Simple Analysis of Lipid Inhibition Activity on an Adipocyte Micro-Cell Pattern Chip
Abstract
:1. Introduction
2. Materials and Methods
2.1. Materials
2.2. Characterization
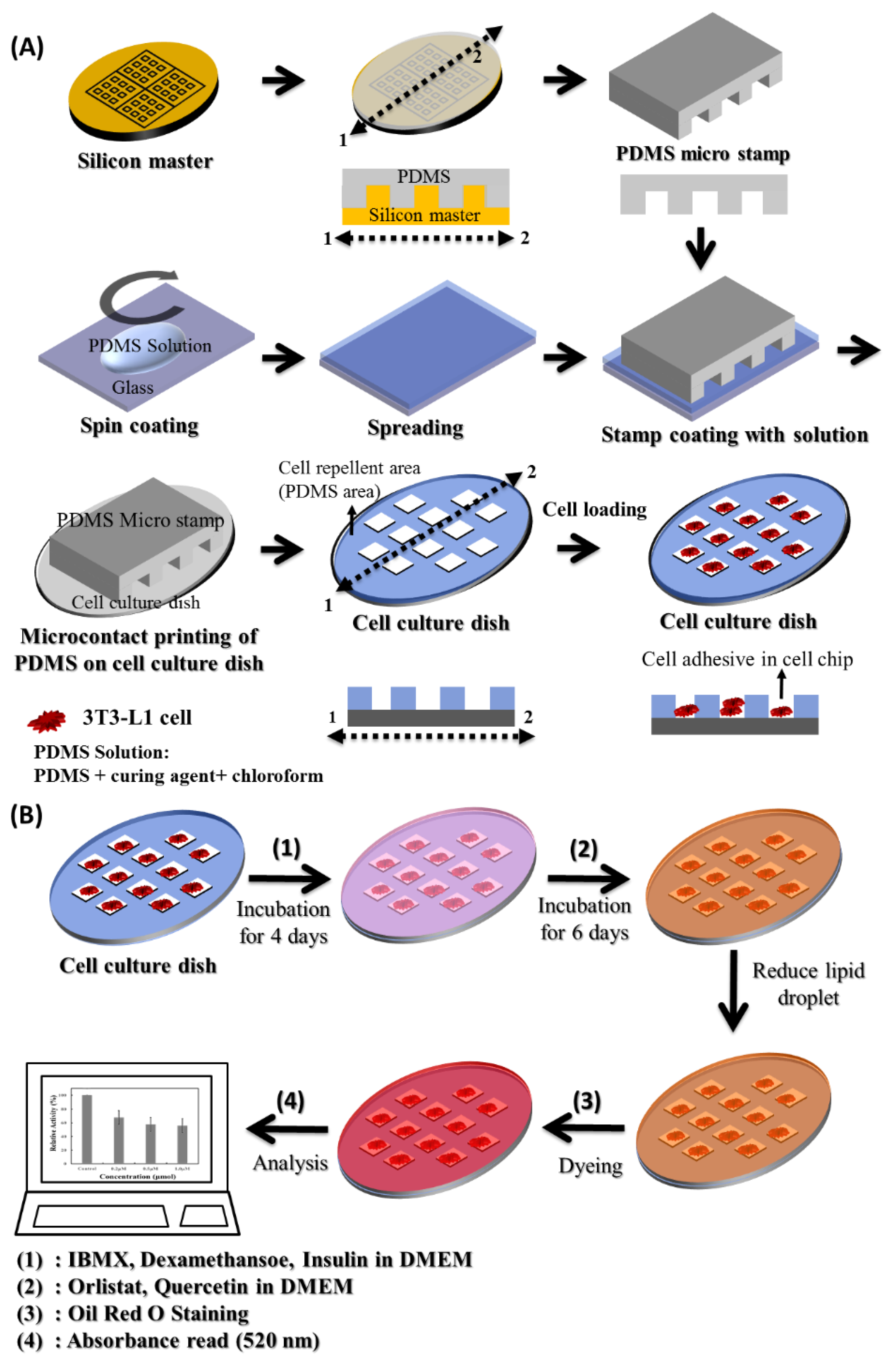
2.3. Fabrication of the Adipocyte Micro-Cell Pattern Chip
2.4. Cell Culture and Patterning
2.5. 3T3-L1 Cell Differentiation on an Adipocyte Micro-Cell Pattern Chip and Effect of Anti-Obesity Agents Using Oil Red O Protocol
2.6. Statistical Analysis
3. Results and Discussion
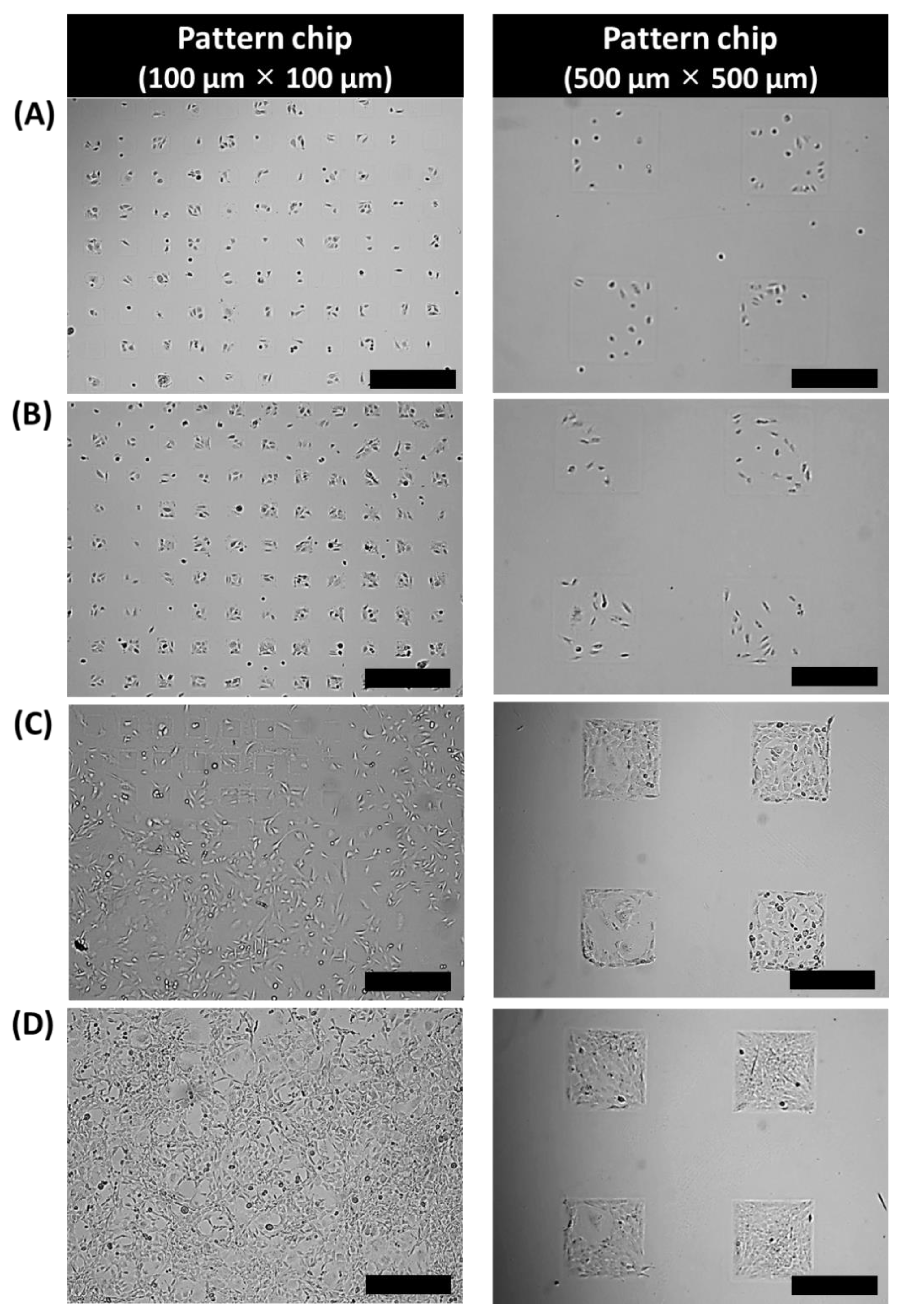
3.1. 3T3-L1 Cell Growth on an Adipocyte Micro-Cell Pattern Chip
3.2. 3T3-L1 Cell Proliferation by Pattern Size of Cell Pattern Chip
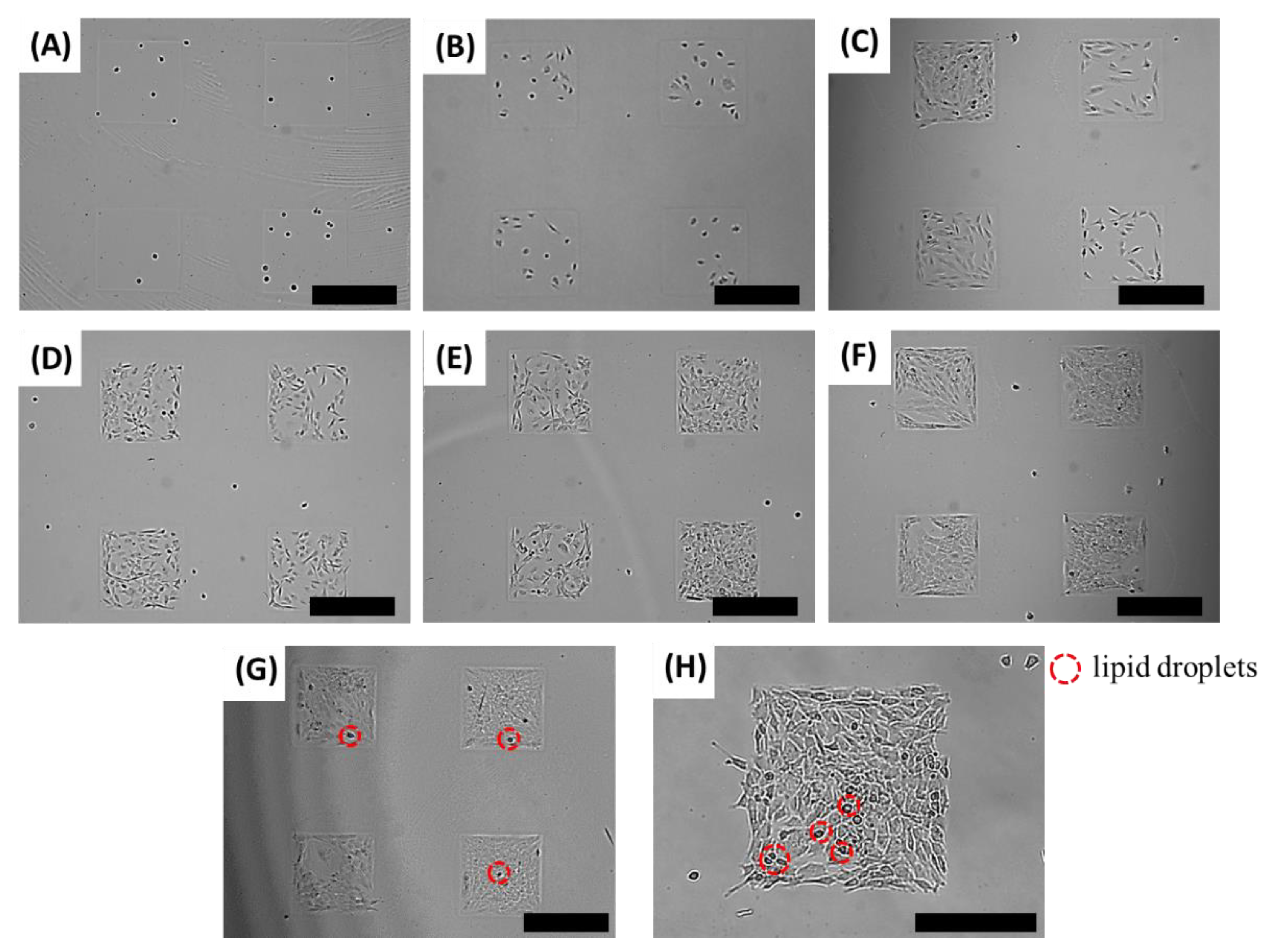
3.3. 3T3-L1 Cell Differentiation on an Adipocyte Micro-Cell Pattern Chip
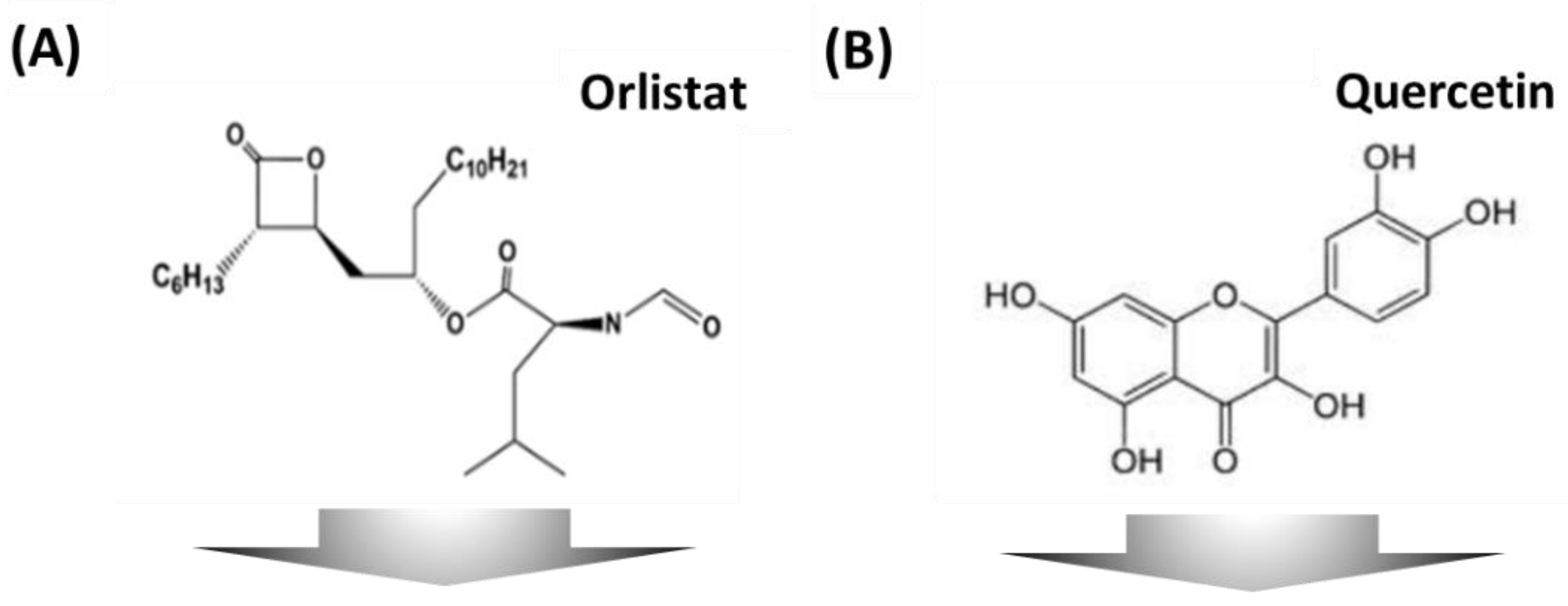
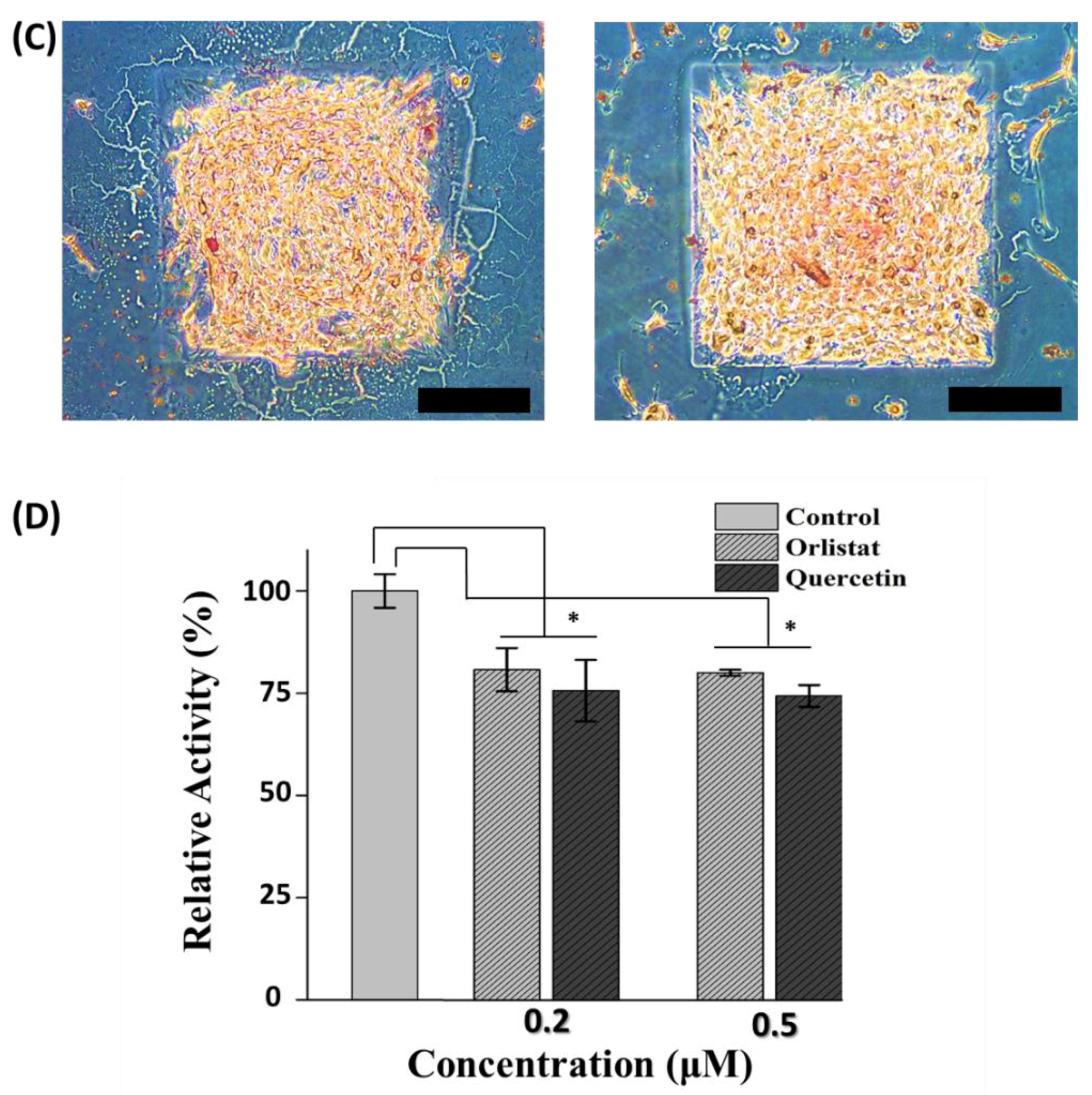
3.4. Inhibition Activity of Anti-Obesity Agents on an Adipocyte Micro-Cell Pattern Chip
4. Conclusions
Author Contributions
Acknowledgments
Conflicts of Interest
References
- Bays, H.; Dujovne, C. Anti-obesity drug development. Expert Opin. Inverstig. Drugs 2002, 11, 1189–1204. [Google Scholar] [CrossRef] [PubMed]
- Kim, H.C.; Kim, M.R. Analysis on awareness and practices for diet according to lifestyles of college students. Korean J. Hum. Ecol. 2010, 19, 157–165. [Google Scholar] [CrossRef]
- Marcelin, G.; Chua, S., Jr. Contributions of adipocyte lipid metabolism to body fat content and implications for the treatment of obesity. Curr. Opin. Pharmacol. 2010, 10, 588–593. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Cooke, D.; Bloom, S. The obesity pipeline: Current strategies in the development of anti-obesity drugs. Nat. Rev. Drug Discov. 2006, 5, 919–931. [Google Scholar] [CrossRef] [PubMed]
- Yang, P.Y.; Liu, K.; Ngai, M.H.; Lear, M.J.; Wenk, M.R.; Yao, S.Q. Activity-based proteome profiling of potential cellular targets of orlistat an FDA-approved drug with anti-tumor activities. J. Am. Chem. Soc. 2009, 132, 656–666. [Google Scholar] [CrossRef] [PubMed]
- Bray, G.A. A concise review on the therapeutics of obesity. Nutrition 2000, 16, 953–960. [Google Scholar] [CrossRef]
- Filippatos, T.D.; Derdemezis, C.S.; Gazi, I.F.; Nakou, E.S.; Mikhailidis, D.P.; Elisaf, M.S. Orlistat-associated adverse effects and drug interactions. Drug Saf. 2008, 31, 53–65. [Google Scholar] [CrossRef] [PubMed]
- Khan, M.A.; Herzog, C.A.; Peter, J.V.; Hartley, G.G.; Madlon-Kay, R.; Dick, C.D. The prevalence of cardiac valvular insufficiency assessed by transthoracic echocardiography in obese patients treated with appetite-suppressant drugs. N. Engl. J. Med. 1998, 339, 713–718. [Google Scholar] [CrossRef] [PubMed]
- Hollman, P.C.; Devries, J.H.; Vanleeuwen, S.D.; Mengelers, M.J.; Katan, M.B. Absorption of dietary quercetin glycosides and quercetin in healthy ileostomy volunteers. Am. J. Clin. Nutr. 1995, 62, 1276–1282. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Namiki, M. Antioxidants/antimutagens in food. Crit. Rev. Food Sci. Nutr. 1990, 294, 273–300. [Google Scholar] [CrossRef] [PubMed]
- Whalley, C.V.; Rankin, S.M.; Hoult, J.R.S.; Jessup, W.; Leake, D.S. Flavonoids inhibit the oxidative modification of low density lipoproteins by macrophages. Biochem. Pharmacol. 1990, 39, 1743–1750. [Google Scholar] [CrossRef]
- Ahn, J.; Lee, H.J.; Kim, S.; Park, J.H.; Ha, T.Y. The anti-obesity effect of quercetin is mediated by the AMPK and MAPK signaling pathways. Biochem. Biophys. Res. Commum. 2008, 373, 545–549. [Google Scholar] [CrossRef] [PubMed]
- Song, K.; Jeong, H.H.; Jin, S.H.; Park, J.S.; Lee, C.S. Optimization of microwell-based cell docking in microvalve integrated microfluidic device. BioChip J. 2014, 8, 227–233. [Google Scholar] [CrossRef]
- Jeong, H.H.; Jin, S.H.; Lee, B.J.; Kim, T.; Lee, C.S. Microfluidic static droplet array for analyzing microbial communication on a population gradient. Lab Chip 2015, 15, 889–899. [Google Scholar] [CrossRef] [PubMed]
- Rosenthal, A.; Macdonald, A.; Voldman, J. Cell patterning chip for controlling the stem cell microenvironment. Biomaterials 2007, 28, 3208–3216. [Google Scholar] [CrossRef] [PubMed]
- Nishizawa, M.; Takahashi, A.; Kaji, H.; Matsue, T. Micropatterned HeLa cell culture on PEG monolayer-coated glass substrates. Chem. Lett. 2002, 31, 904–905. [Google Scholar] [CrossRef]
- Sia, S.K.; Whitesides, G.M. Microfluidic devices fabricated in poly (dimethylsiloxane) for biological studies. Electrophoresis 2003, 24, 3563–3576. [Google Scholar] [CrossRef] [PubMed]
- Whitesides, G.M. The origins and the future of microfluidics. Nature 2006, 442, 368–373. [Google Scholar] [CrossRef] [PubMed]
- Rosen, E.D.; Sarraf, P.; Troy, A.E.; Bradwin, G.; Moore, K.; Milstone, D.S. PPARγ is required for the differentiation of adipose tissue in vivo and in vitro. Mol. Cell 1999, 4, 611–617. [Google Scholar] [CrossRef]
- Zebisch, K.; Voigt, V.; Wabitsch, M.; Brandsch, M. Protocol for effective differentiation of 3T3-L1 cells to adipocytes. Anal. Biochem. 2012, 425, 88–90. [Google Scholar] [CrossRef] [PubMed]
- Kim, G.Y.; Jeong, H.H.; Yeom, S.J.; Lee, C.S.; Roh, C. Efficient and reliable screening of anti-obesity agents on a micro-cell pattern chip. J. Chem. Technol. Biotechnol. 2016, 91, 2688–2692. [Google Scholar] [CrossRef]
- Dowell, P.; Flexner, C.; Kwiterovich, P.O.; Lane, M.D. Suppression of preadipocyte differentiation and promotion of adipocyte death by HIV protease inhibitors. J. Biol. Chem. 2000, 275, 41325–41332. [Google Scholar] [CrossRef] [PubMed]
- Rosen, E.D.; MacDougald, O.A. Adipocyte differentiation from the inside out. Nat. Rev. Mol. Cell Biol. 2006, 7, 885–896. [Google Scholar] [CrossRef] [PubMed]


© 2018 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Kim, G.Y.; Yeom, S.-J.; Jang, S.-C.; Lee, C.-S.; Roh, C.; Jeong, H.-H. Simple Analysis of Lipid Inhibition Activity on an Adipocyte Micro-Cell Pattern Chip. Biomolecules 2018, 8, 37. https://doi.org/10.3390/biom8020037
Kim GY, Yeom S-J, Jang S-C, Lee C-S, Roh C, Jeong H-H. Simple Analysis of Lipid Inhibition Activity on an Adipocyte Micro-Cell Pattern Chip. Biomolecules. 2018; 8(2):37. https://doi.org/10.3390/biom8020037
Chicago/Turabian StyleKim, Gi Yong, Su-Jin Yeom, Sung-Chan Jang, Chang-Soo Lee, Changhyun Roh, and Heon-Ho Jeong. 2018. "Simple Analysis of Lipid Inhibition Activity on an Adipocyte Micro-Cell Pattern Chip" Biomolecules 8, no. 2: 37. https://doi.org/10.3390/biom8020037





