2002, 20, 415. [Google Scholar]
- Sambrook, J; Russell, DW. Molecular Cloning: A Laboratory Manual, 3rd ed; Cold Spring Harbor Laboratory Press: New York, NY, USA, 2001. [Google Scholar]
- Xu, ZF; Qi, WQ; Ouyang, XZ; Yeung, E; Chye, ML. A proteinase inhibitor II of Solanum americanum is expressed in phloem. Plant Mol. Biol 2001, 47, 727–738. [Google Scholar]


© 2009 by the authors; licensee Molecular Diversity Preservation International, Basel, Switzerland. This article is an open-access article distributed under the terms and conditions of the Creative Commons Attribution license (http://creativecommons.org/licenses/by/3.0/).
Share and Cite
Li, A.-M.; Yu, B.-Y.; Chen, F.-H.; Gan, H.-Y.; Yuan, J.-G.; Qiu, R.; Huang, J.-C.; Yang, Z.-Y.; Xu, Z.-F. Characterization of the Sesbania rostrata Phytochelatin Synthase Gene: Alternative Splicing and Function of Four Isoforms. Int. J. Mol. Sci. 2009, 10, 3269-3282. https://doi.org/10.3390/ijms10083269
Li A-M, Yu B-Y, Chen F-H, Gan H-Y, Yuan J-G, Qiu R, Huang J-C, Yang Z-Y, Xu Z-F. Characterization of the Sesbania rostrata Phytochelatin Synthase Gene: Alternative Splicing and Function of Four Isoforms. International Journal of Molecular Sciences. 2009; 10(8):3269-3282. https://doi.org/10.3390/ijms10083269
Chicago/Turabian StyleLi, An-Ming, Bing-Yun Yu, Fu-Hua Chen, Hui-Yan Gan, Jian-Gang Yuan, Rongliang Qiu, Jun-Chao Huang, Zhong-Yi Yang, and Zeng-Fu Xu. 2009. "Characterization of the Sesbania rostrata Phytochelatin Synthase Gene: Alternative Splicing and Function of Four Isoforms" International Journal of Molecular Sciences 10, no. 8: 3269-3282. https://doi.org/10.3390/ijms10083269





