Assessing and Broadening Genetic Diversity of Elymus sibiricus Germplasm for the Improvement of Seed Shattering
Abstract
:1. Introduction
2. Results
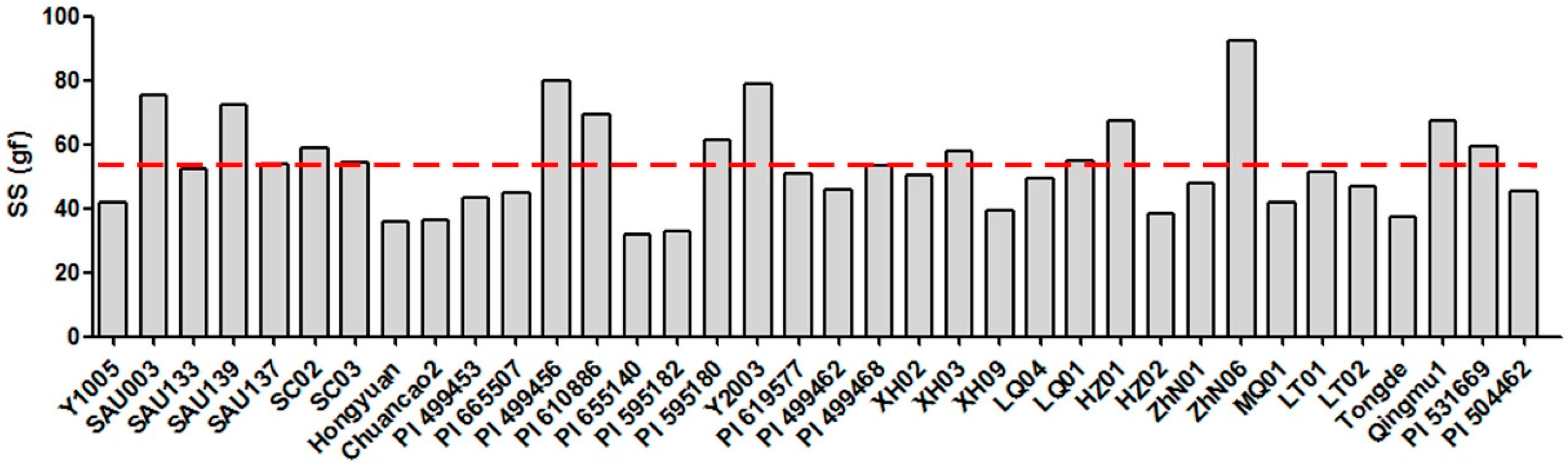
2.1. Seed Shattering Degree of 36 E. sibiricus Accessions
2.2. Polymorphism of EST-SSR Markers and Genetic Relationships of 36 E. sibiricus Accessions
2.3. Genetic and Phenotypic Variation of Hybrid Population
3. Discussion
3.1. Genetic Diversity of E. sibiricus
3.2. Broadening Genetic Diversity for Seed Shattering Improvement
4. Materials and Methods
4.1. Plant Materials
4.2. DNA Extraction and PCR Amplification
4.3. Phenotypic Traits Measurement
4.4. Data Analysis
5. Conclusions
Acknowledgments
Author Contributions
Conflicts of Interest
References
- Dewey, D.R. Cytogenetics of Elymus sibiricus and its hybrids with Agropyron tauri, Elymus canadensis, and Agropyron caninus. Bot. Gaz. 1994, 135, 80–87. [Google Scholar] [CrossRef]
- Xie, W.G.; Zhao, X.H.; Zhang, J.Q.; Wang, Y.R.; Liu, W.X. Assessment of genetic diversity of Siberian wild rye (Elymus sibiricus L.) germplasms with variation of seed shattering and implication for future genetic improvement. Biochem. Syst. Ecol. 2015, 58, 211–218. [Google Scholar] [CrossRef]
- Ma, X.; Chen, S.Y.; Bai, S.Q.; Zhang, X.Q.; Li, D.X.; Zhang, C.B.; Yan, J.J. RAPD analysis of genetic diversity and population structure of Elymus sibiricus (Poaceae) native to the southeastern Qinghai-Tibet Plateau, China. Genet. Mol. Res. 2012, 11, 2708–2718. [Google Scholar] [CrossRef] [PubMed]
- You, M.H.; Liu, J.P.; Bai, S.Q.; Zhang, X.Q.; Yan, J.J. Study on relationship of seed shattering, seed development and yield traits of Elymus sibiricus L. Southwest China J. Agric. Sci. 2011, 24, 1256–1260. [Google Scholar]
- Wu, J.F.; Li, F.; Xu, K.; Gao, G.Z.; Chen, B.Y.; Yan, G.X.; Wang, N.; Qiao, J.W.; Li, J.; Li, H.; et al. Assessing and broadening genetic diversity of a rapeseed germplasm collection. Breeding Sci. 2014, 64, 321–330. [Google Scholar] [CrossRef] [PubMed]
- Jesske, T.; Olberg, B.; Schierholy, A.; Becker, H.C. Resynthesized lines from domesticated and wild Brassica taxa and their hybrids with B. napus L.: Genetic diversity and hybrid yield. Theor. Appl. Genet. 2013, 126, 1053–1065. [Google Scholar] [CrossRef] [PubMed]
- Shen, J.X.; Fu, T.D.; Yang, G.S. Relationship between hybrid performance and genetic diversity based on SSRs and ISSRs in Brassica napus L. Agric. Sci. China 2003, 2, 1083–1090. [Google Scholar]
- Zhao, X.H.; Jiang, X.; Zhao, K.; Zhao, X.H.; Yin, J.; Xie, W.G. Screening of germplasm with low seed shattering rate and evaluation on agronomic traits in Elymus sibiricus L. J. Plant Gen. Resour. 2015, 16, 691–699. [Google Scholar]
- Robins, J.G.; Bushman, B.S.; Jensen, K.B. Dry matter yield combining ability among nine sources of orchardgrass germplasm. Euphytica 2012, 188, 419–428. [Google Scholar] [CrossRef]
- Jia, X.P.; Deng, Y.M.; Sun, X.B.; Liang, L.J.; Ye, X.P. Characterization of the global transcriptome using Illumina sequencing and novel microsatellite marker information in seashore paspalum. Genes Genom. 2015, 37, 77–86. [Google Scholar] [CrossRef]
- Ramu, P.; Billot, C.; Rami, J.F.; Senthilvel, S.; Upadhyaya, H.D.; Reddy, L.A.; Hash, C.T. Assessment of genetic diversity in the sorghum reference set using EST-SSR markers. Theor. Appl. Genet. 2013, 126, 2051–2064. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Xie, W.G.; Robins, J.G.; Bushman, B.S. A genetic linkage map of tetraploid orchardgrass (Dactylis glomerata L.) and quantitative trait loci for heading date. Genome 2012, 55, 360–369. [Google Scholar] [CrossRef] [PubMed]
- Huang, L.K.; Huang, X.; Yan, H.D.; Yin, G.H.; Zhang, X.Q.; Tian, Y.; Zhang, Y.; Jiang, X.M.; Yan, Y.H.; Ma, X.; et al. Constructing DNA fingerprinting of Hemarthria cultivars using EST-SSR and SCoT markers. Genet. Resour. Crop. Evol. 2014, 61, 1047–1055. [Google Scholar] [CrossRef]
- Futuyma, D.J. Evolutionary Biology; Sinauer Associates: Sunderland, MA, USA, 1986; p. 2. [Google Scholar]
- Yan, J.J.; Bai, S.Q.; Zhang, C.B.; You, M.H. A primary investigation report for the wild germplasm of Elymus sibiricus in the Northwest Plateau of Sichuan province. Pruatacult Anim. Husb. 2006, 27, 23–26. [Google Scholar]
- Lei, Y.T.; Zhao, Y.Y.; Yu, F.; Li, Y.; Dou, Q.W. Development and characterization of 53 polymorphic genomic-SSR markers in Siberian wildrye (Elymus sibiricus L.). Conserv. Genet Resour. 2014, 6, 861–864. [Google Scholar] [CrossRef]
- Ma, X.; Zhang, X.Q.; Zhou, Y.H.; Bai, S.Q.; Liu, W. Assessing genetic diversity of Elymus sibiricus (Poaceae: Triticeae) populations from Qinghai-Tibet Plateau by ISSR markers. Biochem. Syst. Ecol. 2008, 36, 514–522. [Google Scholar] [CrossRef]
- Yan, J.J.; Bai, S.Q.; Zhang, X.Q.; You, M.H.; Zhang, C.B.; Li, D.X.; Zeng, Y. Genetic diversity of wild Elymus sibiricus germplasm from the Qinghai-Tibetan Plateau in China detected by SRAP markers. Acta. Prataculturae Sin. 2010, 19, 173–183. [Google Scholar]
- Zhang, J.C.; Xie, W.G.; Wang, Y.R.; Zhao, X.H. Potential of Start Codon Targeted (SCoT) Markers to Estimate Genetic Diversity and Relationships among Chinese Elymus sibiricus Accessions. Molecules 2015, 20, 5987–6001. [Google Scholar] [CrossRef] [PubMed]
- Schoen, D.J.; Brown, A.H.D. Intraspecific variation in population gene diversity and effective population size correlates with the mating system in plants. Proc. Natl. Acad. Sci. USA 1991, 88, 4494–4497. [Google Scholar] [CrossRef] [PubMed]
- Stevens, L.; Salomon, B.; Sun, G. Microsatellite variability and heterozygote excess in Elymus trachycaulus population from British Columbia in Canada. Biochem. Syst. Ecol. 2007, 35, 725–736. [Google Scholar] [CrossRef]
- Slatkin, M. Gene flow and the geographic structure of natural populations. Science 1987, 236, 778–792. [Google Scholar] [CrossRef]
- Schaal, B.A.; Hayworth, D.A.; Olsen, K.M.; Rauscher, J.T.; Smith, W.A. Phylogeographic studies in plants: Problems and prospects. Mol. Ecol. 1998, 7, 465–474. [Google Scholar] [CrossRef]
- Jin, Y.; Lu, B.R. Sampling strategy for genetic diversity. Chin. Biodivers. 2003, 11, 155–161. [Google Scholar]
- Jensen, K.B.; Mott, I.W.; Robins, J.G.; Waldron, B.L.; Nelson, M. Genetic improvement and diversity in Snake River wheatgrass (Elymus wawawaiensis) (Poaceae: Triticeae). Rangeland Ecol. Manag. 2012, 65, 76–84. [Google Scholar] [CrossRef]
- Ozkan, H.; Levy, A.A.; Feldman, M. Allopolyploidy-induced rapid genome evolution in the wheat (Aegilops-Triticum) group. Plant Cell 2001, 13, 1735–1747. [Google Scholar] [CrossRef] [PubMed]
- Levy, A.A.; Feldman, M. The impact of polyploidy on grass genome evolution. Plant Physiol. 2002, 130, 1587–1593. [Google Scholar] [CrossRef] [PubMed]
- Doyle, J.J. DNA protocols for plants-CTAB total DNA isolation. In Molecular Techniques in Taxonomy; Hewit, G.M., Ed.; Springer-Verlag: Berlin, Germany, 1991; pp. 283–293. [Google Scholar]
- Larson, S.R.; Kellogg, E. Genetic dissection of seed production traits and identification of a major-effect seed retention QTL in hybrid Leymus (Triticeae) wildrye. Crop Sci. 2009, 49, 29–40. [Google Scholar] [CrossRef]
- Mott, I.W.; Larson, S.R.; Jones, T.A.; Robins, J.G.; Jensen, K.B.; Peel, M.D. A molecular genetic linkage map identifying the St and H subgenomes of Elymus (Poaceae: Triticeae) wheatgrass. Genome 2011, 54, 819–828. [Google Scholar] [CrossRef] [PubMed]
- Mott, I.W.; Larson, S.R.; Bushman, B.S. Simple sequence repeat (SSR) markers for Elymus, Pseudoroegneria and Pascopyrum species (Triticeae: Gramineae). Plant Genet. Resour. 2011, 9, 489–494. [Google Scholar] [CrossRef]
- Zhou, Q.; Luo, D.; Ma, L.C.; Xie, W.G.; Wang, Y.; Wang, Y.R.; Liu, Z.P. Development and cross-species transferability of EST-SSR markers in Siberian wildrye (Elymus sibiricus L.) using Illumina sequencing. Sci. Rep. 2016, 6, 20549. [Google Scholar] [CrossRef] [PubMed]
- Zhao, Y.F.; Zhang, X.Q.; Ma, X.; Xie, W.G.; Huang, L.K. Morphological and genetic characteristics of hybrid population of Dactylis glomerata. Genet. Mol. Res. 2014, 13, 2491–2503. [Google Scholar] [CrossRef] [PubMed]
- Yeh, F.; Yang, R.; Boyle, T. Quick User Guide. In Popgene; University of Alberta: Edmonton, AB, Canada, 1999. [Google Scholar]
- Ghislain, M.; Zhang, D.P.; Fajardo, D.; Huamán, Z.; Hijmans, R.J. Marker-assisted sampling of the cultivated Andean potato Solanum phureja collection using RAPD markers. Gen. Resour. 1999, 46, 547–555. [Google Scholar]
- Mengoni, A.; Bazzicalupo, M. The statistical treatment of data and the analysis of molecular variance (AMOVA) in molecular microbial ecology. Ann. Microbiol. 2002, 52, 95–102. [Google Scholar]
- Zhang, F.M.; Ge, S. Data analysis in population genetics. I. analysis of RAPD data with AMOVA. Biodivers. Sci. 2002, 10, 438–444. [Google Scholar]
- Pritchard, J.K.; Stephens, M.; Donnelly, P. Inference of population structure from multilocus genotype data. Genetics 2000, 155, 945–959. [Google Scholar] [PubMed]
- Evanno, G.; Regnaut, S.; Goudet, J. Detecting the number of clusters of individuals using the software structure: A simulation study. Mol. Ecol. 2005, 14, 2611–2620. [Google Scholar] [CrossRef] [PubMed]
- Rohlf, F.J. NTSYS-pc: Numerical Taxonomy and Multivariate Analysis System; Exeter Software: New York, NY, USA, 1992. [Google Scholar]
- Sample Availability: Samples of the compounds are available from the authors.



| Population | Code | Accession | Origin | Status |
|---|---|---|---|---|
| SC | 1 | Y1005 | Sichuan, China | Wild |
| SC | 2 | SAU003 | Kangding, Sichuan, China | Wild |
| SC | 3 | SAU133 | Aba, Sichuan, China | Wild |
| SC | 4 | SAU139 | Kangding, Sichuan, China | Wild |
| SC | 5 | SAU137 | Aba, Sichuan, China | Wild |
| SC | 6 | SC02 | Ruoergai, Sichuan, China | Wild |
| SC | 7 | SC03 | Ruoergai, Sichuan, China | Wild |
| SC | 8 | Hongyuan | Hongyuan, Sichuan, China | Breeding line |
| SC | 9 | Chuancao2 | Hongyuan, Sichuan, China | Cultivar |
| NM | 10 | PI 499453 | Inner Mongolia, China | Wild |
| NM | 11 | PI 665507 | Inner Mongolia ,China | Wild |
| NM | 12 | PI 499456 | Inner Mongolia, China | Cultivated |
| NM | 13 | PI 610886 | Inner Mongolia, China | Wild |
| XJ | 14 | PI 655140 | Xinjiang, China | Wild |
| XJ | 15 | PI 595182 | Xinjiang, China | Wild |
| XJ | 16 | PI 595180 | Xinjiang, China | Wild |
| XJ | 17 | Y2003 | Xinjiang, China | Wild |
| XJ | 18 | PI 619577 | Xinjiang, China | Wild |
| XJ | 19 | PI 499462 | Xinjiang, China | Wild |
| XJ | 20 | PI 499468 | Xinjiang, China | Cultivated |
| GS | 21 | XH02 | Xiahe, Gansu, China | Wild |
| GS | 22 | XH03 | Xiahe, Gansu, China | Wild |
| GS | 23 | XH09 | Xiahe, Gansu, China | Wild |
| GS | 24 | LQ04 | Luqu, Gansu, China | Wild |
| GS | 25 | LQ01 | Luqu, Gansu, China | Wild |
| GS | 26 | HZ01 | Hezuo, Gansu, China | Wild |
| GS | 27 | HZ02 | Hezuo, Gansu, China | Wild |
| GS | 28 | ZhN01 | Zhuoni, Gansu, China | Wild |
| GS | 29 | ZhN06 | Zhuoni, Gansu, China | Wild |
| GS | 30 | MQ01 | Maqu, Gansu, China | Wild |
| GS | 31 | LT01 | Lintan, Gansu, China | Wild |
| GS | 32 | LT02 | Lintan, Gansu, China | Wild |
| QH | 33 | Tongde | Qinghai, China | Cultivar |
| QH | 34 | Qingmu1 | Qinghai, China | Cultivar |
| QH | 35 | PI 531669 | Qinghai, China | Wild |
| QH | 36 | PI 504462 | Qinghai, China | Wild |
| Primer ID | Forward Primer (5′-3′) | Reverse Primer (5′-3′) | T | M | TP | %P | PIC |
|---|---|---|---|---|---|---|---|
| Elw0300s019 | TTCATCCATCCAATTCTAGCACAA | GAAGGAGAAGATGGAATCCTTGAA | 8 | 0 | 8 | 100.00 | 0.49 |
| Elw0669s043 | CATCTCACGGCAAGTAAATGAACA | TGCGAGATGGGGTACAATTTTTAT | 12 | 0 | 12 | 100.00 | 0.35 |
| Elw1197s069 | ATGGCCGTAACCCTTTACCTGTAT | TTTCAAAGCCTTTCCAAGTGAATC | 7 | 1 | 6 | 85.71 | 0.48 |
| Elw1420s081 | GGATAGACCCATGAGCTGACTGAT | CTTTCTCCACAAGTTGAACACAAGA | 11 | 0 | 11 | 100.00 | 0.35 |
| Elw1468s087 | TAGCAATAAGTTGCTGCTGCTGTT | CCACCTCTAAATTAATCACCACGAA | 11 | 0 | 11 | 100.00 | 0.47 |
| Elw1675s092 | CAGTTAAAATGCTTGTCCAAATGC | CCATGATTGTTCTGTCAAGAAACG | 10 | 1 | 9 | 90.00 | 0.48 |
| Elw2676s146 | AATTCGAAAGCTGTGGACTTGTCT | CAATTTTGCTCTCAAGAGAACCGT | 10 | 1 | 9 | 90.00 | 0.48 |
| Elw2698s152 | CAAAGCATGTGTAGGCAGTCTTGT | TAACAATGATCAGTTGATCGGACC | 9 | 0 | 9 | 100.00 | 0.50 |
| Elw2807s159 | CCCAAGAAGCAAAAGTGAAGTTGA | ATAATTGCTGTAAAACGGCAGGAA | 11 | 0 | 11 | 100.00 | 0.50 |
| Elw2808s160 | TTTCATATCCGATACCCAGAAAGC | GGGCGACAAGGGTACTACTAACAA | 4 | 0 | 4 | 100.00 | 0.45 |
| Elw3264s184 | TGGACTGCTTTGGGACATAATAGG | CTGAATCATAGCCACCCTGAAAAC | 6 | 0 | 6 | 100.00 | 0.42 |
| Elw3384s187 | AGCTCCTGATAGAAAGAGCCATCA | GGCTGCTGGAACTGAAGACAGTA | 16 | 0 | 16 | 100.00 | 0.36 |
| Elw3492s190 | TGTTGTTGTTCCAGTTCCAGTCTC | AAAAACAACCACACAAGGTTGTCA | 9 | 2 | 7 | 77.78 | 0.43 |
| Elw3995s226 | CTCTAGGGTTTTGGGATTTTAGCC | GTTGTGGAGGTCGGAGAAGGT | 7 | 0 | 7 | 100.00 | 0.49 |
| Elw4419s261 | AGGGTGACTTGTCTTTGGGTGTAA | AGTCAGATGAAGGATGGCTGAAAC | 8 | 1 | 7 | 87.50 | 0.50 |
| Elw5447s306 | TCCTCAAACTCCTCCTCTCTTCG | GAGGTAAGTCTCGACATCCTCGAC | 9 | 0 | 9 | 100.00 | 0.50 |
| Elw5616s393 | TAGTAGCGTGGCACTCCTCTTCTT | GGTACAAACCACCAAAGGTACTGC | 22 | 0 | 22 | 100.00 | 0.42 |
| Elw5627s404 | AGATGAAGCTGGTAACCGAGACAG | ATTTCCTCTAATGGAAGCTCTGGC | 17 | 0 | 17 | 100.00 | 0.46 |
| Ps2283 | GCCACAACAAGAGAAGACCTTGC | GACCTGCATGATGCTCTCGC | 13 | 0 | 13 | 100.00 | 0.40 |
| Ps261 | CTCGAATCCAGCTGAACAATTTCT | AGTCGATCCTCACCTTCATCTCC | 9 | 0 | 9 | 100.00 | 0.45 |
| Ps3447 | AGCTTTATGAAGATCGCCACTCAC | CTGCTGCTGCTACCGTTCTTATTT | 13 | 0 | 13 | 100.00 | 0.50 |
| Ps3577 | CATCTTGCATATAGCTCCTTCGCT | CTCAAGAAACCCACAATCCAATTC | 5 | 0 | 5 | 100.00 | 0.48 |
| Ps938 | TTGCTCCTATGGTTCCACGTAGTT | AAAGTGAAATTCTGCCATCAGAGC | 13 | 1 | 12 | 92.31 | 0.49 |
| Ltc0209 | CAGGAACATGAACAGAAGCCTGTC | GTACTGGTCGAACCACCCAAAGT | 12 | 1 | 11 | 91.67 | 0.50 |
| Ltc0096 | GCGCACTACCGCCTCTTAGTT | GTCCAGGTAGCACACCTCCG | 8 | 2 | 6 | 75.00 | 0.49 |
| ES-7 | CCTCCTCCGTTACCATGTTG | CCCTGCTTTTCCCTCTCTG | 4 | 3 | 1 | 25.00 | 0.27 |
| ES-22 | AAGATATCCTGATGCTGGACAAA | GATCAGATCAATAGCTTGAGCG | 7 | 5 | 2 | 28.57 | 0.23 |
| ES-23 | CGTACTTGCGCCAGAAGTG | AGGTGTCCATCGAAGGGTC | 14 | 2 | 12 | 85.71 | 0.45 |
| ES-51 | GAGCTGAGCTGAGAAGAAAACAG | CACAATCATCTCATCTTCCTTCC | 14 | 0 | 14 | 100.00 | 0.44 |
| ES-97 | ACTGTGGGAGAAGGTGAGAGACT | CTTTCCTCCAGCTCATGGTG | 8 | 4 | 4 | 50.00 | 0.49 |
| ES-123 | AGCATGAAGCTCGACTGTGAGT | GCGAGTACATCTCGTACTTCTGG | 12 | 4 | 8 | 66.67 | 0.49 |
| ES-125 | GAGCATCGACAGATTATTCCTTG | CGAAGGAACCTCTGCAAGAC | 5 | 3 | 2 | 40.00 | 0.23 |
| ES-144 | GGTAGTCGTTGACCCAGATGTC | CACATTGTAAACTGGTCCTCCTC | 5 | 0 | 5 | 100.00 | 0.49 |
| ES-231 | TAGCTGGTCATGCCTAGGAGTAG | CCAGGTGTCAGGATATAGCAAAA | 6 | 1 | 5 | 83.33 | 0.44 |
| ES-236 | TCGCATGCTTATAATCCTTTGAC | TGAGGTCTCTGTCAATACCAACA | 6 | 3 | 3 | 50.00 | 0.35 |
| ES-253 | CATCTCTTCAAACTTGGATTGGT | GTGATCTATACCATTGGCCTCAA | 12 | 4 | 8 | 66.67 | 0.46 |
| ES-259 | CTCCTCTACCTGTCTGCTGCTA | AGATCGTCGACTACGTCAAGAAG | 11 | 0 | 11 | 100.00 | 0.47 |
| ES-310 | CGTAGCAATTCCATTCTATCCAG | TGGTGAGCTAGATTGACACTGAG | 9 | 6 | 3 | 33.33 | 0.33 |
| ES-347 | CATGAAGATGATGCGTGTTTTAAT | CCGACTCCTAATTGAACTCGTAA | 5 | 3 | 2 | 40.00 | 0.47 |
| ES-405 | AGAGAAAAGGAGATTCTCATCCC | GCTGCTCTGCATCCTACTCTATC | 2 | 1 | 1 | 50.00 | 0.43 |
| Mean | 9.5 | 1.2 | 8.3 | 87.11 | 0.44 | ||
| Total | 380 | 49 | 331 |
| POP | NPB | PPB (%) | I | H | Na |
|---|---|---|---|---|---|
| SC | 204 | 73.12 | 0.2650 | 0.1739 | 1.5368 |
| NM | 137 | 56.85 | 0.1947 | 0.1301 | 1.3605 |
| XJ | 177 | 66.79 | 0.2368 | 0.1570 | 1.4658 |
| GS | 299 | 86.17 | 0.3594 | 0.2315 | 1.7868 |
| QH | 70 | 32.71 | 0.0946 | 0.0623 | 1.1842 |
| Mean | 177.4 | 63.13 | 0.2301 | 0.1510 | 1.4668 |
| Source of Variance | Degree of Freedom | Sum of Squares | Variance Component | Total Variation (%) |
|---|---|---|---|---|
| Among geographic regions | 4 | 403.45 | 8.48 | 16.60 |
| Within geographic regions | 31 | 1320.13 | 42.58 | 83.40 |
| Population | SC | NM | XJ | GS | QH |
|---|---|---|---|---|---|
| SC | 0.9460 | 0.9478 | 0.9192 | 0.6453 | |
| NM | 0.0555 | 0.9552 | 0.8783 | 0.6170 | |
| XJ | 0.0536 | 0.0458 | 0.9024 | 0.6514 | |
| GS | 0.0843 | 0.1297 | 0.1027 | 0.7553 | |
| QH | 0.4381 | 0.4829 | 0.4286 | 0.2807 |
| Material Name | PH (cm) | LL (cm) | LW (cm) | FLL (cm) | FLW (cm) | CD (cm) | CN (No.) | TN (No.) | PL (cm) | AL (cm) | 1000-SW (g) | SS (gf) | C.VV (%) |
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Y1005-1 | 56.3 | 12.6 | 0.9 | 9.1 | 0.8 | 0.2 | 3.1 | 183 | 20.0 | 1.1 | 4.4 | 41.9 | 18.37 |
| ZhN06-1 | 98.0 | 16.2 | 0.8 | 9.1 | 0.6 | 0.4 | 3.4 | 152 | 19.2 | 1.1 | 2.4 | 97.2 | 17.09 |
| F1-1 | 69.0 | 18.4 | 0.7 | 13.5 | 0.7 | 0.4 | 3.2 | 135 | 17.8 | 1.0 | 3.4 | 68.2 | 17.82 |
| F1-2 | 79.0 | 20.2 | 1.0 | 15.7 | 0.9 | 0.4 | 3.3 | 111 | 20.0 | 1.1 | 3.5 | 93.0 | 16.03 |
| F1-3 | 88.0 | 21.4 | 1.1 | 15.6 | 1.0 | 0.4 | 3.4 | 114 | 21.7 | 1.1 | 2.8 | 82.1 | 19.83 |
| F1-4 | 82.0 | 22.6 | 1.1 | 17.5 | 1.1 | 0.4 | 3.1 | 95 | 23.3 | 1.3 | 5.1 | 114.9 | 14.46 |
| F1-5 | 76.5 | 23.1 | 1.3 | 18.0 | 1.3 | 0.4 | 4.4 | 95 | 22.5 | 1.2 | 5.4 | 96.4 | 14.32 |
| F1-6 | 81.7 | 22.8 | 0.9 | 18.8 | 1.1 | 0.3 | 3.7 | 132 | 19.4 | 1.1 | 1.8 | 143.4 | 16.82 |
| F1-7 | 77.5 | 20.8 | 1.1 | 15.8 | 1.1 | 0.3 | 4.0 | 165 | 21.3 | 1.4 | 3.7 | 137.5 | 14.62 |
| Max | 98.0 | 23.1 | 1.3 | 18.8 | 1.3 | 0.4 | 4.4 | 183 | 23.3 | 1.4 | 5.4 | 143.4 | 19.83 |
| Min | 56.3 | 12.6 | 0.7 | 9.1 | 0.6 | 0.2 | 3.1 | 95 | 17.8 | 1.0 | 1.8 | 41.9 | 14.32 |
| Mean | 78.7 | 19.8 | 1.0 | 14.8 | 1.0 | 0.3 | 3.5 | 131.3 | 20.6 | 1.2 | 3.6 | 97.2 | 16.59 |
| MPH (%) | 2.5 | 48.4 | 21.8 | 80.9 | 44.0 | 13.3 | 10.3 | -27.8 | 6.4 | 7.4 | 7.6 | 51.1 | |
| HPH (%) | -19.3 | 32.0 | 10.0 | 80.4 | 23.5 | -9.6 | 5.5 | -33.9 | 4.2 | 6.9 | -16.5 | 8.1 | |
| SD | 11.62 | 3.51 | 0.17 | 3.60 | 0.23 | 0.06 | 0.44 | 30.82 | 1.75 | 0.12 | 1.19 | 32.05 | |
| CV (%) | 14.78 | 17.74 | 17.28 | 24.32 | 23.55 | 17.47 | 12.61 | 23.47 | 8.50 | 10.28 | 33.11 | 32.98 |
| Primer ID | Bands Information | BSYF | BSZF | BEPP | BEPF | ||||
|---|---|---|---|---|---|---|---|---|---|
| T | M | TP | % P | PIC | |||||
| Elw0300s019 | 3 | 2 | 1 | 33.33 | 0.15 | 0 | 0 | 0 | 0 |
| Elw0669s043 | 9 | 0 | 9 | 100.00 | 0.42 | 3 | 6 | 0 | 0 |
| Elw1197s069 | 6 | 2 | 4 | 66.67 | 0.30 | 3 | 0 | 0 | 0 |
| Elw1420s081 | 4 | 0 | 4 | 100.00 | 0.42 | 0 | 1 | 0 | 2 |
| Elw1468s087 | 9 | 0 | 9 | 100.00 | 0.38 | 1 | 5 | 0 | 0 |
| Elw1675s092 | 8 | 1 | 7 | 87.50 | 0.30 | 0 | 2 | 0 | 1 |
| Elw2676s146 | 9 | 1 | 8 | 88.89 | 0.33 | 0 | 4 | 0 | 0 |
| Elw2698s152 | 9 | 1 | 8 | 88.89 | 0.34 | 0 | 0 | 0 | 0 |
| Elw2807s159 | 9 | 5 | 4 | 44.44 | 0.14 | 2 | 1 | 0 | 0 |
| Elw2808s160 | 4 | 1 | 3 | 75.00 | 0.35 | 3 | 0 | 0 | 0 |
| Elw3264s184 | 7 | 1 | 6 | 85.71 | 0.37 | 2 | 2 | 0 | 0 |
| Elw3384s187 | 6 | 1 | 5 | 83.33 | 0.41 | 2 | 0 | 0 | 2 |
| Elw3492s190 | 7 | 4 | 3 | 42.86 | 0.16 | 0 | 1 | 0 | 0 |
| Elw3995s226 | 5 | 3 | 2 | 40.00 | 0.08 | 0 | 1 | 0 | 0 |
| Elw4419s261 | 4 | 4 | 0 | 0.00 | 0.00 | 0 | 0 | 0 | 0 |
| Elw5447s306 | 6 | 1 | 5 | 83.33 | 0.31 | 0 | 0 | 0 | 1 |
| Elw5616s393 | 17 | 1 | 16 | 94.12 | 0.40 | 1 | 7 | 0 | 0 |
| Elw5627s404 | 10 | 0 | 10 | 100.00 | 0.39 | 3 | 1 | 0 | 0 |
| Ps2283 | 5 | 0 | 5 | 100.00 | 0.44 | 1 | 0 | 0 | 3 |
| Ps261 | 9 | 3 | 6 | 66.67 | 0.25 | 4 | 0 | 1 | 1 |
| Ps3447 | 5 | 4 | 1 | 20.00 | 0.04 | 1 | 0 | 0 | 0 |
| Ps3577 | 4 | 2 | 2 | 50.00 | 0.10 | 0 | 1 | 1 | 0 |
| Ps938 | 8 | 6 | 2 | 25.00 | 0.09 | 1 | 0 | 0 | 0 |
| Ltc0209 | 8 | 1 | 7 | 87.50 | 0.31 | 2 | 2 | 0 | 0 |
| Ltc0096 | 5 | 3 | 2 | 40.00 | 0.17 | 1 | 0 | 0 | 0 |
| ES-7 | 3 | 3 | 0 | 0.00 | 0.00 | 0 | 0 | 0 | 0 |
| ES-22 | 6 | 6 | 0 | 0.00 | 0.00 | 0 | 0 | 0 | 0 |
| ES-23 | 4 | 3 | 1 | 25.00 | 0.05 | 0 | 0 | 1 | 0 |
| ES-51 | 8 | 1 | 7 | 87.50 | 0.38 | 0 | 4 | 0 | 0 |
| ES-97 | 4 | 4 | 0 | 0.00 | 0.00 | 0 | 0 | 0 | 0 |
| ES-123 | 7 | 5 | 2 | 28.57 | 0.10 | 0 | 1 | 0 | 0 |
| ES-125 | 5 | 5 | 0 | 0.00 | 0.00 | 0 | 0 | 0 | 0 |
| ES-144 | 5 | 4 | 1 | 20.00 | 0.07 | 1 | 0 | 0 | 0 |
| ES-231 | 4 | 4 | 0 | 0.00 | 0.00 | 0 | 0 | 0 | 0 |
| ES-236 | 5 | 4 | 1 | 20.00 | 0.04 | 0 | 0 | 0 | 0 |
| ES-253 | 11 | 6 | 5 | 45.45 | 0.13 | 1 | 0 | 0 | 1 |
| ES-259 | 8 | 0 | 8 | 100.00 | 0.41 | 0 | 2 | 0 | 2 |
| ES-310 | 7 | 7 | 0 | 0.00 | 0.00 | 0 | 0 | 0 | 0 |
| ES-347 | 3 | 3 | 0 | 0.00 | 0.00 | 0 | 0 | 0 | 0 |
| ES-405 | 1 | 1 | 0 | 0.00 | 0.00 | 0 | 0 | 0 | 0 |
| Mean | 6.4 | 2.6 | 3.9 | 59.92 | 0.20 | 0.8 | 1.0 | 0.1 | 0.3 |
| Total | 257 | 103 | 154 | 32 | 41 | 3 | 13 | ||
© 2016 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC-BY) license ( http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Zhang, Z.; Zhang, J.; Zhao, X.; Xie, W.; Wang, Y. Assessing and Broadening Genetic Diversity of Elymus sibiricus Germplasm for the Improvement of Seed Shattering. Molecules 2016, 21, 869. https://doi.org/10.3390/molecules21070869
Zhang Z, Zhang J, Zhao X, Xie W, Wang Y. Assessing and Broadening Genetic Diversity of Elymus sibiricus Germplasm for the Improvement of Seed Shattering. Molecules. 2016; 21(7):869. https://doi.org/10.3390/molecules21070869
Chicago/Turabian StyleZhang, Zongyu, Junchao Zhang, Xuhong Zhao, Wengang Xie, and Yanrong Wang. 2016. "Assessing and Broadening Genetic Diversity of Elymus sibiricus Germplasm for the Improvement of Seed Shattering" Molecules 21, no. 7: 869. https://doi.org/10.3390/molecules21070869
APA StyleZhang, Z., Zhang, J., Zhao, X., Xie, W., & Wang, Y. (2016). Assessing and Broadening Genetic Diversity of Elymus sibiricus Germplasm for the Improvement of Seed Shattering. Molecules, 21(7), 869. https://doi.org/10.3390/molecules21070869





