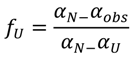2.1. Circular Dichroism of Native Baupain and Secondary Structure Estimation
Circular dichroism and fluorescence spectroscopy of proteins are widely used to monitor conformational changes of proteins with changes in solvent composition. The CD spectrum of baupain in 10 mM PBA solution at pH 7.0 (
Figure 1A, squares) showed two minima, one at 222 nm, related to the strong hydrogen-bonding environment of α-helices, and another at 210 nm. This is similar to the papain spectrum that shows two bands at 222 and 208 nm, characteristic of an α-helix structure [
10]. Analysis of secondary structure content using the CDPro software package with the SELCON3, CONTINLL, CDSSTR programs yielded 44% α-helix, 16% β-sheet, 12% β-turn, and 28% unordered structures, with a standard deviation of 2% [
1]. Cluster analysis showed that baupain belongs to the α+β tertiary structure class, like other cysteine proteases, for example procerain and bromelain [
8,
11]. This classification was corroborated by the results of CDPro estimation (44% α-helix and 16% β-sheet) and spectrum sharpness (with the band at 210 nm slightly more intense than that at 222 nm), which is similar to the papain spectrum that shows two bands (208 and 222 nm) in neutral pH [
12]. Together with biochemical data obtained previously [
1], the current results allow us to classify baupain in the clan CA of the papain family.
2.2. Thermal Denaturation
CD spectroscopy in the far-UV region can monitor conformational changes in the polypeptide backbone. Thermal assays were carried out in the 25 °C to 90 °C range in 5 °C intervals. The spectra in
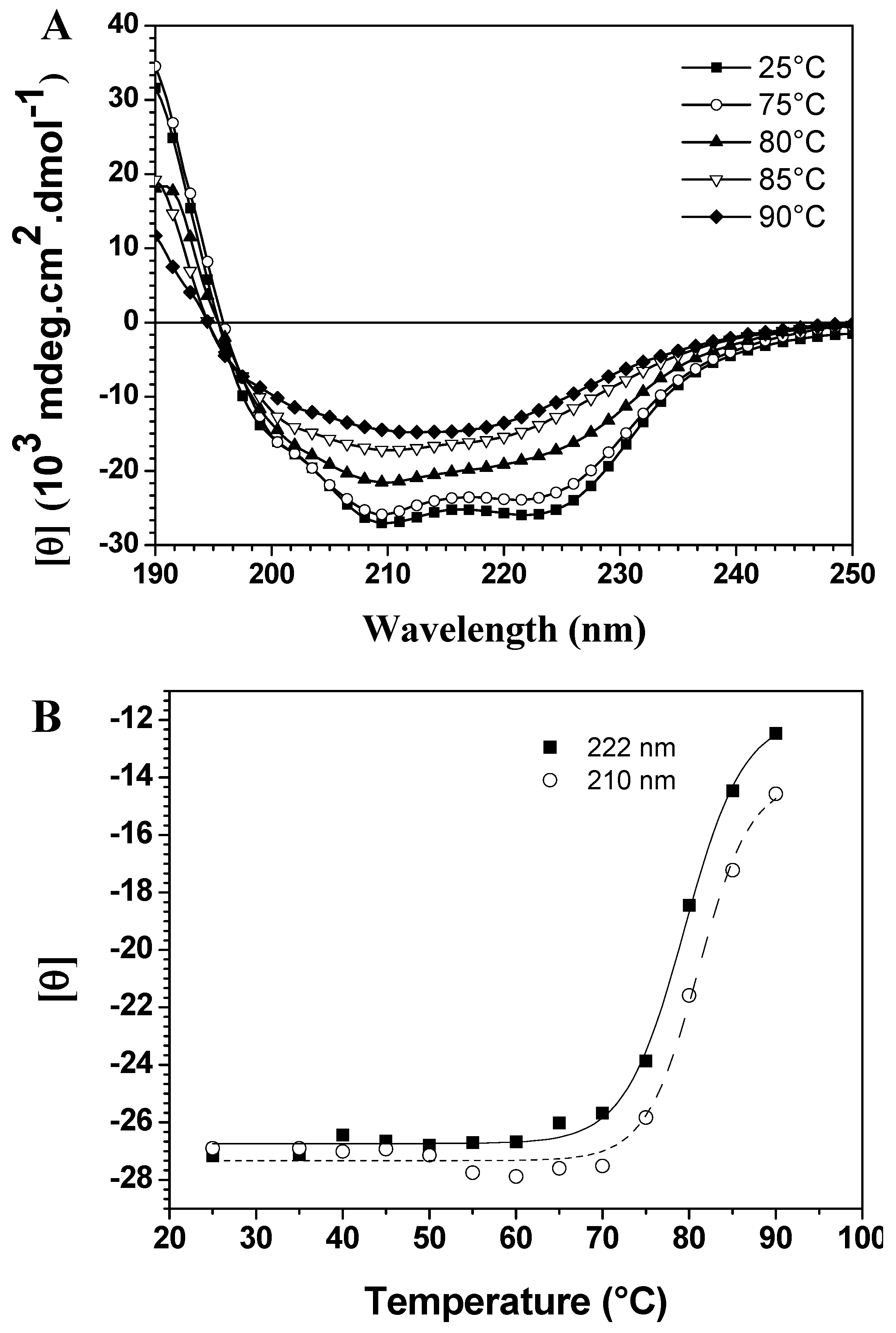
Figure 1A show thermal scans at different temperatures, demonstrating significant differences in thermal stability due to the loss of two typical bands with increased temperature. On the other hand, temperature-induced unfolding of baupain was incomplete since the spectrum at 90 °C still displays a band centered at 215 nm, characteristic of a β-sheet structure, although less intense. To better analyze these conformational changes, the mean residual ellipticities [θ] at 210 nm and 222 nm (MRE
210 and MRE
222) were plotted against temperature, as shown in
Figure 1B. In this case, baupain reveals pronounced structural stability until 70 °C, with only slight fluctuations in the MRE values. A sigmoidal fit to these points yielded a midpoint around 80 °C. Corroborating the spectral changes, a significant variation in secondary structure content occurs in the transition temperature from 80 to 85 °C. Namely, at 80 baupain showed 41% α-helix, 16% β-sheet, 19% β-turn and 26% unordered structure content, whereas at 85 °C the α-helix content decreased to 24% and the β-sheet and unordered structure contents increased to 24% and 33%, respectively. These last values were maintained at 90 °C. In addition, the data show high protein structural stability, possibly assigned to β-sheet structures since they are more resistant and less susceptible to unfolding [
4]. The transition was irreversible, since a refolding attempt by decreasing the temperature under the same unfolding conditions, did not lead to recovery of the original structural characteristics (data not shown).
Figure 1.
Temperature dependent conformational changes of baupain in 10 mM phosphate/borate/acetate (PBA), pH 7.0. (A) Far UV CD spectra of baupain (0.23 μM) at different temperatures. (B) Relative changes of mean residual ellipticity values (MRE) at 210 nm (squares) and 222 nm (circles) as a function of temperature. The sigmoidal fit of these points provides a midpoint around 80 °C for this transition.
Figure 1.
Temperature dependent conformational changes of baupain in 10 mM phosphate/borate/acetate (PBA), pH 7.0. (A) Far UV CD spectra of baupain (0.23 μM) at different temperatures. (B) Relative changes of mean residual ellipticity values (MRE) at 210 nm (squares) and 222 nm (circles) as a function of temperature. The sigmoidal fit of these points provides a midpoint around 80 °C for this transition.
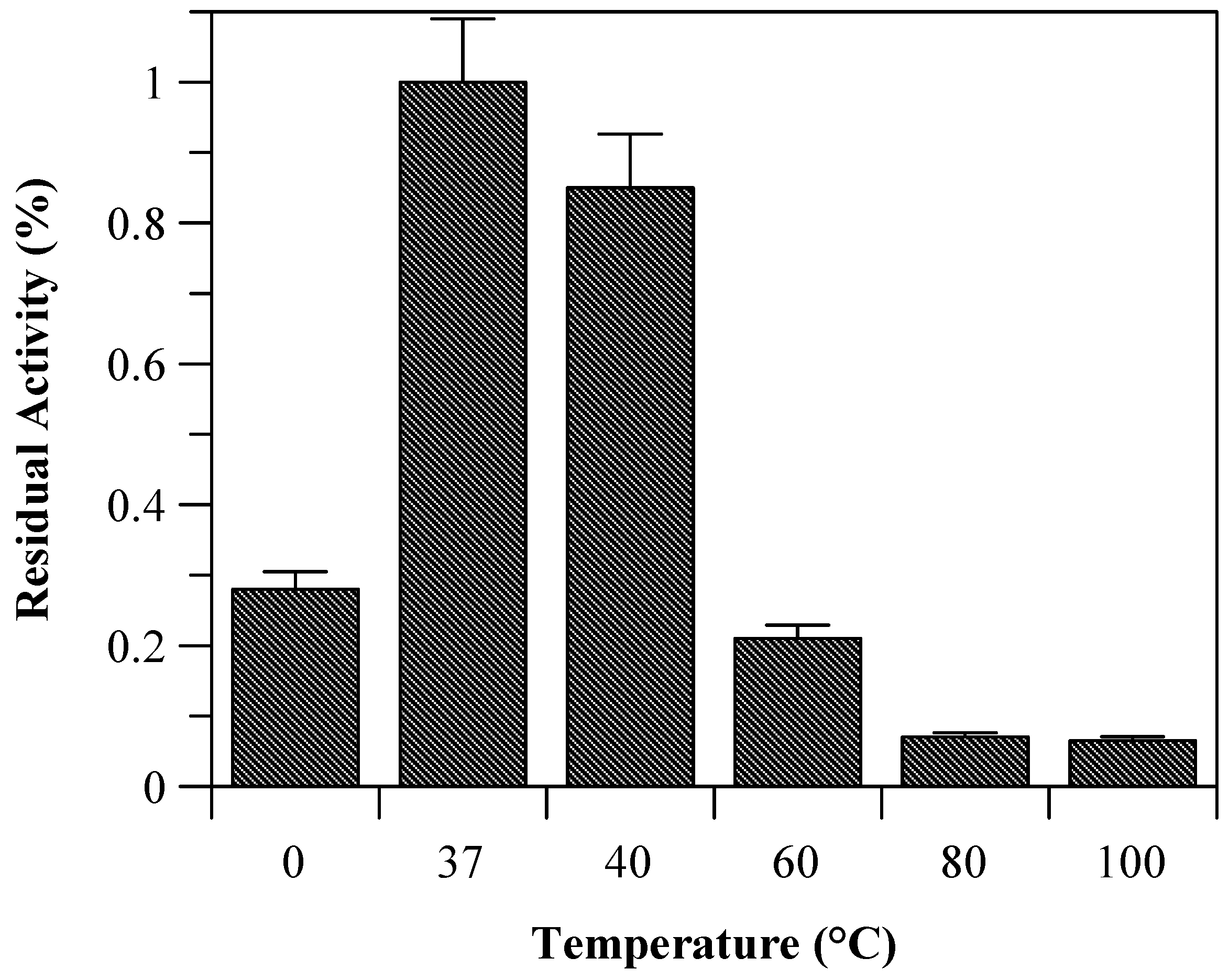
Due to the incomplete thermal unfolding, baupain activity was investigated with regard to its hydrolytic activity on the Z-Phe-Arg-MCA substrate (
Figure 2). Baupain maximum activity was achieved at 37 and 40 °C, and very low activity was observed at extreme temperatures (60, 80, and 100 °C), similar to the behavior observed with papain-like cysteine proteinases [
10]. The data show that although the secondary structure was maintained until 70 °C (
Figure 1), baupain was unable to cleave the substrate at this temperature. These results suggest that changes in tertiary structure led to a modification of the enzyme active site, thus affecting baupain function.
Figure 2.
Residual activity of baupain (0.050 mg/mL) towards Z-Phe-Arg-MCA (0.4 mM) determined at different temperature.
Figure 2.
Residual activity of baupain (0.050 mg/mL) towards Z-Phe-Arg-MCA (0.4 mM) determined at different temperature.
2.3. Guanidine Hydrochloride Unfolding
Chaotropic agents such as GdnHCl or urea are commonly used in unfolding/refolding studies, although the unfolding mechanism is not clear. These studies were followed by far-CD, fluorescence and ANS binding assays.
Intrinsic fluorescence emission provides a sensitive probe to characterize proteins by monitoring tryptophan residues that are very sensitive to the polarity of the environment. The wavelength at the emission maximum (λ
max) and the fluorescence intensity can be used to characterize proteins and to determine conformational changes [
13]. The effects of GdnHCl on fluorescence emission spectra are shown in
Figure 3A. The intrinsic fluorescence emission of baupain at pH 7.0 displayed an emission maximum (λ
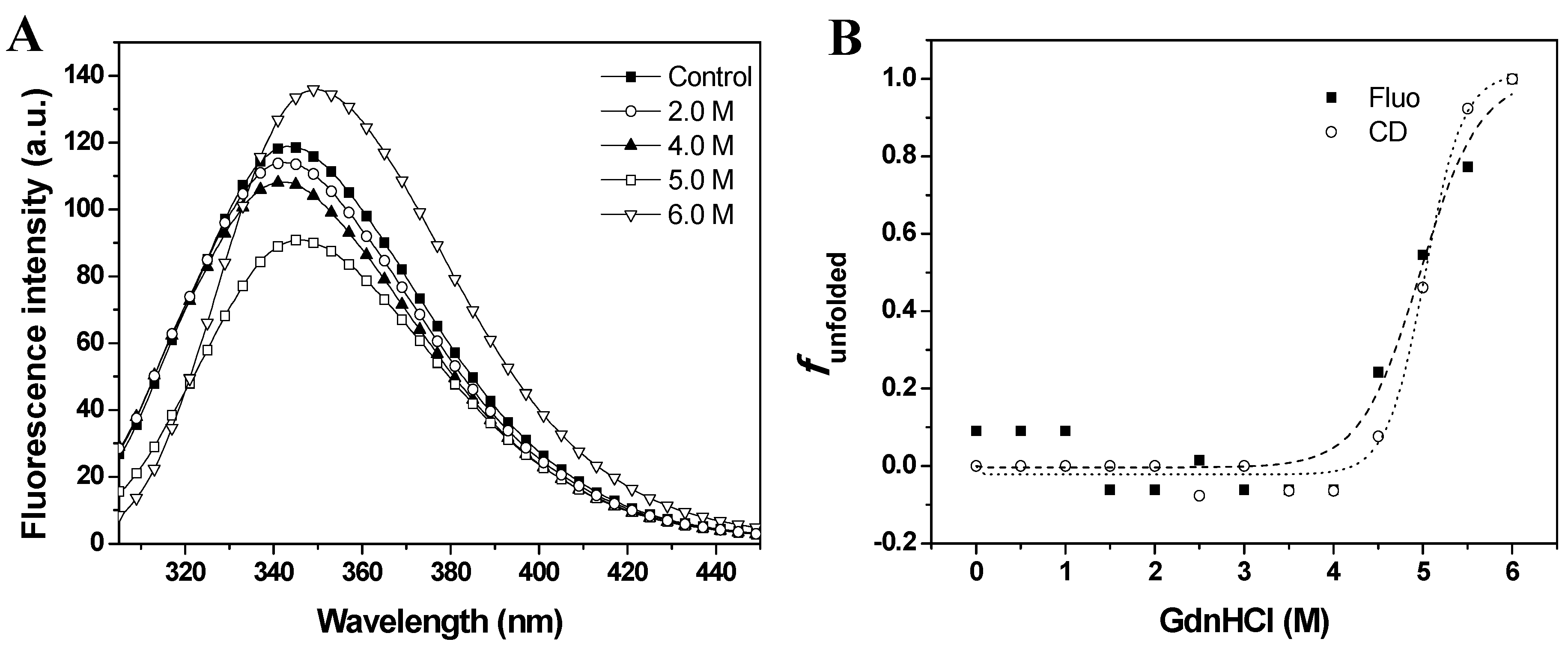
max) at 343 nm, suggesting that tryptophan residues were at least partially exposed to solvent. The fluorescence emission spectra of baupain showed no significant change in intensity or in the emission maxima in 0–3.5 M GdnHCl. However, the fluorescence intensity increased 13% in 6.0 M GdnHCl, along with a red-shift of 7 nm in emission maximum (from 343 to 350 nm), indicating exposure of tryptophan to a more polar environment.
Gradual loss of secondary structure of the enzyme is demonstrated by a marked decrease in the typical CD band at 222 nm (from 24,582 deg·cm
2·dmol
−1 to 4,555 deg·cm
2·dmol
−1) in 6.0 M GdnHCl (data not shown). To better examine the conformational changes induced by GdnHCl on baupain at pH 7.0, the unfolded fractions were calculated by Equation (1) (see Experimental,
Section 3.6) and plotted against GdnHCl concentration values, using as signals the mean residue ellipticity intensities at 222 nm (α
obs, circles) and fluorescence emission maxima (λ
max, squares) (
Figure 3B). The sigmoidal curves suggest the presence of cooperative conformational changes induced by GdnHCl unfolding in a single step from native to unfolded form (N→U), with no detectable intermediates. From the sigmoidal fits we determined the midpoints 4.7 ± 0.2 M (from CD data) and 5.0 ± 0.2 M GdnHCl (from fluorescence data). In order to analyze global conformation changes induced by GdnHCl, the exposure of the hydrophobic surfaces of baupain was monitored by changes in the fluorescence of the ANS dye. This probe has low fluorescence intensity in the presence of a hydrophilic environment, whereas ANS binding to hydrophobic regions leads to a blue-shift of the emission maximum and a pronounced increased in fluorescence intensity. ANS (23 µM) was added to baupain (0.23 µM) in the absence and presence of different concentrations of GdnHCl at pH 7.0
. We noted no significant increase in ANS fluorescence that would indicate the presence of hydrated hydrophobic surfaces for ANS binding (data not shown), as also described by Khurana
et al. [
14].
Figure 3.
GdnHCl-induced unfolding of baupain in 10 mM phosphate/borate/acetate (PBA), pH 7.0, 25 °C, as a function of GdnHCl concentration. (A) Intrinsic fluorescence emission spectra were measured in the absence and in the presence of chaotropic agent. (B) Normalized transition curves for GdnHCl-induced transition. The unfolded fractions were calculated with equation 1 using the fluorescence emission maxima (λmax, squares) and mean residual ellipticity values at 222 nm (circles). The sigmoidal fits for CD (·····) and fluorescence data (----) defined the midpoints at 4.7 ± 0.2 M and 5.0 ± 0.2 M GdnHCl.
Figure 3.
GdnHCl-induced unfolding of baupain in 10 mM phosphate/borate/acetate (PBA), pH 7.0, 25 °C, as a function of GdnHCl concentration. (A) Intrinsic fluorescence emission spectra were measured in the absence and in the presence of chaotropic agent. (B) Normalized transition curves for GdnHCl-induced transition. The unfolded fractions were calculated with equation 1 using the fluorescence emission maxima (λmax, squares) and mean residual ellipticity values at 222 nm (circles). The sigmoidal fits for CD (·····) and fluorescence data (----) defined the midpoints at 4.7 ± 0.2 M and 5.0 ± 0.2 M GdnHCl.
2.4. Acid-Induced Unfolding
An important factor in protein conformational stability is the pH of the environment, which can change the net charge of biomolecules, and thus many proteins denature at extremes pH due to the presence of repulsive forces [
14,
15]. However, the acid denaturation of proteins often results in unfolded states with less denaturation than that obtained by high concentrations of chaotropic agents such as GdnHCl or urea [
8,
9,
10]. This incomplete denaturation may be due to electrostatic repulsion forces that are insufficient to overtake the interactions that favor folding, such as hydrophobic interactions, salt bridges and disulfide bonds. Baupain conformational changes induced by pH variation were monitored by far-CD, fluorescence and ANS binding studies. The effect of pH on the secondary structure of the protein is shown in
Figure 4A (CD spectra of baupain as a function of pH) and 4B (mean residue ellipticity, MRE,
versus pH). The MRE
210 and MRE
222 values for baupain remained unchanged in the pH range 4.0–10.5 (
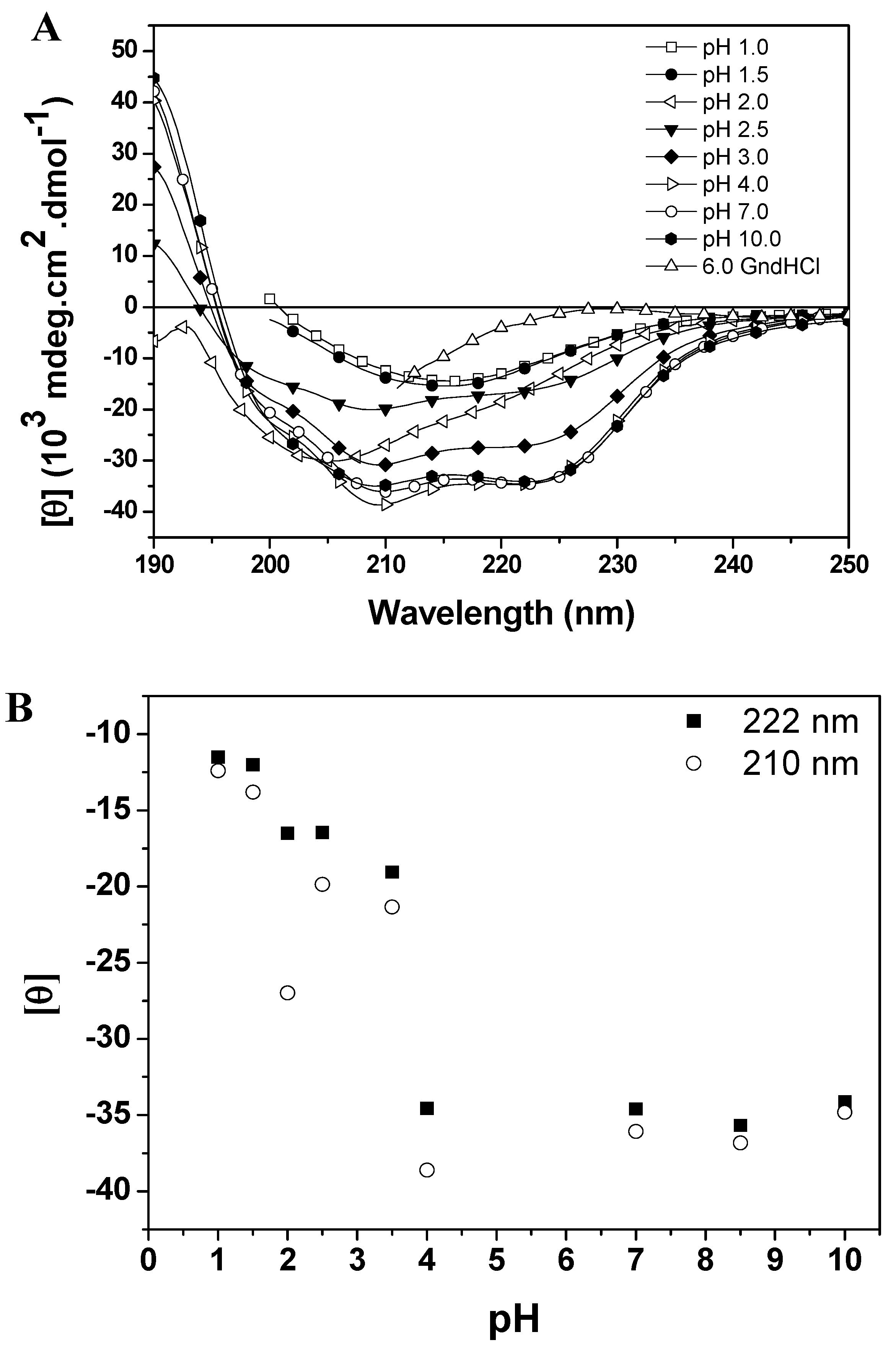
Figure 4B), with the protein retaining most of its α-helices and β structures (44% and 16% at pH 7.0, and 40% and 15% at pH 4.0). For pH values below 3.5 there was a gradual decrease of the characteristic bands of CD (at 210 and 222 nm) with the emergence of a low intensity band at 215 nm at pHs 1.0 and 1.5 (
Figure 4A). On the other hand, at pH 2.0 the protein retained a significant amount of secondary structure as shown by the value of MRE
210 (28,000 deg·cm
2·dmol
−1). Finally, below the pH 2 most of the secondary structure was lost, with a pronounced decrease of MRE values and a partial unfolding of the enzyme. These results have been observed for other cysteine proteases and may suggest the presence of an intermediate state (molten globule) at pH 2.0. In fact, a weak cooperative process is observed in
Figure 4B, suggesting an intermediate state. Baupain also maintains its secondary structure in a wide pH range, which is a peculiar feature also characteristic of other papain-like enzymes [
9,
11]. Thus, the pH-secondary conformational stability of baupain was further studied by evaluating the effect of pH on tertiary structure changes.
Figure 4.
pH dependent conformational changes of baupain at 25 °C. (A) Far UV CD spectra of baupain (0.23 μM) under different pHs. (B) Relative changes of mean residual ellipticity values (MRE) at 210 nm (circles) and 222 nm (squares) as a function of pH. For comparison, the CD spectrum of baupain in 6.0 M GdnHCl is also shown.
Figure 4.
pH dependent conformational changes of baupain at 25 °C. (A) Far UV CD spectra of baupain (0.23 μM) under different pHs. (B) Relative changes of mean residual ellipticity values (MRE) at 210 nm (circles) and 222 nm (squares) as a function of pH. For comparison, the CD spectrum of baupain in 6.0 M GdnHCl is also shown.
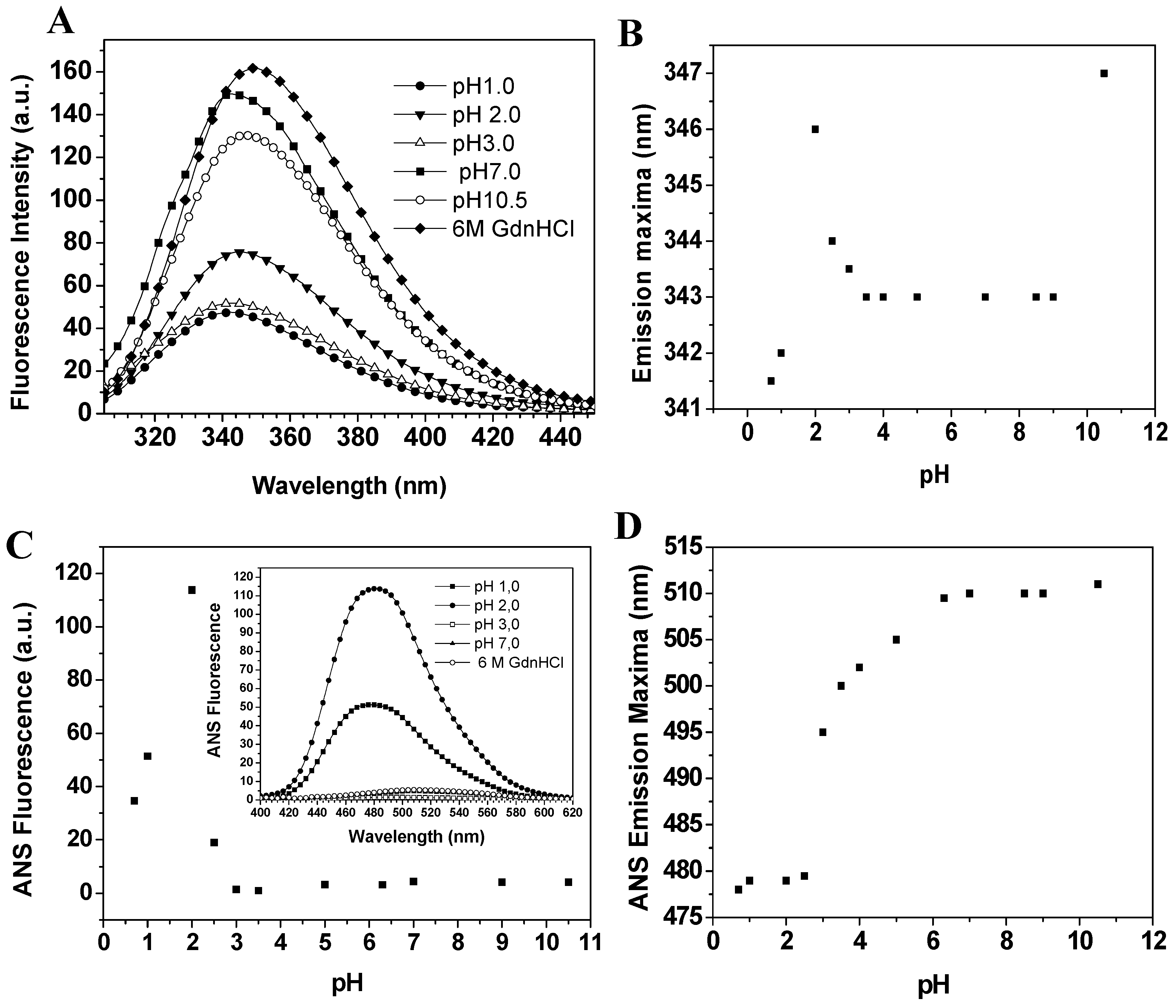
Baupain fluorescence emission shows maximum intensity at pH 7.0, 50% intensity at pH 2.0, and around 67% at pHs 1.0 and 3.0. At basic pH, the fluorescence intensity decreases to 13% (
Figure 5A). It is worth pointing out that the emission maximum (λ
max) was unchanged in the pH range 3.0–10.0 (
Figure 5B). However, at pH 2.0 a slight red shift from 343 to 346 nm was observed for the emission maximum, suggesting that tryptophan residues are exposed to a slightly more hydrophobic environment, which would be compatible with a more unordered structure. Furthermore, the CD spectra still show a significant content of secondary structure at pH 2.0. The effect of pH in the tertiary structure of baupain was also monitored using ANS dye.
Figure 5.
Effect of pH on the fluorescence of baupain (0.23 μM) at 25 °C. (A) Intrinsic fluorescence emission spectra and (B) emission maxima (λmax,) plotted against pH values. (C) ANS fluorescence intensity as a function of pH and ANS fluorescence emission spectra of baupain at pH 2.0, pH 7.0 and GdnHCL 6.0 M (insert). (D) pH dependence of ANS emission maxima (λmax) for baupain.
Figure 5.
Effect of pH on the fluorescence of baupain (0.23 μM) at 25 °C. (A) Intrinsic fluorescence emission spectra and (B) emission maxima (λmax,) plotted against pH values. (C) ANS fluorescence intensity as a function of pH and ANS fluorescence emission spectra of baupain at pH 2.0, pH 7.0 and GdnHCL 6.0 M (insert). (D) pH dependence of ANS emission maxima (λmax) for baupain.
ANS fluorescence intensity changed with pH, reaching a maximum at pH 2.0 (28-fold increase,
Figure 5C). Below pH 2.0, ANS fluorescence intensity was twelve times larger than at pH 7.0, suggesting the structural reorganization of the enzyme to protect its hydrophobic regions from the solvent. The emission maximum shifted to shorter wavelength (478 nm) at pH 2.0, in comparison to the spectrum at pH 7.0 (510 nm,
Figure 5D). This blue shift in emission maximum together with the large increase of fluorescence intensity indicates an extensive solvent exposure of non-polar clusters. Taken together, the CD spectra (
Figure 4A), intrinsic fluorescence spectra (
Figure 5A) and ANS fluorescence spectra (
Figure 5C, insert) at pHs 2.0 and 7.0 and in 6.0 M GdnHCl reveal that the state of baupain molecules under acidic conditions is different from that under neutral pH or relative to the unfolded state.