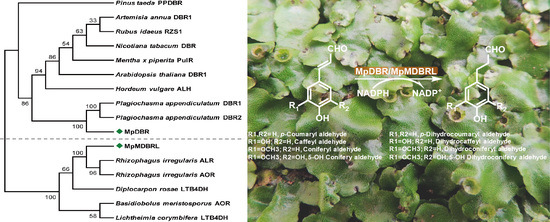
Molecular Diversity of Alkenal Double Bond Reductases in the Liverwort Marchantia paleacea
Abstract
:1. Introduction
2. Results and Discussion
2.1. Isolation and Sequence Analysis of MpDBR and MpMDBRL
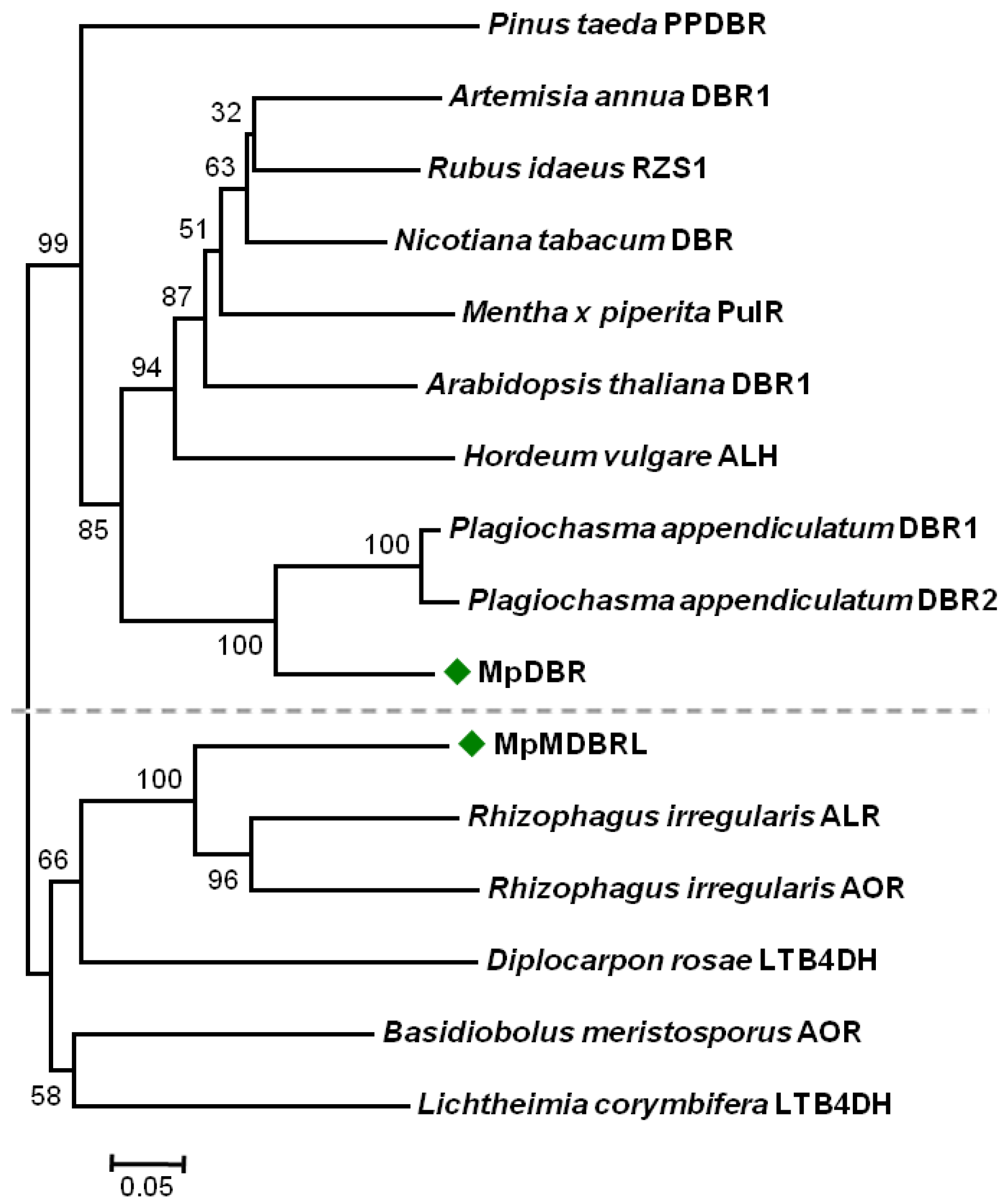
2.2. Sequence Alignment and Phylogenetic Analysis
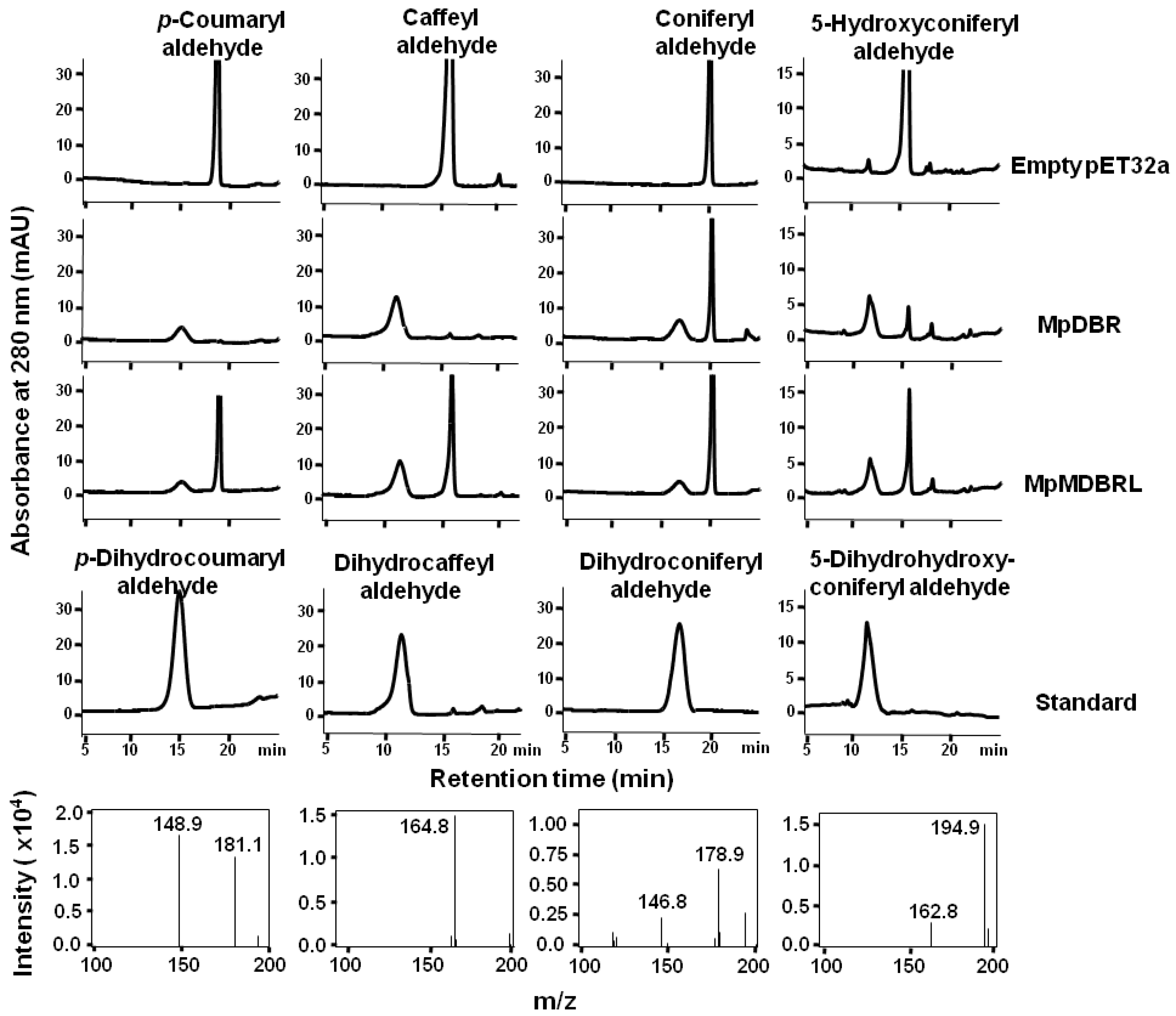
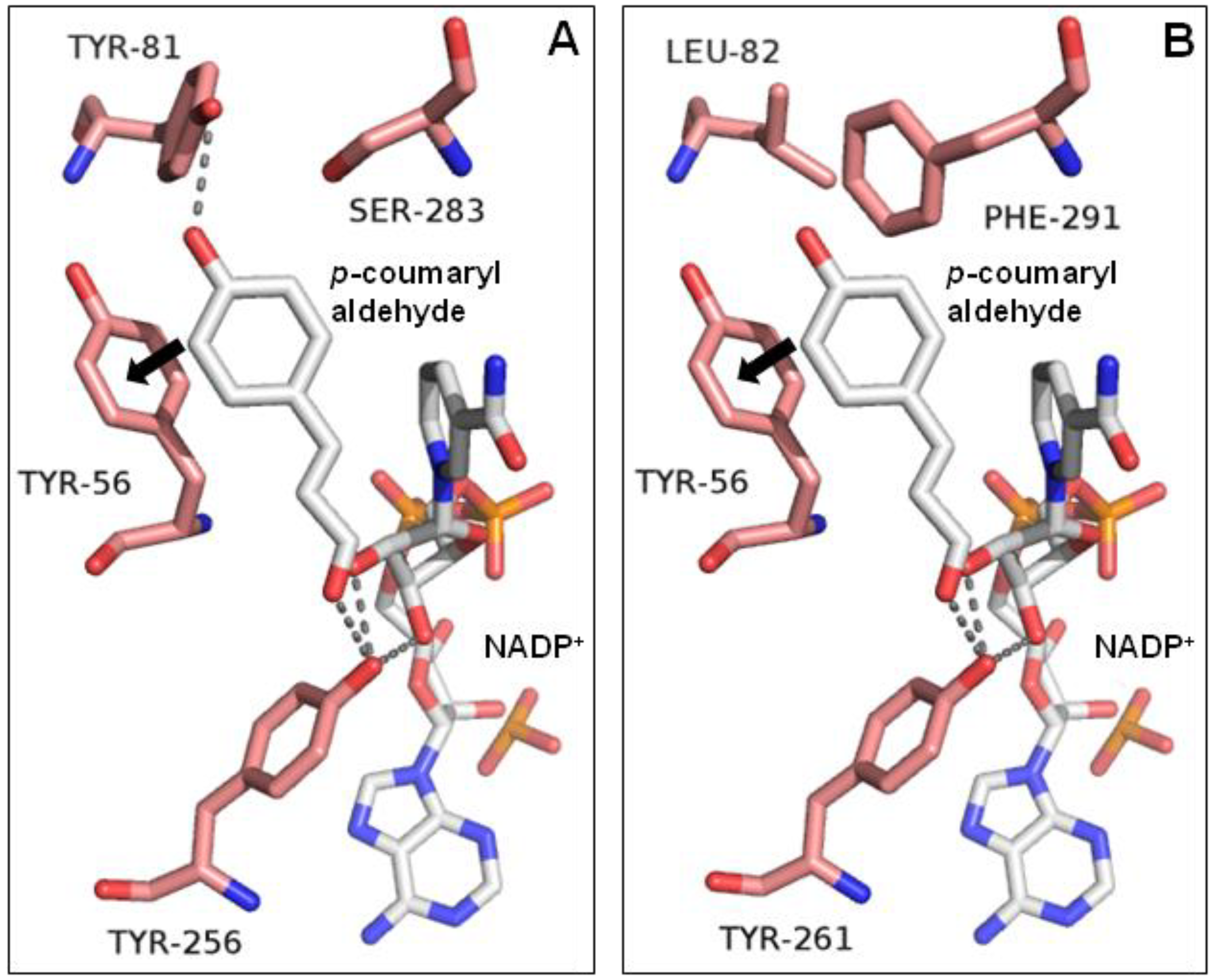
2.3. Functional Analysis
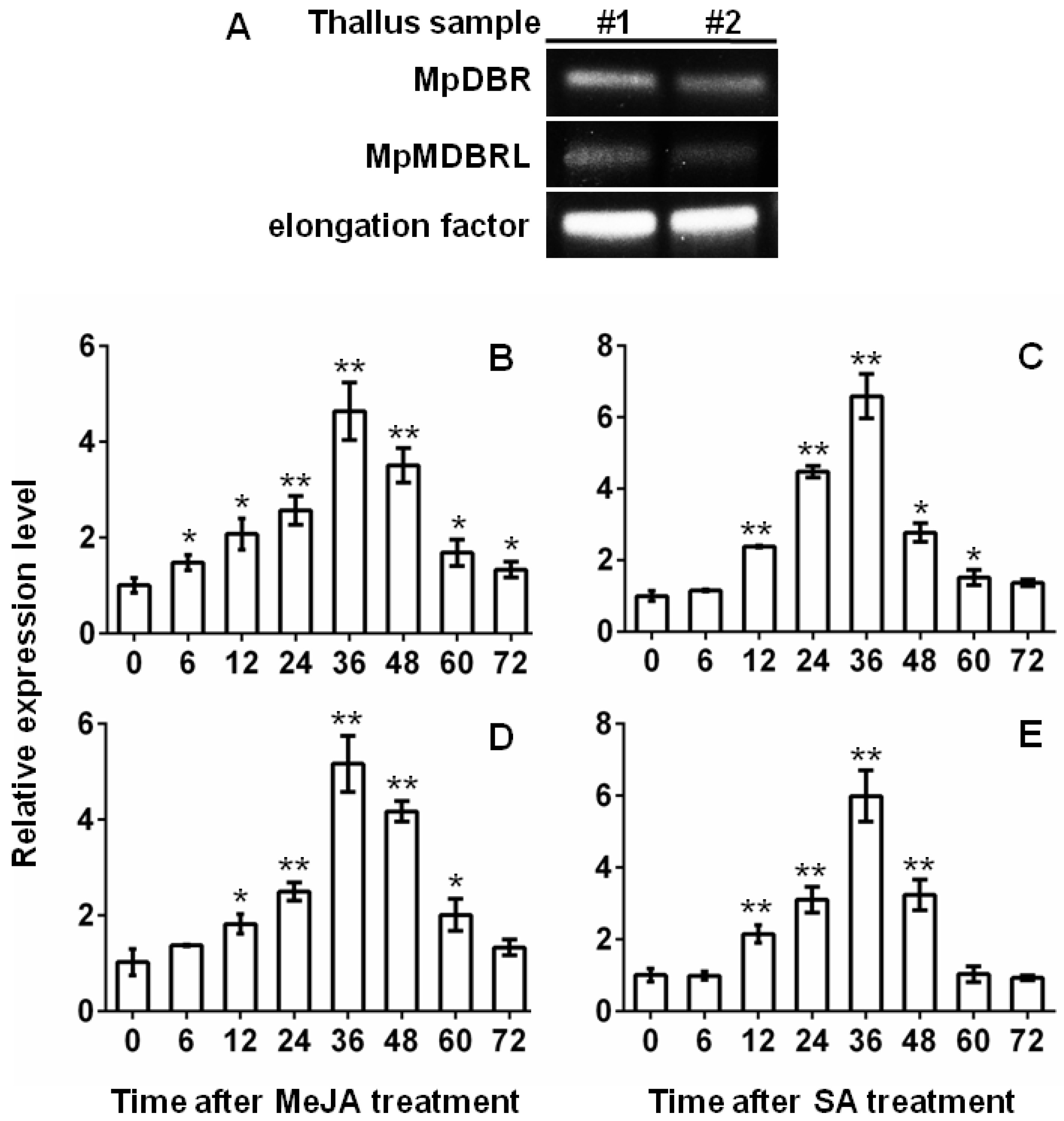
2.4. Transcript Abundance of DBRs and Their Response to Phytohormone Treatment
3. Materials and Methods
3.1. Chemicals and Reagents
3.2. The Isolation and Analysis of DBR Sequences
3.3. Sequence Alignment and Phylogenetic Analysis
3.4. Recombinant Protein Expression and Purification
3.5. Enzyme Assays
3.6. Plant Phytohormone Treatment and Gene Expression Profiling
3.7. Sequence Accession Numbers
4. Conclusions
Supplementary Materials
Author Contributions
Acknowledgments
Conflicts of Interest
References
- Yamauchi, Y.; Hasegawa, A.; Taninaka, A.; Mizutani, M.; Sugimoto, Y. NADPH-dependent reductases involved in the detoxification of reactive carbonyls in plants. J. Biol. Chem. 2011, 286, 6999–7009. [Google Scholar] [CrossRef] [PubMed]
- Esterbauer, H.; Schaur, R.J.; Zollner, H. Chemistry and biochemistry of 4-hydroxynonenal, malonaldehyde and related aldehydes. Free Radic. Biol. Med. 1991, 11, 81–128. [Google Scholar] [CrossRef]
- Mano, J.; Torii, Y.; Hayashi, S.; Takimoto, K.; Matsui, K.; Nakamura, K.; Inzé, D.; Babiychuk, E.; Kushnir, S.; Asada, K. The NADPH: Quinone oxidoreductase P1-zeta-crystallin in Arabidopsis catalyzes the alpha, beta-hydrogenation of 2-alkenals: Detoxication of the lipid peroxide-derived reactive aldehydes. Plant Cell Physiol. 2002, 43, 1445–1455. [Google Scholar] [CrossRef] [PubMed]
- Dick, R.A.; Kwak, M.K.; Sutter, T.R.; Kensler, T.W. Antioxidative function and substrate specificity of NAD(P)H-dependent alkenal/one oxidoreductase. J. Biol. Chem. 2001, 276, 40803–40810. [Google Scholar] [CrossRef] [PubMed]
- Dick, R.A.; Kensler, T.W. The catalytic and kinetic mechanisms of NADPH-dependent alkenal/one oxidoreductase. J. Biol. Chem. 2004, 279, 17269–17277. [Google Scholar] [CrossRef] [PubMed]
- Kasahara, H.; Jiao, Y.; Bedgar, D.L.; Kim, S.J.; Patten, A.M.; Xia, Z.Q.; Davin, L.B.; Lewis, N.G. Pinus taeda phenylpropenal double-bond reductase: Purification, cDNA cloning, heterologous expression in Escherichia coli, and subcellular localization in P. taeda. Phytochemistry 2006, 67, 1765–1780. [Google Scholar] [CrossRef] [PubMed]
- Youn, B.; Kim, S.J.; Moinuddin, S.G.; Lee, C.; Bedgar, D.L.; Harper, A.R.; Davin, L.B.; Lewis, N.G.; Kang, C. Mechanistic and structural studies of apoform, binary, and ternary complexes of the Arabidopsis alkenal double bond reductase At5g16970. J. Biol. Chem. 2006, 281, 40076–40088. [Google Scholar] [CrossRef] [PubMed]
- Wu, Y.; Cai, Y.; Sun, Y.; Xu, R.; Yu, H.; Han, X.; Lou, H.; Cheng, A. A single amino acid determines the catalytic efficiency of two alkenal double bond reductases produced by the liverwort Plagiochasma appendiculatum. FEBS Lett. 2013, 587, 3122–3128. [Google Scholar] [CrossRef] [PubMed]
- Pereira, P.J.B.; Macedoribeiro, S.; Párraga, A.; Pérezluque, R.; Cunningham, O.; Darcy, K.; Mantle, T.J.; Coll, M. Structure of human biliverdin IXβ reductase, an early fetal bilirubin IXβ producing enzyme. Nat. Struct. Mol. Biol. 2001, 8, 215–220. [Google Scholar] [CrossRef] [PubMed]
- Asakawa, Y. Bryophytes: Chemical diversity, synthesis and biotechnology. Flavour Frag. J. 2011, 26, 318–320. [Google Scholar] [CrossRef]
- Kenrick, P.; Crane, P.R. The origin and early evolution of plants on land. Nature 1997, 389, 33–39. [Google Scholar] [CrossRef]
- Edwards, D.; Duckett, J.G.; Richardson, J.B. Hepatic characters in the earliest land plants. Nature 1995, 374, 635–636. [Google Scholar] [CrossRef]
- Delwiche, C.F.; Goodman, C.A.; Chang, C. Land plant model systems branch out. Cell 2017, 171, 265–266. [Google Scholar] [CrossRef] [PubMed]
- Keeling, P.J.; Palmer, J.D. Horizontal gene transfer in eukaryotic evolution. Nat. Rev. Genet. 2008, 9, 605–618. [Google Scholar] [CrossRef] [PubMed]
- Jia, Q.; Li, G.; Köllner, T.G.; Fu, J.; Chen, X.; Xiong, W.; Crandall-Stotler, B.J.; Bowman, J.L.; Weston, D.J.; Zhang, Y.; et al. Microbial-type terpene synthase genes occur widely in nonseed land plants, but not in seed plants. Proc. Natl. Acad. Sci. USA 2016, 113, 12328–12333. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Kumar, S.; Kempinski, C.; Zhuang, X.; Norris, A.; Mafu, S.; Zi, J.; Bell, S.A.; Nybo, S.E.; Kinison, S.E.; Jiang, Z.; et al. Molecular diversity of terpene synthases in the liverwort Marchantia polymorpha. Plant Cell 2016, 28, 2632–2650. [Google Scholar] [CrossRef] [PubMed]
- Bowman, J.L.; Kohchi, T.; Yamato, K.T.; Jenkins, J.; Shu, S.; Ishizaki, K.; Yamaoka, S.; Nishihama, R.; Nakamura, Y.; Berger, F.; et al. Insights into land plant evolution garnered from the Marchantia polymorpha Genome. Cell 2017, 171, 287–304. [Google Scholar] [CrossRef] [PubMed]
- Mansell, D.J.; Toogood, H.S.; Waller, J.; Hughes, J.M.X.; Levy, C.W.; Gardiner, J.M.; Scrutton, N.S. Biocatalytic asymmetric alkene reduction: Crystal structure and characterization of a double bond reductase from Nicotiana tabacum. ACS Catal. 2013, 3, 370–379. [Google Scholar] [CrossRef] [PubMed]
- Koeduka, T.; Watanabe, B.; Suzuki, S.; Hiratake, J.; Mano, J.; Yazaki, K. Characterization of raspberry ketone/zingerone synthase, catalyzing the alpha, beta-hydrogenation of phenylbutenones in raspberry fruits. Biochem. Biophys. Res. Commun. 2011, 412, 104–108. [Google Scholar] [CrossRef] [PubMed]
- Li, G.; Köllner, T.G.; Yin, Y.; Jiang, Y.; Chen, H.; Xu, Y.; Gershenzon, J.; Pichersky, E.; Chen, F. Nonseed plant Selaginella moellendorffii has both seed plant and microbial types of terpene synthases. Proc. Natl. Acad. Sci. USA 2012, 109, 14711–14715. [Google Scholar] [CrossRef] [PubMed]
- Zhang, Y.; Teoh, K.H.; Reed, D.W.; Covello, P.S. Molecular cloning and characterization of Dbr1, a 2-alkenal reductase from Artemisia annua. Botanique 2009, 87, 643–649. [Google Scholar] [CrossRef]
- Kim, S.J.; Kim, M.R.; Bedgar, D.L.; Moinuddin, S.G.; Cardenas, C.L.; Davin, L.B.; Kang, C.; Lewis, N.G. Functional reclassification of the putative cinnamyl alcohol dehydrogenase multigene family in Arabidopsis. Proc. Natl. Acad. Sci. USA 2004, 101, 1455–1460. [Google Scholar] [CrossRef] [PubMed]
- Gao, S.; Yu, H.N.; Wu, Y.F.; Liu, X.Y.; Cheng, A.X.; Lou, H.X. Cloning and functional characterization of a phenolic acid decarboxylase from the liverwort Conocephalum japonicum. Biochem. Biophys. Res. Commun. 2016, 481, 239–244. [Google Scholar] [CrossRef] [PubMed]
- Mano, J.; Bellesboix, E.; Babiychuk, E.; Inzé, D.; Torii, Y.; Hiraoka, E.; Takimoto, K.; Slooten, L.; Asada, K.; Kushnir, S. Protection against photooxidative injury of tobacco leaves by 2-alkenal reductase. Detoxication of lipid peroxide-derived reactive carbonyls. Plant Physiol. 2005, 139, 1773–1783. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Shimizu, T.; Miyamoto, K.; Minami, E.; Nishizawa, Y.; Iino, M.; Nojiri, H.; Yamane, H.; Okada, K. OsJAR1 contributes mainly to biosynthesis of the stress-induced jasmonoyl-isoleucine involved in defense responses in rice. Biosci. Biotechnol. Biochem. 2013, 77, 1556–1564. [Google Scholar] [CrossRef] [PubMed]
- Zhu, Z.; Shi, J.; Cao, J.; He, M.; Wang, Y. VpWRKY3, a biotic and abiotic stress-related transcription factor from the Chinese wild Vitis pseudoreticulata. Plant Cell Rep. 2012, 31, 2109–2120. [Google Scholar] [CrossRef] [PubMed]
- Sun, Y.; Wu, Y.; Zhao, Y.; Han, X.; Lou, H.; Cheng, A. Molecular cloning and biochemical characterization of two cinnamyl alcohol dehydrogenases from a liverwort Plagiochasma appendiculatum. Plant Physiol. Biochem. 2013, 70, 133–141. [Google Scholar] [CrossRef] [PubMed]
- Tamura, K.; Dudley, J.; Nei, M.; Kumar, S. MEGA4: Molecular evolutionary genetics analysis (MEGA) software version 4.0. Mol. Biol. Evol. 2007, 24, 1596–1599. [Google Scholar] [CrossRef] [PubMed]
- Yu, H.N.; Wang, L.; Sun, B.; Gao, S.; Cheng, A.X.; Lou, H.X. Functional characterization of a chalcone synthase from the liverwort Plagiochasma appendiculatum. Plant Cell Rep. 2015, 34, 233–245. [Google Scholar] [CrossRef] [PubMed]
Sample Availability: Samples of the compounds are available from the authors. |


| Substrate | Structure | Specific Activity (nmol mg−1 min−1) | |
|---|---|---|---|
| MpDBR | MpMDBRL | ||
| p-Coumaryl aldehyde |  | 29.08 ± 1.48 | 21.26 ± 1.46 |
| Caffeyl aldehyde |  | 29.60 ± 1.35 | 25.88 ± 0.88 |
| Coniferyl aldehyde |  | 28.61 ± 0.69 | 20.17 ± 0.48 |
| 5-Hydroxyconiferyl aldehyde |  | 26.87 ± 2.21 | 21.34 ± 0.54 |
| Sinapyl aldehyde |  | ND a | ND |
© 2018 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Wu, Y.-F.; Zheng, H.-B.; Liu, X.-Y.; Cheng, A.-X.; Lou, H.-X. Molecular Diversity of Alkenal Double Bond Reductases in the Liverwort Marchantia paleacea. Molecules 2018, 23, 1630. https://doi.org/10.3390/molecules23071630
Wu Y-F, Zheng H-B, Liu X-Y, Cheng A-X, Lou H-X. Molecular Diversity of Alkenal Double Bond Reductases in the Liverwort Marchantia paleacea. Molecules. 2018; 23(7):1630. https://doi.org/10.3390/molecules23071630
Chicago/Turabian StyleWu, Yi-Feng, Hong-Bo Zheng, Xin-Yan Liu, Ai-Xia Cheng, and Hong-Xiang Lou. 2018. "Molecular Diversity of Alkenal Double Bond Reductases in the Liverwort Marchantia paleacea" Molecules 23, no. 7: 1630. https://doi.org/10.3390/molecules23071630






