Micro-RNA Feedback Loops Modulating the Calcineurin/NFAT Signaling Pathway
Abstract
:1. Introduction
2. Results
2.1. miRNAs Targeting the Calcineurin/NFAT Signaling Pathway
2.2. miRNAs Induced by NFAT Transcription Factor Family
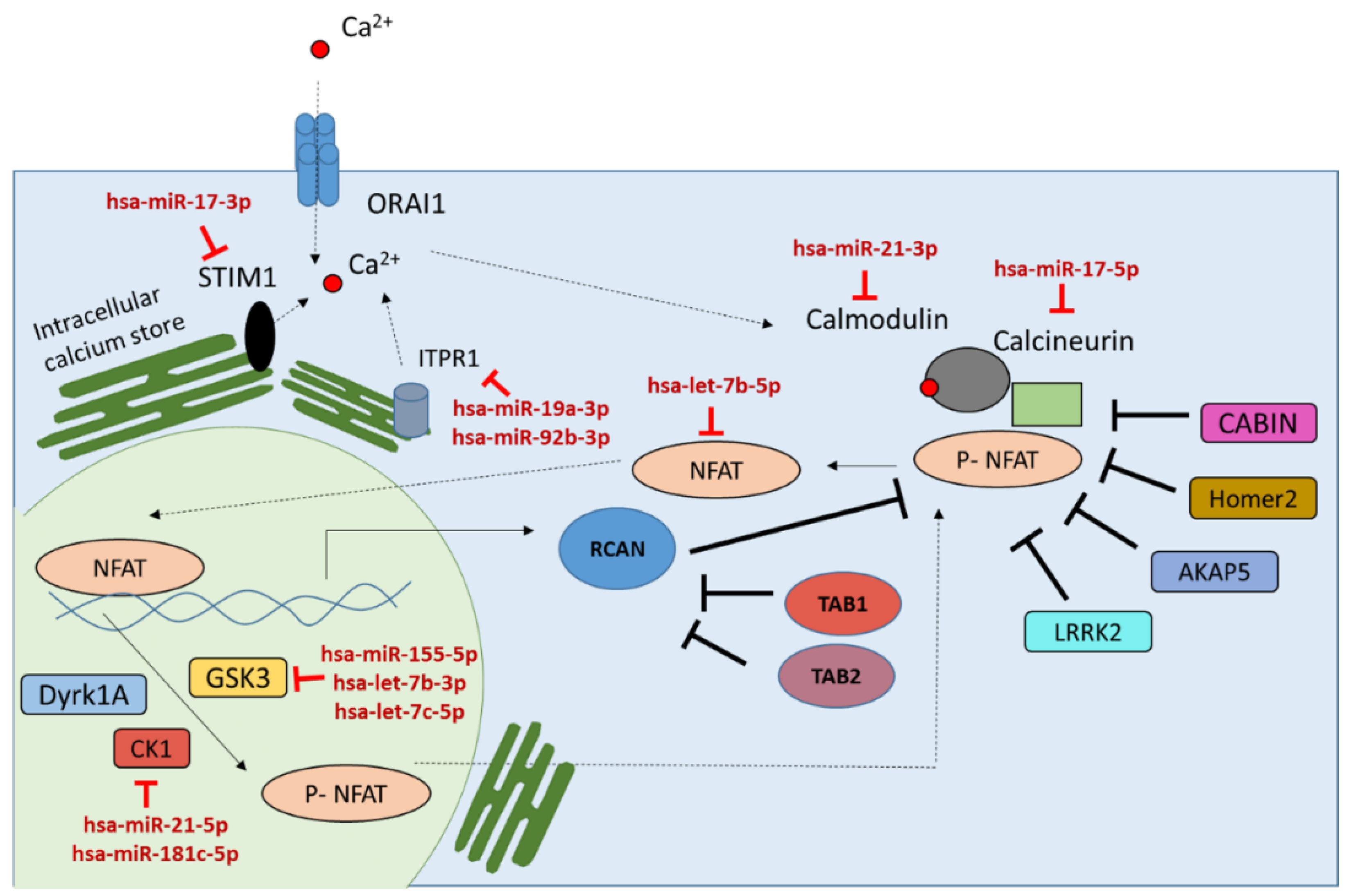
2.3. miRNAs in Negative or Positive Feedback Loop to Modulate the Calcineurin/NFAT Signaling Pathway Activity
3. Discussion
4. Materials and Methods
4.1. Computational Prediction of miRNA Targeting Members of the Calcineurin/NFAT Pathway
4.2. Experimental Validation Data on miRNA Targets
4.3. miRNA Expressed in PBMCs
4.4. miRNA Promoter Sequences and NFAT Transcription Factor Binding Site
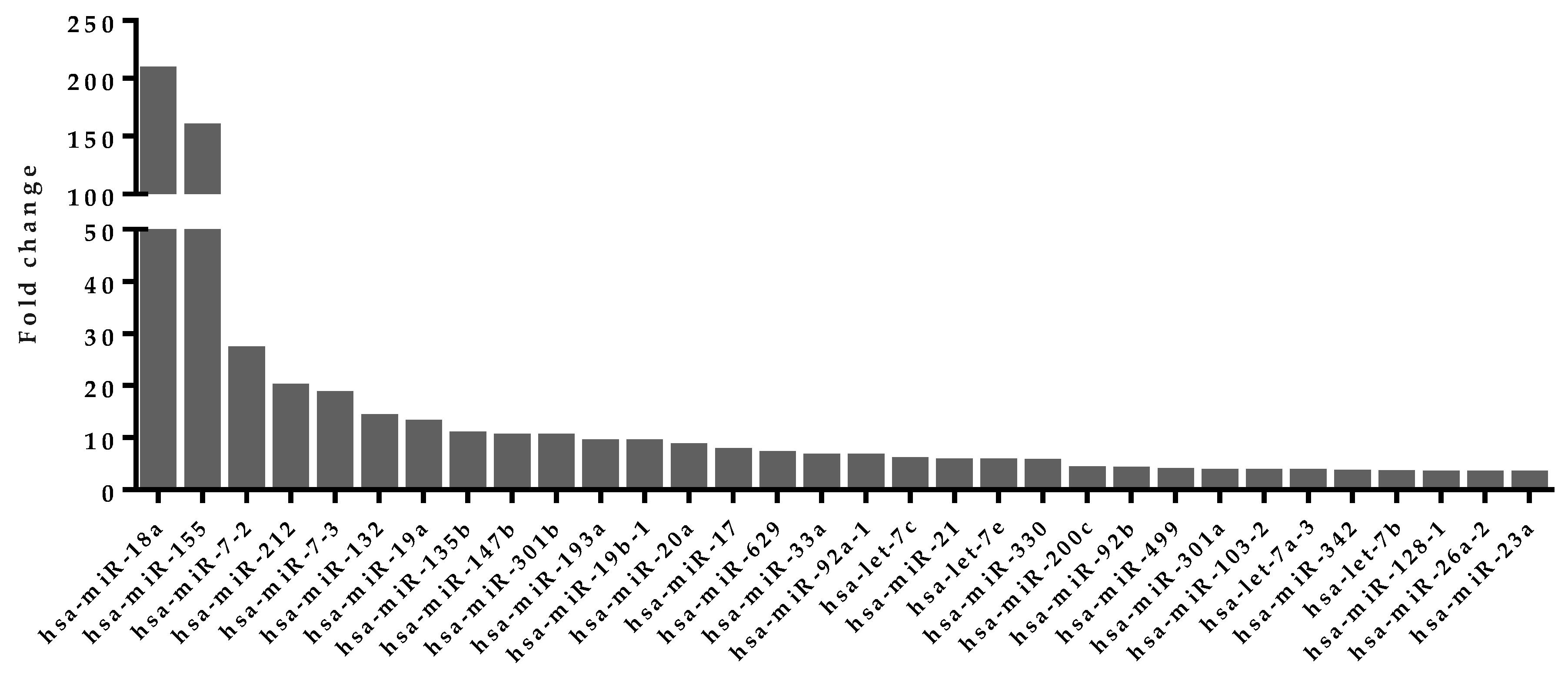
4.5. Upregulated miRNA Selection from Activated Human T Cells
5. Conclusions
Supplementary Materials
Acknowledgments
Author Contributions
Conflicts of Interest
Abbreviations
| NFAT | Nuclear factor of activated T cells |
| PPP3CA | Calmodulin-Dependent Calcineurin A Subunit Alpha Isoform |
| RCAN | RCAN Family Member/ Down Syndrome Candidate Region 1-Like Protein |
| GSK | Glycogen Synthase Kinase |
| DYRK1A | Dual-Specificity Tyrosine-(Y)-Phosphorylation Regulated Kinase 1A |
| HOMER2 | Homer Scaffolding Protein |
| ITPR1 | Inositol 1,4,5-Trisphosphate Receptor, Type 1 |
| MAP3K7 | Mitogen-Activated Protein Kinase Kinase Kinase 7 |
| TAB | TGF-Beta Activated Kinase 1/MAP3K7 Binding Protein |
| CSNK1A1 | Casein Kinase 1, Alpha 1 |
| LRRK | Lrrk Leucine-rich repeat kinase |
| CABIN | Calcineurin Binding Protein |
| AKAP | A-kinase anchor proteins |
| CALM | Calmodulin |
| STIM | stromal interaction molecule |
| ORAI | Calcium Release-Activated Calcium Modulator |
| miRNA | micro RNA |
| RPM | Reads Per Million |
References
- Wu, H.; Peisley, A.; Graef, I.A.; Crabtree, G.R. NFAT signaling and the invention of vertebrates. Trends Cell Biol. 2007, 17, 251–260. [Google Scholar] [CrossRef] [PubMed]
- Rao, A.; Luo, C.; Hogan, P.G. Transcription factors of the NFAT family: Regulation and function. Annu. Rev. Immunol. 1997, 15, 707–747. [Google Scholar] [CrossRef] [PubMed]
- Hogan, P.G.; Chen, L.; Nardone, J.; Rao, A. Transcriptional regulation by calcium, calcineurin, and NFAT. Genes Dev. 2003, 17, 2205–2232. [Google Scholar] [CrossRef] [PubMed]
- Macian, F. NFAT proteins: Key regulators of T-cell development and function. Nat. Rev. Immunol. 2005, 5, 472–484. [Google Scholar] [CrossRef] [PubMed]
- Fric, J.; Zelante, T.; Wong, A.Y.W.; Mertes, A.; Yu, H.-B.; Ricciardi-Castagnoli, P. NFAT control of innate immunity. Blood 2012, 120, 1380–1389. [Google Scholar] [CrossRef] [PubMed]
- Fric, J.; Lim, C.X.F.; Koh, E.G.L.; Hofmann, B.; Chen, J.; Tay, H.S.; Mohammad Isa, S.A.B.; Mortellaro, A.; Ruedl, C.; Ricciardi-Castagnoli, P. Calcineurin/NFAT signalling inhibits myeloid haematopoiesis. EMBO Mol. Med. 2012, 4, 269–282. [Google Scholar] [CrossRef] [PubMed]
- Bartel, D.P. Review MicroRNAs : Target Recognition and Regulatory Functions. Cell 2009, 136, 215–233. [Google Scholar] [CrossRef] [PubMed]
- Kozomara, A.; Griffiths-Jones, S. miRBase: Integrating microRNA annotation and deep-sequencing data. Nucleic Acids Res. 2011, 39, D152–D157. [Google Scholar] [CrossRef] [PubMed]
- Davis, B.N.; Hilyard, A.C.; Lagna, G.; Hata, A. SMAD proteins control DROSHA-mediated microRNA maturation. Nature 2008, 454, 56–61. [Google Scholar] [CrossRef] [PubMed]
- Avraham, R.; Yarden, Y. Feedback regulation of EGFR signalling: Decision making by early and delayed loops. Nat. Rev. Mol. Cell Biol. 2011, 12, 104–117. [Google Scholar] [CrossRef] [PubMed]
- Kang, K.; Peng, X.; Zhang, X.; Wang, Y.; Zhang, L.; Gao, L.; Weng, T.; Zhang, H.; Ramchandran, R.; Raj, J.U.; et al. MicroRNA-124 Suppresses the Transactivation of Nuclear Factor of Activated T Cells by Targeting Multiple Genes and Inhibits the Proliferation of Pulmonary Artery Smooth Muscle Cells. J. Biol. Chem. 2013, 288, 25414–25427. [Google Scholar] [CrossRef] [PubMed]
- Kuchen, S.; Resch, W.; Yamane, A.; Kuo, N.; Li, Z.; Chakraborty, T.; Wei, L.; Laurence, A.; Yasuda, T.; Peng, S.; et al. Regulation of microRNA expression and abundance during lymphopoiesis. Immunity 2010, 32, 828–839. [Google Scholar] [CrossRef] [PubMed]
- Ceppi, M.; Pereira, P.M.; Dunand-Sauthier, I.; Barras, E.; Reith, W.; Santos, M.A.; Pierre, P. MicroRNA-155 modulates the interleukin-1 signaling pathway in activated human monocyte-derived dendritic cells. Proc. Natl. Acad. Sci. USA 2009, 106, 2735–2740. [Google Scholar] [CrossRef] [PubMed]
- Yang, G.; Yang, L.; Zhao, Z.; Wang, J.; Zhang, X. Signature miRNAs Involved in the Innate Immunity of Invertebrates. PLoS ONE 2012, 7, e39015. [Google Scholar] [CrossRef] [PubMed]
- Weitzel, R.P.; Lesniewski, M.L.; Haviernik, P.; Kadereit, S.; Leahy, P.; Greco, N.J.; Laughlin, M.J. MicroRNA 184 regulates expression of NFAT1 in umbilical cord blood CD4 + T cells. Blood 2009, 113, 6648–6657. [Google Scholar] [CrossRef] [PubMed]
- Li, W.; Kong, L.-B.; Li, J.-T.; Guo, Z.-Y.; Xue, Q.; Yang, T.; Meng, Y.-L.; Jin, B.-Q.; Wen, W.-H.; Yang, A.-G. MiR-568 inhibits the activation and function of CD4+ T cells and Treg cells by targeting NFAT5. Int. Immunol. 2013, 1, 1–13. [Google Scholar] [CrossRef] [PubMed]
- Zeng, Y.; Wang, Y.; Wu, Z.; Kang, K.; Peng, X.; Peng, W.; Qu, J.; Liu, L.; Raj, J.U.; Gou, D. MicroRNA-9 enhances the transactivation of nuclear factor of activated T cells by targeting KPNB1 and DYRK1B. Am. J. Physiol. Cell Physiol. 2015. [Google Scholar] [CrossRef] [PubMed]
- Lin, Z.; Murtaza, I.; Wang, K.; Jiao, J.; Gao, J.; Li, P.-F. miR-23a functions downstream of NFATc3 to regulate cardiac hypertrophy. Proc. Natl. Acad. Sci. USA 2009, 106, 12103–12108. [Google Scholar] [CrossRef] [PubMed]
- Ucar, A.; Gupta, S.K.; Fiedler, J.; Erikci, E.; Kardasinski, M.; Batkai, S.; Dangwal, S.; Kumarswamy, R.; Bang, C.; Holzmann, A.; et al. The miRNA-212/132 family regulates both cardiac hypertrophy and cardiomyocyte autophagy. Nat. Commun. 2012, 3, 1078. [Google Scholar] [CrossRef] [PubMed]
- Li, Q.; Lin, X.; Yang, X.; Chang, J. NFATc4 is negatively regulated in miR-133a-mediated cardiomyocyte hypertrophic repression. Am. J. Physiol. Heart Circ. Physiol. 2010, 298, H1340–H1347. [Google Scholar] [CrossRef] [PubMed]
- Yoo, A.S.; Greenwald, I. LIN-12/Notch activation leads to microRNA-mediated down-regulation of Vav in C. elegans. Science 2005, 310, 1330–1333. [Google Scholar] [CrossRef] [PubMed]
- Cook, I.H.; Evans, J.; Maldonado-Pérez, D.; Critchley, H.O.; Sales, K.J.; Jabbour, H.N. Prokineticin-1 (PROK1) modulates interleukin (IL)-11 expression via prokineticin receptor 1 (PROKR1) and the calcineurin/NFAT signalling pathway. Mol. Hum. Reprod. 2009, 16, 158–169. [Google Scholar] [CrossRef] [PubMed]
- Song, D.W.; Ryu, J.Y.; Kim, J.O.; Kwon, E.J.; Kim, D.H. The miR-19a / b family positively regulates cardiomyocyte hypertrophy by targeting atrogin-1 and MuRF-1. Biochem. J. 2014, 162, 151–162. [Google Scholar]
- Dirkx, E.; Gladka, M.M.; Philippen, L.E.; Armand, A.; Kinet, V.; Leptidis, S.; Azzouzi, H.; Salic, K.; Bourajjaj, M.; Silva, G.J.J.; et al. De Nfat and miR-25 cooperate to reactivate the transcription factor Hand2 in heart failure. Nat. Cell Biol. 2013, 15, 1282–1293. [Google Scholar] [CrossRef] [PubMed]
- Da Costa Martins, P.A.; Salic, K.; Gladka, M.M.; Armand, A.-S.; Leptidis, S.; el Azzouzi, H.; Hansen, A.; Coenen-de Roo, C.J.; Bierhuizen, M.F.; van der Nagel, R.; et al. MicroRNA-199b targets the nuclear kinase Dyrk1a in an auto-amplification loop promoting calcineurin/NFAT signalling. Nat. Cell Biol. 2010, 12, 1220–1227. [Google Scholar] [CrossRef] [PubMed]
- Baek, D.; Villén, J.; Shin, C.; Camargo, F.D.; Gygi, S.P.; Bartel, D.P. The impact of microRNAs on protein output. Nature 2008, 455, 64–71. [Google Scholar] [CrossRef] [PubMed]
- Betel, D.; Wilson, M.; Gabow, A.; Marks, D.S.; Sander, C. The microRNA.org resource: Targets and expression. Nucleic Acids Res. 2008, 36, D149–D153. [Google Scholar] [CrossRef] [PubMed]
- Hsu, S.-D.; Lin, F.-M.; Wu, W.-Y.; Liang, C.; Huang, W.-C.; Chan, W.-L.; Tsai, W.-T.; Chen, G.-Z.; Lee, C.-J.; Chiu, C.-M.; et al. miRTarBase: A database curates experimentally validated microRNA–target interactions. Nucleic Acids Res. 2011, 39, D163–D169. [Google Scholar] [CrossRef] [PubMed]
- Lewis, B.P.; Burge, C.B.; Bartel, D.P. Conserved seed pairing, often flanked by adenosines, indicates that thousands of human genes are microRNA targets. Cell 2005, 120, 15–20. [Google Scholar] [CrossRef] [PubMed]
- Yu, S.; Kim, J.; Min, H.; Yoon, S. Ensemble Learning can Significantly Improve Human microRNA Target Prediction. Methods 2014. [Google Scholar] [CrossRef] [PubMed]
- Ahmadi, H.; Ahmadi, A.; Azimzadeh-Jamalkandi, S.; Shoorehdeli, M.A.; Salehzadeh-Yazdi, A.; Bidkhori, G.; Masoudi-Nejad, A. HomoTarget: A new algorithm for prediction of microRNA targets in Homo sapiens. Genomics 2013, 101, 94–100. [Google Scholar] [CrossRef] [PubMed]
- Barski, A.; Jothi, R.; Cuddapah, S.; Cui, K.; Roh, T.-Y.; Schones, D.E.; Zhao, K. Chromatin poises miRNA- and protein-coding genes for expression. Genome Res. 2009, 19, 1742–1751. [Google Scholar] [CrossRef] [PubMed]
- Clipstone, N.A.; Crabtree, G.R. Identification of calcineurin as a key signaling enzyme in lymphocyte-t activation. Nature 1992, 357, 695–697. [Google Scholar] [CrossRef] [PubMed]
- Huang, G.N.; Huso, D.L.; Bouyain, S.; Tu, J.; McCorkell, K.A.; May, M.J.; Zhu, Y.; Lutz, M.; Collins, S.; Dehoff, M.; et al. NFAT binding and regulation of T cell activation by the cytoplasmic scaffolding Homer proteins. Science 2008, 319, 476–481. [Google Scholar] [CrossRef] [PubMed]
- Liu, Q.; Busby, J.C.; Molkentin, J.D. Interaction between TAK1-TAB1-TAB2 and RCAN1-calcineurin defines a signalling nodal control point. Nat. Cell Biol. 2009, 11, 154–161. [Google Scholar] [CrossRef] [PubMed]
- Jang, H.; Cho, E.-J.; Youn, H.-D. A new calcineurin inhibition domain in Cabin1. Biochem. Biophys. Res. Commun. 2007, 359, 129–135. [Google Scholar] [CrossRef] [PubMed]
- Perrino, B.A.; Martin, B.A. Ca(2+)- and myristoylation-dependent association of calcineurin with phosphatidylserine. J. Biochem. 2001, 129, 835–841. [Google Scholar] [CrossRef] [PubMed]
- Crabtree, G.R.; Schreiber, S.L. NIH Public Access. 2010, 138, 1–4. [Google Scholar]
- Yang, J.H.; Li, J.H.; Shao, P.; Zhou, H.; Chen, Y.Q.; Qu, L.H. StarBase: A database for exploring microRNA-mRNA interaction maps from Argonaute CLIP-Seq and Degradome-Seq data. Nucleic Acids Res. 2011, 39, 202–209. [Google Scholar] [CrossRef] [PubMed]
- starBase. Available online: http://starbase.sysu.edu.cn/index.php (accessed on 1 September 2013).
- Agarwal, V.; Bell, G.W.; Nam, J.W.; Bartel, D.P. Predicting effective microRNA target sites in mammalian mRNAs. Elife 2015, 4. [Google Scholar] [CrossRef] [PubMed]
- TargetScanHuman. Available online: http://www.targetscan.org (accessed on 1 August 2015).
- Wong, N.; Wang, X. miRDB: An online resource for microRNA target prediction and functional annotations. Nucleic Acids Res. 2015, 43, D146–D152. [Google Scholar] [CrossRef] [PubMed]
- miRDB. Available online: http://mirdb.org/miRDB/index.html (accessed on 3 May 2015).
- Vlachos, I.S.; Paraskevopoulou, M.D.; Karagkouni, D.; Georgakilas, G.; Vergoulis, T.; Kanellos, I.; Anastasopoulos, I.L.; Maniou, S.; Karathanou, K.; Kalfakakou, D.; et al. DIANA-TarBase v7.0: Indexing more than half a million experimentally supported miRNA:mRNA interactions. Nucleic Acids Res. 2015, 43, D153–D159. [Google Scholar] [CrossRef] [PubMed]
- TarBase. Available online: http://diana.imis.athena-innovation.gr/DianaTools/index.php?r=tarbase/index (accessed on 1 January 2012).
- Maragkakis, M.; Vergoulis, T.; Alexiou, P.; Reczko, M.; Plomaritou, K.; Gousis, M.; Kourtis, K.; Koziris, N.; Dalamagas, T.; Hatzigeorgiou, A.G. DIANA-microT Web server upgrade supports Fly and Worm miRNA target prediction and bibliographic miRNA to disease association. Nucleic Acids Res. 2011, 39. [Google Scholar] [CrossRef] [PubMed]
- DIANA. Available online: http://diana.imis.athena-innovation.gr/DianaTools/index.php (accessed on 30 April 2009).
- Hsu, S.D.; Chu, C.H.; Tsou, A.P.; Chen, S.J.; Chen, H.C.; Hsu, P.W.C.; Wong, Y.H.; Chen, Y.H.; Chen, G.H.; Huang, H. Da miRNAMap 2.0: Genomic maps of microRNAs in metazoan genomes. Nucleic Acids Res. 2008, 36. [Google Scholar]
- miRNAMap. Available online: http://mirnamap.mbc.nctu.edu.tw/index.php (accessed on 19 November 2007).
- Min, H.; Yoon, S. Got target?: Computational methods for microRNA target prediction and their extension. Exp. Mol. Med. 2010, 42, 233–244. [Google Scholar] [CrossRef] [PubMed]
- Shaw, J.P.; Utz, P.J.; Durand, D.B.; Toole, J.J.; Emmel, E.A.; Crabtree, G.R. Identification of a putative regulator of early T cell activation genes. Science 1988, 241, 202–205. [Google Scholar] [CrossRef]
- Baek, K.-H.; Zaslavsky, A.; Lynch, R.C.; Britt, C.; Okada, Y.; Siarey, R.J.; Lensch, M.W.; Park, I.-H.; Yoon, S.S.; Minami, T.; et al. Down’s syndrome suppression of tumour growth and the role of the calcineurin inhibitor DSCR1. Nature 2009, 459, 1126–1130. [Google Scholar] [CrossRef] [PubMed]
- Beurel, E.; Michalek, S.M.; Jope, R.S. Innate and adaptive immune responses regulated by glycogen synthase kinase-3 (GSK3). Trends Immunol 2010, 31, 24–31. [Google Scholar] [CrossRef] [PubMed]
- Becker, W.; Sippl, W. Activation, regulation, and inhibition of DYRK1A. FEBS J. 2011, 278, 246–256. [Google Scholar] [CrossRef] [PubMed]
- Schorge, S.; van de Leemput, J.; Singleton, A.; Houlden, H.; Hardy, J. Human ataxias: A genetic dissection of inositol triphosphate receptor (ITPR1)-dependent signaling. Trends Neurosci. 2010, 33, 211–219. [Google Scholar] [CrossRef] [PubMed]
- Sharma, S.; Findlay, G.M.; Bandukwala, H.S.; Oberdoerffer, S.; Baust, B.; Li, Z.; Schmidt, V.; Hogan, P.G.; Sacks, D.B.; Rao, A. Dephosphorylation of the nuclear factor of activated T cells (NFAT) transcription factor is regulated by an RNA-protein scaffold complex. Proc. Natl. Acad. Sci. USA 2011, 108, 11381–11386. [Google Scholar] [CrossRef] [PubMed]
- Liu, Z.; Lee, J.; Krummey, S.; Lu, W.; Cai, H.; Lenardo, M.J. The kinase LRRK2 is a regulator of the transcription factor NFAT that modulates the severity of inflammatory bowel disease. Nat. Immunol. 2011, 12, 1063–1070. [Google Scholar] [CrossRef] [PubMed]
- Li, H.; Stein, A.; Dell’Acqua, M.L.; Pink, M.D.; Murphy, J.G.; Hogan, P.G. Balanced interactions of calcineurin with AKAP79 regulate Ca2+–calcineurin–NFAT signaling. Nat. Struct. Mol. Biol. 2012, 19, 337–345. [Google Scholar] [CrossRef] [PubMed]
- Hou, X.; Chen, J.; Luo, Y.; Liu, F.; Xu, G.; Gao, Y. Silencing of STIM1 attenuates hypoxia-induced PASMCs proliferation via inhibition of the SOC/Ca2+/NFAT pathway. Respir. Res. 2013, 14, 2. [Google Scholar] [CrossRef] [PubMed]
- Srikanth, S.; Gwack, Y. Orai1-NFAT signalling pathway triggered by T cell receptor stimulation. Mol. Cells 2013, 35, 182–194. [Google Scholar] [CrossRef] [PubMed]
- miRTarBase. Available online: http://mirtarbase.mbc.nctu.edu.tw/index.php (accessed on 30 October 2015).
- miRmine. Available online: http://guanlab.ccmb.med.umich.edu/mirmine (accessed on 1 January 2016).
- Lee, Y.; Jeon, K.; Lee, J.-T.; Kim, S.; Kim, V.N. MicroRNA maturation: Stepwise processing and subcellular localization. EMBO J. 2002, 21, 4663–4670. [Google Scholar] [CrossRef] [PubMed]
- Lee, Y.; Kim, M.; Han, J.; Yeom, K.-H.; Lee, S.; Baek, S.H.; Kim, V.N. MicroRNA genes are transcribed by RNA polymerase II. EMBO J. 2004, 23, 4051–4060. [Google Scholar] [CrossRef] [PubMed]
- Messeguer, X.; Escudero, R.; Farré, D.; Núñez, O.; Martínez, J.; Albà, M.M. PROMO: Detection of known transcription regulatory elements using species-tailored searches. Bioinformatics 2002, 18, 333–334. [Google Scholar] [CrossRef] [PubMed]
- Yang, J.H.; Li, J.H.; Jiang, S.; Zhou, H.; Qu, L.H. ChIPBase: A database for decoding the transcriptional regulation of long non-coding RNA and microRNA genes from ChIP-Seq data. Nucleic Acids Res. 2013, 41, D177–D187. [Google Scholar] [CrossRef] [PubMed]
- ChiPBase. Available online: http://deepbase.sysu.edu.cn/chipbase/index.php (accessed on 1 November 2012).
- Kent, W.J.; Sugnet, C.W.; Furey, T.S.; Roskin, K.M.; Pringle, T.H.; Zahler, A.M.; Haussler, D. The Human Genome Browser at UCSC. Genome Res. 2002, 12, 996–1006. [Google Scholar] [CrossRef] [PubMed]
- UCSC Genome Bioinformatics. Available online: http://genome.ucsc.edu/index.html (accessed on 1 May 2005).
- TFSEARCH. Available online: http://diyhpl.us/~bryan/irc/protocol-online/protocol-cache/TFSEARCH.html (accessed on 1 April 1998).
- PROMO. Available online: http://alggen.lsi.upc.es/cgi-bin/promo_v3/promo/promoinit.cgi?dirDB=TF_8.3 (accessed on 18 February 2002).
- Becskei, A.; Serrano, L. Engineering stability in gene networks by autoregulation. Nature 2000, 405, 590–593. [Google Scholar] [CrossRef] [PubMed]
- Harris, S.L.; Levine, A.J. The p53 pathway: Positive and negative feedback loops. Oncogene 2005, 24, 2899–2908. [Google Scholar] [CrossRef] [PubMed]
- Rosenfeld, N.; Elowitz, M.B.; Alon, U. Negative autoregulation speeds the response times of transcription networks. J. Mol. Biol. 2002, 323, 785–793. [Google Scholar] [CrossRef]
- Kumar, M.; Sahu, S.K.; Kumar, R.; Subuddhi, A.; Maji, R.K.; Jana, K.; Gupta, P.; Raffetseder, J.; Lerm, M.; Ghosh, Z.; et al. MicroRNA let-7 modulates the immune response to mycobacterium tuberculosis infection via control of A20, an inhibitor of the NF-κB pathway. Cell Host Microbe 2015, 17, 345–356. [Google Scholar] [CrossRef] [PubMed]
- Jacob, F.; Monod, J. Genetic regulatory mechanisms in the synthesis of proteins. J. Mol. Biol. 1961, 3, 318–356. [Google Scholar] [CrossRef]
- Helwak, A.; Kudla, G.; Dudnakova, T.; Tollervey, D. Mapping the human miRNA interactome by CLASH reveals frequent noncanonical binding. Cell 2013, 153, 654–665. [Google Scholar] [CrossRef] [PubMed]
- Rey-Giraud, F.; Hafner, M.; Ries, C.H. In vitro generation of monocyte-derived macrophages under serum-free conditions improves their tumor promoting functions. PLoS ONE 2012, 7, e42656. [Google Scholar] [CrossRef] [PubMed]
- Tsitsiou, E.; Lindsay, M.A. microRNAs and the immune response. Curr. Opin. Pharmacol. 2009, 9, 514–520. [Google Scholar] [CrossRef] [PubMed]
- Bazzoni, F.; Rossato, M.; Fabbri, M.; Gaudiosi, D.; Mirolo, M.; Mori, L.; Tamassia, N.; Mantovani, A.; Cassatella, M.A.; Locati, M. Induction and regulatory function of miR-9 in human monocytes and neutrophils exposed to proinflammatory signals. Proc. Natl. Acad. Sci. USA 2009, 106, 5282–5287. [Google Scholar] [CrossRef] [PubMed]
- Thai, T.-H.; Calado, D.P.; Casola, S.; Ansel, K.M.; Xiao, C.; Xue, Y.; Murphy, A.; Frendewey, D.; Valenzuela, D.; Kutok, J.L.; et al. Regulation of the germinal center response by microRNA-155. Science 2007, 316, 604–608. [Google Scholar] [CrossRef] [PubMed]
- Rothenberg Melissa, K.; Mingler, M.E.; Cole, E.T.; Orkin, S.H.; Thomas, X.; Lu, B.J.; Hartner, J.; Lim, E.-J.; Fabry, V.; Lu, T.X.; et al. Delayed-Type Hypersensitivity MicroRNA-21 Limits in Vivo Immune Response-Mediated Activation of the IL-12/IFN-g Pathway, Th1 Polarization, and the Severity of Delayed-Type Hypersensitivity. J. Immunol. 2011, 187, 3362–3373. [Google Scholar]
- Gabriely, G.; Wurdinger, T.; Kesari, S.; Esau, C.C.; Burchard, J.; Linsley, P.S.; Krichevsky, A.M. MicroRNA 21 promotes glioma invasion by targeting matrix metalloproteinase regulators. Mol. Cell. Biol. 2008, 28, 5369–5380. [Google Scholar] [CrossRef] [PubMed]
- Liu, G.; Min, H.; Yue, S.; Chen, C.Z. Pre-miRNA loop nucleotides control the distinct activities of mir-181a-1 and mir-181c in early T cell development. PLoS ONE 2008, 3. [Google Scholar] [CrossRef] [PubMed]
- Kishore, S.; Jaskiewicz, L.; Burger, L.; Hausser, J.; Khorshid, M.; Zavolan, M. A quantitative analysis of CLIP methods for identifying binding sites of RNA-binding proteins. Nat. Methods 2011, 8, 559–564. [Google Scholar] [CrossRef] [PubMed]

| Gene | Description | |
|---|---|---|
| NFATc1 | Family of transcription factors that are expressed in the vertebrates. These transcription factors induce expression of cytokine genes important for immune response. | [38,52] |
| NFATc2 | ||
| NFATc3 | ||
| NFATc4 | ||
| PPP3CA | Calcium-dependent, calmodulin-stimulated protein phosphatase. It dephosphorylates the NFAT transcription factors and facilitates nuclear transport of NFAT. | [33] |
| PPP3CB | ||
| PPP3R1 | ||
| PPP3R2 | ||
| RCAN1 | Transcribed by NFAT transcription factor and interacts with calcineurin to inhibit calcineurin-dependent signaling pathways. | [38,53] |
| GSK3B | Phosphorylates NFAT into its inactive form and facilitates the cytoplasmic transport of NFAT. | [38,54,55] |
| DYRK1A | ||
| HOMER2 | Is a negative regulator of T cell activation and binds with NFAT to compete with calcineurin binding. | [34] |
| ITPR1 | Intracellular channel that mediates calcium release from the endoplasmic reticulum. | [38,56] |
| MAP3K7 | Selectively induces calcineurin-NFAT signaling through direct phosphorylation of RCAN1. | [35] |
| TAB1 | Phosphorylated RCAN1 and inhibits RCAN1 from binding to calcineurin. | [35] |
| TAB2 | ||
| CSNK1A1/CK1 | Works as a negative inhibitor of the NFAT by phosphorylation and mediates its cytoplasmic translocation. | [57] |
| LRRK2 | Negative regulator of the transcription factor NFAT and is a component of a complex that includes the large non-coding RNA NRON (an NFAT repressor). | [58] |
| CABIN1 | Binds specifically to the activated form of calcineurin and inhibits calcineurin-mediated signal transduction and NFAT nuclear translocation. | [36] |
| AKAP5 | Inhibits calcineurin-dependent dephosphorylation of NFAT | [59] |
| CALM1 | Calmodulin is a calcium binding protein that activates the calcineurin. | [37] |
| STIM1 | Ca2+ sensor proteins on the endoplasmic reticulum, which is an indispensable part in the activation of store-operated Ca2+ channels (SOC). | [60] |
| ORAI1 | Calcium channel subunit that is activated by the calcium sensor STIM1 when calcium stores are depleted. | [61] |
| miRNAs Involved in Feedback Loops | Target Genes | miRNA Average Expression in Activated T Cells (RPM) |
|---|---|---|
| hsa-miR-21-3p | CALM1 | 191 |
| hsa-let-7b-5p | NFATC1 | 8617 |
| hsa-miR-17-5p | PPP3R1 | 778 |
| hsa-miR-17-3p | STIM1 | 859 |
| hsa-miR-19a-3p | ITPR1 | 4379 |
| hsa-miR-92b-3p | ITPR1 | 254 |
| hsa-miR-21-5p | CSNK1A1 | 372,670 |
| hsa-miR-181c-5p | CSNK1A1 | 321 |
| hsa-let-7c-5p | GSK3A | 889 |
| hsa-let-7b-3p | GSK3B | 77 |
| hsa-miR-155-5p | GSK3B | 282,932 |
© 2016 by the author; licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC-BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Kannambath, S. Micro-RNA Feedback Loops Modulating the Calcineurin/NFAT Signaling Pathway. Non-Coding RNA 2016, 2, 3. https://doi.org/10.3390/ncrna2020003
Kannambath S. Micro-RNA Feedback Loops Modulating the Calcineurin/NFAT Signaling Pathway. Non-Coding RNA. 2016; 2(2):3. https://doi.org/10.3390/ncrna2020003
Chicago/Turabian StyleKannambath, Shichina. 2016. "Micro-RNA Feedback Loops Modulating the Calcineurin/NFAT Signaling Pathway" Non-Coding RNA 2, no. 2: 3. https://doi.org/10.3390/ncrna2020003
APA StyleKannambath, S. (2016). Micro-RNA Feedback Loops Modulating the Calcineurin/NFAT Signaling Pathway. Non-Coding RNA, 2(2), 3. https://doi.org/10.3390/ncrna2020003





