Secondary Metabolite Profiling of Species of the Genus Usnea by UHPLC-ESI-OT-MS-MS
Abstract
1. Introduction
2. Results and Discussion
2.1. Metabolomics in Usnea barbata
Depsides
Depsidones
Lipids
Diphenylethers
Dibenzofurans
Unknown Compounds
2.2. Metabolomics in Usnea antarctica
Depsides
Depsidones
Lipids
Other Compounds
2.3. Metabolomics in Usnea rubicunda
Depsides
Depsidones
Lipids
Unknown Compounds
Other Compounds
2.4. Metabolomics in Usnea subfloridana
Depsides
Depsidones
Lipids
Unknown Compounds
Other Compounds
3. Materials and Methods
3.1. Lichen Material
3.2. UHPLC-Orbitrap-ESI-MS-MS
3.2.1. Sample Preparation
3.2.2. Instruments
3.2.3. LC Parameters
3.2.4. MS Parameters
4. Conclusions
Supplementary Materials
Acknowledgments
Author Contributions
Conflicts of Interest
References
- Upreti, D.K.; Divakar, P.K.; Shukla, V.; Bajpai, R. Recent Advances on Lichenology; Springer: New Delhi, India, 2015. [Google Scholar]
- Paliya, B.S.; Bajpai, R.; Jadaun, V.; Kumar, J.; Kumar, S.; Upreti, D.K.; Singh, B.R.; Nayaka, S.; Joshi, Y.; Singh, B.N.; et al. The genus Usnea: A potent phytomedicine with multifarious ethnobotany, phytochemistry and pharmacology. RSC Adv. 2016, 6, 21672–21696. [Google Scholar]
- Huneck, S.; Yoshimura, I. Identification of Lichen Substances; Springer: Berlin, Germany, 1996. [Google Scholar]
- Rankovic, B. (Ed.) Lichen Secondary Metabolites: Bioactive Properties and Pharmaceutical Potential; Springer: New York, NY, USA, 2015. [Google Scholar]
- Simirgiotis, M.J.; Quispe, C.; Areche, C.; Sepúlveda, B. Phenolic compounds in Chilean Mistletoe (Quintral, Tristerix tetrandus) analyzed by UHPLC-Q/Orbitrap/MS/MS and its antioxidant properties. Molecules 2016, 21, 245. [Google Scholar] [CrossRef] [PubMed]
- Cornejo, A.; Salgado, F.; Caballero, J.; Vargas, R.; Simirgiotis, M.; Areche, C. Secondary metabolites in Ramalina terebrata detected by UHPLC/ESI/MS/MS and identificaction of parietin as tau protein inhibitor. Int. J. Mol. Sci. 2016, 17, E1303. [Google Scholar] [CrossRef] [PubMed]
- Castro, O.N.; Benites, J.; Rodilla, J.; Santiago, J.C.; Simirgiotis, M.; Sepúlveda, B.; Areche, C. Metabolomic analysis of the lichen Everniopsis trulla using ultra high performance liquid chromatography-quadrupole-orbitrap mass spectrometry (UHPLC-Q-OT-MS). Chromatographia 2017, 80, 967–973. [Google Scholar] [CrossRef]
- Le Pogam, P.; Schinkovitz, A.; Legouin, B.; Le Lamer, A.C.; Boustie, J.; Richomme, P. Matrix-Free UV-laser desorption ionization mass spectrometry as a versatile approach for accelerating dereplication studies on Lichens. Anal. Chem. 2015, 87, 10421–10428. [Google Scholar] [CrossRef] [PubMed]
- Musharraf, S.G.; Kanwal, N.; Thadhani, V.M.; Choudhary, M.I. Rapid identification of lichen compounds based on the structure-fragmentation relationship using ESI-MS/MS analysis. Anal. Methods 2015, 7, 6066–6076. [Google Scholar] [CrossRef]
- Parrot, D.; Jan, S.; Baert, N.; Guyot, S.; Tomasi, S. Comparative metabolite profiling and chemical study of Ramalina siliquosa complex using LC–ESI-MS/MS approach. Phytochemistry 2013, 89, 114–124. [Google Scholar] [CrossRef] [PubMed]
- Holzmann, G.; Leuckert, C. Applications of negative fast atom bombardment and MS/MS to screening of lichen compounds. Phytochemistry 1990, 29, 2277–2283. [Google Scholar] [CrossRef]
- Parrot, D.; Peresse, A.T.; Hitti, E.; Carrie, D.; Grube, M.; Tomasi, S. Qualitative and spatial metabolite profiling of lichens by a LC-MS approach combined with optimised extraction. Phytochem. Anal. 2015, 26, 23–33. [Google Scholar] [CrossRef] [PubMed]
Sample Availability: Samples of the lichens are available from the authors. |
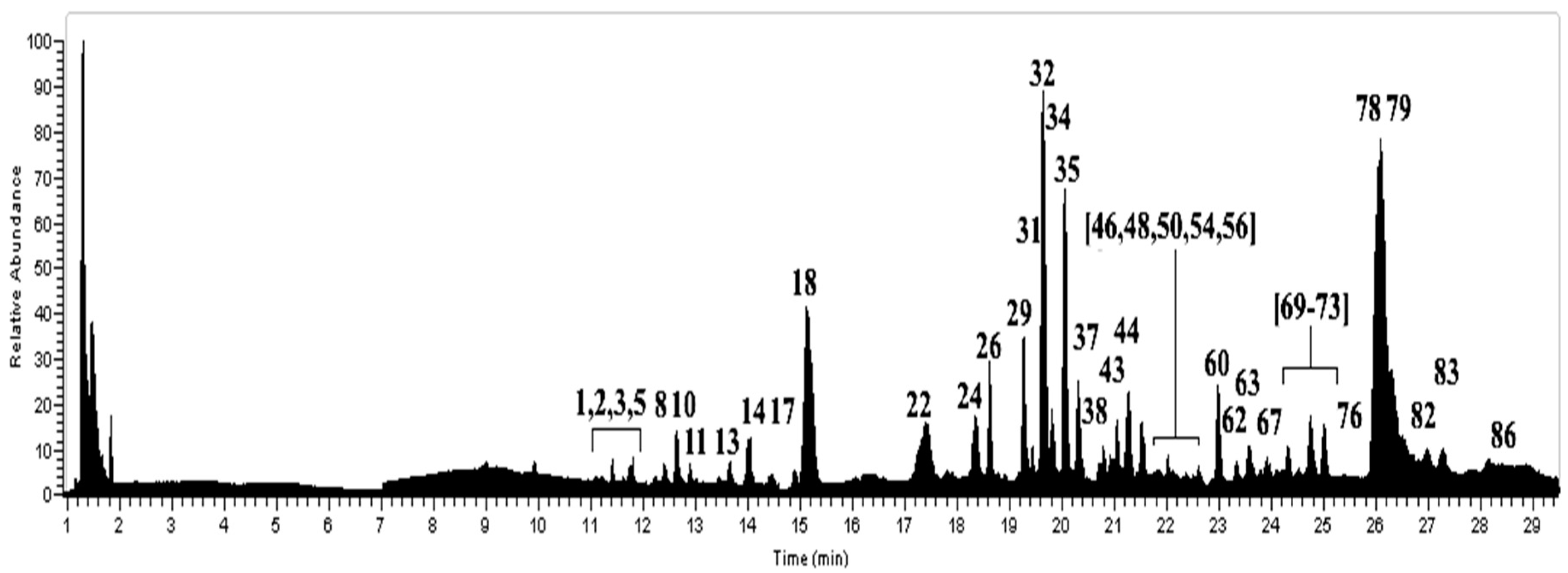
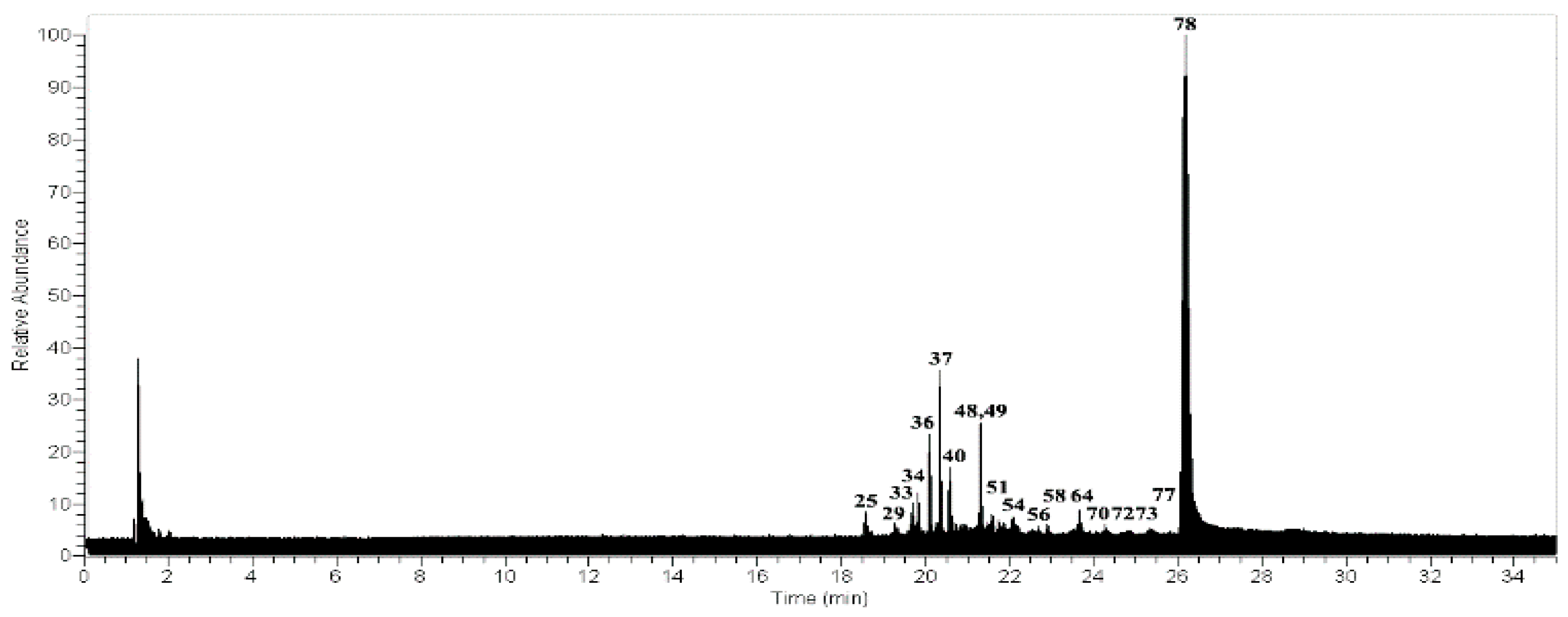
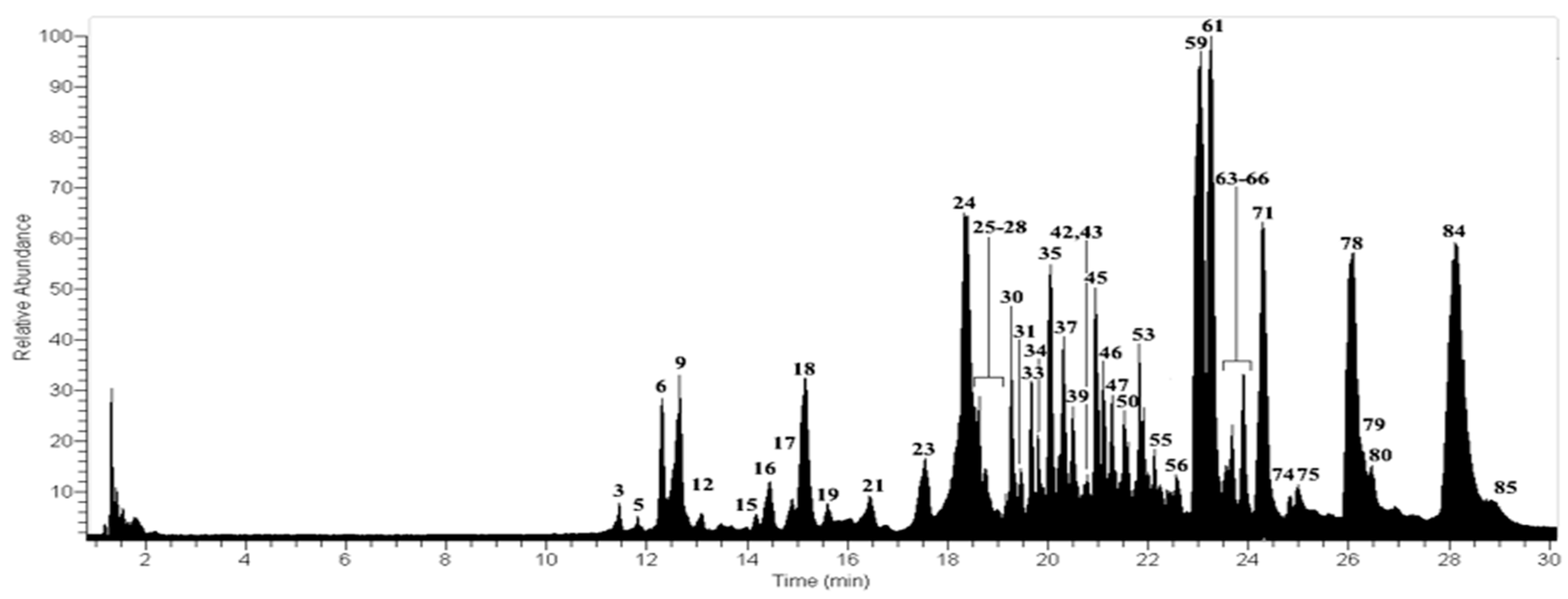
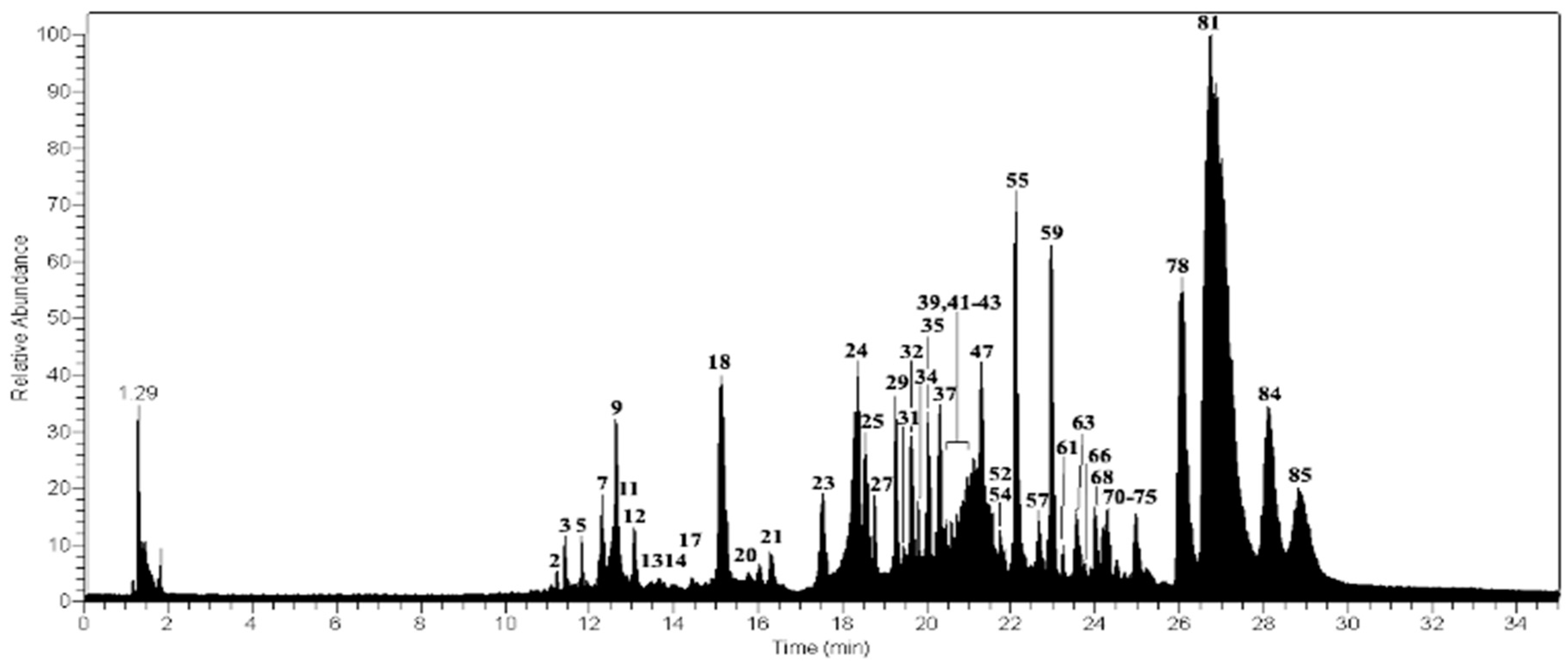
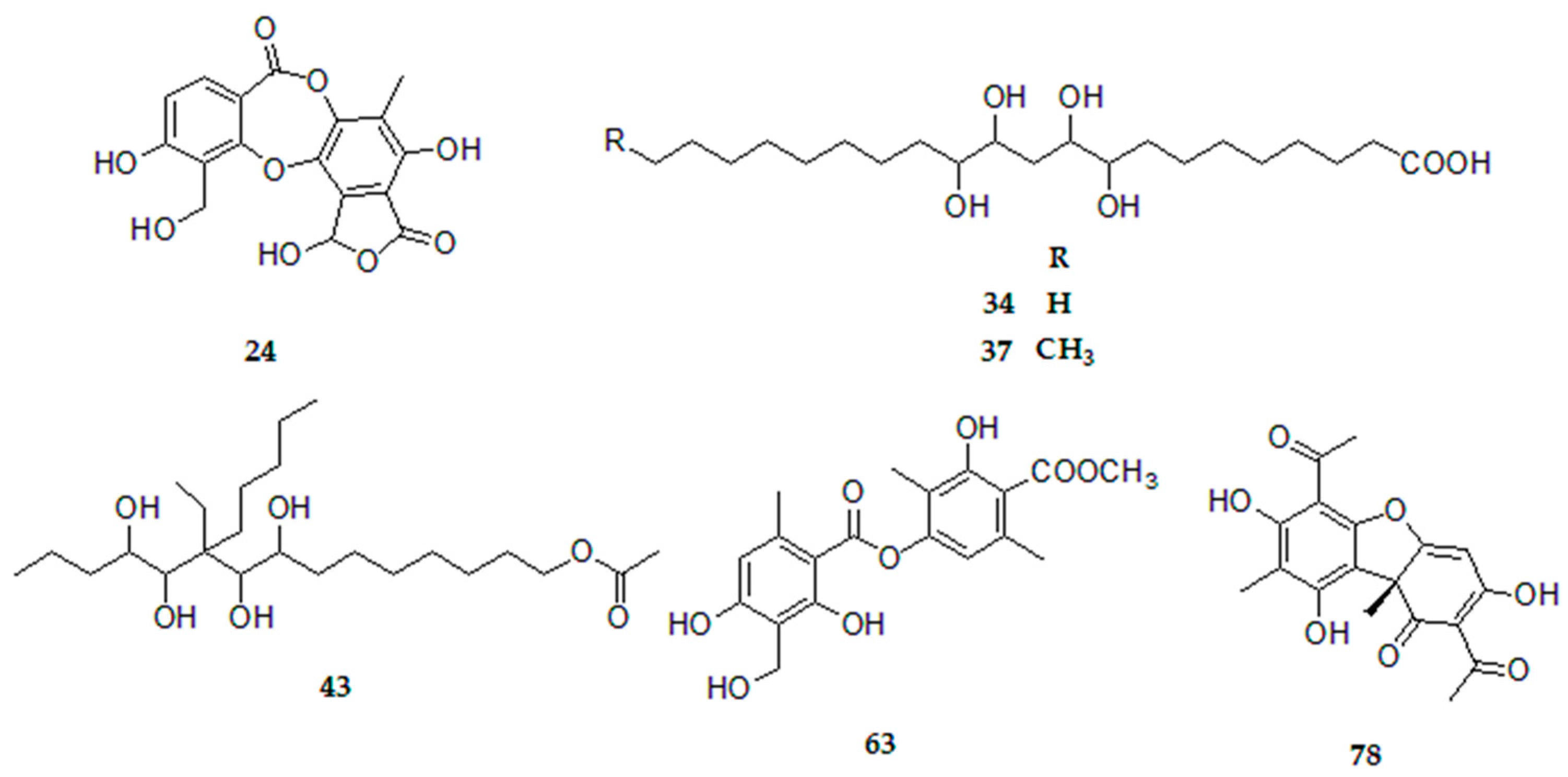
| Peak | Tentative Identification | [M − H]− | Retention Time (min) | Theoretical Mass (m/z) | Measured Mass (m/z) | Accuracy (ppm) | Metabolite Type | MS2 Ions (ppm) | Lichens |
|---|---|---|---|---|---|---|---|---|---|
| 1 | Unknown | C21H19O12 | 11.09 | 463.0882 | 463.0883 | 0.4 | - | - | UB |
| 2 | Unknown | C22H19O10N2 | 11.21 | 471.1040 | 471.1045 | 1.1 | - | - | UB, US, UR |
| 3 | Unknown | C18H13O10 | 11.42 | 389.0514 | 389.0515 | 0.2 | - | - | UB, US, UR |
| 4 | Unknown | C24H25O15 | 11.74 | 553.1193 | 553.1195 | 0.4 | - | - | UB |
| 5 | Unknown | C22H18O11N | 11.80 | 472.0880 | 472.0886 | 1.3 | - | - | UB, US, UR |
| 6 | Conprotocetraric acid | C18H15O9 | 12.27 | 375.0722 | 375.0724 | 0.5 | d | 357.0618; 313.0722 295.0611; 251.0710 | UR |
| 7 | Haemathamnolic acid | C19H15O10 | 12.29 | 403.0671 | 403.0675 | 0.9 | d | 209.0002 | US |
| 8 | Barbatolic acid | C18H13O10 | 12.40 | 389.0514 | 389.0516 | 0.2 | d | 167.0342; 121.0285 | UB |
| 9 | Siphulellic acid isomer | C19H13O10 | 12.62 | 401.0509 | 401.0518 | 0.7 | D | 359.0410; 240.0427 | US, UR |
| 10 | Thamnolic acid | C19H15O11 | 12.64 | 419.0620 | 419.0619 | 0.2 | d | 375.0725; 167.0344 | UB |
| 11 | Unknown | C29H21O9 | 12.88 | 513.1186 | 513.1149 | 7.2 | - | - | UB, US |
| 12 | Unknown | C21H19O11 | 13.06 | 447.0927 | 447.0935 | 1.8 | - | - | UR |
| 13 | Unknown | C20H15O11 | 13.64 | 431.0620 | 431.0620 | 0.0 | - | - | UB, US |
| 14 | Unknown | C21H22O15 | 14.01 | 514.0959 | 514.0988 | 5.6 | - | - | UB, US |
| 15 | Squamatic acid | C19H17O9 | 14.17 | 389.0878 | 389.0880 | 0.5 | d | 211.0260 | UR |
| 16 | Protocetraric acid | C18H13O9 | 14.43 | 373.0565 | 373.0567 | 0.5 | D | 311.0559; 267.0657 | UR |
| 17 | Unknown | C22H18O10N | 14.88 | 456.0931 | 456.0936 | 1.1 | - | - | UB, US, UR |
| 18 | Salazinic acid | C18H11O10 | 15.11 | 387.0358 | 387.0352 | 0.2 | D | 269.0464; 241.0504 151.0394; 123.0445 | UB, US, UR |
| 19 | Conhypoprotocetraric acid | C18H15O8 | 15.60 | 359.0772 | 359.0775 | 0.8 | d | 253.0870 | UR |
| 20 | Physodalic acid | C20H15O10 | 15.79 | 415.0671 | 415.0674 | 0.7 | D | 387.0712; 343.0815 | US |
| 21 | Constictic acid | C19H13O10 | 16.32 | 401.0514 | 401.0516 | 0.5 | D | 357.0618; 313.0718 | US, UR |
| 22 | Haemathamnolic acid isomer | C19H15O10 | 17.40 | 403.0665 | 403.0672 | 1.7 | d | 373.0568 | UB |
| 23 | Hypoconstictic acid derivative | C19H15O9 | 17.55 | 387.0722 | 387.0724 | 0.5 | D | 343.0824; 299.0925 | US, UR |
| 24 | Connorstictic acid | C18H13O9 | 18.35 | 373.0565 | 373.0568 | 0.8 | D | 329.0666; 181.0555 | UB, UA, US, UR |
| 25 | Menegazziaic acid | C18H13O9 | 18.53 | 373.0565 | 373.0567 | 0.7 | D | 329.0667; 167.0344 151.0395 | US, UR |
| 26 | tetrahydroxyeicosanoic acid | C20H39O6 | 18.64 | 375.2752 | 375.2752 | 0.0 | L | - | UB, UR |
| 27 | Unknown | C21H19O10 | 18.77 | 431.0978 | 431.0986 | 1.8 | - | - | US, UR |
| 28 | Pentahydroxytricosanoic acid | C23H45O7 | 19.16 | 433.3171 | 433.3174 | 0.7 | L | - | UR |
| 29 | Siphulellic acid | C19H13O10 | 19.27 | 401.0509 | 401.0514 | 1.6 | D | 253.0505; 149.0238 123.0444 | UB, US |
| 30 | Tetrahydroxyheneicosanoic | C21H41O6 | 19.27 | 389.2909 | 389.2010 | 0.3 | L | - | UA, UR |
| 31 | Lecanoric acid | C16H13O7 | 19.44 | 317.0667 | 317.0661 | 1.9 | d | 167.0343; 149.0237 123.0444 | UB, US, UR |
| 32 | Galbinic acid | C20H13O11 | 19.66 | 429.0458 | 429.0463 | 1.4 | D | 269.0456; 149.0238 123.0443 | UB, US |
| 33 | Pentahydroxytetracosanoic acid | C24H47O7 | 19.69 | 447.3327 | 447.3307 | 4.5 | L | - | UA, UR |
| 34 | tetrahydroxydocosanoic acid | C22H43O6 | 19.80 | 403.3065 | 403.3067 | 0.6 | L | - | UB, UA, US, UR |
| 35 | Norstictic acid | C18H11O9 | 20.09 | 371.0409 | 371.0403 | 1.6 | D | 327.0509 151.0393; 123.0444 | UB, US, UR |
| 36 | Fumarprotocetraric acid | C22H15O12 | 20.11 | 471.0569 | 471.0547 | 4.7 | D | 355.0441; 311.0545 115.0023 | UA |
| 37 | tetrahydroxytricosanoic acid | C23H45O6 | 20.33 | 417.3222 | 417.3216 | 0.4 | L | - | UB, UA, US, UR |
| 38 | tetrahydroxydocosanoic acid | C22H43O6 | 20.38 | 403.3065 | 403.3066 | 0.6 | L | - | UB |
| 39 | Cryptostictic acid | C19H15O9 | 20.44 | 387.0716 | 387.0726 | 2.6 | D | 343.0826; 311.0566 267.0661 | US, UR |
| 40 | Hypoconstictic acid | C19H15O9 | 20.57 | 387.0716 | 387.0705 | 2.8 | D | 343.0808; 299.0923 | UA |
| 41 | Tetrahydroxydioxoheneicosanoic acid | C21H37O8 | 20.58 | 417.2494 | 417.2498 | 0.9 | L | - | US |
| 42 | Tetrahydroxytricosanoic acid | C23H45O6 | 20.74 | 417.3222 | 417.3225 | 0.7 | L | - | US, UR |
| 43 | 6-ethyl-6-n-pentylpentadecan-4,5,7,8,15-pentol-15-acetate | C24H47O6 | 20.80 | 431.3373 | 431.3379 | 1.4 | L | - | UB, UA, US, UR |
| 44 | Unknown | C18H11O8 | 20.93 | 355.0459 | 355.0462 | 0.8 | - | - | UB |
| 45 | Unknown | C29H59O7N9 | 20.95 | 645.4537 | 645.4937 | 0.0 | - | - | UR |
| 46 | α-acetylconstictic acid | C21H15O11 | 21.05 | 443.0620 | 443.0620 | 0.0 | D | 383.0418; 343.0425 269.0457 | UB |
| 47 | Trihydroxytrioxodocosanoic acid | C22H37O8 | 21.29 | 429.2494 | 429.2496 | 0.4 | L | - | US, UR |
| 48 | Gyrophoric acid * | C24H19O10 | 21.32 | 467.0978 | 467.0985 | 1.3 | d | 317.0667; 167.0345 149.0238; 123.0443 | UB, UA |
| 49 | Tetrahydroxyhexacosenoic acid | C26H49O6 | 21.32 | 457.3535 | 457.3510 | 4.4 | L | - | UA |
| 50 | Stictic acid | C19H13O9 | 21.54 | 385.0565 | 385.0567 | 0.5 | D | 357.0617; 313.0720 179.0347 | UB, UR |
| 51 | Tetrahydroxypentacosanoic acid | C25H49O6 | 21.59 | 445.3535 | 445.3515 | 4.5 | L | - | UA |
| 52 | Tetrahydroxydioxotricosanoic acid | C23H41O8 | 21.73 | 445.2807 | 445.2812 | 1.1 | L | - | US |
| 53 | 4-O-methylnorsekikaic acid | C21H23O8 | 21.85 | 403.1398 | 403.1400 | 0.4 | d | 209.0416; 193.0803 165.0916 | UR |
| 54 | Tetrahydroxyhexacosanoic acid | C26H51O6 | 22.03 | 459.3691 | 459.3693 | 0.4 | L | - | UB, UA, US |
| 55 | Methyl 3,4-dicarboxy-3-hydroxy-19-oxoeicosanoate | C23H39O8 | 22.14 | 443.2650 | 443.2653 | 0.4 | L | - | US, UR |
| 56 | Nonahydroxyoctacosanoic acid | C28H55O11 | 22.65 | 567.3650 | 567.3667 | 3.0 | L | - | UB, UA |
| 57 | Trihydroxytrioxotetracosanoic acid | C24H41O8 | 22.72 | 457.2807 | 457.2810 | 0.6 | L | - | US |
| 58 | Baeomycesic acid | C19H17O8 | 22.88 | 373.0923 | 373.0915 | 2.1 | d | - | UA |
| 59 | Neodihydromurolic acid | C21H37O5 | 22.94 | 369.2646 | 369.2650 | 1.1 | L | - | US, UR |
| 60 | Diffractaic acid | C20H21O7 | 22.98 | 373.1293 | 373.1294 | 0.3 | d | 297.1149; 181.0499 177.0549; 137.0600 | UB |
| 61 | Murolic acid | C21H35O5 | 23.23 | 367.2490 | 367.2494 | 0.4 | L | - | US, UR |
| 62 | β-Alectoronic acid | C28H31O9 | 23.43 | 511.1968 | 511.1976 | 1.3 | DE | 369.1339; 247.0969 163.0396 | UB |
| 63 | Methyl 8-hydroxy-4-O-demethylbarbatate | C19H19O8 | 23.58 | 375.1080 | 375.1085 | 1.6 | d | 343.0818; 181.0500 | UB, UA, US, UR |
| 64 | Placodiolic acid | C19H19O8 | 23.65 | 375.1080 | 375.1069 | 2.9 | DBF | 343.0806; 259.0597 231.0648 | UA |
| 65 | Hydroxyeicosatrienoic acid | C20H33O3 | 23.65 | 321.2435 | 321.2437 | 0.6 | L | - | UR |
| 66 | muronic acid | C21H33O5 | 23.89 | 365.2333 | 365.2338 | 0.6 | L | - | US, UR |
| 67 | Tetrahydroxytrioxotricosanoic acid | C23H39O9 | 23.98 | 459.2594 | 459.2602 | 1.7 | L | - | UB |
| 68 | Trihydroxytrioxopentacosanoic acid | C25H43O8 | 24.00 | 471.2963 | 471.2970 | 1.5 | L | - | US |
| 69 | Divaricatic acid | C21H23O7 | 24.09 | 387.1449 | 387.1452 | 0.7 | d | 195.0661; 177.0551 151.0759 | UB |
| 70 | Barbatic acid | C19H19O7 | 24.20 | 359.1136 | 359.1139 | 0.8 | d | 181.0503; 163.0394 137.0601 | UB, UA, US |
| 71 | Sekikaic acid | C22H25O8 | 24.31 | 417.1549 | 417.1556 | 1.7 | d | 225.0764; 209.0815 165.0915 | UB, US, UR |
| 72 | 8-hydroxybarbatic acid | C19H19O8 | 24.53 | 375.1080 | 375.1088 | 2.4 | d | 195.0663; 181.0506 | UB, UA, US |
| 73 | Lobaric acid | C25H27O8 | 24.73 | 455.1711 | 455.1712 | 0.4 | D | 411.1815; 367.1909 352.1681; 296.1048 | UB, UA, US |
| 74 | Boninic acid | C25H31O8 | 24.83 | 459.2024 | 459.2027 | 0.6 | d | 209.1180 | UR |
| 75 | Norcaperatic acid | C20H35O7 | 24.95 | 387.2388 | 387.2392 | 0.8 | L | - | US, UR |
| 76 | Tetrahydroxytrioxotetracosanoic acid | C24H41O9 | 25.01 | 473.2756 | 473.2756 | 0.0 | L | - | UB |
| 77 | Ethyl-4-O-methylolivetolcarboxylate | C15H21O4 | 25.34 | 265.1445 | 265.1466 | 7.9 | A | - | UA |
| 78 | Usnic acid * | C18H15O7 | 26.04 | 343.0818 | 343.0823 | 1.5 | DBF | 328.0591; 259.0609; 231.0661 | UB, UA, US, UR |
| 79 | Atranorin | C19H17O8 | 26.31 | 373.0923 | 373.0929 | 1.6 | d | 177.0187; 163.0394 | UB, UR |
| 80 | trioxotricosanoic acid | C23H39O5 | 26.45 | 395.2803 | 395.2805 | 0.5 | L | - | UR |
| 81 | Perlatolic acid | C25H31O7 | 26.77 | 443.2075 | 443.2077 | 0.5 | d | 223.0973; 205.0867 179.1073 | US |
| 82 | Hexadecadienoic acid | C16H27O2 | 26.96 | 251.2017 | 251.2016 | 0.1 | L | - | UB |
| 83 | Dihydroxyheptadecatrienoic acid | C17H27O4 | 27.31 | 295.1915 | 295.1916 | 0.3 | L | - | UB |
| 84 | Caperatic acid | C21H37O7 | 28.10 | 401.2545 | 401.2548 | 0.7 | L | - | US, UR |
| 85 | Hydroxytrioxotricosanoic acid | C23H39O6 | 28.87 | 411.2752 | 411.2754 | 0.4 | L | - | US, UR |
| 86 | Chloroatranorin | C19H16O8Cl | 28.96 | 407.0534 | 407.0541 | 1.5 | d | 228.9906; 210.9800 163.0394 | UB |
© 2017 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Salgado, F.; Albornoz, L.; Cortéz, C.; Stashenko, E.; Urrea-Vallejo, K.; Nagles, E.; Galicia-Virviescas, C.; Cornejo, A.; Ardiles, A.; Simirgiotis, M.; et al. Secondary Metabolite Profiling of Species of the Genus Usnea by UHPLC-ESI-OT-MS-MS. Molecules 2018, 23, 54. https://doi.org/10.3390/molecules23010054
Salgado F, Albornoz L, Cortéz C, Stashenko E, Urrea-Vallejo K, Nagles E, Galicia-Virviescas C, Cornejo A, Ardiles A, Simirgiotis M, et al. Secondary Metabolite Profiling of Species of the Genus Usnea by UHPLC-ESI-OT-MS-MS. Molecules. 2018; 23(1):54. https://doi.org/10.3390/molecules23010054
Chicago/Turabian StyleSalgado, Francisco, Laura Albornoz, Carmen Cortéz, Elena Stashenko, Kelly Urrea-Vallejo, Edgar Nagles, Cesar Galicia-Virviescas, Alberto Cornejo, Alejandro Ardiles, Mario Simirgiotis, and et al. 2018. "Secondary Metabolite Profiling of Species of the Genus Usnea by UHPLC-ESI-OT-MS-MS" Molecules 23, no. 1: 54. https://doi.org/10.3390/molecules23010054
APA StyleSalgado, F., Albornoz, L., Cortéz, C., Stashenko, E., Urrea-Vallejo, K., Nagles, E., Galicia-Virviescas, C., Cornejo, A., Ardiles, A., Simirgiotis, M., García-Beltrán, O., & Areche, C. (2018). Secondary Metabolite Profiling of Species of the Genus Usnea by UHPLC-ESI-OT-MS-MS. Molecules, 23(1), 54. https://doi.org/10.3390/molecules23010054