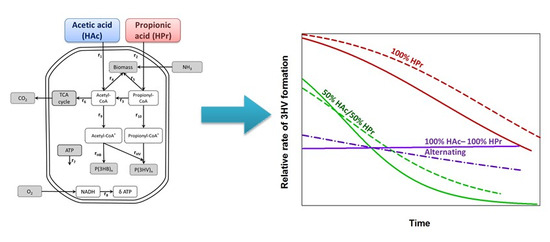
The Evolution of Polymer Composition during PHA Accumulation: The Significance of Reducing Equivalents
Abstract
:1. Introduction
2. Materials and Methods
2.1. Experimental Set-Up
2.2. Fed-Batch PHA Production
2.3. Analytical Methods
2.4. Experimental Design for PHA Accumulations
2.5. Rate and Yield Calculations
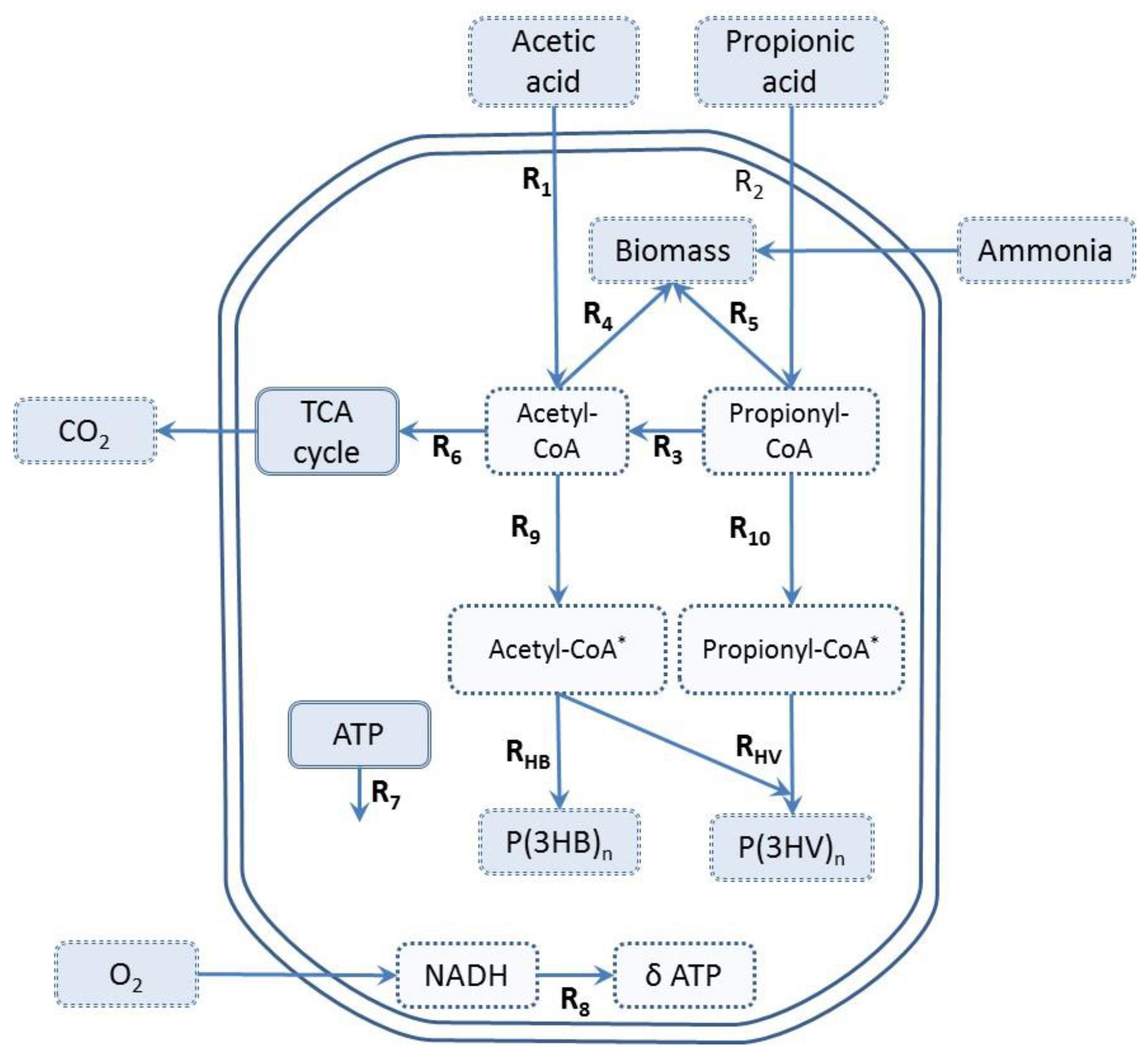
2.6. Metabolic Flux Analysis (MFA)
- -
- Active biomass can be formed either from acetyl-CoA or propionyl-CoA. Previous models were performed under ammonia limiting conditions with negligible cellular growth [4,13]. In the present work, we assumed that fluxes of acetyl-CoA and propionyl-CoA used for active biomass () synthesis are proportional to the consumption rate of acetic and propionic acid () [16].
- -
- Reactions R9 and R10 are reversible, the rest are irreversible reactions.
- -
- The maintenance requirement () was an estimated flux while the P/O ratio was fixed (δ = 3).
- -
- PHA depolymerization was not considered.
3. Results and Discussion
3.1. Biomass Growth and PHA Content
3.2. Monomer Development
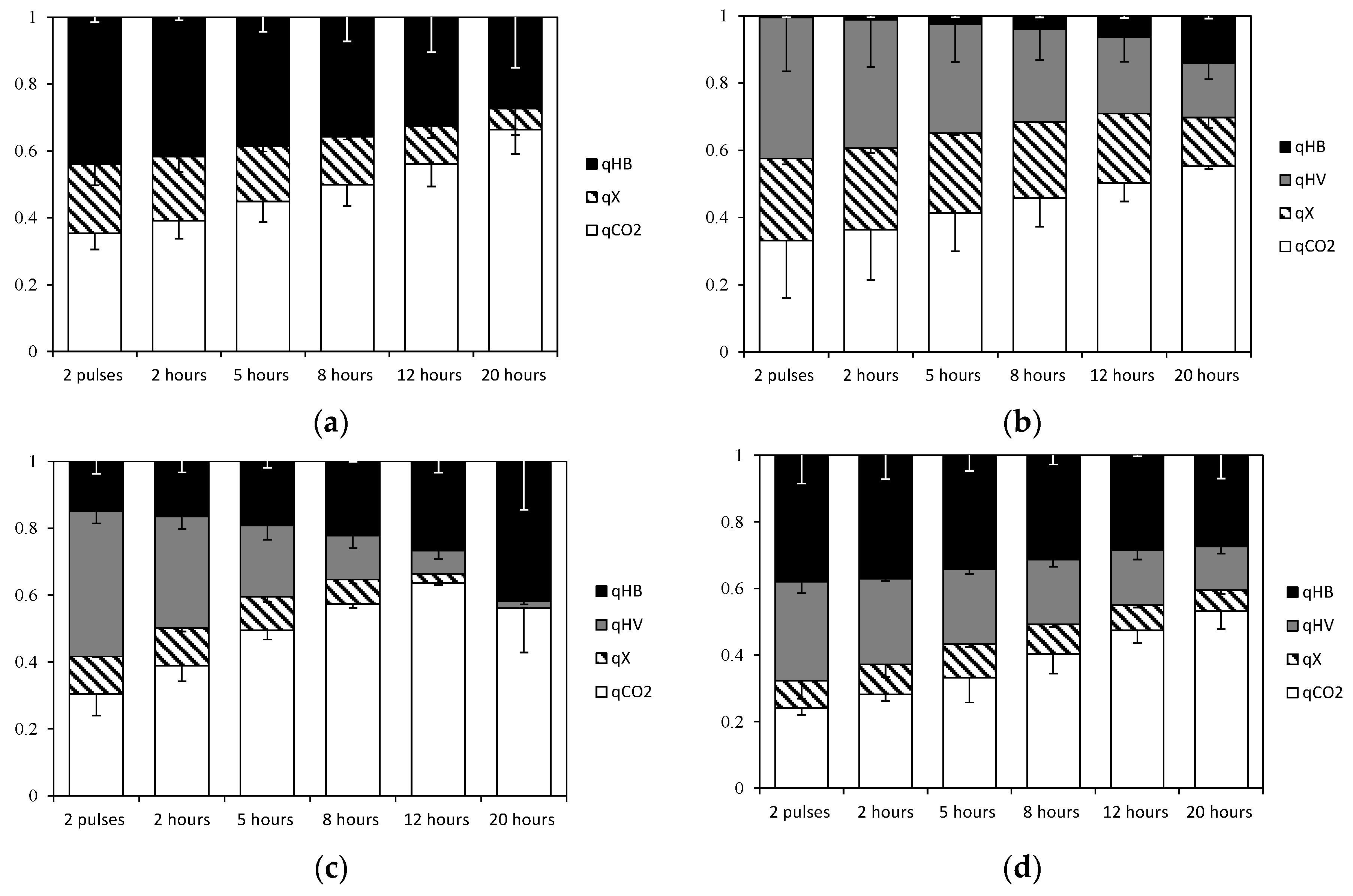
3.3. Carbon Flux to PHA, Biomass and CO2
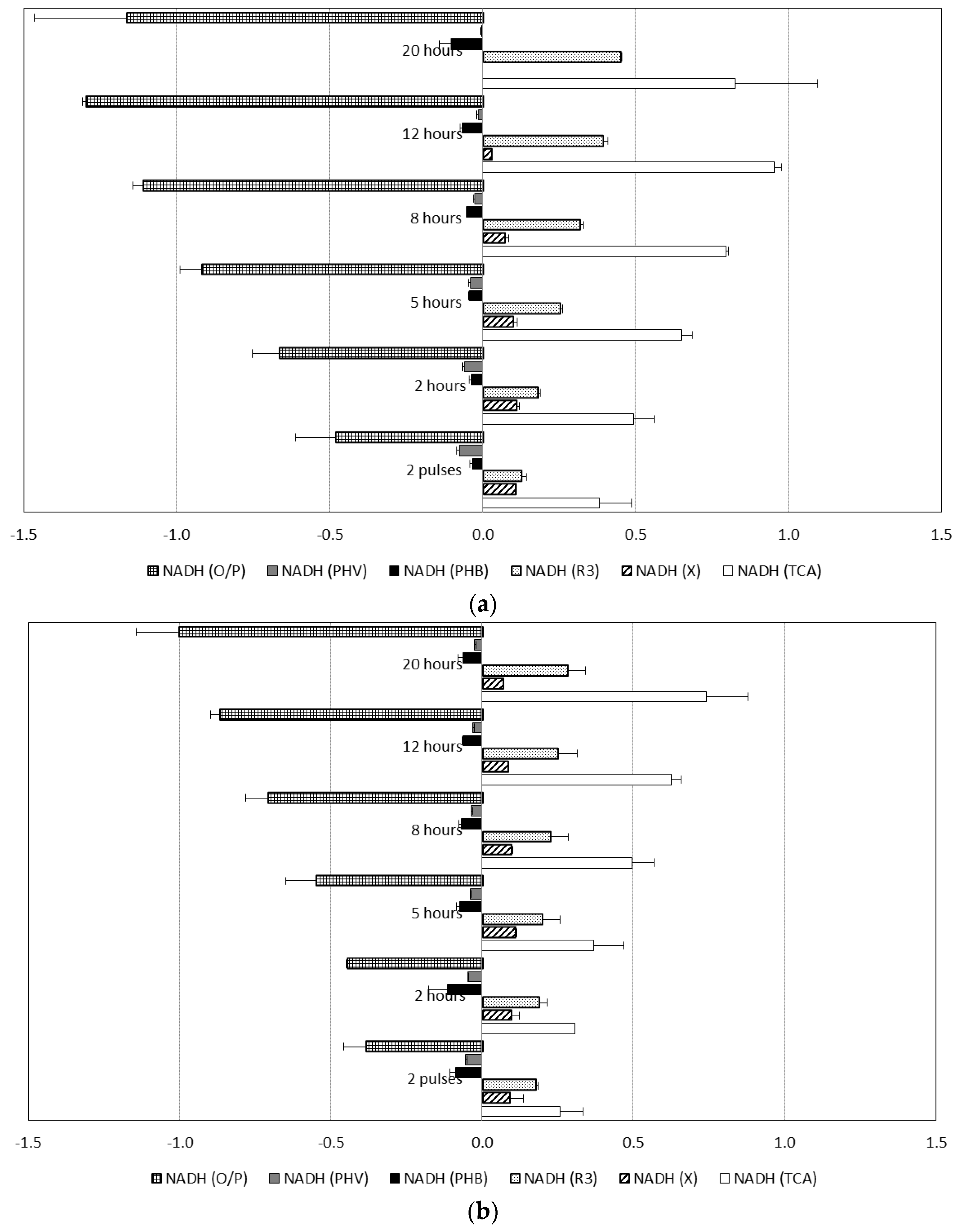
3.4. Pathways for Generation of Reducing Power
4. Conclusions
Supplementary Materials
Acknowledgments
Author Contributions
Conflicts of Interest
Abbreviations
| %3HV | 3HV fraction in total PHA (mol 3HV/mol PHA) |
| %3HVinst | Instantaneous 3HV content (mol 3HV/mol PHA) |
| %PHA | PHA intracellular content (gPHA/gVSS) |
| %PHA0 | Initial PHA intracellular content (gPHA/gVSS) |
| Fraction of propionic acid uptake to total carbon uptake flux | |
| 3HB | 3-Hydroxybutyrate |
| 3HV | 3-Hydroxyvalerate |
| AcCoA | Acetyl-CoA |
| ATP | Adenosine triphosphate |
| CO2 | Carbon dioxide |
| CoA | Coenzyme A |
| COD | Chemical oxygen demand |
| fPHA | PHA fraction with respect to active biomass (PHA/X) |
| HAc | Acetic acid |
| HPr | Propionic acid |
| NAD+ | Nicotinamide adenine dinucleotide (oxidised) |
| NADH | Nicotinamide adenine dinucleotide (reduced) |
| NADP+ | Nicotinamide adenine dinucleotide phosphate (oxidised) |
| NADPH | Nicotinamide adenine dinucleotide phosphate (reduced) |
| PHA | Poly(3-hydroxyalkanoate) or PHA concentration at certain time |
| PHA0 | Initial PHA concentration |
| PHB | Poly(3-hydroxybutyrate) |
| PHV | Poly(3-hydroxyvalerate) |
| PrCoA | Propionyl-CoA |
| qHAc | Acetic acid specific consumption rate (Cmol HAc/Cmol X·h) |
| qHB | Specific 3HB monomer synthesis rate (Cmol 3HB/Cmol X·h) |
| qHPr | Propionic acid specific consumption rate (Cmol HPr/Cmol X·h) |
| qHV | Specific 3HV monomer synthesis rate (Cmol 3HV/Cmol X·h) |
| qPHA | Specific PHA synthesis rate (Cmol PHA/Cmol X·h) |
| qS | Specific VFA consumption rate (Cmol VFA/Cmol X·h) |
| RQ | Respiratory quotient |
| TCA | Tricarboxylic acid cycle |
| v | Reaction rate (Cmol/Cmol X·h) |
| VFA | Volatile fatty acid |
| VSS | Volatile suspended solids |
| X | Active biomass |
| X0 | Initial active biomass |
References
- Laycock, B.; Halley, P.; Pratt, S.; Werker, A.; Lant, P. The chemomechanical properties of microbial polyhydroxyalkanoates. Prog. Polym. Sci. 2013, 38, 536–583. [Google Scholar] [CrossRef]
- Filipe, C.D.M.; Daigger, G.T.; Grady, C.P.L. A metabolic model for acetate uptake under anaerobic conditions by glycogen accumulating organisms: Stoichiometry, kinetics, and the effect of pH. Biotechnol. Bioeng. 2001, 76, 17–31. [Google Scholar] [CrossRef] [PubMed]
- Taguchi, K.; Taguchi, S.; Sudesh, K.; Maehara, A.; Tsuge, T.; Doi, Y. Metabolic pathways and engineering of polyhydroxyalkanoate biosynthesis. Biopolym. Online 2005, 3. [Google Scholar] [CrossRef]
- Dias, J.M.L.; Oehmen, A.; Serafim, L.S.; Lemos, P.C.; Reis, M.A.M.; Oliveira, R. Metabolic modelling of polyhydroxyalkanoate copolymers production by mixed microbial cultures. BMC Syst. Biol. 2008, 2, 59. [Google Scholar] [CrossRef] [PubMed]
- Lemos, P.C.; Serafim, L.S.; Santos, M.M.; Reis, M.A.M.; Santos, H. Metabolic pathway for propionate utilization by phosphorus-accumulating organisms in activated sludge: C-13 labeling and in vivo nuclear magnetic resonance. Appl. Environ. Microb. 2003, 69, 241–251. [Google Scholar] [CrossRef]
- Grousseau, E.; Blanchet, E.; Deleris, S.; Albuquerque, M.G.E.; Paul, E.; Uribelarrea, J.L. Impact of sustaining a controlled residual growth on polyhydroxybutyrate yield and production kinetics in Cupriavidus necator. Bioresour. Technol. 2013, 148, 30–38. [Google Scholar] [CrossRef] [PubMed]
- Gurieff, N. Production of Biodegradable Polyhydroxyalkanoate Polymers Using Advanced Biological Wastewater Treatment Process Technology. Ph.D. Thesis, The University of Queensland, Brisbane, Australia, 2007. [Google Scholar]
- Serafim, L.S.; Lemos, P.C.; Torres, C.; Reis, M.A.M.; Ramos, A.M. The influence of process parameters on the characteristics of polyhydroxyalkanoates produced by mixed cultures. Macromol. Biosci. 2008, 8, 355–366. [Google Scholar] [CrossRef] [PubMed]
- Albuquerque, M.G.E.; Martino, V.; Pollet, E.; Averous, L.; Reis, M.A.M. Mixed culture polyhydroxyalkanoate (PHA) production from volatile fatty acid (VFA)-rich streams: Effect of substrate composition and feeding regime on pha productivity, composition and properties. J. Biotechnol. 2011, 151, 66–76. [Google Scholar] [CrossRef] [PubMed]
- Ivanova, G.; Serafim, L.S.; Lemos, P.C.; Ramos, A.M.; Reis, M.A.M.; Cabrita, E.J. Influence of feeding strategies of mixed microbial cultures on the chemical composition and microstructure of copolyesters p(3HB-co-3HV) analyzed by NMR and statistical analysis. Magn. Reson. Chem. 2009, 47, 497–504. [Google Scholar] [CrossRef] [PubMed]
- Arcos-Hernandez, M.V.; Laycock, B.; Donose, B.C.; Pratt, S.; Halley, P.; Al-Luaibi, S.; Werker, A.; Lant, P.A. Physicochemical and mechanical properties of mixed culture polyhydroxyalkanoate (PHBV). Eur. Polym. J. 2013, 49, 904–913. [Google Scholar] [CrossRef]
- Johnson, K.; Van Loosdrecht, M.C.M.; Kleerebezem, R. Influence of ammonium on the accumulation of polyhydroxybutyrate (PHB) in aerobic open mixed cultures. J. Biotechnol. 2010, 147, 73–79. [Google Scholar] [CrossRef] [PubMed]
- Pardelha, F.; Albuquerque, M.G.E.; Reis, M.A.M.; Dias, J.M.L.; Oliveira, R. Flux balance analysis of mixed microbial cultures: Application to the production of polyhydroxyalkanoates from complex mixtures of volatile fatty acids. J. Biotechnol. 2012, 162, 336–345. [Google Scholar] [CrossRef] [PubMed]
- Serafim, L.S.; Lemos, P.C.; Oliveira, R.; Ramos, A.M.; Reis, M.A.M. High storage of PHB by mixed microbial cultures under aerobic dynamic feeding conditions. Eur. Symp. Environ. Biotechnol. 2004, 479–482. [Google Scholar]
- Valentino, F.; Karabegouic, L.; Majone, M.; Morgan-Sagastume, F.; Werker, A. Polyhydroxyalkanoate (PHA) storage within a mixed-culture biomass with simultaneous growth as a function of accumulation substrate nitrogen and phosphorus levels. Water Res. 2015, 77, 49–63. [Google Scholar] [CrossRef] [PubMed]
- Jiang, Y.; Hebly, M.; Kleerebezem, R.; Muyzer, G.; van Loosdrecht, M.C.M. Metabolic modeling of mixed substrate uptake for polyhydroxyalkanoate (PHA) production. Water Res. 2011, 45, 1309–1321. [Google Scholar] [CrossRef] [PubMed]
- Pardelha, F.; Albuquerque, M.G.E.; Reis, M.A.M.; Oliveira, R.; Dias, J.M.L. Dynamic metabolic modelling of volatile fatty acids conversion to polyhydroxyalkanoates by a mixed microbial culture. New Biotechnol. 2014, 31, 335–344. [Google Scholar] [CrossRef] [PubMed]
- Murugan Janarthanan, O.; Laycock, B.; Montano-Herrera, L.; Lu, Y.; Arcos-Hernandez, M.V.; Werker, A.; Pratt, S. Fluxes in PHA-storing microbial communities during enrichment and biopolymer accumulation processes. New Biotechnol. 2016, 33, 61–72. [Google Scholar] [CrossRef] [PubMed]
- Werker, A.G.; Bengtsson, S.O.H.; Karlsson, C.A.B. Method for Accumulation of Polyhydroxyalkanoates in Biomass with on-Line Monitoring for Feed Rate Control and Process Termination. WO 2011070544 A2, 16 June 2011. [Google Scholar]
- Morgan-Sagastume, F.; Pratt, S.; Karlsson, A.; Cirne, D.; Lant, P.; Werker, A. Production of volatile fatty acids by fermentation of waste activated sludge pre-treated in full-scale thermal hydrolysis plants. Bioresour. Technol. 2011, 102, 3089–3097. [Google Scholar] [CrossRef] [PubMed]
- American Public Health Association (APHA). Standard Methods for the Examination of Water and Wastewater; American Public Health Association: Washington, DC, USA, 1995. [Google Scholar]
- Motulsky, H.J. Prism5 Statistics Guide; Graphpad Software Inc.: San Diego, CA, USA, 2007. [Google Scholar]
- Zwietering, M.H.; Jongenburger, I.; Rombouts, F.M.; Vantriet, K. Modeling of the bacterial growth curve. Appl. Environ. Microb. 1990, 56, 1875–1881. [Google Scholar]
- Third, K.A.; Newland, M.; Cord-Ruwisch, R. The effect of dissolved oxygen on PHB accumulation in activated sludge cultures. Biotechnol. Bioeng. 2003, 82, 238–250. [Google Scholar] [CrossRef] [PubMed]
- Villadsen, J.; Nielsen, J.; Lidén, G. Bioreaction Engineering Principles, 3rd ed.; Springer: New York, NY, USA, 2011. [Google Scholar]
- Klamt, S.; Saez-Rodriguez, J.; Gilles, E.D. Structural and functional analysis of cellular networks with cellnetanalyzer. BMC Syst. Biol. 2007. [Google Scholar] [CrossRef] [PubMed]
- Stephanopolulos, G.N.; Aristidou, A.A.; Nielsen, J. Metabolic Engineering: Principles and Methodologies; Academic Press: San Diego, CA, USA, 1998. [Google Scholar]
- Escapa, I.F.; Garcia, J.L.; Buhler, B.; Blank, L.M.; Prieto, M.A. The polyhydroxyalkanoate metabolism controls carbon and energy spillage in Pseudomonas putida. Environ. Microbiol. 2012, 14, 1049–1063. [Google Scholar] [CrossRef] [PubMed]
- Ren, Q.; de Roo, G.; Ruth, K.; Witholt, B.; Zinn, M.; Thony-Meyer, L. Simultaneous accumulation and degradation of polyhydroxyalkanoates: Futile cycle or clever regulation? Biomacromolecules 2009, 10, 916–922. [Google Scholar] [CrossRef] [PubMed]
- Arias, S.; Bassas-Galia, M.; Molinari, G.; Timmis, K.N. Tight coupling of polymerization and depolymerization of polyhydroxyalkanoates ensures efficient management of carbon resources in Pseudomonas putida. Microb. Biotechnol. 2013, 6, 551–563. [Google Scholar] [CrossRef] [PubMed]
- Lefebvre, G.; Rocher, M.; Braunegg, G. Effects of low dissolved-oxygen concentrations on poly(3-hydroxybutyrate-co-3-hydroxyvalerate) production by Alcaligenes eutrophus. Appl. Environ. Microb. 1997, 63, 827–833. [Google Scholar]
- Lemos, P.C.; Serafim, L.S.; Reis, M.A.M. Synthesis of polyhydroxyalkanoates from different short-chain fatty acids by mixed cultures submitted to aerobic dynamic feeding. J. Biotechnol. 2006, 122, 226–238. [Google Scholar] [CrossRef] [PubMed]
- Yu, J.; Si, Y.T. Metabolic carbon fluxes and biosynthesis of polyhydroxyalkanoates in Ralstonia eutropha on short chain fatty acids. Biotechnol. Prog. 2004, 20, 1015–1024. [Google Scholar] [CrossRef] [PubMed]
- Kim, J.I.; Varner, J.D.; Ramkrishna, D. A hybrid model of anaerobic E. Coli GJT001: Combination of elementary flux modes and cybernetic variables. Biotechnol. Prog. 2008, 24, 993–1006. [Google Scholar] [CrossRef] [PubMed]
- Zeng, A.P.; Ross, A.; Deckwer, W.D. A method to estimate the efficiency of oxidative-phosphorylation and biomass yield from ATP of a facultative anaerobe in continuous culture. Biotechnol. Bioeng. 1990, 36, 965–969. [Google Scholar] [CrossRef] [PubMed]
- Shimizu, H.; Kozaki, Y.; Kodama, H.; Shioya, S. Maximum production strategy for biodegradable copolymer P(HB-co-HV) in fed-batch culture of Alcaligenes eutrophus. Biotechnol. Bioeng. 1999, 62, 518–525. [Google Scholar] [CrossRef]


| Experiment set | Substrate Composition and Feeding Strategy (gCOD Basis) | Experiment Label | Process Time (h) | Initial VSS (g·L−1) | Total substrate added (gCOD) | Feed Concentration (gCOD·L−1) | Total Number of Pulses | ||
|---|---|---|---|---|---|---|---|---|---|
| HAc | HPr | HAc | HPr | ||||||
| 1 | 100% Acetic acid | Exp 1 | 23.2 | 1.2 | 1374 | - | 96 | 138 | - |
| Exp 1′ | 21.9 | 1.4 | 1684 | - | 98 | 147 | - | ||
| 2 | 100% Propionic acid | Exp 2 | 23.3 | 1.2 | - | 939 | 106 | - | 77 |
| Exp 2′ | 24.6 | 1.7 | - | 1439 | 102 | - | 137 | ||
| 3 | 50% Acetic/50% propionic acid | Exp 3 | 24.2 | 1.5 | 600 | 600 | 98 | 95 | |
| Exp 3′ | 20.4 | 1.9 | 817 | 817 | 98 | 140 | |||
| 4 | 100% Acetic acid—100% propionic acid (alternating) | Exp 4 | 22.3 | 1.4 | 1429 | 982 | 101/103 | 106 | 105 |
| Exp 4′ | 22.5 | 1.7 | 570 | 552 | 94/97 | 57 | 56 | ||
| Experiment Label | Consumed (mol HAc/mol VFA) | Consumed (mol HPr/mol VFA) | %PHA Plateau (gPHA/gVSS) | %3HV (mol 3HV/mol PHA) at 20 h | YPHA/S (gCOD PHA/gCOD VFA) | YX/S (gCOD X/gCOD VFA) | μmax (h−1) | Final X/X0 (gCOD/gCOD) | (Cmol VFA/Cmol X·h) | (Cmol PHA/Cmol X·h) |
|---|---|---|---|---|---|---|---|---|---|---|
| Exp 1 | 1.0 | 0 | 0.56 ± 0.04 | 0 | 0.48 ± 0.02 | 0.17 ± 0.03 | 0.27 ± 0.03 | 2.20 | 0.75 | 0.33 |
| Exp 1′ | 1.0 | 0 | 0.48 ± 0.06 | 0 | 0.38 ± 0.04 | 0.15 ± 0.01 | 0.21 ± 0.02 | 2.28 | 0.80 | 0.36 |
| Exp 2 | 0 | 1.0 | 0.40 ± 0.04 | 74 | 0.31 ± 0.03 | 0.16 ± 0.03 | 0.18 ± 0.02 | 2.13 | 0.33 | 0.11 |
| Exp 2′ | 0 | 1.0 | 0.48 ± 0.03 | 80 | 0.40 ± 0.07 | 0.18 ± 0.03 | 0.13 ± 0.02 | 2.09 | 0.33 | 0.18 |
| Exp 3 | 0.64 | 0.36 | 0.48 ± 0.06 | 40 | 0.39 ± 0.03 | 0.17 ± 0.05 | 0.35 ± 0.05 | 1.88 | 0.63 | 0.19 |
| Exp 3′ | 0.64 | 0.36 | 0.52 ± 0.03 | 42 | 0.45 ± 0.03 | 0.12 ± 0.05 | 0.27 ± 0.05 | 1.75 | 0.40 | 0.19 |
| Exp 4 | 0.72 | 0.28 | 0.59 ± 0.03 | 34 | 0.52 ± 0.03 | 0.18 ± 0.02 | 0.22 ± 0.02 | 3.08 | 0.37 | 0.23 |
| Exp 4′ | 0.64 | 0.36 | 0.52 ± 0.06 | 36 | 0.49 ± 0.03 | 0.20 ± 0.02 | 0.19 ± 0.02 | 1.98 | 0.45 | 0.20 |
| Experiment Set | Elapsed Duration | %Conversion PrCoA to AcCoA (mmol/mmol) | RQ (Cmmol CO2/mol O2) | ATP Dissipated (molATP/Cmmol VFA Consumed) | |||
|---|---|---|---|---|---|---|---|
| 1 | 2 pulses | - | - | 1.19 | (0.03) | 0.57 | (0.45) |
| 2 h | - | - | 1.16 | (0.03) | 0.87 | (0.46) | |
| 5 h | - | - | 1.13 | (0.03) | 1.32 | (0.46) | |
| 8 h | - | - | 1.10 | (0.04) | 1.71 | (0.45) | |
| 12 h | - | - | 1.07 | (0.03) | 2.34 | (0.24) | |
| 20 h | - | - | 1.05 | (0.04) | 3.11 | (0.44) | |
| 2 | 2 pulses | 0.49 | (0.11) | 0.79 | (0.03) | 1.83 | (1.36) |
| 2 h | 0.51 | (0.10) | 0.80 | (0.02) | 2.09 | (1.19) | |
| 5 h | 0.55 | (0.07) | 0.81 | (0.01) | 2.50 | (0.90) | |
| 8 h | 0.59 | (0.05) | 0.81 | (0.01) | 2.84 | (0.67) | |
| 12 h | 0.65 | (0.03) | 0.82 | (0.01) | 3.21 | (0.43) | |
| 20 h | 0.75 | (0.01) | 0.83 | (0.01) | 3.68 | (0.08) | |
| 3 | 2 pulses | 0.27 | (0.04) | 1.08 | (0.06) | 0.62 | (0.38) |
| 2 h | 0.39 | (0.02) | 1.01 | (0.03) | 1.17 | (0.26) | |
| 5 h | 0.55 | (0.02) | 0.99 | (0.02) | 1.99 | (0.18) | |
| 8 h | 0.69 | (0.02) | 0.95 | (0.01) | 2.66 | (0.08) | |
| 12 h | 0.85 | (0.03) | 0.95 | (0.00) | 3.37 | (0.04) | |
| 20 h | 0.97 | (0.01) | 1.01 | (0.08) | 0.46 | (0.28) | |
| 4 | 2 pulses | 0.44 | (0.05) | 1.14 | (0.11) | 0.27 | (0.04) |
| 2 h | 0.46 | (0.01) | 1.10 | (0.10) | 0.46 | (0.19) | |
| 5 h | 0.48 | (0.07) | 1.05 | (0.08) | 0.76 | (0.59) | |
| 8 h | 0.54 | (0.07) | 1.01 | (0.05) | 1.30 | (0.49) | |
| 12 h | 0.60 | (0.07) | 0.99 | (0.03) | 1.83 | (0.35) | |
| 20 h | 0.67 | (0.05) | 0.97 | (0.01) | 2.22 | (0.68) | |
© 2017 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license ( http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Montano-Herrera, L.; Laycock, B.; Werker, A.; Pratt, S. The Evolution of Polymer Composition during PHA Accumulation: The Significance of Reducing Equivalents. Bioengineering 2017, 4, 20. https://doi.org/10.3390/bioengineering4010020
Montano-Herrera L, Laycock B, Werker A, Pratt S. The Evolution of Polymer Composition during PHA Accumulation: The Significance of Reducing Equivalents. Bioengineering. 2017; 4(1):20. https://doi.org/10.3390/bioengineering4010020
Chicago/Turabian StyleMontano-Herrera, Liliana, Bronwyn Laycock, Alan Werker, and Steven Pratt. 2017. "The Evolution of Polymer Composition during PHA Accumulation: The Significance of Reducing Equivalents" Bioengineering 4, no. 1: 20. https://doi.org/10.3390/bioengineering4010020