1. IMGT®: The Birth of Immunoinformatics
IMGT
®, the international ImMunoGeneTics information system
® [
1,
2], was created in 1989 by Marie-Paule Lefranc at Montpellier, France (CNRS and Montpellier University). The founding of IMGT
® marked the advent of immunoinformatics, a new science, which emerged at the interface between immunogenetics and bioinformatics. For the first time, immunoglobulin (IG) or antibody and T cell receptor (TR) variable (V), diversity (D), joining (J) and constant (C) genes were officially recognized as “genes” as well as the conventional genes [
3,
4,
5,
6]. This major breakthrough allowed genes and data of the complex and highly diversified adaptive immune responses to be managed in genomic databases and tools.
The adaptive immune response was acquired by jawed vertebrates (or
gnathostomata) more than 450 million years ago and is found in all extant jawed vertebrate species from fishes to humans. It is characterized by a remarkable immune specificity and memory, which are properties of the B and T cells owing to an extreme diversity of their antigen receptors. The specific antigen receptors comprise the IG or antibodies of the B cells and plasmacytes [
3], and the TR [
4]. The IG recognize antigens in their native (unprocessed) form, whereas the TR recognize processed antigens, which are presented as peptides by the highly polymorphic major histocompatibility (MH, in humans HLA for human leucocyte antigens) proteins.
The potential antigen receptor repertoire of each individual is estimated to comprise about 2 × 10
12 different IG and TR, and the limiting factor is only the number of B and T cells that an organism is genetically programmed to produce [
3,
4]. This huge diversity results from the complex molecular synthesis of the IG and TR chains and, more particularly, of their variable domains (V-DOMAIN) which, at their N-terminal end, recognize and bind the antigens [
3,
4]. The IG and TR synthesis includes several unique mechanisms that occur at the DNA level: combinatorial rearrangements of the V, D and J genes that code the V-DOMAIN (the V-(D)-J being spliced to the C gene that encodes the C-REGION in the transcript); exonuclease trimming at the ends of the V, D and J genes; and random addition of nucleotides by the terminal deoxynucleotidyl transferase (TdT) that creates the junctional N-diversity regions, and later during B cell differentiation, for the IG, somatic hypermutations, gene conversion (e.g., in birds), and class or subclass switch in higher vertebrates [
3,
4].
IMGT
® manages the diversity and complexity of the IG and TR and the polymorphism of the MH of humans and other vertebrates. IMGT
® is also specialized in the other proteins of the immunoglobulin superfamily (IgSF) and MH superfamily (MhSF) and related proteins of the immune system (RPI) of vertebrates and invertebrates [
2]. IMGT
® provides a common access to standardized data from genome, proteome, genetics, and two-dimensional (2D) and three-dimensional (3D) structures. IMGT
® is the acknowledged high-quality integrated knowledge resource in immunogenetics for exploring immune functional genomics. IMGT
® comprises seven databases (for sequences, genes, and 3D structures) [
7,
8,
9,
10,
11,
12] and 17 online tools [
13,
14,
15,
16,
17,
18,
19,
20,
21,
22,
23,
24,
25,
26,
27,
28] (
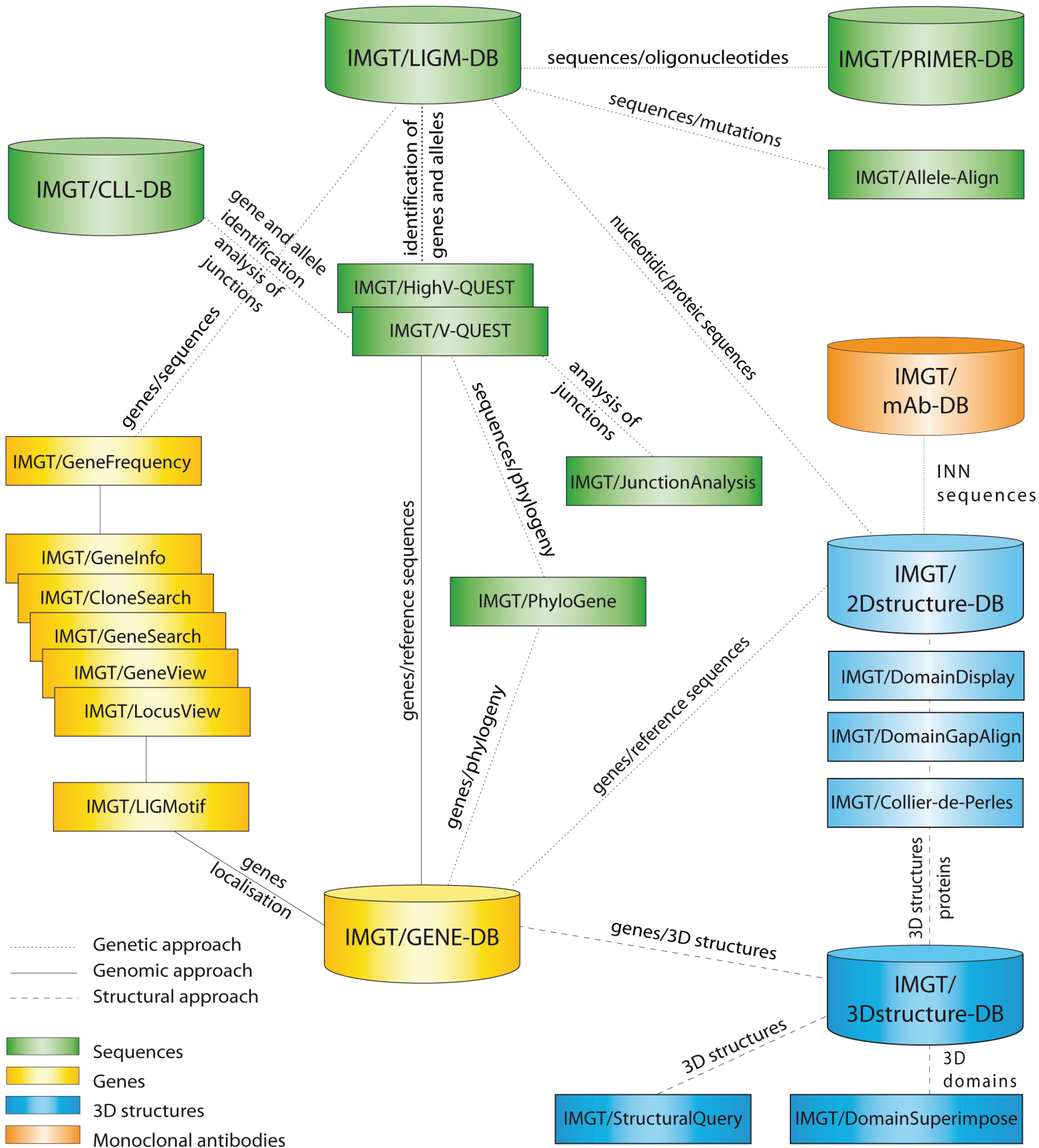
Figure 1), as well as more than 15,000 pages of web resources (e.g., IMGT Scientific chart, IMGT Repertoire, IMGT Education > Aide-mémoire [
29], The IMGT Medical page, The IMGT Veterinary page, The IMGT Biotechnology page, The IMGT Immunoinformatics page) [
2]. IMGT
® is the global reference in immunogenetics and immunoinformatics [
30,
31,
32,
33,
34,
35,
36,
37,
38,
39,
40,
41,
42,
43,
44,
45]. Its standards have been endorsed by the World Health Organization-International Union of Immunological Societies (WHO-IUIS) Nomenclature Committee since 1995 (first IMGT
® online access at the 9th International Congress of Immunology, San Francisco, CA, USA) [
46,
47] and the WHO International Nonproprietary Names (INN) Programme [
48,
49].
Figure 1.
IMGT
®, the international ImMunoGeneTics information system
® [
1,
2]. Databases are shown as cylinders and tools as rectangles. The web resources are not shown.
Figure 1.
IMGT
®, the international ImMunoGeneTics information system
® [
1,
2]. Databases are shown as cylinders and tools as rectangles. The web resources are not shown.
The accuracy and the consistency of the IMGT
® data are based on IMGT-ONTOLOGY [
50,
51,
52], the first, and so far unique, ontology for immunogenetics and immunoinformatics [
50,
51,
52,
53,
54,
55,
56,
57,
58,
59,
60,
61,
62,
63,
64,
65,
66,
67,
68,
69]. IMGT-ONTOLOGY manages the immunogenetics knowledge through diverse facets that rely on seven axioms: IDENTIFICATION, DESCRIPTION, CLASSIFICATION, NUMEROTATION, LOCALIZATION, ORIENTATION, and OBTENTION [
51,
52,
56]. The concepts generated from these axioms led to the elaboration of the IMGT
® standards that constitute the IMGT Scientific chart: e.g., IMGT
® standardized keywords (IDENTIFICATION) [
57], IMGT
® standardized labels (DESCRIPTION) [
58], IMGT
® standardized gene and allele nomenclature (CLASSIFICATION) [
59], IMGT unique numbering [
60,
61,
62,
63,
64,
65] and its standardized graphical 2D representation or IMGT Colliers de Perles [
66,
67,
68,
69] (NUMEROTATION).
With a focus on IG, we first review the fundamental information generated from these IMGT-ONTOLOGY concepts which led to the IMGT Scientific chart rules. The major IMGT
® tools and databases used for IG repertoire analysis, antibody engineering and humanization, and IG/Ag structures are then briefly presented: IMGT/V-QUEST [
13,
14,
15,
16,
17,
18] for the analysis of rearranged nucleotide sequence with the results of the integrated IMGT/JunctionAnalysis [
19,
20]; IMGT/Automat [
21,
22] and IMGT/Collier-de-Perles tool [
27]; IMGT/HighV-QUEST, the high-throughput version for Next-Generation Sequencing (NGS) [
23,
24]; IMGT/DomainGapAlign [
10,
25,
26] for amino acid sequence analysis; IMGT/3Dstructure-DB for 3D structures [
9,
10,
11]; and its extension, IMGT/2Dstructure-DB (for antibodies and other proteins for which the 3D structure is not available). IMGT
® tools and databases run against IMGT reference directories built from sequences annotated in IMGT/LIGM-DB [
7], the IMGT
® nucleotide database (176,806 sequences from 346 species in October 2014), and from IMGT/GENE-DB [
8], the IMGT
® gene database (3464 genes and 5118 alleles from 21 species, of which there were 710 genes and 1439 alleles for
Homo sapiens and 868 genes and 1318 alleles for
Mus musculus in October 2014).
An interface, IMGT/mAb-DB [
12], has been developed to provide an easy access to therapeutic antibody amino acid sequences (links to IMGT/2Dstructure-DB) and structures (links to IMGT/3Dstructure-DB, if 3D structures are available). IMGT/mAb-DB data include monoclonal antibodies (mAb, INN suffix –mab) (a –mab is defined by the presence of at least an IG variable domain) and fusion proteins for immune applications (FPIA, INN suffix –cept) (a –cept is defined by a receptor fused to an Fc) from the WHO-INN programme [
48,
49]. This database also includes a few composite proteins for clinical applications (CPCA) (e.g., protein or peptide fused to an Fc for only increasing their half-life, identified by the INN prefix ef–) and some RPI used, unmodified, for clinical applications.
The unified IMGT
® approach is of major interest for bridging knowledge from IG repertoire in normal and pathological situations [
70,
71,
72,
73,
74,
75], IG allotypes and immunogenicity [
76,
77,
78], NGS repertoire [
23,
24], antibody engineering and humanization [
33,
40,
41,
42,
79,
80,
81,
82,
83,
84,
85,
86].
4. IMGT® Databases for IG V-DOMAIN and C-DOMAIN Analysis
4.1. IMGT/3Dstructure-DB
4.1.1. IMGT/3Dstructure-DB card
IMGT/3Dstructure-DB [
9,
10,
11]; the IMGT
® structure database; provides IMGT
® annotation and contact analysis of IG 3D structures; and paratope/epitope description of IG/antigen complexes (
Table 5). There is one “IMGT/3Dstructure-DB card” per IMGT/3Dstructure-DB entry and this card provides access to all data related to that entry. The “PDB code” (4 letters and/or numbers; e.g., 1n0x) is used as “IMGT entry ID” for the 3D structures obtained from the Research Collaboratory for Structural Bioinformatics (RCSB) Protein Data Bank (PDB) [
98]. The IMGT/3Dstructure-DB card provides eight search/display options: “Chain details”; “Contact analysis”; “Paratope and epitope”; “3D visualization Jmol or QuickPDB”; “Renumbered IMGT files”; “IMGT numbering comparison”; “References and links”; and “Printable card” [
9,
10,
11].
4.1.2. IMGT Chain and Domain Annotation
The “Chain details” section comprises information first on the chain itself, then per domain [
9,
10,
11]. Chain and domain annotation includes the IMGT gene and allele names (CLASSIFICATION), region and domain delimitations (DESCRIPTION) and domain AA positions according to the IMGT unique numbering (NUMEROTATION) [
60,
61,
62,
63,
64,
65] (
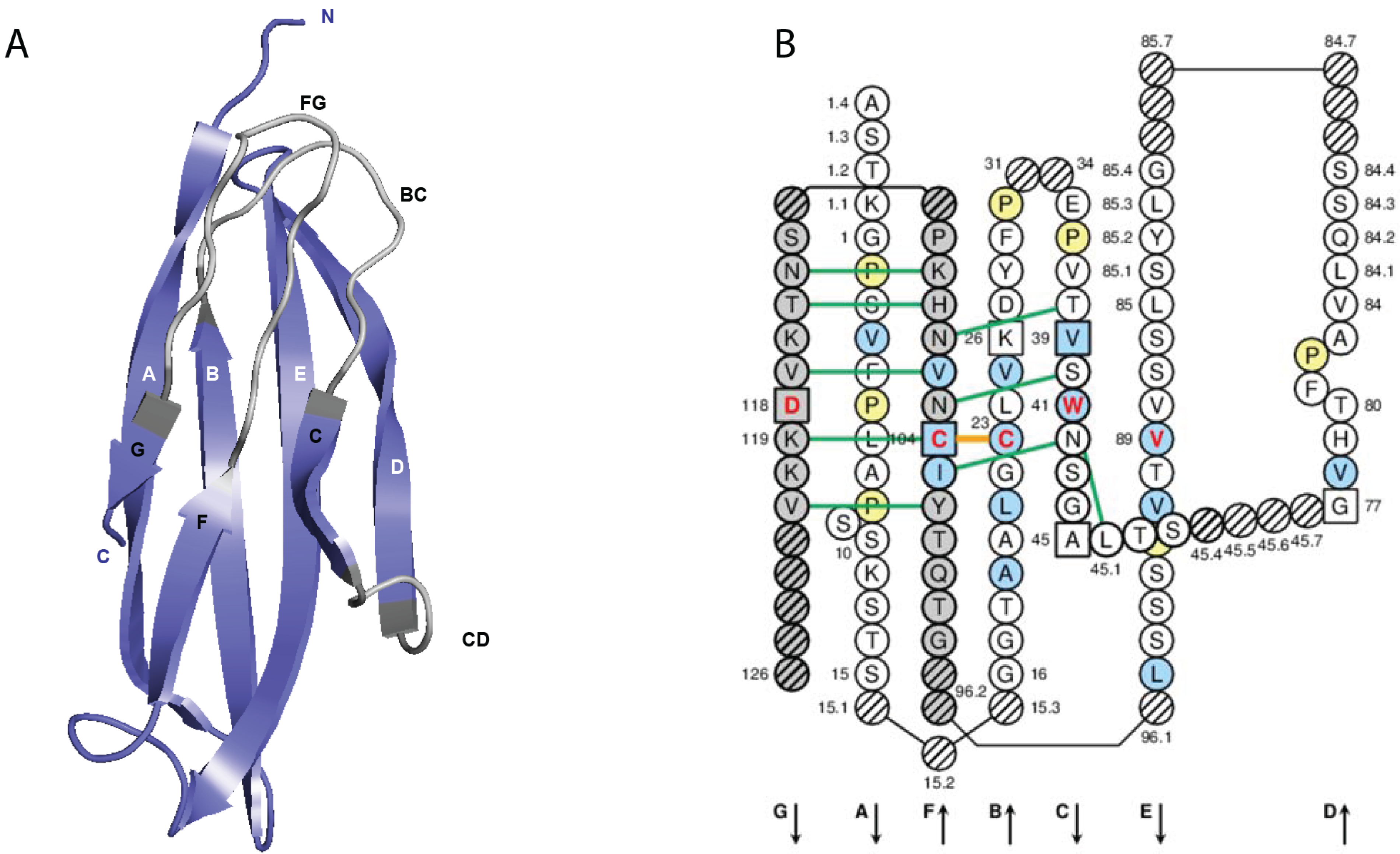
Figure 5A). The closest IMGT
® genes and alleles (found expressed in each domain of a chain) are identified with the integrated IMGT/DomainGapAlign [
10,
25,
26], which aligns the amino acid sequences of the 3D structures with the IMGT/DomainSeq reference directory.
4.1.4. Paratope and Epitope
In an IG/Ag complex, the amino acids in contact at the interface between the IG and the Ag constitute the paratope on the IG V-DOMAIN surface and the epitope on the Ag surface. For IG/Ag, the paratope and epitope are displayed in Contact analysis (
Figure 5B,C), but for each V domain separately. Clicking on the “Paratope and epitope” tag (displayed in the IMGT/3Dstructure-DB card, only if relevant), gives access to “IMGT paratope and epitope details”, which are described in a standardized way. Each amino acid which belongs to the paratope is defined by its position in an IG V-DOMAIN. Each amino acid that belongs to the epitope is defined by its position in the chain in the 3D structure or, if the antigen belongs to an IgSF or MhSF protein and if the epitope is part of a characterized V, C, or G domain, by its position in the domain according to the IMGT unique numbering.
Figure 5.
IMGT/3Dstructure-DB. (
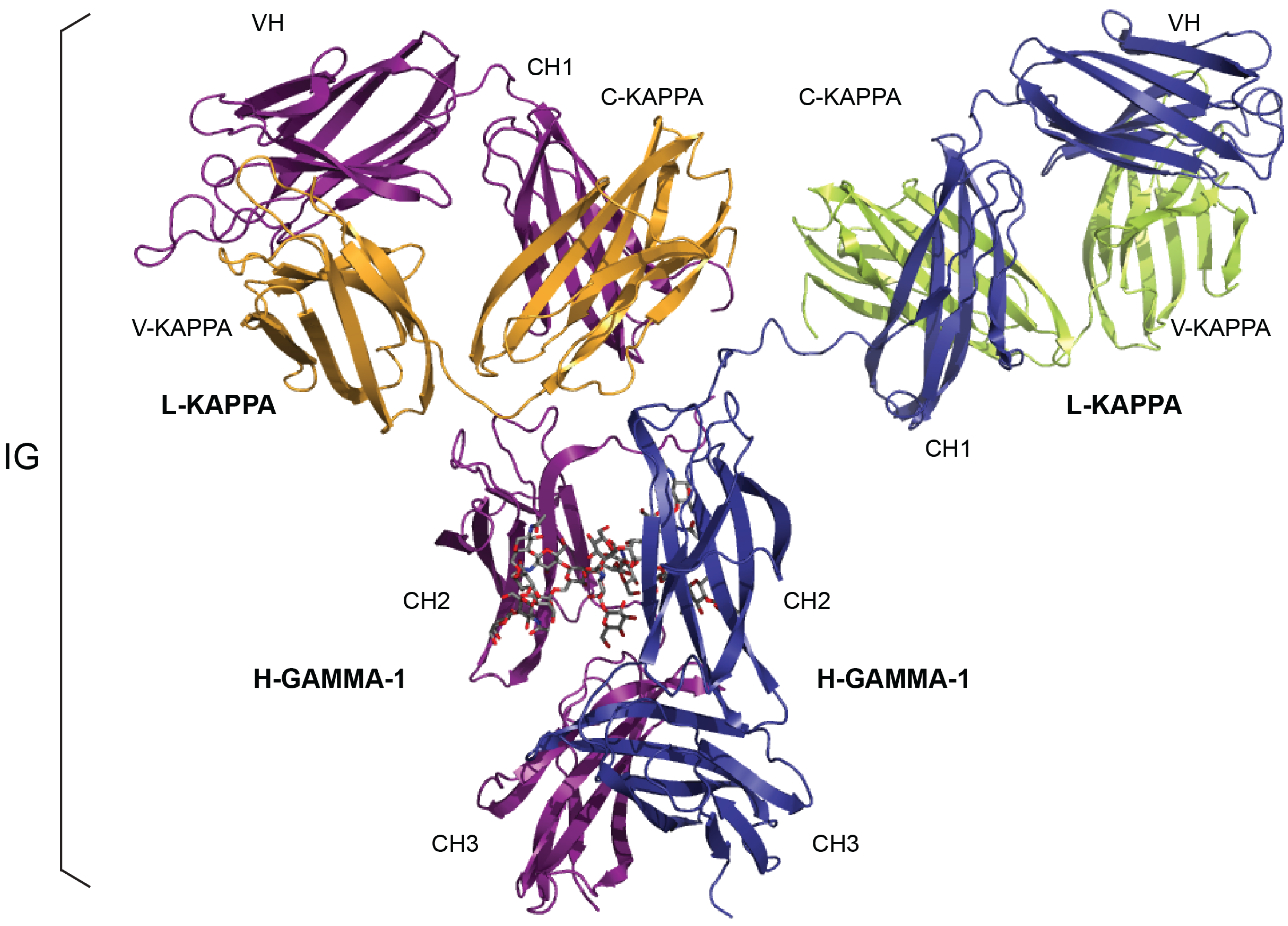
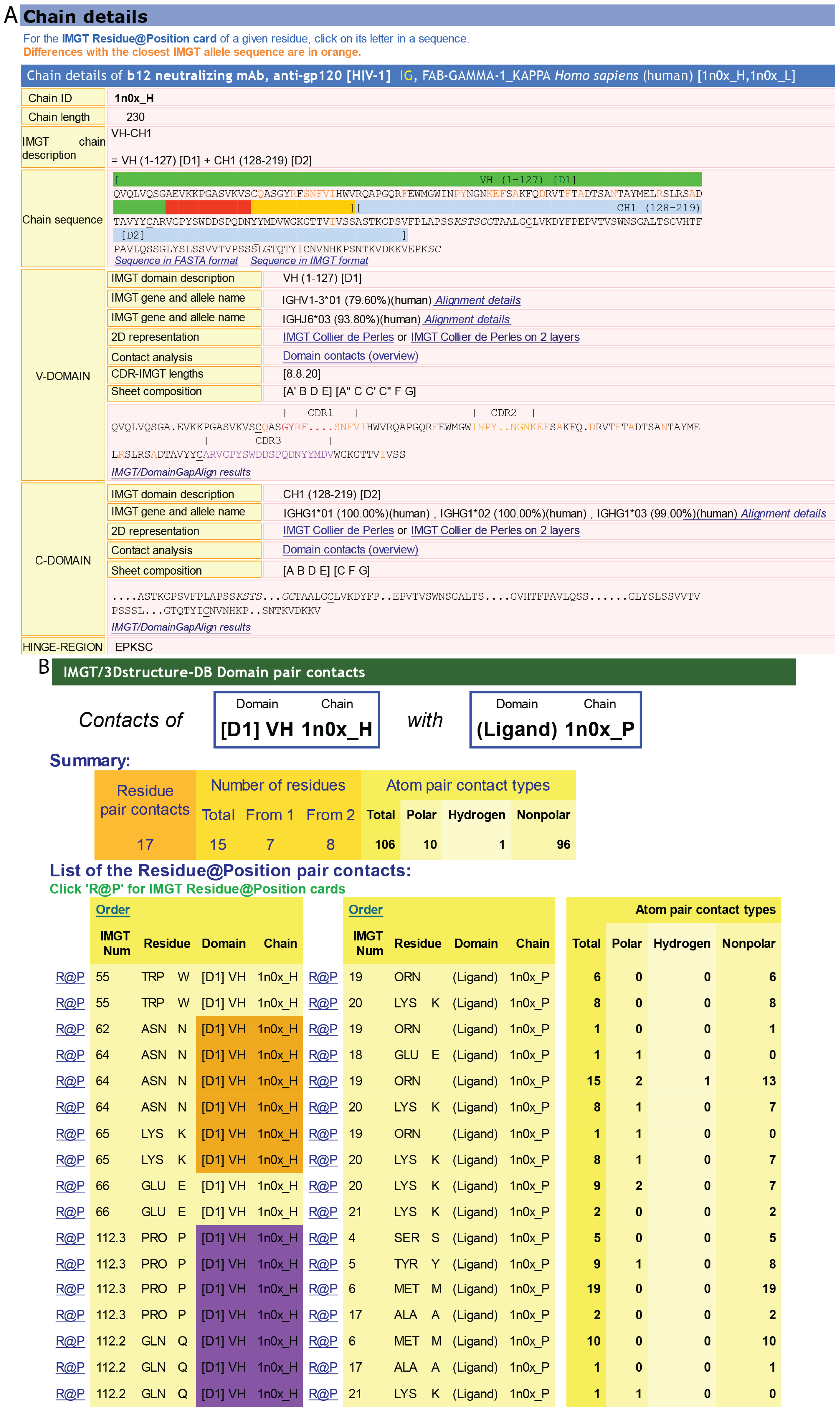
A) IMGT/3Dstructure-DB card. The “IMGT/3Dstructure-DB card” is available for each entry of the database. The “Chain details” shows, first, information on the chain (Chain ID, Chain length, IMGT chain description, Chain sequence), then a detailed description of each domain of the chain. The description of the V-DOMAIN (VH) and C-DOMAIN (CH1) of the VH-CH1 chain (1n0x_H) of the b12 Fab is shown. A similar result display interface is provided in IMGT/2Dstructure-DB cards but without “Contact analysis” (and without hydrogen bonds in IMGT Collier de Perles on 2 layers); (
B) IMGT/3Dstructure-DB Domain pair contacts between the “VH” and the “Ligand” (antigen, Ag) of an IG/Ag complex. The VH is from the VH-CH1 chain (1n0x_H) of the b12 Fab and the ligand is a synthetic peptide (1n0x_P). The VH is in contact with the ligand by three AA of the CDR2-IMGT (orange online) (N62, N64 and K65) and two AA of the CDR3-IMGT (purple online) (P112.3 and Q112.2). The two AA which interact with the ligand but do not belong to the CDR-IMGT are the anchors W55 and E66. These contacts are not unexpected given by the small size (peptide) of the ligand; (
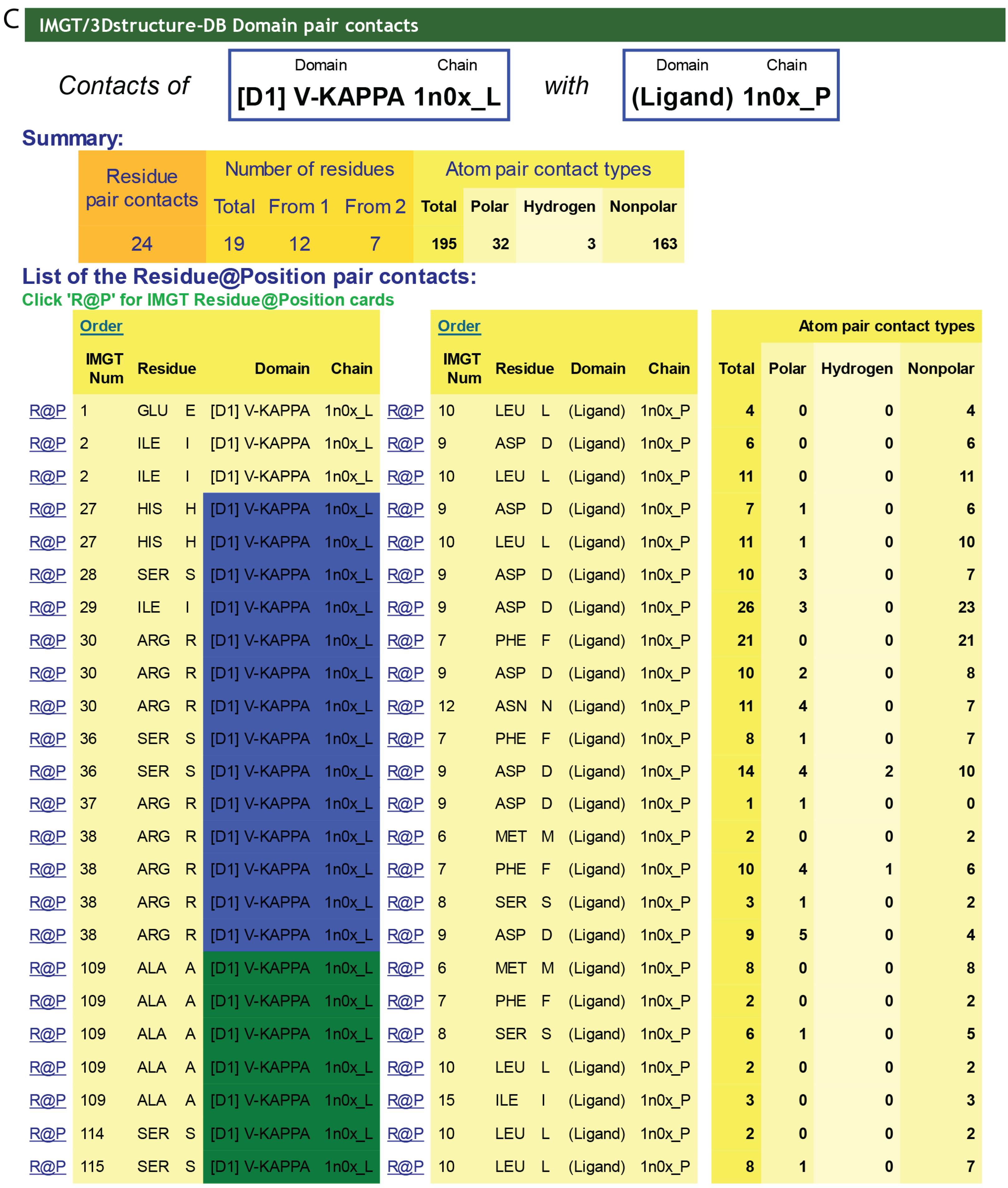
C) IMGT/3Dstructure-DB Domain pair contacts between the “V-KAPPA” and the “Ligand” (Ag) of an IG/Ag complex. The V-KAPPA is from the L-KAPPA chain (1n0x_L) of the b12 Fab and the ligand is the peptide (1n0x_P) as in (
B). The V-KAPPA is in contact with the ligand by seven AA of the CDR1-IMGT (blue online) (H27, S28, I29, R30, S36, R37 and R38) and three AA of the CDR3-IMGT (greenblue online) (A109, S114 and S115). “Polar”, “Hydrogen bond” and “Nonpolar” are selected by default in “Atom pair contact types” options at the bottom of the page (not shown). The user can also choose to display these contacts by “Atom pair contact categories” (BB), (SS), (BS) and (SB). Clicking on R@P gives access to the IMGT Residue@Position card. The IG/Ag complex structure is 1n0x from IMGT/3Dstructure-DB [
1,
9,
10,
11].
Figure 5.
IMGT/3Dstructure-DB. (
A) IMGT/3Dstructure-DB card. The “IMGT/3Dstructure-DB card” is available for each entry of the database. The “Chain details” shows, first, information on the chain (Chain ID, Chain length, IMGT chain description, Chain sequence), then a detailed description of each domain of the chain. The description of the V-DOMAIN (VH) and C-DOMAIN (CH1) of the VH-CH1 chain (1n0x_H) of the b12 Fab is shown. A similar result display interface is provided in IMGT/2Dstructure-DB cards but without “Contact analysis” (and without hydrogen bonds in IMGT Collier de Perles on 2 layers); (
B) IMGT/3Dstructure-DB Domain pair contacts between the “VH” and the “Ligand” (antigen, Ag) of an IG/Ag complex. The VH is from the VH-CH1 chain (1n0x_H) of the b12 Fab and the ligand is a synthetic peptide (1n0x_P). The VH is in contact with the ligand by three AA of the CDR2-IMGT (orange online) (N62, N64 and K65) and two AA of the CDR3-IMGT (purple online) (P112.3 and Q112.2). The two AA which interact with the ligand but do not belong to the CDR-IMGT are the anchors W55 and E66. These contacts are not unexpected given by the small size (peptide) of the ligand; (
C) IMGT/3Dstructure-DB Domain pair contacts between the “V-KAPPA” and the “Ligand” (Ag) of an IG/Ag complex. The V-KAPPA is from the L-KAPPA chain (1n0x_L) of the b12 Fab and the ligand is the peptide (1n0x_P) as in (
B). The V-KAPPA is in contact with the ligand by seven AA of the CDR1-IMGT (blue online) (H27, S28, I29, R30, S36, R37 and R38) and three AA of the CDR3-IMGT (greenblue online) (A109, S114 and S115). “Polar”, “Hydrogen bond” and “Nonpolar” are selected by default in “Atom pair contact types” options at the bottom of the page (not shown). The user can also choose to display these contacts by “Atom pair contact categories” (BB), (SS), (BS) and (SB). Clicking on R@P gives access to the IMGT Residue@Position card. The IG/Ag complex structure is 1n0x from IMGT/3Dstructure-DB [
1,
9,
10,
11].
![Biomolecules 04 01102 g005a]()
![Biomolecules 04 01102 g005b]()
4.1.5. Renumbered Flat File and IMGT Numbering Comparison
“Renumbered IMGT file” allows viewing (or downloading) of an IMGT coordinate file renumbered according to the IMGT unique numbering, and with added IMGT specific information on chains and domains (added in the “REMARK 410” lines (blue online) and identical to the “Chain details” annotation).
“IMGT numbering comparison” provides, per domain, the IMGT DOMAIN numbering by comparison with the PDB numbering and the residue (3-letter and 1-letter names), which allows standardized IMGT representations using generic tools.
4.2. IMGT/2Dstructure-DB
IMGT/2Dstructure-DB was created as an extension of IMGT/3Dstructure-DB [
9,
10,
11] to describe and analyze amino acid sequences of chains and domains for which no 3D structures were available (
Table 5). IMGT/2Dstructure-DB uses the IMGT/3Dstructure-DB informatics frame and interface, which allow one to analyze, manage and query IG (and also TR and MH, as well as other IgSF and MhSF) and engineered proteins (FPIA, CPCA) as polymeric receptors made of several chains, in contrast to the IMGT/LIGM-DB sequence database that analyzes and manages sequences individually [
7]. The amino acid sequences are analysed with the IMGT
® criteria of standardized identification [
57], description [
58], nomenclature [
59] and numerotation [
60,
61,
62,
63,
64,
65].
The current IMGT/2Dstructure-DB entries include amino acid sequences of antibodies from Kabat [
95] (those for which there were no available nucleotide sequences) and amino acid sequences of mAb and FPIA from the WHO-INN programme [
12,
48,
49]. Queries can be made on an individual entry using the “Entry ID” or the “Molecule name”. The same query interface is used for IMGT/2Dstructure-DB and IMGT/3Dstructure-DB. Thus a “trastuzumab’ query in “Molecule name” allows retrieval of six results: two INN (“trastuzumab” and “trastuzumab emtansine”) from IMGT/2Dstructure-DB and four 3D structures from IMGT/3Dstructure-DB. For mAb and FPIA results, INN sequences represent the reference sequences [
12,
48,
49] as sequences of the 3D structures may have been engineered or may contain experimental errors.
The IMGT/2Dstructure-DB cards provide standardized IMGT information on IG chains and domains and IMGT Colliers de Perles on one or two layers, in a format identical to that provided for the sequence analysis in IMGT/3Dstructure-DB; however, the information on experimental structural data (hydrogen bonds in IMGT Collier de Perles on two layers, Contact analysis) is only available in the corresponding IMGT/3Dstructure-DB cards if the antibodies have been crystallized.
5. IMGT® IG V-DOMAIN and C-DOMAIN Analysis for Antibody Humanization and Engineering
5.1. CDR-IMGT Delimitation for Grafting
The objective of antibody humanization is to graft at the DNA level the CDR of an antibody V domain, from mouse (or other species) and of a given specificity, onto a human V domain framework, thus preserving the specificity of the original (murine or other species) antibody while decreasing its immunogenicity [
99]. IMGT/DomainGapAlign [
10,
25,
26] is the reference tool for antibody humanization design based on CDR grafting. Indeed, it precisely defines the CDR-IMGT to be grafted and helps in selecting the most appropriate human FR-IMGT by providing the alignment of the amino acid sequences between the mouse (or other species) and the closest human V-DOMAIN.
Analyses performed on humanized therapeutic antibodies underline the importance of a correct delimitation of the CDR and FR. As an example, two amino acid changes were required in the first version of the humanized VH of alemtuzumab, in order to restore the specificity and affinity of the original rat antibody. The positions of these amino acid changes (S28 > F and S35 > F) are now known to be located in the CDR1-IMGT and should have been directly grafted, but at the time of this mAb humanization they were considered as belonging to the FR according to the Kabat numbering [
95]. In contrast, positions 66–74 were, at the same time, considered as belonging to the CDR according to the Kabat numbering, whereas they clearly belong to the FR2-IMGT and the corresponding sequence should have been “human” instead of being grafted from the “rat” sequence (IMGT
® [
1], The IMGT Biotechnology page > Antibody humanization > Alemtuzumab).
5.2. Evaluation of the Degree of “Humanization” of an IG V Sequence
IMGT/DomaingapAlign is used to evaluate the degree of “humanization” of an IG V sequence, either obtained from a species other than human (e.g., mouse or rat), or obtained from engineered human sequences (e.g., selected from combinatorial library or mutated). IMGT/DomaingapAlign provides an objective assessment of the degree of humanization of the user sequence, based on sequence alignments, independently on the source of the starting sequence (e.g., species) and independently on the experimental methodology that was used with the objective of humanizing it. A query of the user sequence against “V” of “any” species will display “Homo sapiens” IG V genes at the top of the results, in the case of a successfully “humanized” V. In contrast, the query will display V genes of species other than Homo sapiens for an unsuccessful humanization: in that case the V gene is “non-human” and the IG chain to which it belongs is “chimeric”.
5.3. IGHG1 Alleles and G1m Allotypes
Allotypes are polymorphic markers of an IG subclass that correspond to amino acid changes and are detected serologically by antibody reagents [
77]. In therapeutic antibodies (human, humanized or chimeric) [
12], allotypes may represent potential immunogenic residues [
76], as demonstrated by the presence of antibodies in individuals immunized against these allotypes [
77]. The allotypes of the human heavy gamma chains of the IgG are designated as Gm (for gamma marker).
The allotypes G1m, G2m and G3m are carried by the constant region of the gamma1, gamma2 and gamma3 chains, encoded by the IGHG1, IGHG2 and IGHG3 genes, respectively [
77]. The gamma1 chains express different combinations of G1m allotypes or G1m alleles: G1m3, G1m3,1, G1m17,1, and G1m17,1,2 (
Table 6). The C region of the G1m3,1, G1m17,1 and G1m17,1,2 chains differ from that of the G1m3 chains by two, three and four amino acids, respectively [
77]. Two additional G1m alleles (G1m17,1,28 and G1m17,1,27,28) have been identified by serology in the Negroid populations, whereas another allele (G1m17,1,27) was deduced from a sequence with the AA change expected for the Gm27 allotype [
77]. The correspondence between the G1m alleles and IGHG1 alleles is shown in
Table 6.
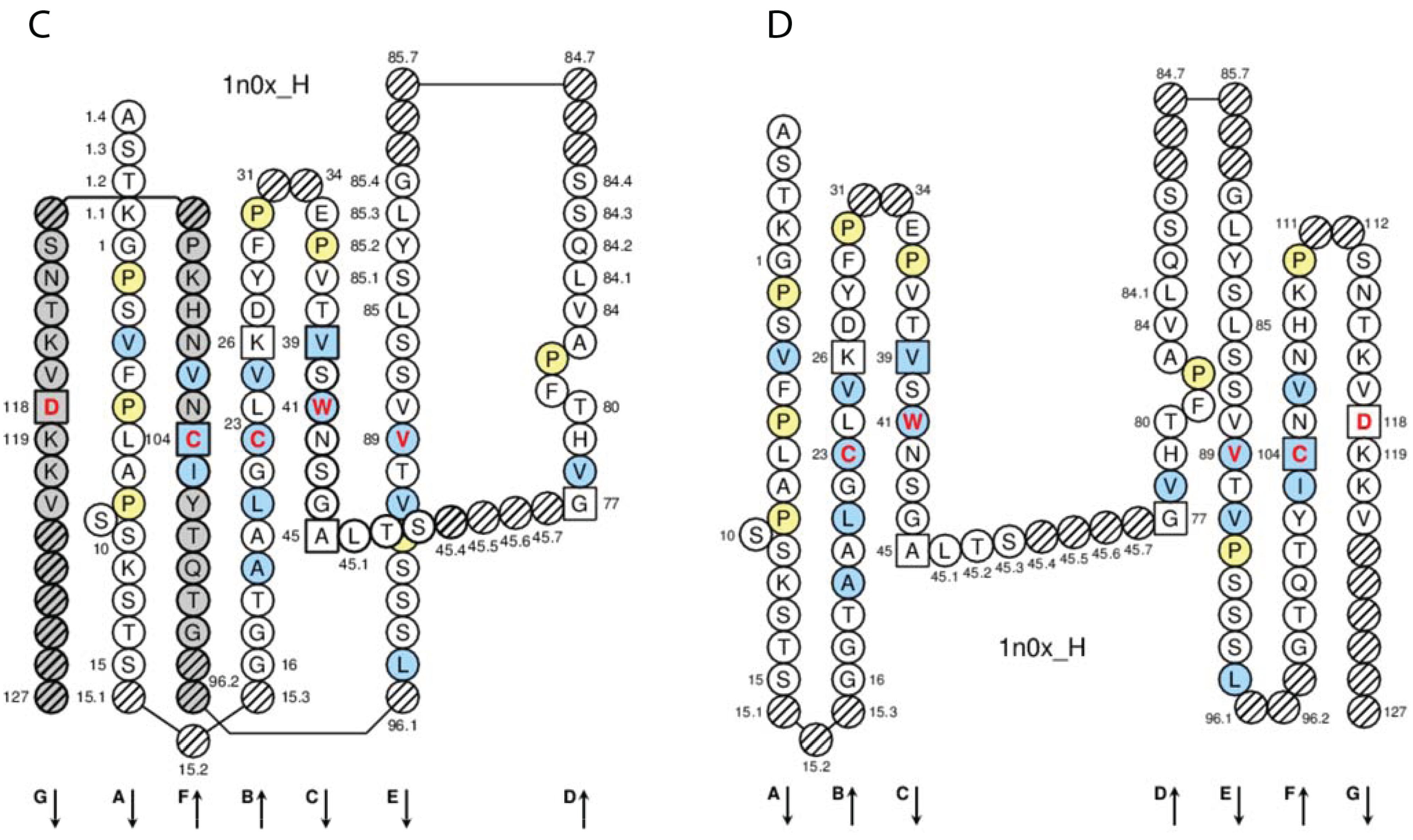
In the IGHG1 CH1, the lysine at position 120 (K120) in strand G corresponds to the G1m17 allotype [
77] (
Figure 4D). The isoleucine I103 (strand F) is specific of the gamma1 chain isotype. If an arginine is expressed at position 120 (R120), the simultaneous presence of R120 and I103 corresponds to the expression of the G1m3 allotype [
77]. For the gamma3 and gamma4 isotypes (which also have R120 but T in 103), R120 only corresponds to the expression of the nG1m17 isoallotype (an isoallotype or nGm is detected by antibody reagents that identify this marker as an allotype in one IgG subclass and as an isotype for other subclasses) [
77].
Table 6.
Correspondence between the IGHG1 alleles and G1m alleles.
Table 6.
Correspondence between the IGHG1 alleles and G1m alleles.
| IGHG1 Alleles | G1m Alleles a | IMGT Amino acid Positions b | Populations [77] |
|---|
| allotypes | Isoallotypes c | CH1 | CH3 | |
|---|
| 103 | 120 | 12 | 14 | 110 |
|---|
| | G1m17/nG1m1 | G1m1/nG1m1 | G1m2/- |
|---|
| G1m3 d |
|---|
| IGHG1*01 e, IGHG1*02 e | G1m17,1 | | I | K | D | L | A | Caucasoid Negroid Mongoloid |
| IGHG1*04 | G1m17,1,27 | |
| IGHG1*05p | G1m17,1,28 | Negroid |
| IGHG1*06p | G1m17,1,27,28 | Negroid |
| IGHG1*03 | G1m3 | nG1m1, nG1m17 | I | R | E | M | A | Caucasoid |
| IGHG1*07p f | G1m17,1,2 | | I | K | D | L | G | Caucasoid Mongoloid |
| IGHG1*08p f | G1m3,1 | nG1m17 | I | R | D | L | A | Mongoloid |
In the IGHG1 CH3, the aspartate D12 and leucine L14 (strand A) correspond to G1m1, whereas glutamate E12 and methionine M14 correspond to the nG1m1 isoallotype [
77] (
Table 6). A glycine at position 110 corresponds to G1m2, whereas an alanine does not correspond to any allotype (G1m2-negative chain). Therapeutic antibodies are most frequently of the IgG1 isotype, and to avoid a potential immunogenicity, the constant region of the gamma1 chains are often engineered to replace the G1m3 allotype by the less immunogenic G1m17 (CH1 R120 > K) (G1m17 is more extensively found in different populations) [
77].
5.4. IGHG N-Linked Glycosylation Site CH2 N84.4
A N-linked glycosylation site is present in the CH2 domain of the constant region of the human IG heavy chains of the four IgG isotypes. The N-linked glycosylation site belongs to the classical N-glycosylation motif N-X-S/T (where N is asparagine, X any amino acid except proline, S serine, T threonine) and is defined as CH2 N84.4. As shown in the IMGT Collier de Perles, this asparagine is localized at the DE turn. The IMGT unique numbering has the advantage of identifying the C domain (here, CH2) and, in the domain, the amino acid and its localization (here, N84.4) which can be visualized in the IMGT Collier de Perles and correlated with the 3D structure [
70,
84,
85].
6. Conclusions
IMGT-ONTOLOGY and the IMGT
® information system, which are at the origin of immunoinformatics [
45], have provided the concepts, the knowledge environment and the informatics frame for a standardized and integrated analysis of the IG, from gene to structure and function. IG repertoire analysis, therapeutic antibody engineering and humanization, paratope/epitope characterization, immunotherapy represent major current fields of immunoinformatics at the forefront of basic, pharmaceutical and clinical research owing to major methodological and medical advances.
The IMGT
® standards for IG are used in clinical applications. Thus, IMGT/V-QUEST is frequently used by clinicians for the analysis of IG somatic hypermutations in leukemia, lymphoma and myeloma, and more particularly in chronic lymphocytic leukemia (CLL) [
16,
72,
73,
74,
75] in which the percentage of mutations of the rearranged IGHV gene in the VH of the leukemic clone has a prognostic value for the patients. For this evaluation, IMGT/V-QUEST is the standard recommended by the European Research Initiative on CLL (ERIC) for comparative analysis between laboratories [
72]. The sequences of the V-(D)-J junctions determined by IMGT/JunctionAnalysis [
19,
20] are also used in the characterization of stereotypic patterns in CLL [
73,
74] and for the synthesis of probes specific of the junction for the detection and follow-up of minimal residual diseases (MRD) in leukemias and lymphomas. A new era is opening in hemato-oncology with the use of NGS for analysis of the clonality and MRD identification, making IMGT
® standards use more needed as ever. More generally, the IMGT/HighV-QUEST web portal is a paradigm for identification of IMGT clonotype diversity and expression in NGS immune repertoire analysis of the adaptive immune response in infectious diseases, in vaccination, and for next generation repertoire immunoprofiling [
24].
The therapeutic monoclonal antibody engineering field represents the most promising potential in medicine. A standardized analysis of IG genomic and expressed sequences, structures and interactions is crucial for a better molecular understanding and comparison of the mAb specificity, affinity, half-life, Fc effector properties and potential immunogenicity. IMGT-ONTOLOGY concepts have become a necessity for IG loci description of newly sequenced genomes, antibody structure/function characterization, antibody engineering (single chain Fragment variable (scFv), phage displays, combinatorial libraries) and antibody humanization (chimeric, humanized and human antibodies) [
33,
40,
82,
83,
84,
85,
86]. IMGT
® standardization allows repertoire analysis and antibody humanization studies to move to novel high-throughput methodologies with the same high-quality criteria. The CDR-IMGT lengths are now required for mAb INN applications and are included in the WHO-INN definitions [
49], bringing a new level of standardized information in the comparative analysis of therapeutic antibodies.