Identification of Oxygenated Fatty Acid as a Side Chain of Lipo-Alkaloids in Aconitum carmichaelii by UHPLC-Q-TOF-MS and a Database
Abstract
:1. Introduction
2. Results and Discussion
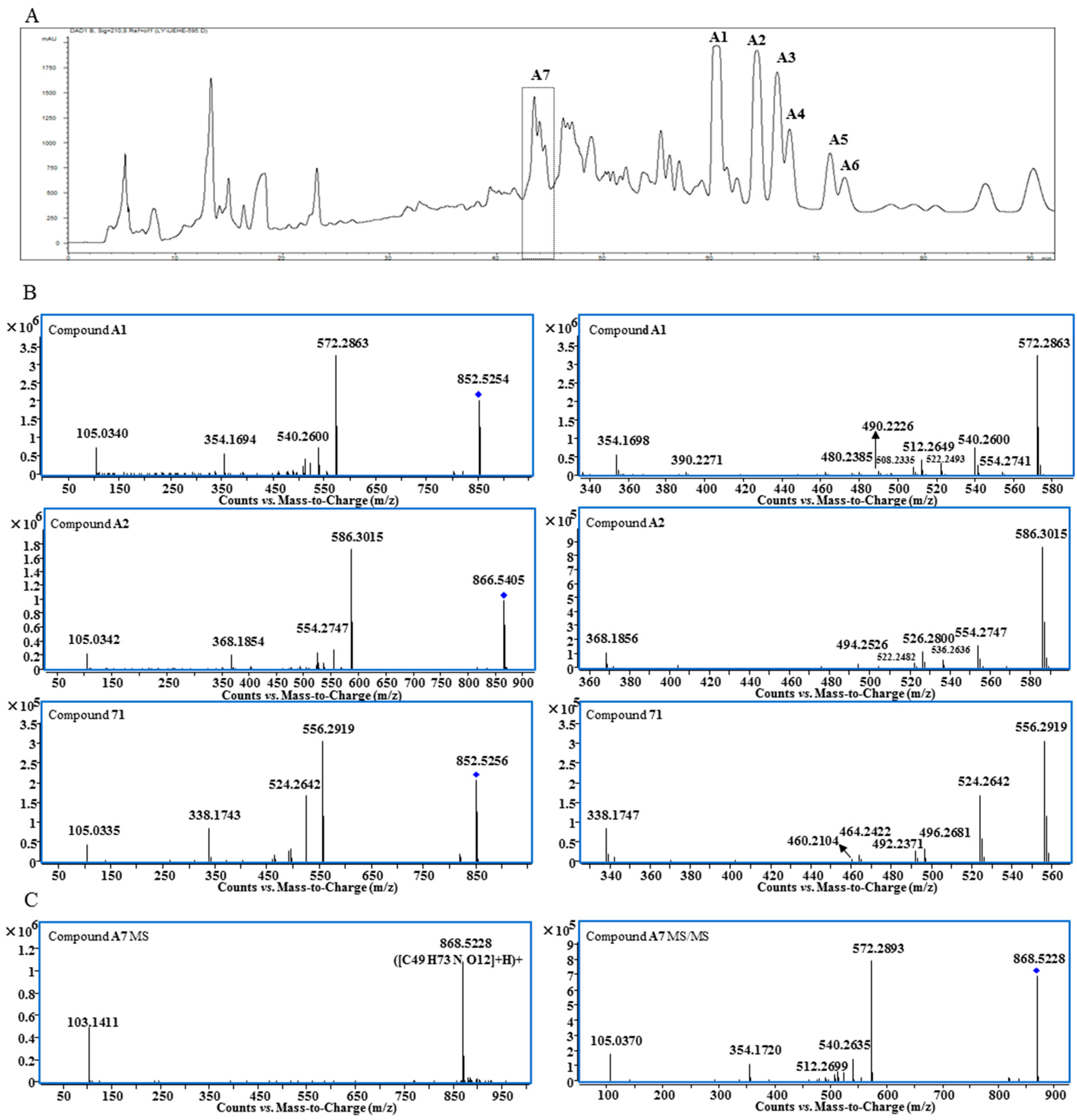
2.1. Isolation and Structural Elucidation of Lipo-Alkaloids A1–A7
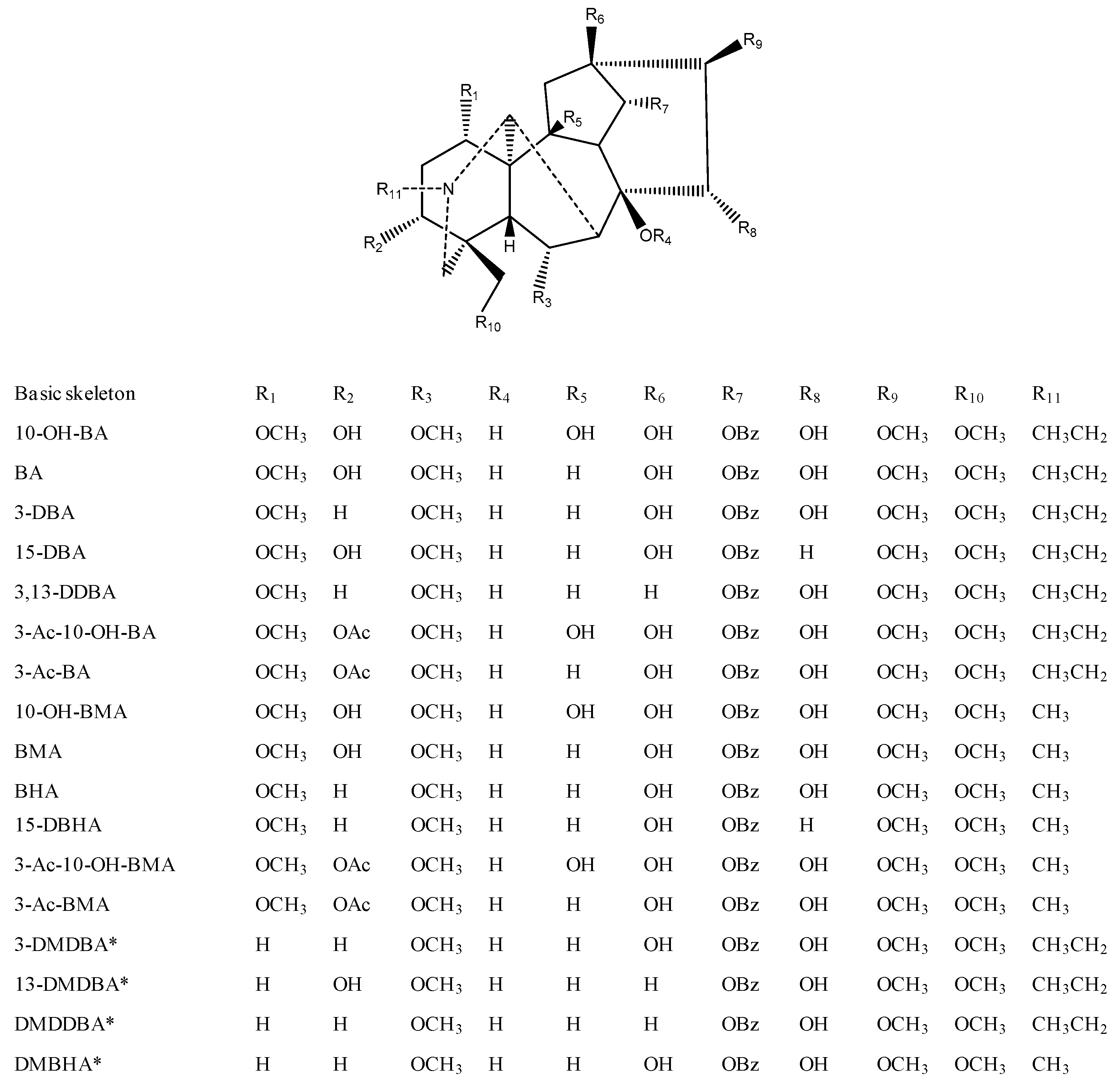
2.2. Establishment of Lipo-Alkaloids Database
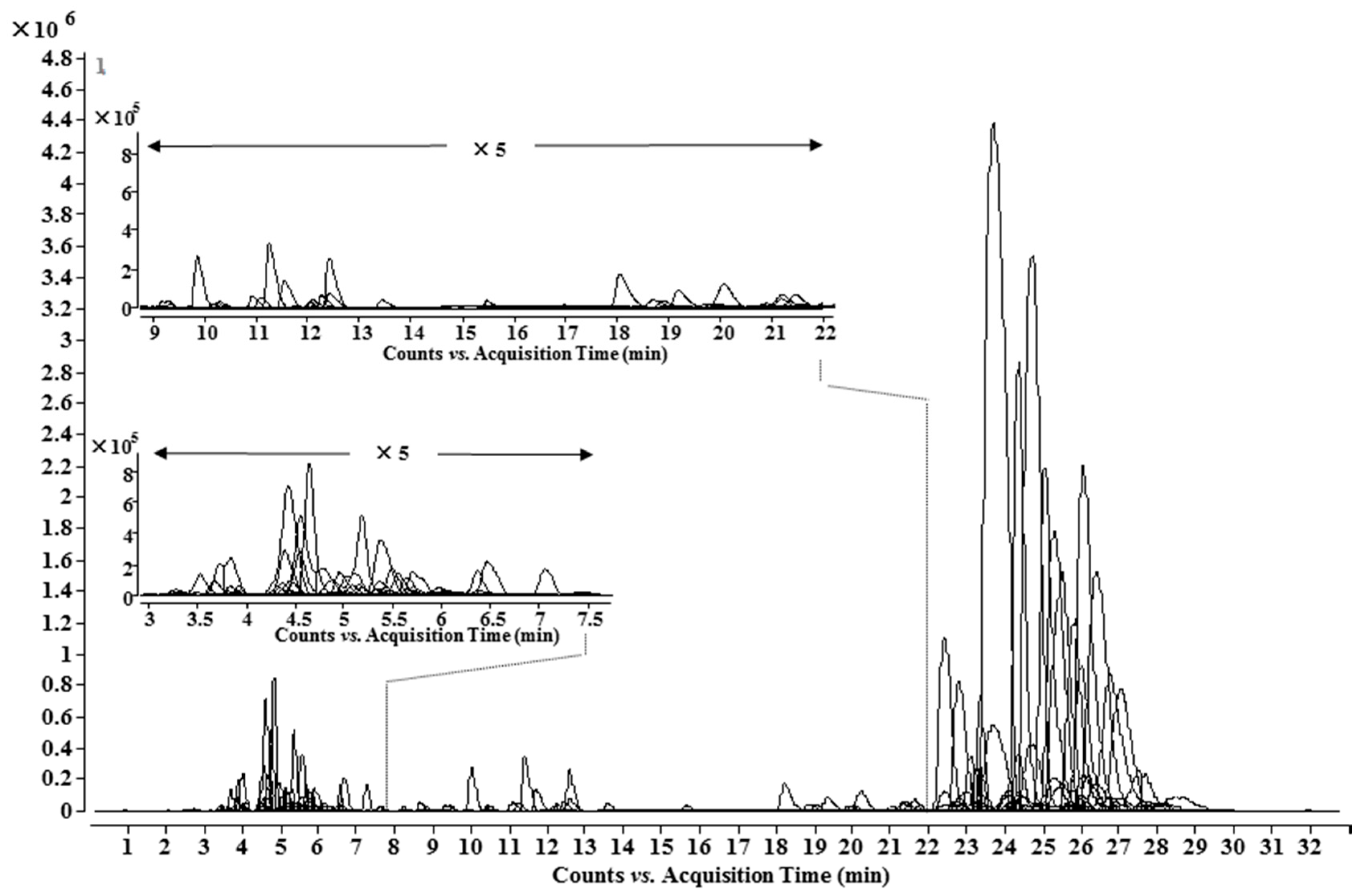
2.3. Determination of Lipo-Alkaloids by Combination of Database, UHPLC-Q-TOF-MS and MS/MS Analysis
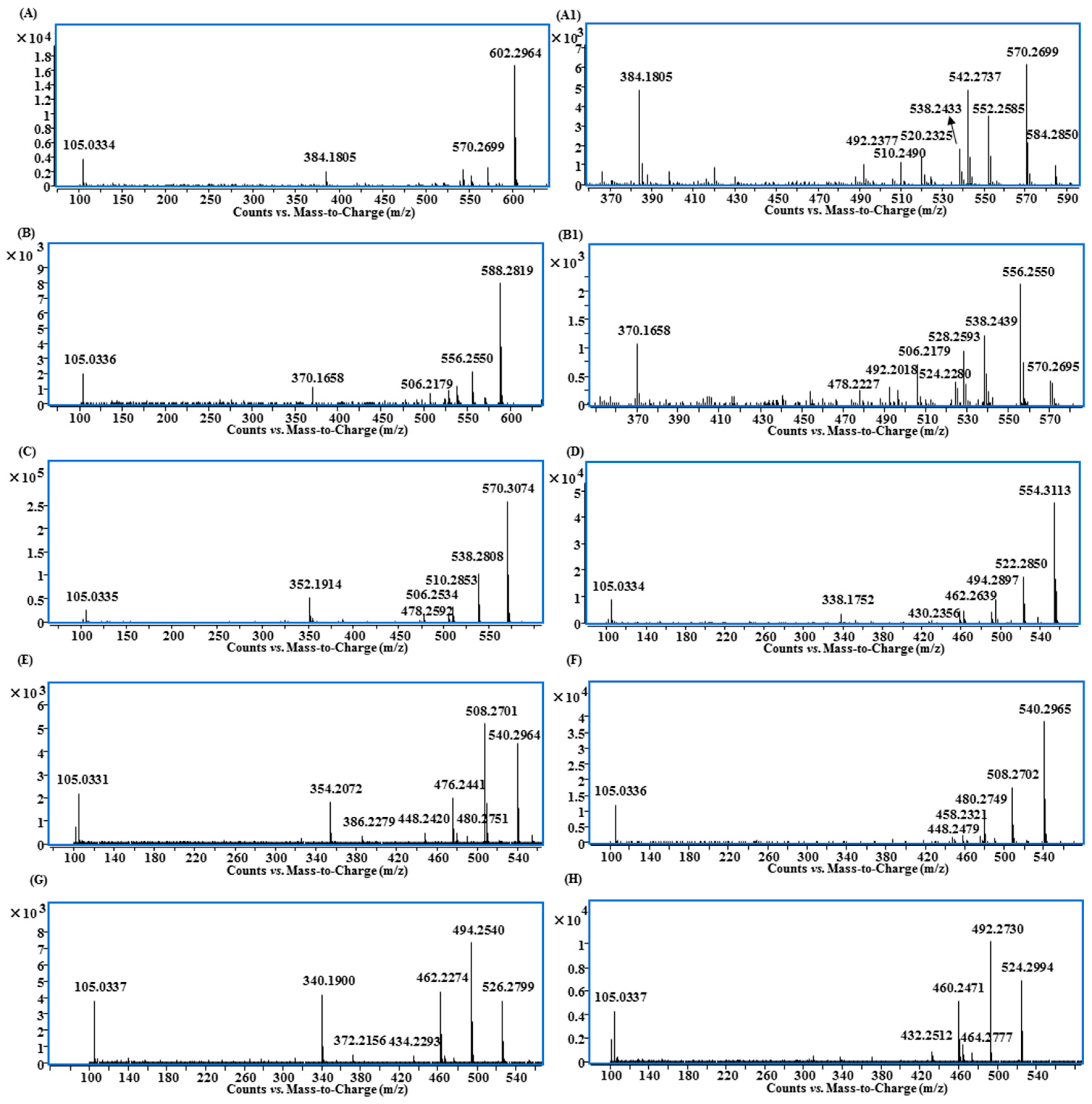
2.4. MS/MS Characterizations of Aconitane Skeletons in Lipo-Alkaloids
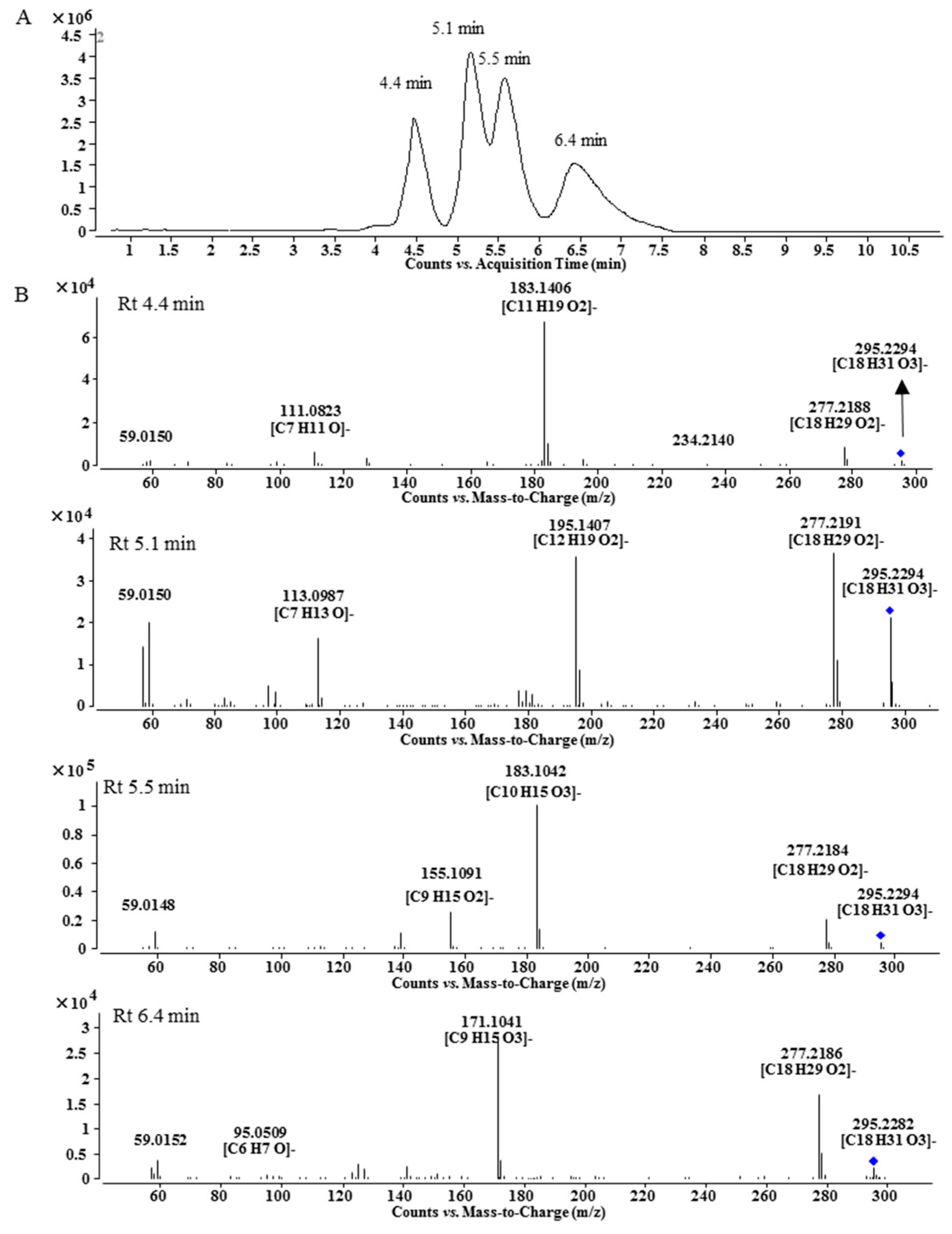
2.5. Fatty Acid Side Chains in Lipo-Alkaloids
3. Materials and Methods
3.1. Chemicals and Reagents
3.2. Plant Materials
3.3. Separation of Lipo-Alkaloids
3.4. Alkaline Hydrolysis of Peak A7
3.5. Preparation of Methanol Extracts of Herbal Sample
3.6. UHPLC-Q-TOF-MS Analysis
3.7. Establishment of the Lipo-Alkaloids Database
4. Conclusions
Supplementary Materials
Acknowledgments
Author Contributions
Conflicts of Interest
Abbreviation
| UHPLC-Q-TOF-MS | ultra-high performance liquid chromatography-quadrupole-time of flight-mass spectrometry |
| BA | 14-benzoylaconine |
| BMA | 14-benzoylmesaconine |
| BHA | 14-benzoylhypaconine |
| 10-OH-BA | 10-hydroxy-14-benzoylaconine |
| 3-DBA | 3-deoxy-14-benzoylaconine |
| 15-DBA | 15-deoxy-14-benzoylaconine |
| 3,13-DDBA | 3,13-dideoxy-14-benzoylaconine |
| 10-OH-BMA | 10-hydroxy-14-benzoylmesaconine |
| 15-DBHA | 15-deoxy-14-benzoylhypaconine |
| 3-Ac-10-OH-BA | 3-acetyl-10-hydroxy-14-benzoylaconine |
| 3-Ac-10-OH-BMA | 3-acetyl-10-hydroxy-14-benzoylmesaconine |
| 3-Ac-BA | 3-acetyl-14-benzoylaconine |
| 3-Ac-BMA | 3-acetyl-14-benzoylmesaconine |
| 3-DMDBA | demethoxy-3-deoxy-14-benzoylaconine |
| 13-DMDBA | demethoxy-13-deoxy-14-benzoylaconine |
| DMDDBA | demethoxy-3,13-dideoxy-14-benzoylaconine |
| DMBHA | demethoxy-14-benzoylhypaconine |
References
- Borcsa, B.; Csupor, D.; Forgo, P.; Widowitz, U.; Bauer, R.; Hohmann, J. Aconitum lipo-alkaloids-semisynthetic products of the traditional medicine. Nat. Prod. Commun. 2011, 6, 527–536. [Google Scholar] [PubMed]
- Chodoeva, A.; Bosc, J.J.; Guillon, J.; Costet, P.; Decendit, A.; Merillon, J.M.; Leger, J.M.; Jarry, C.; Robert, J. Hemisynthesis and antiproliferative properties of mono-[O-(14-benzoylaconine-8-yl)]esters and bis-[O-(14-benzoylaconine-8-yl)]esters. Eur. J. Med. Chem. 2012, 54, 343–351. [Google Scholar] [CrossRef] [PubMed]
- Zhao, Y.F.; Song, F.R.; Yue, H.; Guo, X.H.; Li, H.L.; Liu, Z.Q.; Liu, S.Y. Biotransformation of Deoxyaconitine of Metabolite of Aconitine by Human Intestinal Bacteria and Electrospray Ionization Tandem Mass Spectrometry. Chem. J. Chin. Univ. 2007, 28, 2051–2055. [Google Scholar]
- Zhao, Y.F.; Song, F.R.; Guo, X.H.; Liu, S.Y. Studies on the biotransformation of aconitine in human intestinal bacteria using soft-ionization mass spectrometry. Chem. J. Chin. Univ. 2008, 29, 55–59. [Google Scholar] [CrossRef]
- Zhao, Y.F.; Song, F.R.; Wang, X.Y.; Guo, X.H.; Liu, Z.Q.; Liu, S.Y. Studies on the biotransformation of 16-O-demethylaconitine and electrospray ionization tandem mass spectrometry. Acta Chim. Sin. 2008, 66, 525–530. [Google Scholar]
- Wang, Y.; Liu, Z.; Song, F.; Liu, S. Electrospray ionization tandem mass spectrometric study of the aconitines in the roots of aconite. Rapid Commun. Mass Spectrom. 2002, 16, 2075–2082. [Google Scholar] [CrossRef] [PubMed]
- Wu, W.; Liang, Z.; Zhao, Z.; Cai, Z. Direct analysis of alkaloid profiling in plant tissue by using matrix-assisted laser desorption/ionization mass spectrometry. J. Mass Spectrom. 2007, 42, 58–69. [Google Scholar] [CrossRef] [PubMed]
- Yue, H.; Pi, Z.; Song, F.; Liu, Z.; Cai, Z.; Liu, S. Studies on the aconitine-type alkaloids in the roots of Aconitum carmichaeli Debx. by HPLC/ESIMS/MS(n). Talanta 2009, 77, 1800–1807. [Google Scholar] [CrossRef] [PubMed]
- Sun, W.; Song, F.; Cui, M.; Liu, S. Simultaneous determination of lipo-alkaloids extracted from Aconitum carmiechaeli using electrospray ionization mass spectrometry and multiple tandem mass spectrometry. Planta Med. 1999, 65, 432–436. [Google Scholar] [CrossRef] [PubMed]
- Csupor, D.; Wenzig, E.M.; Zupko, I.; Wolkart, K.; Hohmann, J.; Bauer, R. Qualitative and quantitative analysis of aconitine-type and lipo-alkaloids of Aconitum carmichaelii roots. J. Chromatogr. A 2009, 1216, 2079–2086. [Google Scholar] [CrossRef] [PubMed]
- Zhang, J.; Huang, Z.H.; Qiu, X.H.; Yang, Y.M.; Zhu da, Y.; Xu, W. Neutral fragment filtering for rapid identification of new diester-diterpenoid alkaloids in roots of Aconitum carmichaeli by ultra-high-pressure liquid chromatography coupled with linear ion trap-orbitrap mass spectrometry. PLoS ONE 2012, 7, e52352. [Google Scholar] [CrossRef] [PubMed]
- Chodoeva, A.; Bosc, J.J.; Guillon, J.; Decendit, A.; Petraud, M.; Absalon, C.; Vitry, C.; Jarry, C.; Robert, J. 8-O-Azeloyl-14-benzoylaconine: A new alkaloid from the roots of Aconitum karacolicum Rapcs and its antiproliferative activities. Bioorg. Med. Chem. 2005, 13, 6493–6501. [Google Scholar] [CrossRef] [PubMed]
- Wang, F.P.; Chen, Q.H. The C19-diterpenoid alkaloids. Alkaloids. Chem. Biol. 2010, 69, 1–577. [Google Scholar] [PubMed]
- Levison, B.S.; Zhang, R.; Wang, Z.; Fu, X.; DiDonato, J.A.; Hazen, S.L. Quantification of fatty acid oxidation products using online high-performance liquid chromatography tandem mass spectrometry. Free Radic. Biol. Med. 2013, 59, 2–13. [Google Scholar] [CrossRef] [PubMed]
- Garscha, U.; Oliw, E.H. Steric analysis of 8-hydroxy- and 10-hydroxyoctadecadienoic acids and dihydroxyoctadecadienoic acids formed from 8R-hydroperoxyoctadecadienoic acid by hydroperoxide isomerases. Anal. Biochem. 2007, 367, 238–246. [Google Scholar] [CrossRef] [PubMed]
- Wang, Y.; Song, F.; Xu, Q.; Liu, Z.; Liu, S. Characterization of aconitine-type alkaloids in the flowers of Aconitum kusnezoffii by electrospray ionization tandem mass spectrometry. J. Mass Spectrom. 2003, 38, 962–970. [Google Scholar] [CrossRef] [PubMed]
- Liu, W.; Pi, Z.; Wang, X.; Song, F.; Liu, S. HPLC/ESI-MSn and ESI-MS studies on the Aconitum alkaloids in three Chinese medicinal herbs. J. Sep. Sci. 2010, 33, 2898–2906. [Google Scholar] [CrossRef] [PubMed]
- Yue, H.; Pi, Z.; Song, F.; Liu, Z.; Liu, S. Studies on the Components of Aconitum carmichaeli by Using HPLC/ESI-MSn. Acta Chem. Sin. 2008, 66, 211–215. [Google Scholar]
- Yue, H.; Pi, Z.F.; Zhao, Y.F.; Song, F.R.; Liu, Z.Q.; Liu, S.Y. Investigation of Aconitine-type Alkaloids from Processed Tuber of Aconitum carmiechaeli by HPLC-ESI-MS/MSn. Chem. Res. Chin. Univ. 2007, 23, 625–627. [Google Scholar] [CrossRef]
- Bai, Y.; Desai, H.K.; Pelletier, S.W. Long-chain fatty acid esters of some norditerpenoid alkaloids. J. Nat. Prod. 1994, 57, 963–970. [Google Scholar] [CrossRef] [PubMed]
- Liu, S.Y.; Liu, Z.Q.; Wang, Y.; Song, F.R.; Xu, Q.X.; Liu, N. Synthesis of Aconitum Alkaloid Type Fatty Acid Ester. Patent CN 1,580,047 A, 16 February 1995. [Google Scholar]
- hen, L.H.; Jian, L.J.; Su, Z.M.; Qiu, Y.Q.; Wang, Y.; Liu, S.Y. ESI-MSn Behavior and Quantum Chemistry Calculation of Stability of Fragment Ions of Diester-diterpenoid Alkaloids (DDA). Chem. J. Chin. Univ. 2005, 26, 2340–2344. [Google Scholar]
- Gobel, C.; Feussner, I. Methods for the analysis of oxylipins in plants. Phytochemistry 2009, 70, 1485–1503. [Google Scholar] [CrossRef] [PubMed]
- Lipidomics Gateway. Available online: http://www.lipidmaps.org (accessed on 21 March 2016).
- Blee, E. Impact of phyto-oxylipins in plant defense. Trends Plant Sci. 2002, 7, 315–322. [Google Scholar] [CrossRef]
- Martinez, J.; Vogler, O.; Casas, J.; Barcelo, F.; Alemany, R.; Prades, J.; Nagy, T.; Baamonde, C.; Kasprzyk, P.G.; Teres, S.; et al. Membrane structure modulation, protein kinase C alpha activation, and anticancer activity of minerval. Mol. Pharmacol. 2005, 67, 531–540. [Google Scholar] [CrossRef] [PubMed]
- Choi, H.G.; Park, Y.M.; Lu, Y.; Chang, H.W.; Na, M.; Lee, S.H. Inhibition of prostaglandin D(2) production by trihydroxy fatty acids isolated from Ulmus davidiana var. Japonica. Phytother. Res. 2013, 27, 1376–1380. [Google Scholar] [CrossRef] [PubMed]
- Sample Availability: Not available.

| Basic Skeleton | [M + H − FA]+ | [M + H − FA − CH3OH]+ | [M + H − FA − CH3OH − H2O]+ | [M + H − FA − CH3OH − CO]+ | [M + H − FA − 2CH3OH]+ | [M + H − FA − 2CH3OH − H2O]+ | [M + H − FA − 2CH3OH − CO]+ | [M + H − FA − 3CH3OH]+ | [M + H − FA − 2CH3OH − BzOH]+ | [M + H − FA − 3CH3OH − BzOH]+ | [M + H − FA − 3CH3OH − BzOH − AcOH]+ |
|---|---|---|---|---|---|---|---|---|---|---|---|
| 3-Ac-BMA | 614.2965 | 582.2703 | - | 554.2754 | 550.2441 | - | 522.2492 | - | - | 396.1811 | 336.1600 |
| 10-OH-BA | 602.2965 | 570.2703 | 552.2597 | 542.2754 | 538.2441 | 520.2335 | 510.2492 | 506.2179 | weak | 384.1811 | - |
| 10-OH-BMA | 588.2809 | 556.2547 | 538.2441 | 528.2597 | 524.2284 | 506.2179 | 496.2335 | 492.2022 | weak | 370.1654 | - |
| BA | 586.3016 | 554.2754 | 536.2648 | 526.2805 | 522.2492 | 504.2386 | 494.2543 | 490.2230 | weak | 368.1862 | - |
| BMA | 572.2860 | 540.2597 | 522.2492 | 512.2648 | 508.2335 | 490.2230 | 480.2386 | 476.2073 | weak | 354.1705 | - |
| DBA | 570.3067 | 538.2805 | - | 510.2856 | 506.2543 | - | 478.2593 | - | weak | 352.1913 | - |
| BHA | 556.2910 | 524.2648 | - | 496.2704 | 492.2386 | - | 464.2437 | 460.2124 | weak | 338.1756 | - |
| 3,13-DDBA | 554.3118 | 522.2856 | - | 494.2906 | 490.2593 | - | 462.2644 | 458.2331 | - | - | - |
| 3-DMDBA * | 540.2961 | 508.2699 | - | 480.2750 | 476.2437 | - | 448.2448 | - | 354.2069 | - | - |
| 13-DMDBA * | 540.2961 | 508.2699 | 490.2593 | 480.2750 | 476.2437 | 458.2331 | 448.2448 | - | - | - | - |
| DMBHA * | 526.2805 | 494.2543 | - | - | 462.2280 | - | 434.2331 | - | 340.1913 | - | - |
| 3,13-DMDDBA * | 524.3014 | 492.2750 | - | 464.2801 | 460.2488 | - | 432.2539 | - | - | - | - |
| No. | Rt (min) | Calcd. [M + H]+ | [M + H]+ | Alkaloids | Ref. | No. | Rt (min) | Calcd. [M + H]+ | [M + H]+ | Alkaloids | Ref. |
|---|---|---|---|---|---|---|---|---|---|---|---|
| 1 | 3.4 | 692.3277 | 692.3285 | 8-dhbtn-BMA * | 75 | 21.8 | 884.5519 | 884.5494 | 8-hode-BA * | ||
| 2 | 3.4 | 762.2968 | 762.2968 | 8-act-10-OH-BMA * | 76 | 22.4 | 866.5049 | 866.5061 | 8-linolen-10-OH-BMA | [16] | |
| 3 | 3.6 | 902.3805 | 902.3777 | 8-gvl-BMA * | 77 | 22.5 | 880.5206 | 880.5193 | 8-linolen-10-OH-BA | [16] | |
| 4 | 3.8 | 746.3018 | 746.3002 | 8-act-BMA * | 78 | 22.5 | 824.4943 | 824.4888 | 8-pmde-BMA | [8] | |
| 5 | 4.0 | 692.3277 | 692.3258 | 8-hbtn-10-OH-BMA * | 79 | 22.7 | 854.5413 | 854.5409 | 8-hode-BHA | [8] | |
| 6 | 4.2 | 914.3805 | 914.3784 | 8-gcf-BMA * | 80 | 22.9 | 850.5100 | 850.5105 | 8-linolen-BMA | [6] | |
| 7 | 4.5 | 928.3961 | 928.3946 | 8-gfr-BMA * | 81 | 22.9 | 812.4943 | 812.4874 | 8-ptde-BMA | [6] | |
| 8 | 4.6 | 730.3069 | 730.3041 | 8-act-BHA * | 82 | 23.0 | 772.4630 | 772.4618 | 8-laur-BMA * | ||
| 9 | 4.7 | 734.3382 | 734.3324 | 8-adp-10-OH-BMA * | 83 | 23.1 | 864.5256 | 864.5252 | 8-linolen-BA | [8] | |
| 10 | 4.7 | 734.3382 | 734.3324 | 8-hadp-BMA * | 84 | 23.2 | 826.5100 | 826.5052 | 8-pme-BMA | [7] | |
| 11 | 4.8 | 676.3328 | 676.3321 | 8-hbtn-BMA * | 85 | 23.3 | 818.5202 | 818.5209 | 8-linolen-3-DMDBA * | ||
| 12 | 5.0 | 760.3175 | 760.3189 | 8-act-BA * | 86 | 23.4 | 848.5307 | 848.5310 | 8-linolen-DBA | [6] | |
| 13 | 5.3 | 760.3175 | 760.3176 | 8-dohpnd-BMA * | 87 | 23.6 | 868.5206 | 868.5195 | 8-lino-10-OH-BMA | [17] | |
| 14 | 5.6 | 672.3015 | 672.3020 | 8-fmr-BHA * | 88 | 23.8 | 838.5100 | 838.5105 | 8-hpdde-BMA * | ||
| 15 | 5.8 | 660.3384 | 660.3380 | 8-hbtn-BHA * | 89 | 23.8 | 840.5256 | 840.5184 | 8-pme-BA | [18] | |
| 16 | 5.8 | 744.3226 | 744.3244 | 8-act-DBA * | 90 | 23.9 | 868.5569 | 868.5559 | 8-hode-DBA * | ||
| 17 | 5.9 | 942.4118 | 942.4084 | 8-gfr-BA * | 91 | 23.9 | 882.5362 | 882.5317 | 8-lino-10-OH-BA | [8] | |
| 18 | 6.2 | 704.3277 | 704.3249 | 8-scn-BA | [2] | 92 | 24.1 | 822.5151 | 822.5140 | 8-hpdde-BHA * | |
| 19 | 6.3 | 674.3173 | 674.3172 | 8-scn-BHA * | 93 | 24.2 | 834.5151 | 834.5136 | 8-linolen-BHA | [6] | |
| 20 | 6.6 | 732.3590 | 732.3601 | 8-adp-BA | [2] | 94 | 24.2 | 844.5206 | 844.5218 | 8-pal-10-OH-BMA | [6] |
| 21 | 6.7 | 702.3484 | 702.3473 | 8-adp-BHA * | 95 | 24.3 | 806.5202 | 806.5190 | 8-lino-DMBHA * | ||
| 22 | 6.8 | 674.3535 | 674.3499 | 8-hbte-DBA * | 96 | 24.3 | 784.4994 | 784.4973 | 8-myr-BHA | [19] | |
| 23 | 6.9 | 686.3171 | 686.3170 | 8-fmr-DBA * | 97 | 24.4 | 820.5358 | 820.5348 | 8-lino-13-DMDBA * | ||
| 24 | 7.2 | 688.3691 | 688.3682 | 8-hvlr-DBA * | 98 | 24.5 | 840.5256 | 840.5249 | 8-hpde-BMA | [8] | |
| 25 | 7.6 | 686.3171 | 686.3173 | 8-gtn-BHA * | 99 | 24.6 | 814.5100 | 814.5087 | 8-ptdn-BMA | [6] | |
| 26 | 7.7 | 716.3641 | 716.3633 | 8-adp-DBA * | 100 | 24.6 | 796.4994 | 796.4960 | 8-ptde-BHA | [8] | |
| 27 | 8.2 | 718.3433 | 718.3443 | 8-gtr-BA | [2] | 101 | 24.7 | 854.5413 | 854.5778 | 8-hstr-DMA * | |
| 28 | 8.7 | 658.3222 | 658.3244 | 8-bte-BMA * | 102 | 24.8 | 852.5256 | 852.5254 | 8-lino-BMA | [6] | |
| 29 | 9.8 | 760.3903 | 760.3903 | 8-azl-BMA * | 103 | 24.9 | 820.5358 | 820.5359 | 8-lino-3-DMDBA * | ||
| 30 | 10.3 | 744.4059 | 774.4051 | 8-sbc-BMA * | 104 | 25.0 | 832.5358 | 832.5355 | 8-linolen-3,13-DDBA * | ||
| 31 | 10.8 | 644.3429 | 644.3423 | 8-btn-BHA * | 105 | 25.0 | 802.5252 | 802.5237 | 8-linolen-DMDDBA * | ||
| 32 | 10.8 | 674.3535 | 674.3529 | 8-btn-BA * | 106 | 25.1 | 836.5307 | 836.5293 | 8-lino-BHA | [6] | |
| 33 | 10.8 | 674.3535 | 674.3529 | 8-vlr-BMA* | 107 | 25.1 | 810.5151 | 810.5145 | 8-pme-BHA | [8] | |
| 34 | 10.9 | 744.3954 | 744.3960 | 8-hnne-BMA * | 108 | 25.3 | 866.5413 | 866.5408 | 8-lino-BA | [20] | |
| 35 | 11.4 | 902.5266 | 902.5268 | 8-thode-BMA * | 109 | 25.3 | 870.5362 | 870.5372 | 8-ole-10-OH-BMA | [6] | |
| 36 | 11.5 | 758.4110 | 758.4119 | 8-azl-DBA * | 110 | 25.5 | 808.5358 | 808.5306 | 8-ole-DMBHA * | ||
| 37 | 11.6 | 758.4110 | 758.4107 | 8-hnne-BA * | 111 | 25.5 | 828.5256 | 828.5264 | 8-ptdn-BA | [6] | |
| 38 | 11.8 | 688.3691 | 688.3678 | 8-vlr-BA | [4] | 112 | 25.7 | 858.5362 | 858.5352 | 8-pal-10-OH-BA | [6] |
| 39 | 12.0 | 658.3586 | 658.3558 | 8-btn-DBA | [3] | 113 | 25.9 | 798.5151 | 798.5128 | 8-ptdn-BHA | [6] |
| 40 | 12.0 | 702.312 | 702.3845 | 8-hxn-BA | [4] | 114 | 25.9 | 824.5307 | 824.5305 | 8-pme-DBA * | |
| 41 | 12.0 | 658.3586 | 658.3599 | 8-vlr-BHA * | 115 | 26.0 | 822.5515 | 822.5527 | 8-ole-13-DMDBA * | ||
| 42 | 12.2 | 916.5417 | 916.5399 | 8-thode-BA * | 116 | 26.0 | 796.5358 | 796.5371 | 8-pal-DMDBA * | ||
| 43 | 12.3 | 886.5311 | 886.5662 | 8-thode-BHA * | 117 | 26.1 | 822.5515 | 822.5511 | 8-ole-3-DMDBA * | ||
| 44 | 12.5 | 728.4004 | 728.3993 | 8-hnne-BHA * | 118 | 26.1 | 828.5256 | 828.5251 | 8-pal-BMA | [20] | |
| 45 | 13.0 | 900.5468 | 900.5474 | 8-thode-DBA * | 119 | 26.1 | 796.5358 | 796.5368 | 8-pal-13-DMDBA * | ||
| 46 | 13.2 | 790.4372 | 790.4367 | 8-dhudn-BMA | 120 | 26.2 | 804.5409 | 804.5387 | 8-lino-DMDDBA * | ||
| 47 | 13.3 | 672.3742 | 672.3752 | 8-vlr-DBA * | 121 | 26.2 | 884.5519 | 884.5494 | 8-ole-10-OH-BA | [6] | |
| 48 | 13.3 | 916.5417 | 916.5401 | 8-thnde-BMA * | 122 | 26.3 | 854.5413 | 854.5403 | 8-ole-BMA | [6] | |
| 49 | 13.5 | 800.4216 | 800.4234 | 8-dded-BMA * | 123 | 26.6 | 850.5464 | 850.5459 | 8-lino-DBA | [8] | |
| 50 | 13.6 | 774.4423 | 774.4400 | 8-dhudn-BHA | 124 | 26.8 | 880.5569 | 880.5540 | 8-ecde-BMA | [12] | |
| 51 | 13.6 | 782.5202 | 782.5161 | 8-pal-DMBHA * | 125 | 26.9 | 834.5515 | 834.5494 | 8-lino-3,13-DDBA | [18] | |
| 52 | 15.1 | 716.4004 | 716.3982 | 8-otn-BMA * | 126 | 26.9 | 842.5413 | 842.5396 | 8-pal-BA | [20] | |
| 53 | 15.2 | 884.5155 | 884.5174 | 8-dhodd-BMA * | 127 | 26.9 | 812.5307 | 812.5292 | 8-ptdn-DBA | [8] | |
| 54 | 15.9 | 730.4161 | 730.4154 | 8-nnn-BMA * | 128 | 27.0 | 894.5362 | 894.5336 | 3-Acetyl-8-lino-BMA | [16] | |
| 55 | 16.2 | 898.5311 | 898.5307 | 8-dhodd-BA * | 129 | 27.1 | 868.5569 | 868.5576 | 8-ole-BA | [20] | |
| 56 | 16.3 | 886.5311 | 886.5318 | 8-dhode-BMA* | 130 | 27.2 | 838.5464 | 838.5443 | 8-ole-BHA | [6] | |
| 57 | 16.8 | 900.5468 | 900.5438 | 8-dhode-BA * | 131 | 27.4 | 806.5565 | 806.5512 | 8-ole-DMDDBA * | ||
| 58 | 17.0 | 866.5049 | 866.5054 | 8-hodt-BMA * | 132 | 27.5 | 812.5307 | 812.5300 | 8-pal-BHA | [6] | |
| 59 | 17.1 | 868.5206 | 868.5191 | 8-dhodd-BHA * | 133 | 27.6 | 856.5569 | 856.5553 | 8-str-BMA | [10] | |
| 60 | 17.3 | 888.5468 | 888.5438 | 8-dhstr-BMA * | 134 | 27.7 | 836.5619 | 836.5712 | 8-ole-3,13-DDBA * | ||
| 61 | 17.4 | 870.5362 | 870.5335 | 8-dhode-BHA * | 135 | 28.0 | 826.5464 | 826.5465 | 8-pal-DBA | [6] | |
| 62 | 18.1 | 884.5313 | 884.5292 | 8-hodd-10-OH-BMA * | 136 | 28.2 | 926.6352 | 926.6308 | 8-tcn-BMA * | ||
| 63 | 18.1 | 884.5519 | 884.5486 | 8-dhode-DBA * | 137 | 28.3 | 780.5409 | 780.5384 | 8-pal-DMDDBA * | ||
| 64 | 18.4 | 880.5206 | 880.5199 | 8-hodt-BA * | 138 | 28.4 | 842.5413 | 842.5399 | 8-hpdn-BMA | [6] | |
| 65 | 18.4 | 850.500 | 850.5096 | 8-hodt-BHA * | 139 | 28.5 | 852.5620 | 852.5620 | 8-ole-DBA | [6] | |
| 66 | 18.5 | 872.5519 | 872.5518 | 8-dhstr-BHA * | 140 | 28.9 | 912.6195 | 912.6194 | 8-dcn-BMA * | ||
| 67 | 18.8 | 868.5206 | 868.5196 | 8-hodd-BMA * | 141 | 29.0 | 810.5515 | 810.5490 | 8-pal-3,13-DDBA | [8] | |
| 68 | 18.8 | 910.5311 | 910.5304 | 8-dhecte-BMA * | 142 | 29.7 | 870.5726 | 870.5569 | 8-str-BA | [20] | |
| 69 | 19.4 | 882.5368 | 882.5349 | 8-hodd-BA * | 143 | 29.7 | 954.6665 | 954.6648 | 8-ttcn-BA | [5] | |
| 70 | 19.8 | 924.5548 | 924.5455 | 8-dhhctte-BA * | 144 | 30.5 | 840.5620 | 840.5583 | 8-hpdn-DBA | [8] | |
| 71 | 20.1 | 852.5256 | 852.5264 | 8-hodd-BHA * | 145 | 30.5 | 840.5620 | 840.5636 | 8-str-BHA | [8] | |
| 72 | 20.1 | 870.5362 | 870.5359 | 8-hode-BMA * | 146 | 30.5 | 940.6508 | 940.6506 | 8-ttcn-BMA | [21] | |
| 73 | 21.0 | 864.5256 | 864.5255 | 8-hodt-DBA * | 147 | 30.9 | 926.6279 | 926.6308 | 8-dcn-BA | [8] | |
| 74 | 21.2 | 866.5413 | 866.5410 | 8-hodd-DBA * | 148 | 31.6 | 896.6246 | 896.6225 | 8-dcn-BHA | [21] |
| No. | MF | Possible Structure | No. of LAs | Ref. | No. | MF | Possible Structure | No. of LAs | Ref. |
|---|---|---|---|---|---|---|---|---|---|
| 1 | C3H6O2 | Propanoic acid | ND | [1] | 40 | C17H30O2 * | Heptadecadienoic acid | 2 | |
| 2 | C4H4O4 | Fumaric acid | 2 | [11] | 41 | C17H32O2 | Heptadecenoic acid | 1 | [1] |
| 3 | C4H6O2 * | Butenoic acid | 1 | 42 | C17H34O2 | Heptadecanoic acid | 2 | [1] | |
| 4 | C4H6O3 * | Hydroxybutenoic acid | 1 | 43 | C18H30O2 | Linolenic acid | 9 | [1] | |
| 5 | C4H6O4 | Succinic acid | 2 | [1,2] | 44 | C18H30O3 * | Hydroxyoctadecatrienoic acid | 4 | |
| 6 | C4H6O5 | Malic acid | ND | 45 | C18H32O2 | Linoleic acid | 12 | [1] | |
| 7 | C4H8O2 | Butanoic acid | 3 | [11] | 46 | C18H32O3 * | Hydroxyoctadecadienoic acid | 5 | |
| 8 | C4H8O3 | Hydroxybutanoic acid | 3 | [11] | 47 | C18H32O4 * | Dihydroxyoctadecadienoic acid | 3 | |
| 9 | C4H8O4 * | Dihydroxybutanoic acid | 1 | 48 | C18H34O2 | Oleic acid | 11 | [1] | |
| 10 | C5H6O4 | Glutaconic acid | 2 | [11] | 49 | C18H34O3 * | Hydroxyoctadecenoic acid | 4 | |
| 11 | C5H6O5 * | Oxoglutaric acid | 1 | 50 | C18H34O4 * | Dihydroxyoctadecenoic acid | 4 | ||
| 12 | C5H8O4 | Glutaric acid | 1 | [2] | 51 | C18H34O5 * | Trihydroxyoctadecenoic acid | 4 | |
| 13 | C5H10O2 | Valeric acid | 4 | 52 | C18H36O2 | Stearic acid | 3 | [1] | |
| 14 | C5H10O3 * | Hydroxyvaleric acid | 1 | 53 | C18H36O3 * | Hydroxystearic acid | 1 | ||
| 15 | C6H6O6 * | Aconitic acid | 5 | 54 | C18H36O4 * | Dihydroxy stearic acid | 2 | ||
| 16 | C6H10O4 | Adipic acid | 3 | [2,11] | 55 | C19H32O2 | Nonadecatrienoic acid | ND | [8] |
| 17 | C6H10O5 * | Hydroxyadipic acid | 5 | 56 | C19H34O2 | Nonadecadienoic acid | ND | [1] | |
| 18 | C6H12O2 | Hexanoic acid | 1 | 57 | C19H36O2 | Nonadecenoic acid | ND | [1] | |
| 19 | C7H8O4 * | Heptadienedioic acid | 1 | 58 | C19H36O5 * | Trihydroxynonadecenoic acid | 1 | ||
| 20 | C7H8O6 * | Dioxoheptanedioic acid | 1 | 59 | C19H38O2 | Nonadecanoic acid | ND | [1] | |
| 21 | C7H12O4 | Pimelic acid | ND | [2] | 60 | C20H30O2 | Eicosapentaenoic acid | ND | [1] |
| 22 | C8H14O4 | Suberic acid | ND | [2] | 61 | C20H32O2 | Eicosatetraenoic acid | ND | [1] |
| 23 | C8H16O2 * | Octanoic acid | 1 | 62 | C20H34O2 | Eicosatrienoic acid | ND | [1] | |
| 24 | C9H16O3 * | Hydroxynonenoic acid | 3 | 63 | C20H34O4 * | Dihydroxyeicosatrienoic acid | 1 | ||
| 25 | C9H16O4 | Azelaic acid | 2 | [12] | 64 | C20H36O2 | Eicosadienoic acid | 1 | [1] |
| 26 | C9H18O2 * | Nonanoic acid | 1 | 65 | C20H38O2 | Eicosenoic acid | ND | [1] | |
| 27 | C10H18O4 | Sebacic acid | 1 | [2] | 66 | C20H40O2 | Eicosanoic acid | ND | [1] |
| 28 | C11H20O4 | Undecanedioic acid | ND | [2] | 67 | C21H32O2 | Henicosapentaenoic acid | ND | [1] |
| 29 | C11H22O4 * | Dihydroxyundecanoic acid | 2 | 68 | C21H34O2 | Henicosatetraenoic acid | ND | [1] | |
| 30 | C12H20O4 * | Dodecenedioic acid | 1 | 69 | C21H34O4 * | Dihydroxyhenicosatetrenoic acid | 1 | ||
| 31 | C12H24O2 | Lauric acid | 1 | [1] | 70 | C22H32O2 | Docosahexaenoic acid | ND | [1] |
| 32 | C14H24O2 | Tetradecadienoic acid | ND | [1] | 71 | C22H44O2 | Docosanoic acid | 3 | [1] |
| 33 | C14H26O2 | Tetradecenoic acid | ND | [1] | 72 | C23H42O2 | Tricosadienoic Acid | ND | [1] |
| 34 | C14H28O2 | Myristic acid | 1 | [1] | 73 | C23H44O2 | Tricosenoic Acid | ND | [1] |
| 35 | C15H28O2 | Pentadecenoic acid | 2 | [1] | 74 | C23H46O2 * | Tricosanoic Acid | 1 | |
| 36 | C15H30O2 | Pentadecanoic acid | 4 | [1] | 75 | C24H46O2 | Tetracosenoic acid | ND | [1] |
| 37 | C16H28O2 | Palmitadienoic acid | 1 | [1] | 76 | C24H48O2 | Tetrecosanoic acid | 2 | [1] |
| 38 | C16H30O2 | Palmitoleic acid | 4 | [1] | 77 | C25H50O2 | Pentacosanoic acid | ND | [1] |
| 39 | C16H32O2 | Palmitic acid | 11 | [1] |
© 2016 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons by Attribution (CC-BY) license ( http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Liang, Y.; Wu, J.-L.; Leung, E.L.-H.; Zhou, H.; Liu, Z.; Yan, G.; Liu, Y.; Liu, L.; Li, N. Identification of Oxygenated Fatty Acid as a Side Chain of Lipo-Alkaloids in Aconitum carmichaelii by UHPLC-Q-TOF-MS and a Database. Molecules 2016, 21, 437. https://doi.org/10.3390/molecules21040437
Liang Y, Wu J-L, Leung EL-H, Zhou H, Liu Z, Yan G, Liu Y, Liu L, Li N. Identification of Oxygenated Fatty Acid as a Side Chain of Lipo-Alkaloids in Aconitum carmichaelii by UHPLC-Q-TOF-MS and a Database. Molecules. 2016; 21(4):437. https://doi.org/10.3390/molecules21040437
Chicago/Turabian StyleLiang, Ying, Jian-Lin Wu, Elaine Lai-Han Leung, Hua Zhou, Zhongqiu Liu, Guanyu Yan, Ying Liu, Liang Liu, and Na Li. 2016. "Identification of Oxygenated Fatty Acid as a Side Chain of Lipo-Alkaloids in Aconitum carmichaelii by UHPLC-Q-TOF-MS and a Database" Molecules 21, no. 4: 437. https://doi.org/10.3390/molecules21040437
APA StyleLiang, Y., Wu, J.-L., Leung, E. L.-H., Zhou, H., Liu, Z., Yan, G., Liu, Y., Liu, L., & Li, N. (2016). Identification of Oxygenated Fatty Acid as a Side Chain of Lipo-Alkaloids in Aconitum carmichaelii by UHPLC-Q-TOF-MS and a Database. Molecules, 21(4), 437. https://doi.org/10.3390/molecules21040437







