Replacing Standard Reporters from Molecular Cloning Plasmids with Chromoproteins for Positive Clone Selection
Abstract
:1. Introduction
2. Results
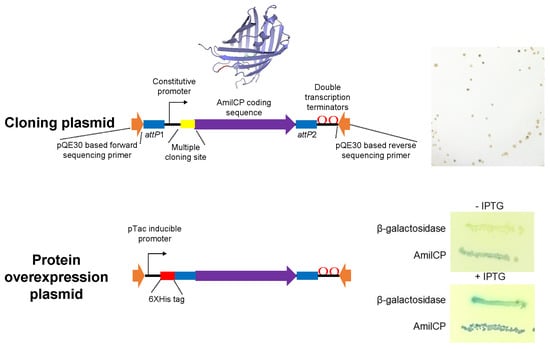
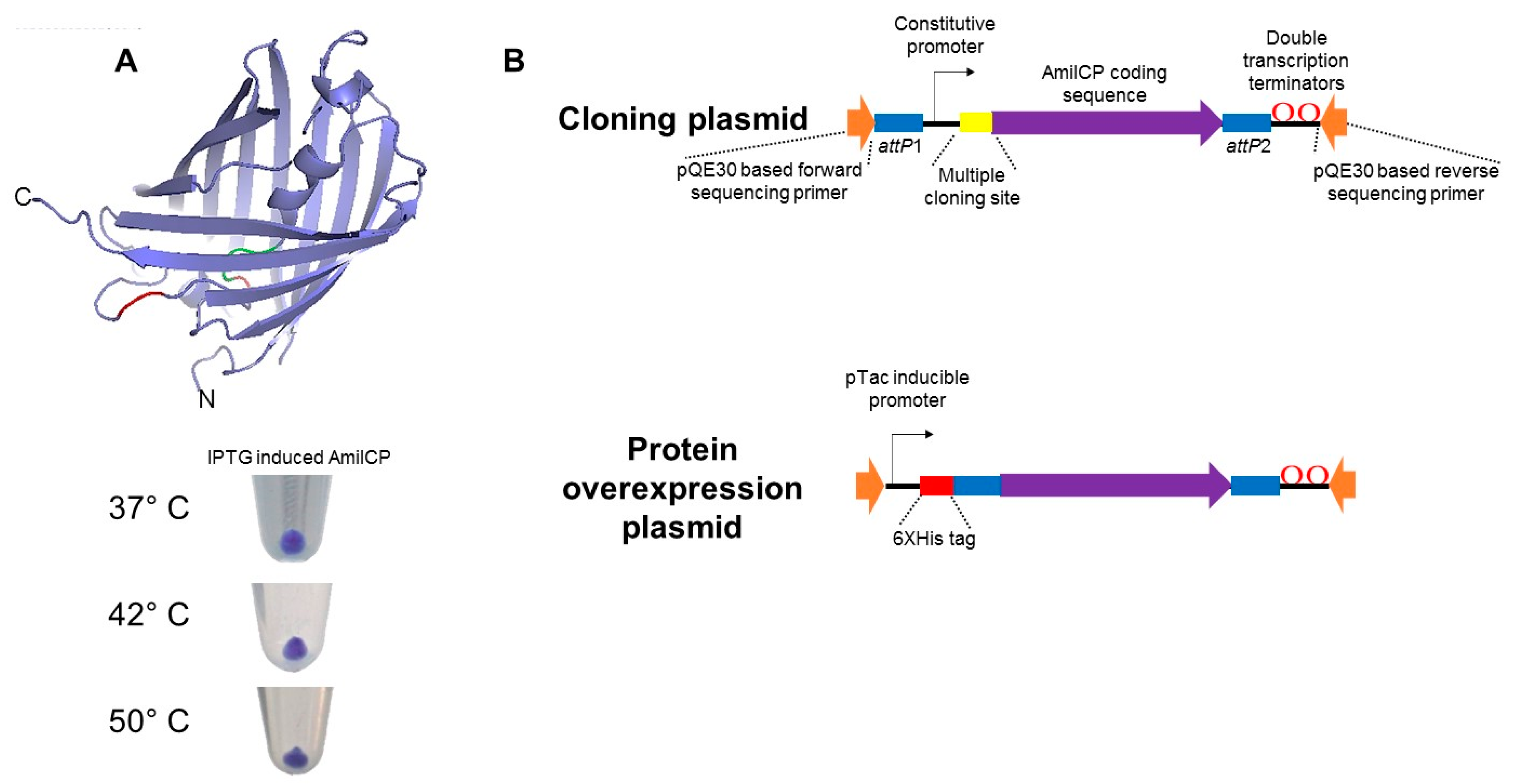
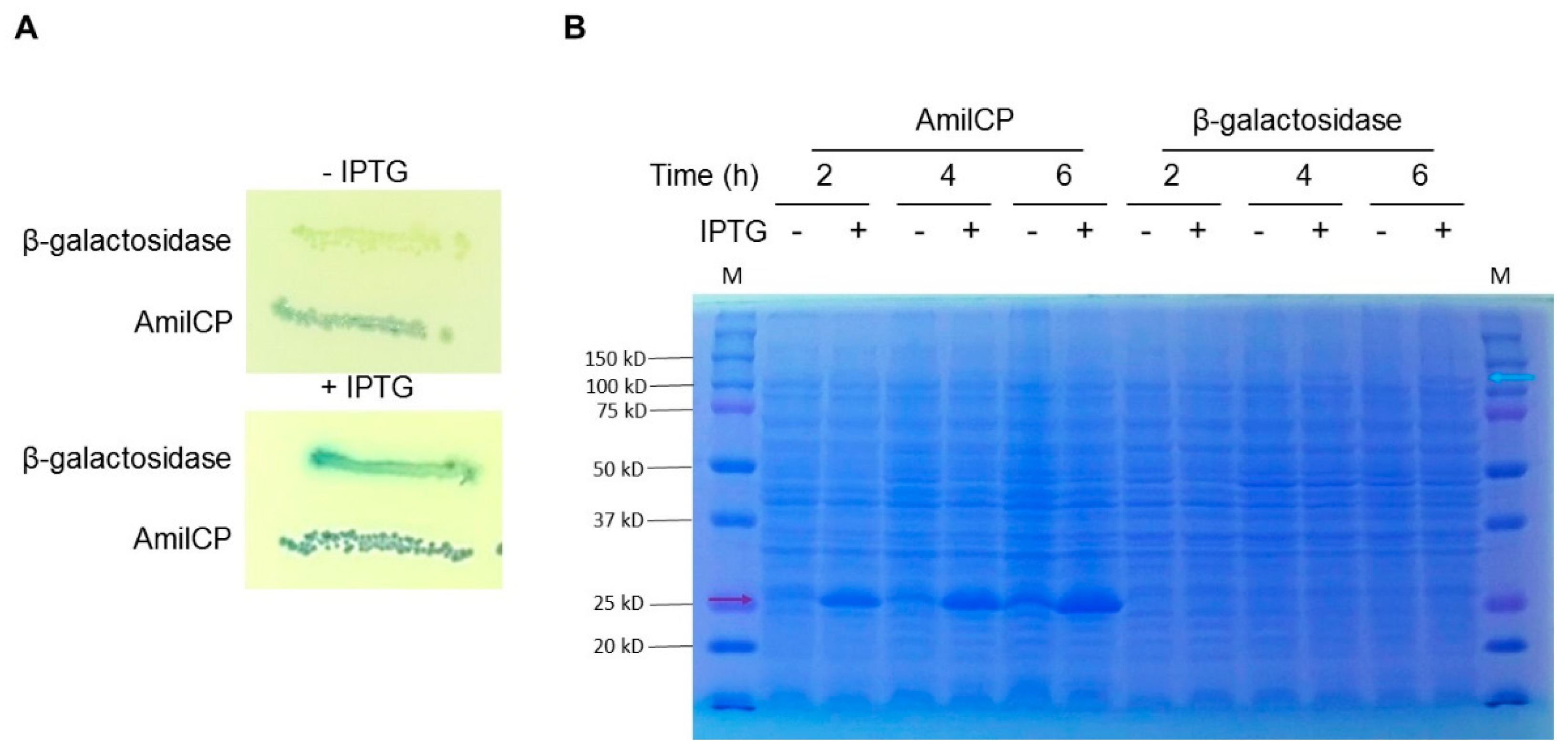
2.1. AmilCP as a Reporter for Cloning
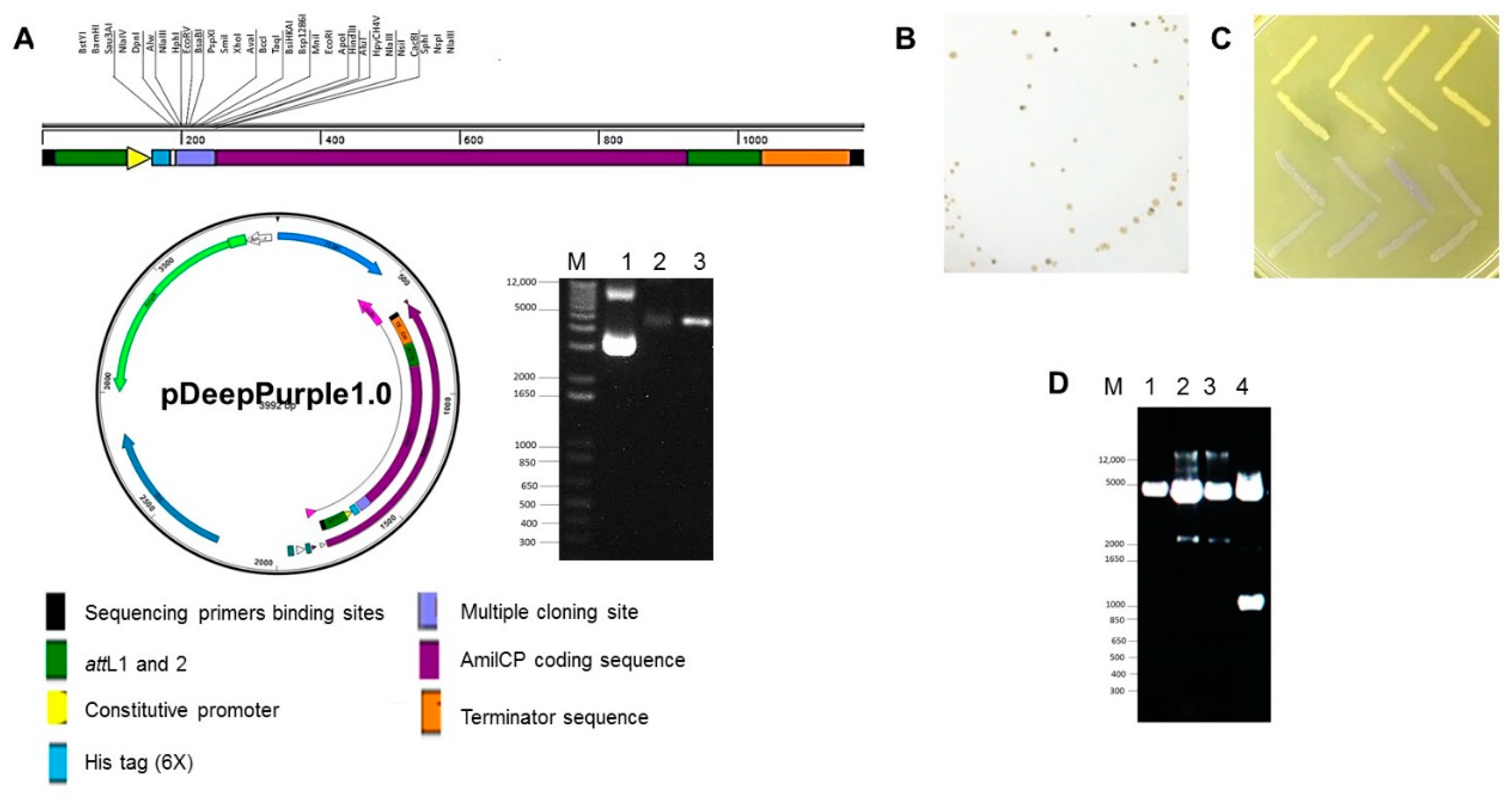
2.2. Generation of Cloning Plasmids Using AmilCP Reporter
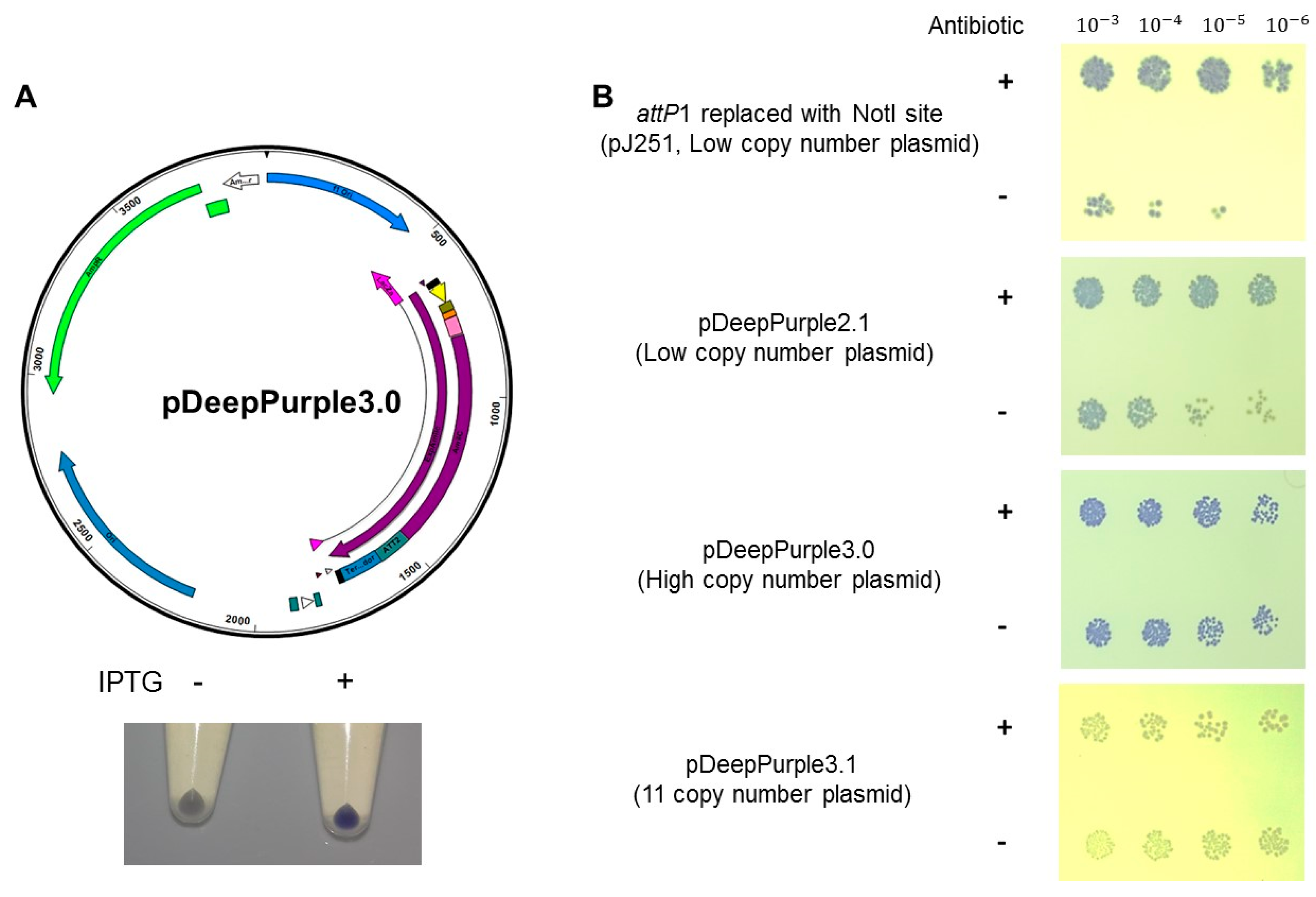
2.3. Protein Expression Plasmids Based on AmilCP Cloning Selection
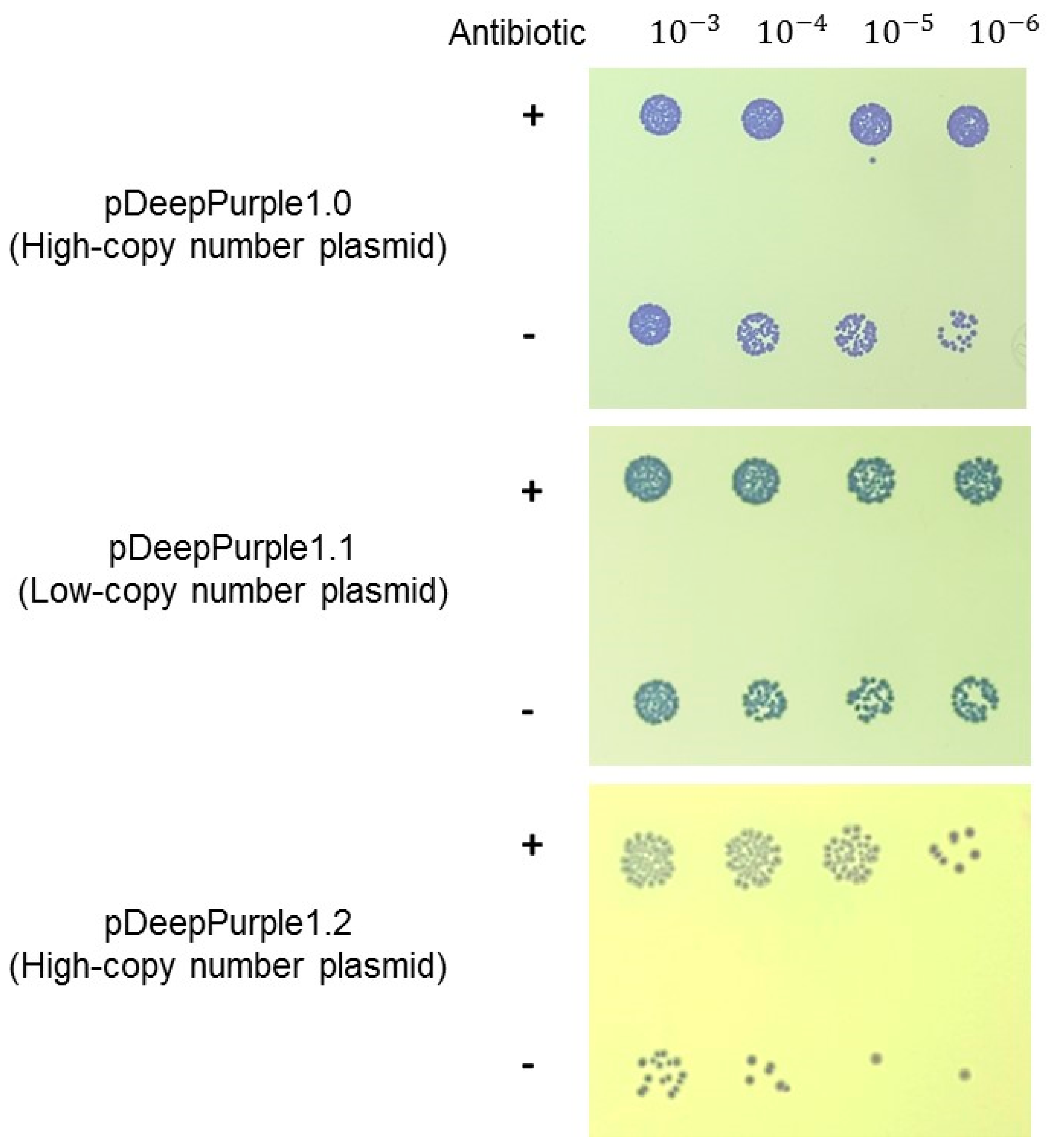
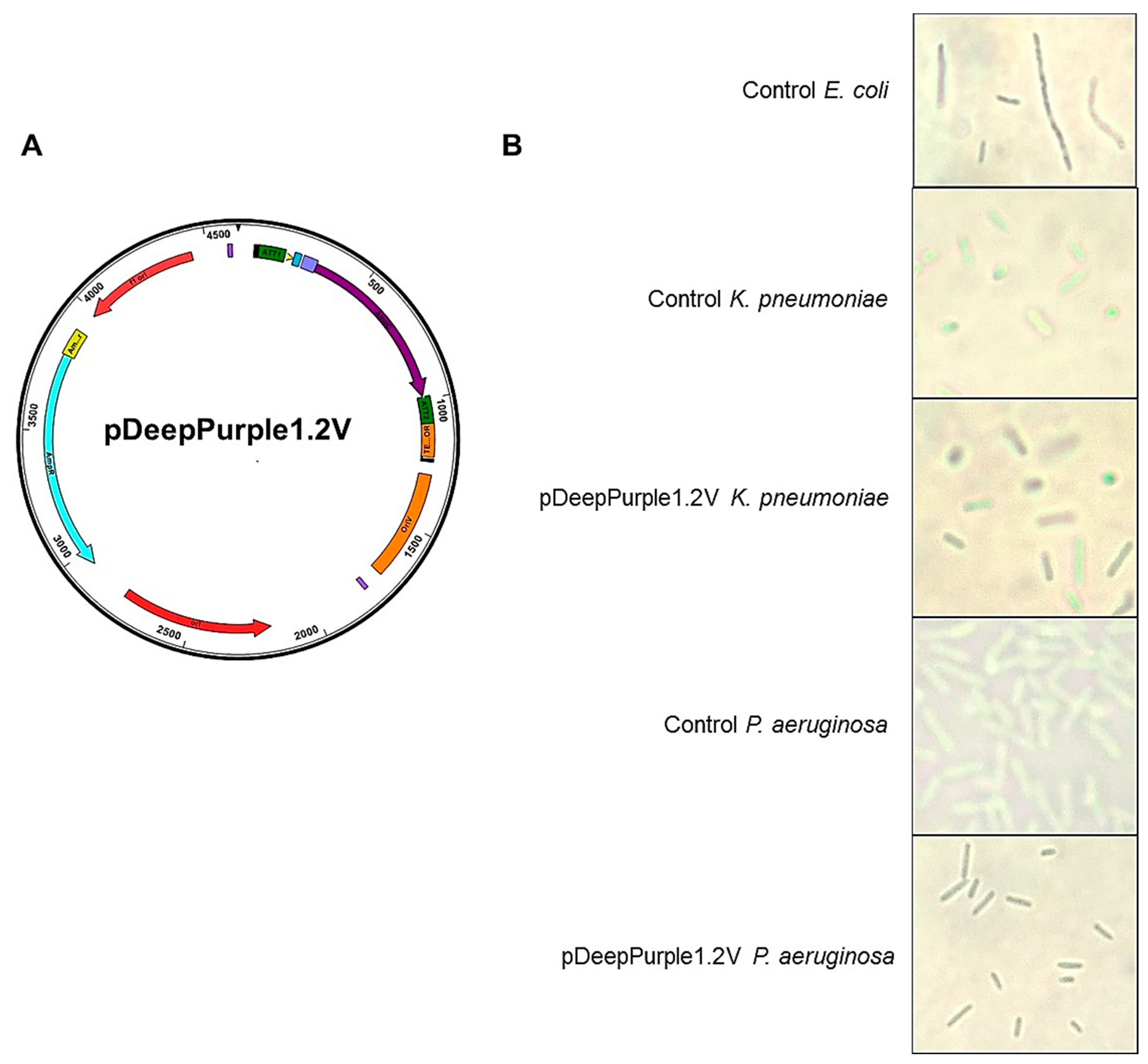
2.4. Expanding the Application of pDeepPurple1.2 Plasmid
3. Discussion
4. Materials and Methods
4.1. Cell Culture and Media
4.2. Synthetic Cassettes and Plasmid Constructions
4.3. Chromogenic Plasmid Stability Analysis
4.4. Cloning Test
4.5. Protein Expression Test in pDeepPurple 3.0
4.6. Plasmid Mutagenesis
4.7. Protein Sequence and Structural Analysis
4.8. K. preumoniae and P. aeruginosa Transformation
Supplementary Materials
Author Contributions
Acknowledgments
Conflicts of Interest
References
- Padmanabhan, S.; Banerjee, S.; Mandi, N. Screening of Bacterial Recombinants: Strategies and Preventing False Positives, Molecular Cloning—Selected Applications in Medicine and Biology; Brown, G., Ed.; InTech: London, UK, 2011; ISBN 978-953-307-398-9. [Google Scholar]
- Jackson, D.A.; Symons, R.H.; Berg, P. Biochemical method for inserting new genetic information into DNA of Simian Virus 40: Circular SV40 DNA molecules containing lambda phage genes and the galactose operon of Escherichia coli. Proc. Natl. Acad. Sci. USA 1972, 69, 2904–2909. [Google Scholar] [CrossRef] [PubMed]
- Kado, C.I. Historical Events That Spawned the Field of Plasmid Biology. Microbiol. Spectr. 2014, 2. [Google Scholar] [CrossRef]
- Baek, C.H.; Liss, M.; Clancy, K.; Chesnut, J.; Katzen, F. DNA Assembly Tools and Strategies for the Generation of Plasmids. Microbiol. Spectr. 2014, 2. [Google Scholar] [CrossRef]
- Hülter, N.; Ilhan, J.; Wein, T.; Kadibalban, A.S.; Hammerschmidt, K.; Dagan, T. An evolutionary perspective on plasmid lifestyle modes. Curr. Opin. Microbiol. 2017, 38, 74–80. [Google Scholar] [CrossRef] [PubMed]
- Chaffin, D.O.; Rubens, C.E. Blue/white screening of recombinant plasmids in Gram-positive bacteria by interruption of alkaline phosphatase gene (phoZ) expression. Gene 1998, 219, 91–99. [Google Scholar] [CrossRef]
- Ullmann, A.; Jacob, F.; Monod, J. Characterization by in vitro complementation of a peptide corresponding to an operator-proximal segment of the beta-galactosidase structural gene of Escherichia coli. J. Mol. Biol. 1967, 24, 339–343. [Google Scholar] [CrossRef]
- Langley, K.E.; Villarejo, M.R.; Fowler, A.V.; Zamenhof, P.J.; Zabin, I. Molecular basis of beta-galactosidase alpha-complementation. Proc. Natl. Acad. Sci. USA 1975, 72, 1254–1257. [Google Scholar] [CrossRef] [PubMed]
- Juers, D.H.; Jacobson, R.H.; Wigley, D.; Zhang, X.J.; Huber, R.E.; Tronrud, D.E.; Matthews, B.W. High resolution refinement of beta-galactosidase in a new crystal form reveals multiple metal-binding sites and provides a structural basis for alpha-complementation. Protein Sci. 2000, 9, 1685–1699. [Google Scholar] [CrossRef] [PubMed]
- Jobling, M.G.; Holmes, R.K. Construction of vectors with the p15a replicon, kanamycin resistance, inducible lacZa and pUC18 or pUC19 multiple cloning sites. Nucleic Acids Res. 1990, 18, 5315–5316. [Google Scholar] [CrossRef] [PubMed]
- Slilaty, S.N.; Lebel, S. Accurate insertional inactivation of lacZalpha: Construction of pTrueBlue and M13TrueBlue cloning vectors. Gene 1998, 213, 83–91. [Google Scholar] [CrossRef]
- Karlovský, P.; Prell, H.H. Screening of pUC plasmid clones for inserts based on growth rate (without X-gal). Nucleic Acids Res. 1987, 15, 6753. [Google Scholar] [CrossRef] [PubMed]
- Speltz, E.B.; Regan, L. White and green screening with circular polymerase extension cloning for easy and reliable cloning. Protein Sci. 2013, 22, 859–864. [Google Scholar] [CrossRef] [PubMed]
- Heuermann, K.; Cosgrove, J. S-Gal: An autoclavable dye for color selection of cloned DNA inserts. Biotechniques 2001, 30, 1142–1147. [Google Scholar] [PubMed]
- Reddy, M. Positive selection system for identification of recombinants using alpha-complementation plasmids. Biotechniques 2004, 37, 948–952. [Google Scholar] [PubMed]
- Li, P.; Li, J.; Guo, Z.; Tang, W.; Han, J.; Meng, X.; Hao, T.; Zhu, Y.; Zhang, L.; Chen, Y. An efficient blue-white screening based gene inactivation system for Streptomyces. Appl. Microbiol. Biotechnol. 2015, 99, 1923–1933. [Google Scholar] [CrossRef] [PubMed]
- Kayo, I.; Koji, S.; Hideki, Y.; Katsushiro, M.; Tsutomu, M. Masanori Sugiyama, Construction of a new cloning vector utilizing a cryptic plasmid and the highly expressed melanin-synthesizing gene operon from Streptomyces castaneoglobisporus. FEMS Microbiol. Lett. 1998, 168, 195–199. [Google Scholar] [CrossRef]
- Bardonnet, N.; Trautwetter, A.; Couchoux-Luthaud, G.; Blanco, C. Plasmids with the uidA reporter gene for the detection of promoters and transcription signals. Mol. Gen. Genet. 1988, 212, 390–392. [Google Scholar] [CrossRef] [PubMed]
- Dahlén, G.; Linde, A. Screening plate method for detection of bacterial beta-glucuronidase. Appl. Microbiol. 1973, 26, 863–866. [Google Scholar] [PubMed]
- Roessner, C.A.; Scott, A.I. Fluorescence-based method for selection of recombinant plasmids. Biotechniques 1995, 19, 760–764. [Google Scholar] [PubMed]
- Bernard, P. Positive selection of recombinant DNA by CcdB. Biotechniques 1996, 21, 320–323. [Google Scholar] [PubMed]
- Zhang, A.K. Chromogenic Assay for Lung Cancer-Related EGFR Exon 19 Hotspot Deletion Mutations. Genet. Test. Mol. Biomark. 2016, 20, 18–23. [Google Scholar] [CrossRef] [PubMed]
- Tran, D.T.M.; Phan, T.T.P.; Huynh, T.K.; Dang, N.T.K.; Huynh, P.T.K.; Nguyen, T.M.; Truong, T.T.T.; Tran, T.L.; Schumann, W.; Nguyen, H.D. Development of inducer-free expression plasmids based on IPTG-inducible promoters for Bacillus subtilis. Microb. Cell Fact. 2017, 16, 130. [Google Scholar] [CrossRef] [PubMed]
- Alieva, N.O.; Konzen, K.A.; Field, S.F.; Meleshkevitch, E.A.; Hunt, M.E.; Beltran-Ramirez, V.; Miller, D.J.; Wiedenmann, J.; Salih, A.; Matz, M.V. Diversity and evolution of coral fluorescent proteins. PLoS ONE 2008, 3, e2680. [Google Scholar] [CrossRef] [PubMed]
- Levy, O.; Appelbaum, L.; Leggat, W.; Gothlif, Y.; Hayward, D.C.; Miller, D.J.; Hoegh-Guldberg, O. Light-responsive cryptochromes from a simple multicellular animal, the coral Acropora millepora. Science 2007, 318, 467–470. [Google Scholar] [CrossRef] [PubMed]
- Prescott, M.; Ling, M.; Beddoe, T.; Oakley, A.J.; Dove, S.; Hoegh-Guldberg, O.; Devenish, R.J.; Rossjohn, J. The 2.2 A crystal structure of a pocilloporin pigment reveals a nonplanar chromophore conformation. Structure 2003, 11, 275–284. [Google Scholar] [CrossRef]
- Zawada, D.G.; Mazel, C.H. Fluorescence-based classification of Caribbean coral reef organisms and substrates. PLoS ONE 2014, 9, e84570. [Google Scholar] [CrossRef] [PubMed]
- Battad, J.M.; Wilmann, P.G.; Olsen, S.; Byres, E.; Smith, S.C.; Dove, S.G.; Turcic, K.N.; Devenish, R.J.; Rossjohn, J.; Prescott, M.A. Structural basis for the pH-dependent increase in fluorescence efficiency of chromoproteins. J. Mol. Biol. 2007, 368, 998–1010. [Google Scholar] [CrossRef] [PubMed]
- Simeon, S.; Shoombuatong, W.; Anuwongcharoen, N.; Preeyanon, L.; Prachayasittikul, V.; Wikberg, J.E.; Nantasenamat, C. osFP: A web server for predicting the oligomeric states of fluorescent proteins. J. Cheminform. 2016, 8, 72. [Google Scholar] [CrossRef] [PubMed]
- Inouye, S.; Ogawa, H.; Yasuda, K.; Umesono, K.; Tsuji, F.I. A bacterial cloning vector using a mutated Aequorea green fluorescent protein as an indicator. Gene 1997, 189, 159–162. [Google Scholar] [CrossRef]
- Shinzato, C.; Shoguchi, E.; Tanaka, M.; Satoh, N. Fluorescent protein candidate genes in the coral Acropora digitifera genome. Zool. Sci. 2012, 29, 260–264. [Google Scholar] [CrossRef] [PubMed]
- Zhang, Y.S. Blue-white screening liquid can eliminate false positives in blue-white colony screening. Genet. Mol. Res. 2016, 15. [Google Scholar] [CrossRef] [PubMed]
- Dykxhoorn, D.M.; St Pierre, R.; Linn, T. A set of compatible tac promoter expression vectors. Gene 1996, 177, 133–136. [Google Scholar] [CrossRef]
- Smith, M.A.; Bidochka, M.J. Bacterial fitness and plasmid loss: The importance of culture conditions and plasmid size. Can. J. Microbiol. 1998, 44, 351–355. [Google Scholar] [CrossRef] [PubMed]
- De Boer, H.A.; Comstock, L.J.; Vasser, M. The tac promoter: A functional hybrid derived from the trp and lac promoters. Proc. Natl. Acad. Sci. USA 1983, 80, 21–25. [Google Scholar] [CrossRef] [PubMed]
- Parr, R.D.; Ball, J.M. New donor vector for generation of histidine-tagged fusion proteins using the Gateway Cloning System. Plasmid 2003, 49, 179–183. [Google Scholar] [CrossRef]
- Andreou, A.I.; Nakayama, N. Mobius Assembly: A versatile Golden-Gate framework towards universal DNA assembly. PLoS ONE 2018, 13, e0189892. [Google Scholar] [CrossRef] [PubMed]
- Choe, J.; Guo, H.H.; van den Engh, G. A dual-fluorescence reporter system for high-throughput clone characterization and selection by cell sorting. Nucleic Acids Res. 2005, 33, e49. [Google Scholar] [CrossRef] [PubMed]
- Sleight, S.C.; Bartley, B.A.; Lieviant, J.A.; Sauro, H.M. In-Fusion BioBrick assembly and re-engineering. Nucleic Acids Res. 2010, 38, 2624–2636. [Google Scholar] [CrossRef] [PubMed]
- Yamaguchi, K.; Yamaguchi, M. The replication origin of pSC101: The nucleotide sequence and replication functions of the ori region. Gene 1984, 29, 211–219. [Google Scholar] [PubMed]
- Ohansson, M.U.; Zoete, V.; Michielin, O.; Guex, N. Defining and searching for structural motifs using DeepView/Swiss-PdbViewer. BMC Bioinform. 2012, 13, 173. [Google Scholar] [CrossRef] [PubMed]
- Kelley, L.A.; Mezulis, S.; Yates, C.M.; Wass, M.N.; Sternberg, M.J. The Phyre2 web portal for protein modeling, prediction and analysis et al. Nat. Protoc. 2015, 10, 845–858. [Google Scholar] [PubMed]
- Chudakov, D.M.; Matz, M.V.; Lukyanov, S.; Lukyanov, K.A. Fluorescent proteins and their applications in imaging living cells and tissues. Physiol. Rev. 2010, 90, 1103–1163. [Google Scholar] [CrossRef] [PubMed]
- Schrodinger LLC. The PyMOL Molecular Graphics System, Version 1.8; Shrodinger LLC.: New York, NY, USA, 2015. [Google Scholar]
Sample Availability: Samples of the compounds are available from the authors. |

| Plasmid name | Backbone | Features | Antibiotic Selection Marker |
|---|---|---|---|
| pDeepPurple1.0 | pTZ19R | High copy number cloning plasmid | Ampicillin |
| pDeepPurple1.1 | pEXT20 | 40 copy cloning plasmid | Ampicillin |
| pDeepPurple1.2 | pGEM T-easy | High copy number cloning plasmid | Ampicillin |
| pDeepPurple1.2V | pGEM T-easy | High copy number cloning plasmid, oriV sequence present rendering a plasmid selectable in other bacteria besides E. coli | Ampicillin |
| pJ251 attP1 replaced with a NotI restriction site | pJ251 | low copy number cloning plasmid modified from the original design for protein expression, only NotI restriction site present for cloning desired gene | Kanamycin |
| pDeepPurple2.1 | pJ251 | Low copy number cloning plasmid for protein expression, multiple cloning site present | Kanamycin |
| pDeepPurple3.0 | pTZ19R | High copy number cloning plasmid for protein expression, multiple cloning site present | Ampicillin |
| pDeepPurple3.1 | pACT3 | 11 copy cloning plasmid for protein expression, multiple cloning site present | Chloramphenicol |
| Part Name | Accession Number or Identification Number | Features | Purpose |
|---|---|---|---|
| Promoter sequence (Constitutive) | BBa_J23100 | 35 bp strong constitutive promoter | Constitutive expression of AmilCP protein for clone selection (Blue/White screening). |
| Promoter sequence (tac inducible promoter, functional hybrid derived from the trp and lac promoters) | BBa_K864400 | 61 bp inducible promoter | Inducible expression of proteins in expression plasmids [35]. |
| RBS | BBa_B0035 | 27 bp ribosome binding site, optimal for prokaryotic organisms | Constitutive expression of AmilCP protein for clone selection (Blue/White screening). |
| Terminator sequence | BBa_B0015 | 129 bp terminator sequence | Constitutive expression of AmilCP protein for clone selection (Blue/White screening) and protein expression plasmids. |
| att sequences | Invitrogen Gateway® | att sequences for recombination cloning (Gateway® system) | Cloning in vectors containing att sequences. Although, expression plasmid is limited due to toxicity generated by attL2 sequence (see text). |
| OriV | BBa_K125340 | 424 bp fragment, origin of replication. Broad host capabilities | Generate a plasmid capable of replicating in other hosts besides E. coli. |
| Primer Name | 5’ to 3’ Sequence | Application |
|---|---|---|
| F forward | CCCCGAAAAGTGCCACCT | Primer pair used to amplify all the designed cassettes and cloning them into different plasmid backbones |
| F reverse | GTTCTGAGGTCATTACTGGTATAAACG | |
| OriV Fwd | AACCCCTGCAATAACTGTC | Primer pair used to amplify the oriV replication origin from a gBlock synthesized fragment |
| OriV Rv | GCTGAATGATCGACCGAG | |
| NotIinsertionFw | ACTAGATGCATCACCATCACCATCACGCG GCCGCTGGCTTCCAGCAGTTCGGTGATTGCG | Primer pair to replace the attP1 sequence in the protein expression cassette and replace it with a multiple cloning site |
| NotIinsertionRv | CGCAATCACCGAACTGCTGGAAGCCAGCGG CCGCGTGATGGTGATGGTGATGCATCTAGT | |
| MCS1 | GGCCGCGGATCCCATGATATCCATCACCTCGA GCACCACCACAAGCTTCATGCATGCGC | Primer pair to generate a multiple cloning site for the protein expression cassette. Once aligned, the ends are compatible to a NotI digested plasmid. |
| MCS2 | CGCCGGGCATGCATGAAGCTTGTGGTGGTG CTCGAGGTGATGGATATCATGGGATCCGC | |
| MutsspIpact3 | CCTTGCGTATAAAATTTGCCCATGGTG | Primer pair to eliminate SspI restriction site from the cat gene in pACT3 plasmid. |
| MutsspIpact3compl | CACCATGGGCAAATTTTATACGCAAGG | |
| CaroFw | GGGGCCAACTTTGTACAAAGAAGCA GGCTATGCCCTTTACCATTGATAG | Primer pair to amplify and clone the water-soluble carotenoid protein sequence using the Gateway® cloning system. |
| CaroRv | GGGGACCACTTTGTACAAGAAAGCTG GGTCTAACGGGCAAAATTCAAAAG |
© 2018 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Tafoya-Ramírez, M.D.; Padilla-Vaca, F.; Ramírez-Saldaña, A.P.; Mora-Garduño, J.D.; Rangel-Serrano, Á.; Vargas-Maya, N.I.; Herrera-Gutiérrez, L.J.; Franco, B. Replacing Standard Reporters from Molecular Cloning Plasmids with Chromoproteins for Positive Clone Selection. Molecules 2018, 23, 1328. https://doi.org/10.3390/molecules23061328
Tafoya-Ramírez MD, Padilla-Vaca F, Ramírez-Saldaña AP, Mora-Garduño JD, Rangel-Serrano Á, Vargas-Maya NI, Herrera-Gutiérrez LJ, Franco B. Replacing Standard Reporters from Molecular Cloning Plasmids with Chromoproteins for Positive Clone Selection. Molecules. 2018; 23(6):1328. https://doi.org/10.3390/molecules23061328
Chicago/Turabian StyleTafoya-Ramírez, Margarita Daniela, Felipe Padilla-Vaca, Ana Patricia Ramírez-Saldaña, Josué Daniel Mora-Garduño, Ángeles Rangel-Serrano, Naurú Idalia Vargas-Maya, Luz Janeth Herrera-Gutiérrez, and Bernardo Franco. 2018. "Replacing Standard Reporters from Molecular Cloning Plasmids with Chromoproteins for Positive Clone Selection" Molecules 23, no. 6: 1328. https://doi.org/10.3390/molecules23061328