Function Annotation of an SBP-box Gene in Arabidopsis Based on Analysis of Co-expression Networks and Promoters
Abstract
:1. Introduction
2. Materials and Methods
2.2. Arabidopsis interactome network and clustering analysis
2.3. Promoter analysis
3. Results
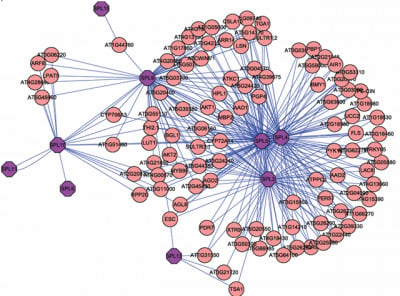
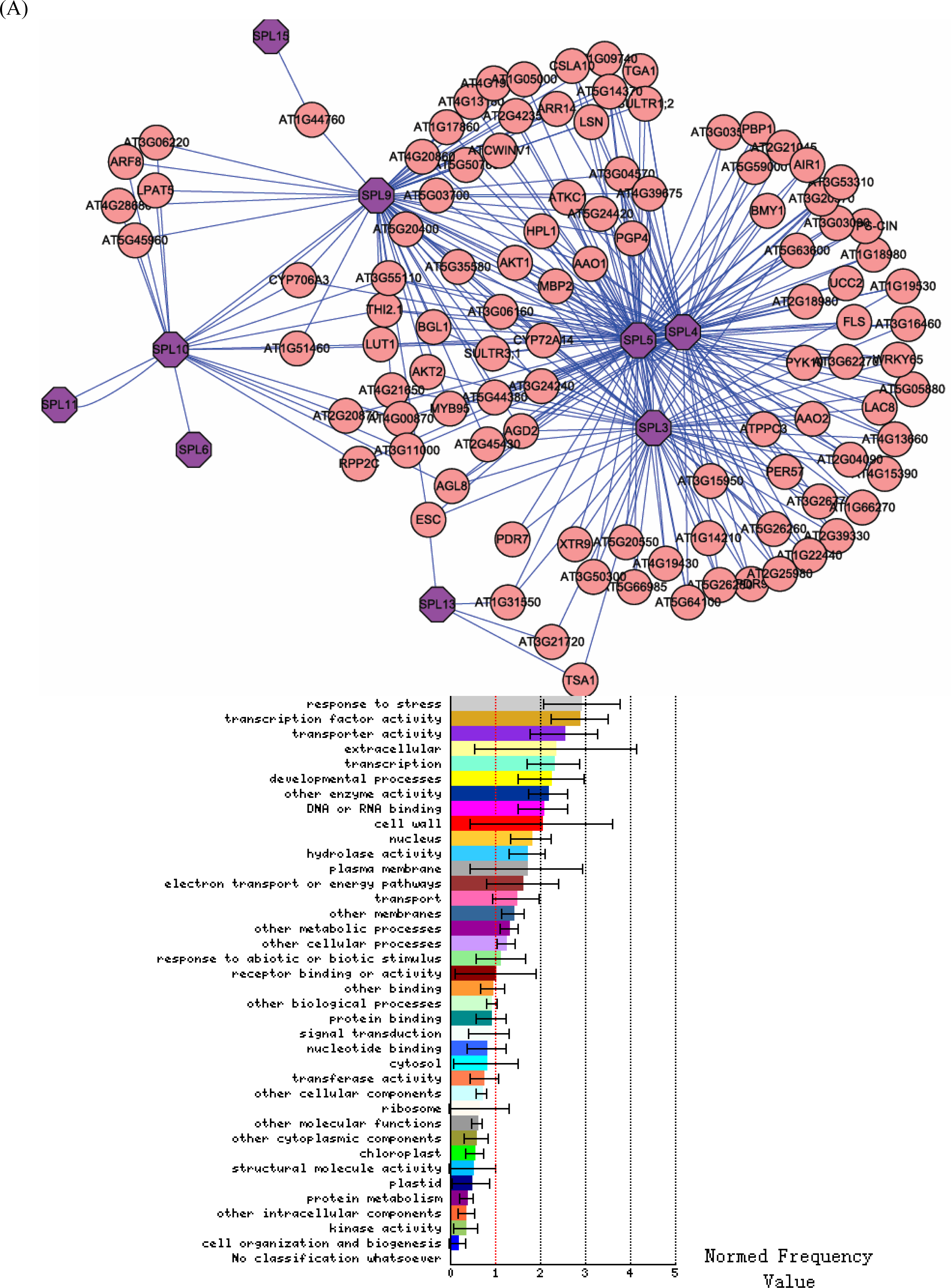
3.1. Expression correlation and gene ontology (GO) analyses
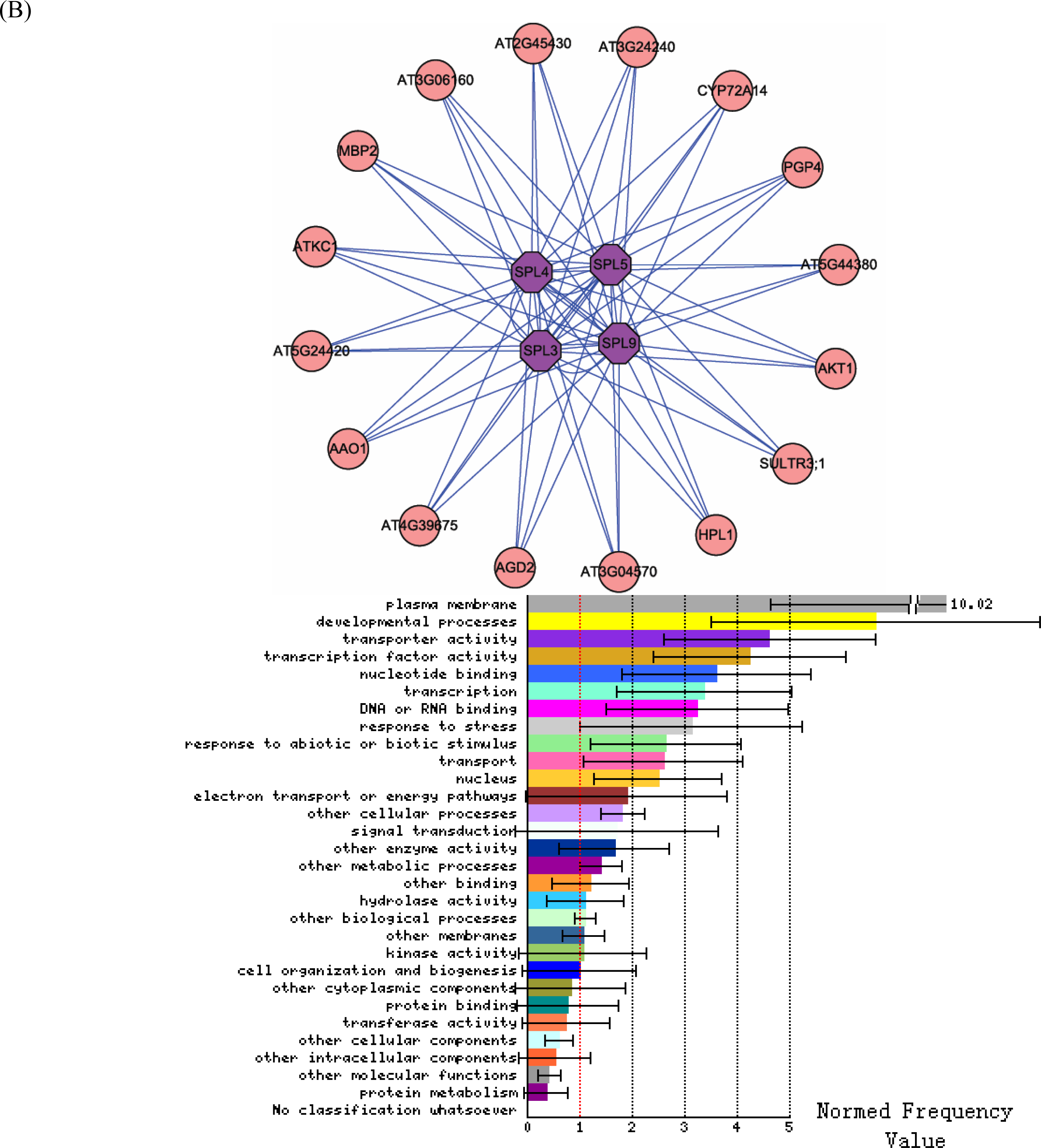
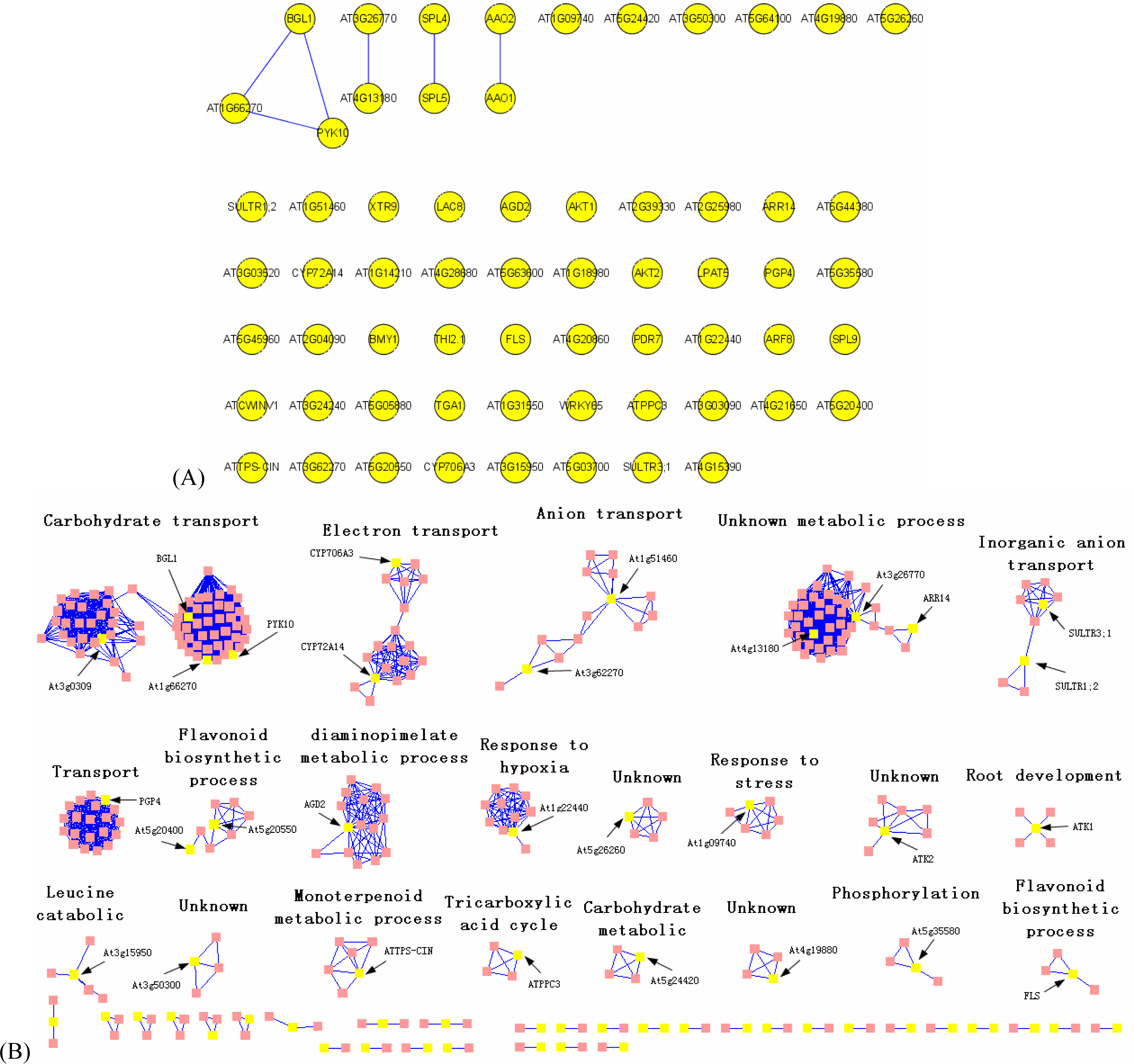
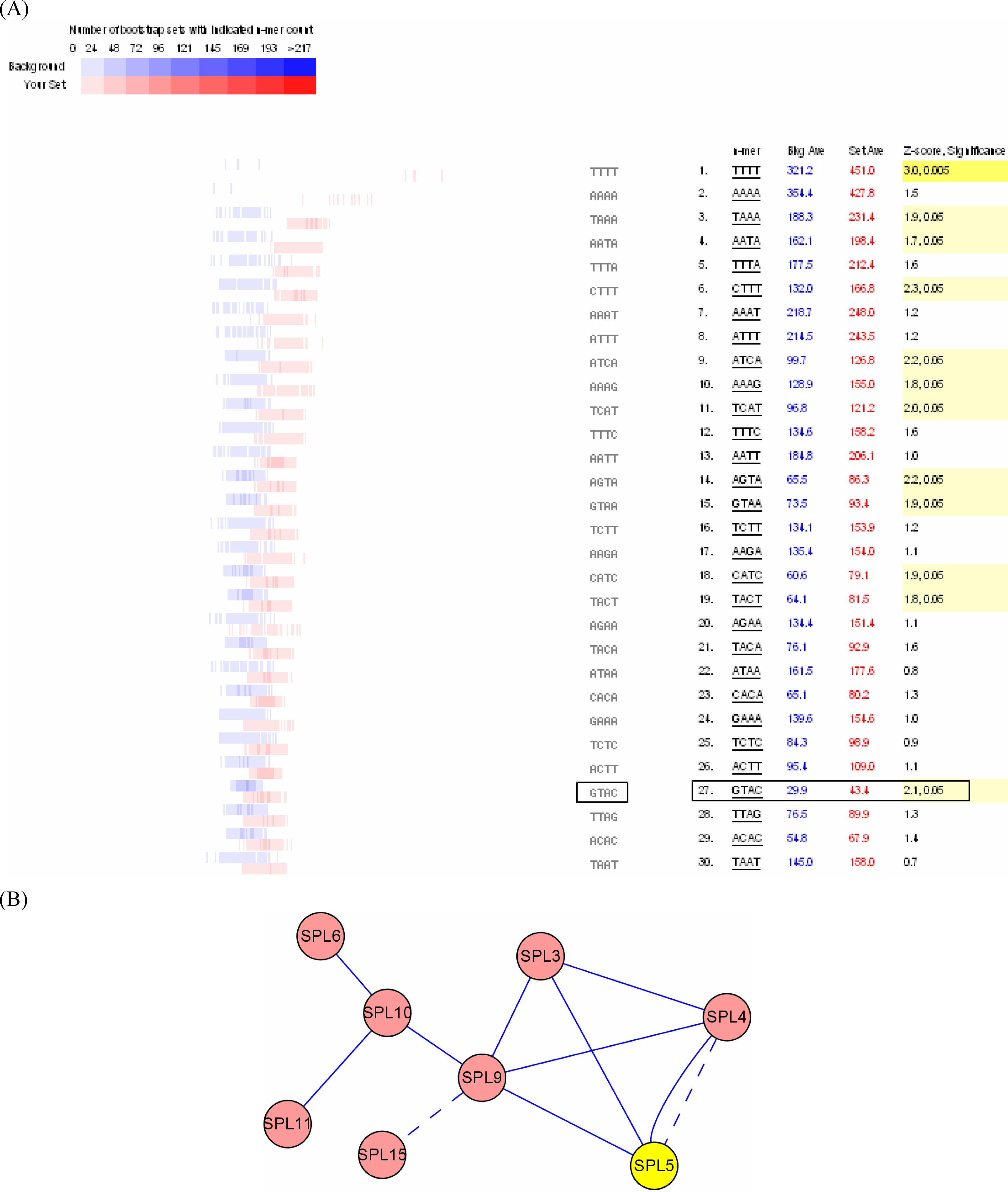
3.2. Interactome analysis
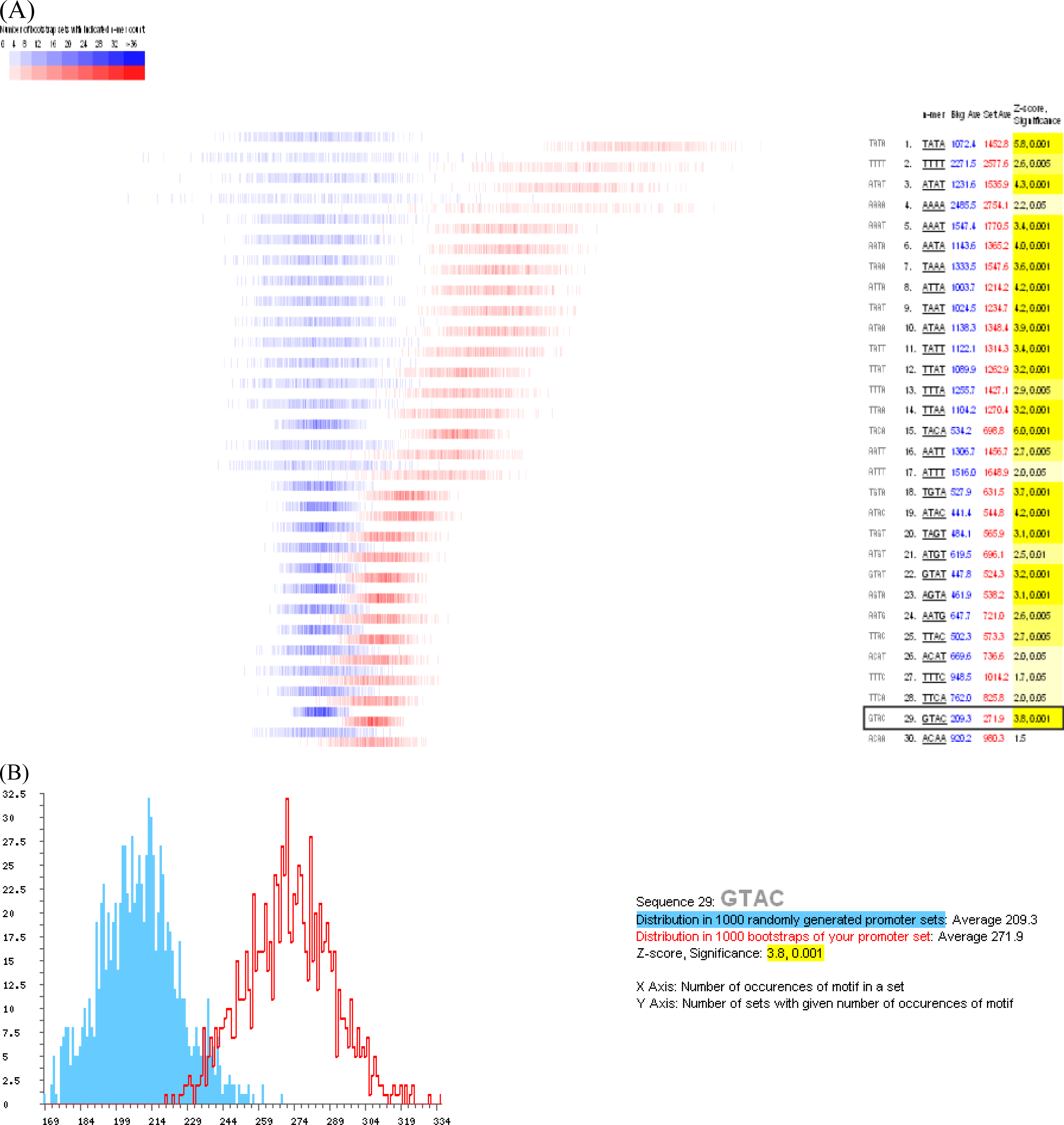
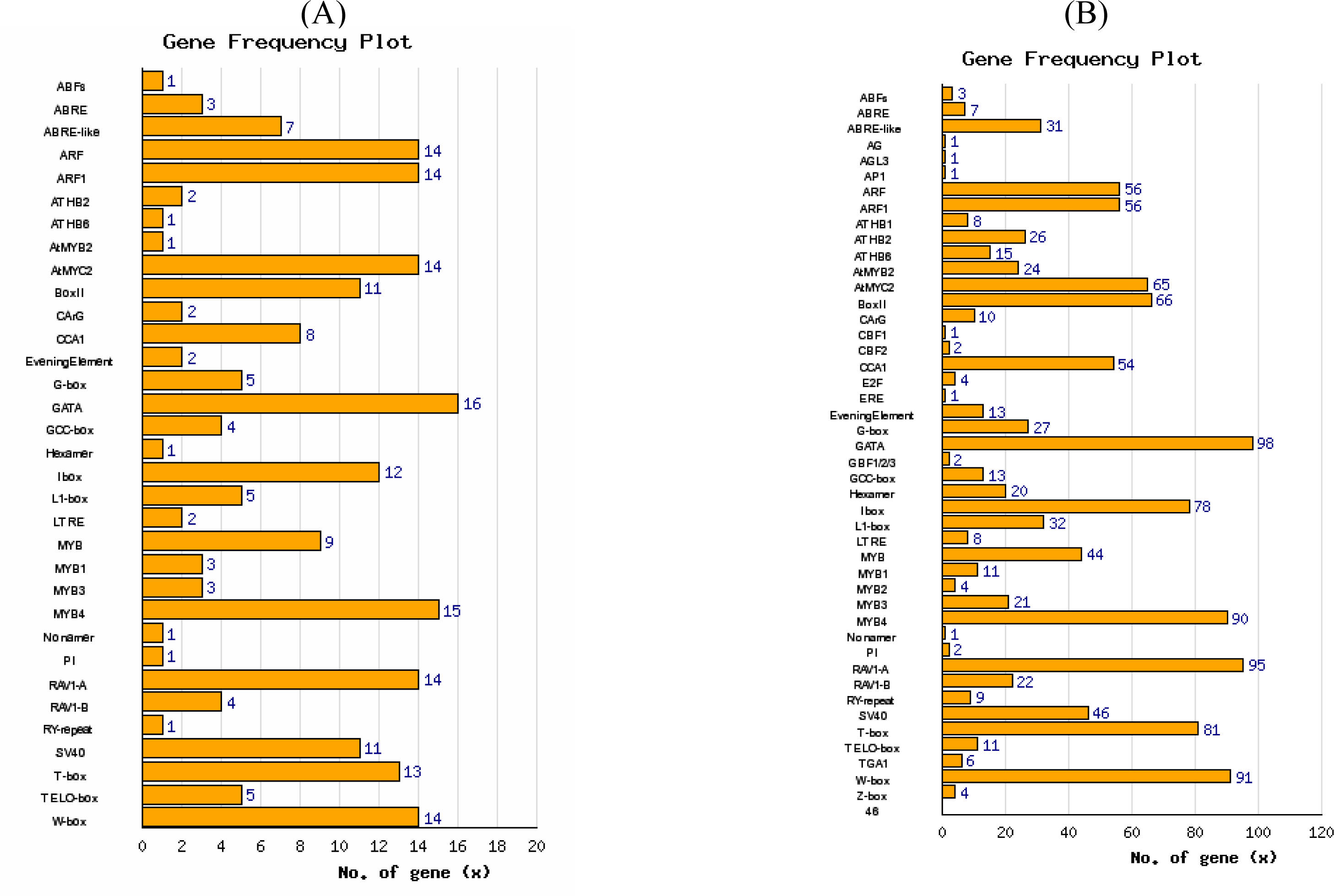
3.3. Promoter analysis
4. Discussion
4.1. SPL genes may play a key role in the stress response
4.2. A complex control network linking SPL genes and other transcription families
4.3. SPL genes and integral membrane proteins
4.4. SPL genes may regulate themselves
4.5. SPL genes have varied functions
5. Conclusion
Appendix: Supplementary Material
Acknowledgments
References
- Riechmann, JL; Heard, J; Martin, G; Reuber, L; Jiang, C; Keddie, J; Adam, L; Pineda, O; Ratcliffe, OJ; Samaha, RR; et al. Arabidopsis transcription factors: genome-wide comparative analysis among eukaryotes. Science 2000, 290, 2105–2110. [Google Scholar]
- Cardon, GH; Hohmann, S; Nettesheim, K; Saedler, H; Huijser, P. Functional analysis of the Arabidopsis thaliana SBP-box gene SPL3: a novel gene involved in the floral transition. Plant J 1997, 12, 367–77. [Google Scholar]
- Cardon, G; Hohmann, S; Klein, J; Nettesheim, K; Saedler, H; Huijser, P. Molecular characterisation of the Arabidopsis SBP-box genes. Gene 1999, 237, 91–104. [Google Scholar]
- Klein, J; Saedler, H; Huijser, P. A new family of DNA binding proteins includes putative transcriptional regulators of the Antirrhinum majus floral meristem identity gene SQUAMOSA. Mol. Gen. Genet 1996, 250, 7–16. [Google Scholar]
- Huijser, P; Klein, J; Lonnig, WE; Meijer, H; Saedler, H; Sommer, H. Bracteomania, an inflorescence anomaly, is caused by the loss of function of the MADS-box gene squamosa in Antirrhinum majus. EMBO J 1992, 11, 1239–49. [Google Scholar]
- Unte, US; Sorensen, AM; Pesaresi, P; Gandikota, M; Leister, D; Saedler, H; Huijser, P. SPL8, an SBP-box gene that affects pollen sac development in Arabidopsis. Plant Cell 2003, 15, 1009–1019. [Google Scholar]
- Stone, JM; Liang, X; Nekl, ER; Stiers, JJ. Arabidopsis AtSPL14, a plant-specific SBP-domain transcription factor, participates in plant development and sensitivity to fumonisin B1. Plant J 2005, 41, 744–754. [Google Scholar]
- Wang, H; Nussbaum-Wagler, T; Li, B; Zhao, Q; Vigouroux, Y; Faller, M; Bomblies, K; Lukens, L; Doebley, JF. The origin of the naked grains of maize. Nature 2005, 436, 714–719. [Google Scholar]
- Manning, K; Tor, M; Poole, M; Hong, Y; Thompson, AJ; King, GJ; Giovannoni, JJ; Seymour, GB. A naturally occurring epigenetic mutation in a gene encoding an SBP-box transcription factor inhibits tomato fruit ripening. Nat. Genet 2006, 38, 948–952. [Google Scholar]
- Schwarz, S; Grande, AV; Bujdoso, N; Saedler, H; Huijser, P. The microRNA regulated SBP-box genes SPL9 and SPL15 control shoot maturation in Arabidopsis. Plant Mol. Biol 2008, 67, 183–95. [Google Scholar]
- Schmid, M; Davison, TS; Henz, SR; Pape, UJ; Demar, M; Vingron, M; Scholkopf, B; Weigel, D; Lohmann, JU. A gene expression map of Arabidopsis thaliana development. Nat. Genet 2005, 37, 501–506. [Google Scholar]
- Barrett, T; Troup, DB; Wilhite, SE; Ledoux, P; Rudnev, D; Evangelista, C; Kim, IF; Soboleva, A; Tomashevsky, M; Edgar, R. NCBI GEO: mining tens of millions of expression profiles—database and tools update. Nucleic Acids Res 2007, 35, D760–D765. [Google Scholar]
- Kilian, J; Whitehead, D; Horak, J; Wanke, D; Weinl, S; Batistic, O; D’Angelo, C; Bornberg-Bauer, E; Kudla, J; Harter, K. The AtGenExpress global stress expression data set: Protocols, evaluation and model data analysis of UV-B light, drought and cold stress responses. Plant J 2007, 50, 347–363. [Google Scholar]
- Irizarry, RA; Bolstad, BM; Collin, F; Cope, LM; Hobbs, B; Speed, TP. Summaries of Affymetrix GeneChip probe level data. Nucleic Acids Res 2003, 31, e15. [Google Scholar]
- Qin, LX; Beyer, RP; Hudson, FN; Linford, NJ; Morris, DE; Kerr, KF. Evaluation of methods for oligonucleotide array data via quantitative real-time PCR. BMC Bioinformatics 2006, 7, 23. [Google Scholar]
- Wei, H; Persson, S; Mehta, T; Srinivasasainagendra, V; Chen, L; Page, GP; Somerville, C; Loraine, A. Transcriptional coordination of the metabolic network in Arabidopsis. Plant Physiol 2006, 142, 762–774. [Google Scholar]
- Maere, S; Heymans, K; Kuiper, M. BiNGO: A Cytoscape plugin to assess overrepresentation of gene ontology categories in biological networks. Bioinformatics 2005, 21, 3448–3449. [Google Scholar]
- Garcia, O; Saveanu, C; Cline, M; Fromont-Racine, M; Jacquier, A; Schwikowski, B; Aittokallio, T. GOlorize: A Cytoscape plug-in for network visualization with Gene Ontology-based layout and coloring. Bioinformatics 2007, 23, 394–396. [Google Scholar]
- Toufighi, K; Brady, SM; Austin, R; Ly, E; Provart, NJ. The Botany Array Resource: e-Northerns, Expression Angling, and promoter analyses. Plant J 2005, 43, 153–163. [Google Scholar]
- Kropat, J; Tottey, S; Birkenbihl, RP; Depege, N; Huijser, P; Merchant, S. A regulator of nutritional copper signaling in Chlamydomonas is an SBP domain protein that recognizes the GTAC core of copper response element. Proc. Natl. Acad. Sci. USA 2005, 102, 18730–18735. [Google Scholar]
- Birkenbihl, RP; Jach, G; Saedler, H; Huijser, P. Functional dissection of the plant-specific SBP-domain: Overlap of the DNA-binding and nuclear localization domains. J. Mol. Biol 2005, 352, 585–596. [Google Scholar]
- Liang, X; Nazarenus, TJ; Stone, JM. Identification of a consensus DNA-binding site for the Arabidopsis thaliana SBP domain transcription factor, AtSPL14, and binding kinetics by surface plasmon resonance. Biochemistry 2008, 47, 3645–3653. [Google Scholar]
- Chen, W; Provart, NJ; Glazebrook, J; Katagiri, F; Chang, HS; Eulgem, T; Mauch, F; Luan, S; Zou, G; Whitham, SA; et al. Expression profile matrix of Arabidopsis transcription factor genes suggests their putative functions in response to environmental stresses. Plant Cell 2002, 14, 559–574. [Google Scholar]
- Jain, M; Nijhawan, A; Arora, R; Agarwal, P; Ray, S; Sharma, P; Kapoor, S; Tyagi, AK; Khurana, JP. F-box proteins in rice. Genome-wide analysis, classification, temporal and spatial gene expression during panicle and seed development, and regulation by light and abiotic stress. Plant Physiol 2007, 143, 1467–1483. [Google Scholar]
- Arora, R; Agarwal, P; Ray, S; Singh, AK; Singh, VP; Tyagi, AK; Kapoor, S. MADS-box gene family in rice: Genome-wide identification, organization and expression profiling during reproductive development and stress. BMC Genomics 2007, 8, 242. [Google Scholar]
- Nijhawan, A; Jain, M; Tyagi, AK; Khurana, JP. Genomic survey and gene expression analysis of the basic leucine zipper transcription factor family in rice. Plant Physiol 2008, 146, 333–350. [Google Scholar]
- Egea-Cortines, M; Saedler, H; Sommer, H. Ternary complex formation between the MADS-box proteins SQUAMOSA, DEFICIENS and GLOBOSA is involved in the control of floral architecture in Antirrhinum majus. EMBO J 1999, 18, 5370–5379. [Google Scholar]
- Honma, T; Goto, K. Complexes of MADS-box proteins are sufficient to convert leaves into floral organs. Nature 2001, 409, 525–529. [Google Scholar]
- Immink, RG; Ferrario, S; Busscher-Lange, J; Kooiker, M; Busscher, M; Angenent, GC. Analysis of the petunia MADS-box transcription factor family. Mol. Genet. Genomics 2003, 268, 598–606. [Google Scholar]
- de Folter, S; Immink, RG; Kieffer, M; Parenicova, L; Henz, SR; Weigel, D; Busscher, M; Kooiker, M; Colombo, L; Kater, MM; et al. Comprehensive interaction map of the Arabidopsis MADS Box transcription factors. Plant Cell 2005, 17, 1424–1433. [Google Scholar]
- Laitinen, RA; Broholm, S; Albert, VA; Teeri, TH; Elomaa, P. Patterns of MADS-box gene expression mark flower-type development in Gerbera hybrida (Asteraceae). BMC Plant Biol 2006, 6, 11. [Google Scholar]
- Jakoby, M; Weisshaar, B; Droge-Laser, W; Vicente-Carbajosa, J; Tiedemann, J; Kroj, T; Parcy, F. bZIP transcription factors in Arabidopsis. Trends Plant Sci 2002, 7, 106–111. [Google Scholar]
- Lebel, E; Heifetz, P; Thorne, L; Uknes, S; Ryals, J; Ward, E. Functional analysis of regulatory sequences controlling PR-1 gene expression in Arabidopsis. Plant J 1998, 16, 223–233. [Google Scholar]
- Chen, W; Singh, KB. The auxin, hydrogen peroxide and salicylic acid induced expression of the Arabidopsis GST6 promoter is mediated in part by an ocs element. Plant J 1999, 19, 667–677. [Google Scholar]
- Eulgem, T; Rushton, PJ; Robatzek, S; Somssich, IE. The WRKY superfamily of plant transcription factors. Trends Plant Sci 2000, 5, 199–206. [Google Scholar]
- Mandel, MA; Gustafson-Brown, C; Savidge, B; Yanofsky, MF. Molecular characterization of the Arabidopsis floral homeotic gene APETALA1. Nature 1992, 360, 273–237. [Google Scholar]
- Ferrandiz, C; Gu, Q; Martienssen, R; Yanofsky, MF. Redundant regulation of meristem identity and plant architecture by FRUITFULL, APETALA1 and CAULIFLOWER. Development 2000, 127, 725–734. [Google Scholar]
- Cui, J; Li, P; Li, G; Xu, F; Zhao, C; Li, Y; Yang, Z; Wang, G; Yu, Q; Shi, T. AtPID: Arabidopsis thaliana protein interactome database--an integrative platform for plant systems biology. Nucleic Acids Res 2008, 36, D999–1008. [Google Scholar]
- Guo, AY; Zhu, QH; Gu, X; Ge, S; Yang, J; Luo, J. Genome-wide identification and evolutionary analysis of the plant specific SBP-box transcription factor family. Gene 2008, 418, 1–8. [Google Scholar]
- Becraft, PW; Bongard-Pierce, DK; Sylvester, AW; Poethig, RS; Freeling, M. The liguleless-1 gene acts tissue specifically in maize leaf development. Dev. Biol 1990, 141, 220–232. [Google Scholar]
- Eriksson, M; Moseley, JL; Tottey, S; Del Campo, JA; Quinn, J; Kim, Y; Merchant, S. Genetic dissection of nutritional copper signaling in chlamydomonas distinguishes regulatory and target genes. Genetics 2004, 168, 795–807. [Google Scholar]
- Zhang, H; Rong, H; Pilbeam, D. Signalling mechanisms underlying the morphological responses of the root system to nitrogen in Arabidopsis thaliana. J. Exp. Bot 2007, 58, 2329–2338. [Google Scholar]
| AGI ID | Annotation | AGI ID | Annotation |
|---|---|---|---|
| AT5G20960 | ALDEHYDE OXIDASE 1 | AT3G18850 | LPAT5__LPAT5 |
| AT1G52030 | MYROSINASE-BINDING PROTEIN 2 | AT4G00870 | basic helix-loop-helix (bHLH) family protein |
| AT2G26650 | ARABIDOPSIS K TRANSPORTER 1 | AT5G37020 | ARF8 |
| AT3G55110 | ABC transporter family protein | AT3G09260 | phosphate starvation-response 3.1 oxidoreductase, 2OG-Fe(II) oxygenase family |
| AT4G15440 | HYDROPEROXIDE LYASE 1 | AT5G20400 | protein |
| AT3G06160 | transcriptional factor B3 family protein | AT4G19430 | unknown protein |
| AT5G44380 | FAD-binding domain-containing protein | AT5G17820 | peroxidase 57 (PER57) (P57) (PRXR10) |
| AT5G60910 | FUL | AT3G06220 | DNA binding / transcription factor |
| AT2G42200 | SPL9 | AT5G45960 | GDSL-motif lipase/hydrolase family protein |
| AT2G45430 | DNA-binding protein-related | AT4G21650 | subtilase family protein UCC2__UCC2 (UCLACYANIN 2); copper |
| AT3G04570 | DNA-binding protein-related | AT2G44790 | ion binding |
| AT4G32650 | ARABIDOPSIS THALIANA K+ RECTIFYING CHANNEL 1 | AT3G28500 | 60S acidic ribosomal protein P2 (RPP2C) |
| AT3G14680 | CYP72A14 | AT4G12550 | Auxin-Induced in Root cultures 1 |
| AT1G20900 | ESCAROLA | AT4G20860 | FAD-binding domain-containing protein |
| AT1G60680 | ARF-GAP DOMAIN 2 | AT4G28680 | tyrosine decarboxylase, putative |
| AT3G53130 | LUTEIN DEFICIENT 1 | AT5G63600 | flavonol synthase, putative |
| AT3G51895 | SULFATE TRANSPORTER 1 | AT2G01760 | ARABIDOPSIS RESPONSE REGULATOR 14 |
| AT2G47000 | P-GLYCOPROTEIN 4 | AT5G59000 | zinc finger (C3HC4-type RING finger) family protein |
| AT4G39675 | unknown protein | AT5G03700 | PAN domain-containing protein |
| AT5G24420 | glucosamine/galactosamine-6-phosphate isomerase-related | AT2G21045 | similar to unknown protein |
| AT3G24240 | leucine-rich repeat transmembrane protein kinase, putative | AT5G50760 | auxin-responsive family protein |
| AT1G72260 | toxin receptor binding | AT3G15950 | (TSA1-LIKE); unknown protein |
| AT3G15270 | SPL5 | AT1G17860 | trypsin and protease inhibitor family protein |
| AT2G33810 | SPL3 | AT2G20870 | cell wall protein precursor, putative |
| AT1G53160 | SPL4 | AT5G20550 | oxidoreductase |
| AT5G05880 | UDP-glucosyl transferase family protein | AT1G05000 | tyrosine specific protein phosphatase family protein |
| AT1G24070 | Cellulose synthase-like A10 | AT3G43600 | AAO2 |
| AT3G20370 | MATH domain-containing protein | AT4G22200 | Arabidopsis K+ transporter 2 |
| AT5G26280 | MATH domain-containing protein | AT3G21720 | isocitrate lyase, putative |
| AT1G51460 | ABC transporter family protein | AT1G52410 | TSK-ASSOCIATING PROTEIN 1 |
| AT3G16460 | jacalin lectin family protein | AT4G19880 | similar to unknown protein |
| AT3G53480 | PLEIOTROPIC DRUG RESISTANCE 9 | AT2G18980 | peroxidase, putative |
| AT5G44620 | CYP706A3 | AT1G18980 | germin-like protein, putative |
| AT5G01040 | laccase 8 | AT4G13180 | short-chain dehydrogenase |
| AT5G65210 | TGA1 | AT5G63590 | Flavonol synthase |
| AT1G52400 | BGL1 | AT3G03090 | ATVGT1__sugar transporter family protein |
| AT5G14370 | similar to CIL | AT3G25820 | TERPENE SYNTHASE-LIKE SEQUENCE-1,8-CINEOLE |
| AT1G66270 | beta-glucosidase | AT4G15210 | BETA-AMYLASE |
| AT5G02030 | HB-6_RPL_LSN_BLH9_BLR_PNY_RPL_VAN__LSN | AT2G42350 | zinc finger family protein |
| AT4G13660 | pinoresinol-lariciresinol reductase, putative | AT3G50300 | transferase family protein |
| AT4G15390 | transferase family protein | AT3G03520 | phosphoesterase family protein |
| AT5G64100 | peroxidase, putative | AT1G15210 | PLEIOTROPIC DRUG RESISTANCE 7 |
| AT1G78000 | SULFATE TRANSPORTER 1;2 | AT3G16420 | PYK10-BINDING PROTEIN 1 |
| AT1G29280 | WRKY DNA-binding protein 65 | AT3G13790 | ARABIDOPSIS THALIANA CELL WALL INVERTASE 1 |
| AT2G39330 | jacalin lectin family protein | AT3G62270 | anion exchange family protein |
| AT1G09740 | ethylene-responsive protein, putative | AT4G25820 | XYLOGLUCAN ENDOTRANSGLYCOSYLASE 9 |
| AT1G22440 | alcohol dehydrogenase, putative | AT1G27370 | SPL10 |
| AT5G26260 | MATH domain-containing protein | AT5G66985 | unknown protein |
| AT2G25980 | jacalin lectin family protein | AT3G26770 | short-chain dehydrogenase |
| AT1G31550 | carboxylic ester hydrolase/lipase | AT3G11000 | similar to kelch repeat-containing protein |
| AT1G19530 | unknown protein | AT3G14940 | PHOSPHOENOLPYRUVATE CARBOXYLASE 3 |
| AT1G74430 | AtMYB95 | AT1G44760 | universal stress protein (USP) family protein |
| AT2G04090 | MATE efflux family protein | AT1G14210 | ribonuclease T2 family protein |
| AT5G35580 | kinase | AT1G27360 | SPL11 |
| AT3G53310 | transcriptional factor B3 family protein | AT1G69170 | SPL6 |
| AT5G50570 | SPL13 | AT3G57920 | SPL15 |
© 2009 by the authors; licensee Molecular Diversity Preservation International, Basel, Switzerland. This article is an open-access article distributed under the terms and conditions of the Creative Commons Attribution license ( http://creativecommons.org/licenses/by/3.0/). This article is an open-access article distributed under the terms and conditions of the Creative Commons Attribution license ( http://creativecommons.org/licenses/by/3.0/).
Share and Cite
Wang, Y.; Hu, Z.; Yang, Y.; Chen, X.; Chen, G. Function Annotation of an SBP-box Gene in Arabidopsis Based on Analysis of Co-expression Networks and Promoters. Int. J. Mol. Sci. 2009, 10, 116-132. https://doi.org/10.3390/ijms10010116
Wang Y, Hu Z, Yang Y, Chen X, Chen G. Function Annotation of an SBP-box Gene in Arabidopsis Based on Analysis of Co-expression Networks and Promoters. International Journal of Molecular Sciences. 2009; 10(1):116-132. https://doi.org/10.3390/ijms10010116
Chicago/Turabian StyleWang, Yi, Zongli Hu, Yuxin Yang, Xuqing Chen, and Guoping Chen. 2009. "Function Annotation of an SBP-box Gene in Arabidopsis Based on Analysis of Co-expression Networks and Promoters" International Journal of Molecular Sciences 10, no. 1: 116-132. https://doi.org/10.3390/ijms10010116
APA StyleWang, Y., Hu, Z., Yang, Y., Chen, X., & Chen, G. (2009). Function Annotation of an SBP-box Gene in Arabidopsis Based on Analysis of Co-expression Networks and Promoters. International Journal of Molecular Sciences, 10(1), 116-132. https://doi.org/10.3390/ijms10010116