The Role of Functional Amyloids in Multicellular Growth and Development of Gram-Positive Bacteria
Abstract
:1. Introduction
2. Functional Amyloid Proteins in Streptomyces coelicolor
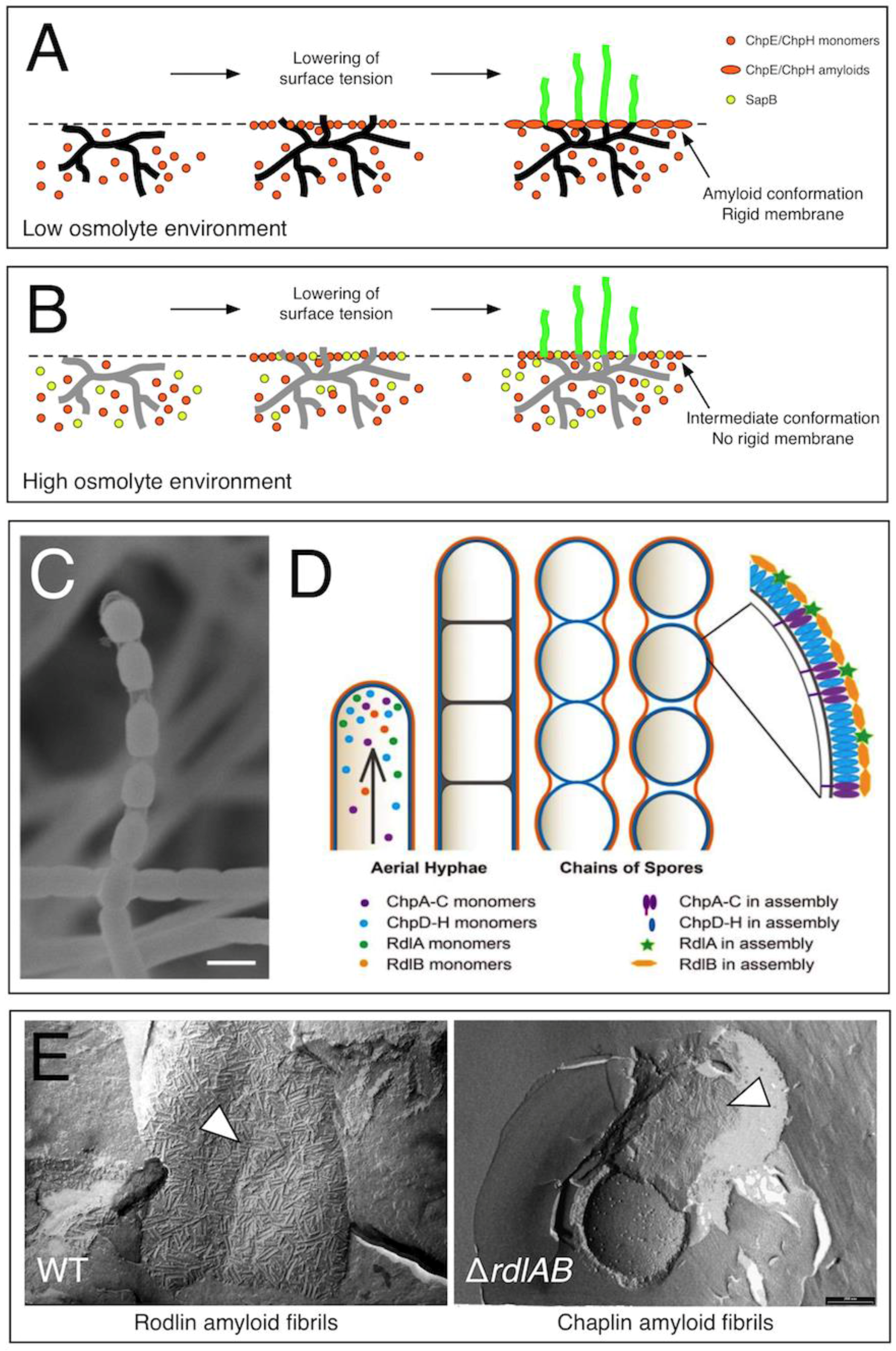
2.1. Chaplins Are Surfactants Mediating the Escape of Hyphae into the Air
2.2. Functional Homologues with Chaplin-Like Functions in Streptomycetes
2.3. Developing Aerial Hyphae Are Surrounded by a Second Amyloidal Layer Formed by the Rodlin Proteins
3. Diverse Functions of the TasA Amyloid in Bacillus subtilis
3.1. TasA Amyloid Fibers as a Key Structural Element of the B. subtilis Extracellular Matrix
3.2. Role of TasA in Surface Spreading
3.3. TasA as a Weapon
3.4. Role of TasA in Sporulation
3.5. Conserved and Less Conserved Features of TasA
4. Conclusions
Acknowledgments
Author Contributions
Conflicts of Interest
References
- Ballard, C.; Gauthier, S.; Corbett, A.; Brayne, C.; Aarsland, D.; Jones, E. Alzheimer’s disease. Lancet 2011, 377, 1019–1031. [Google Scholar] [CrossRef]
- Eisenberg, D.; Jucker, M. The amyloid state of proteins in human diseases. Cell 2012, 148, 1188–1203. [Google Scholar] [CrossRef] [PubMed]
- Larsen, P.; Nielsen, J.L.; Otzen, D.; Nielsen, P.H. Amyloid-like adhesins produced by floc-forming and filamentous bacteria in activated sludge. Appl. Environ. Microbiol. 2008, 74, 1517–1526. [Google Scholar] [CrossRef] [PubMed]
- Gebbink, M.F.B.G.; Claessen, D.; Bouma, B.; Dijkhuizen, L.; Wösten, H.A.B. Amyloids—A functional coat for microorganisms. Nat. Rev. Microbiol. 2005, 3, 333–341. [Google Scholar] [CrossRef] [PubMed]
- Blanco, L.P.; Evans, M.L.; Smith, D.R.; Badtke, M.P.; Chapman, M.R. Diversity, biogenesis and function of microbial amyloids. Trends. Microbiol. 2012, 20, 66–73. [Google Scholar] [CrossRef] [PubMed]
- Dueholm, M.S.; Larsen, P.; Finster, K.; Stenvang, M.R.; Christiansen, G.; Vad, B.S.; Boggild, A.; Otzen, D.E.; Nielsen, P.H. The tubular sheaths encasing Methanosaeta thermophila filaments are functional amyloids. J. Biol. Chem. 2015, 290, 20590–20600. [Google Scholar] [CrossRef] [PubMed]
- Fowler, D.M.; Koulov, A.V.; Alory-Jost, C.; Marks, M.S.; Balch, W.E.; Kelly, J.W. Functional amyloid formation within mammalian tissue. PLoS Biol. 2006, 4, e6. [Google Scholar] [CrossRef] [PubMed]
- Van der Meij, A.; Worsley, S.F.; Hutchings, M.I.; van Wezel, G.P. Chemical ecology of antibiotic production by actinomycetes. FEMS Microbiol. Rev. 2017, 41, 392–416. [Google Scholar] [CrossRef] [PubMed]
- Chater, K.F.; Losick, R. Mycelial life style of Streptomyces coelicolor A3(2) and its relatives. In Bacteria as Multicellular Organisms; Shapiro, J., Dworkin, M., Eds.; Oxford University Press: Oxford, UK, 1997; pp. 149–182. [Google Scholar]
- Manteca, A.; Fernandez, M.; Sanchez, J. A death round affecting a young compartmentalized mycelium precedes aerial mycelium dismantling in confluent surface cultures of Streptomyces antibioticus. Microbiology 2005, 151, 3689–3697. [Google Scholar] [CrossRef] [PubMed]
- Claessen, D.; Rink, R.; de Jong, W.; Siebring, J.; de Vreugd, P.; Boersma, F.G.H.; Dijkhuizen, L.; Wösten, H.A.B. A novel class of secreted hydrophobic proteins is involved in aerial hyphae formation in Streptomyces coelicolor by forming amyloid-like fibrils. Genes Dev. 2003, 17, 1714–1726. [Google Scholar] [CrossRef] [PubMed]
- Elliot, M.A.; Karoonuthaisiri, N.; Huang, J.; Bibb, M.J.; Cohen, S.N.; Kao, C.M.; Buttner, M.J. The chaplins: A family of hydrophobic cell-surface proteins involved in aerial mycelium formation in Streptomyces coelicolor. Genes Dev. 2003, 17, 1727–1740. [Google Scholar] [CrossRef] [PubMed]
- Elliot, M.A.; Talbot, N.J. Building filaments in the air: Aerial morphogenesis in bacteria and fungi. Curr. Opin. Microbiol. 2004, 7, 594–601. [Google Scholar] [CrossRef] [PubMed]
- Duong, A.; Capstick, D.S.; Di Berardo, C.; Findlay, K.C.; Hesketh, A.; Hong, H.J.; Elliot, M.A. Aerial development in Streptomyces coelicolor requires sortase activity. Mol. Microbiol. 2012, 83, 992–1005. [Google Scholar] [CrossRef] [PubMed]
- Bokhove, M.; Claessen, D.; de Jong, W.; Dijkhuizen, L.; Boekema, E.J.; Oostergetel, G.T. Chaplins of Streptomyces coelicolor self-assemble into two distinct functional amyloids. J. Struct. Biol. 2013, 184, 301–309. [Google Scholar] [CrossRef] [PubMed]
- Ekkers, D.M.; Claessen, D.; Galli, F.; Stamhuis, E.J. Surface modification using interfacial assembly of the Streptomyces chaplin proteins. Appl. Microbiol. Biotechnol. 2014, 98, 4491–4501. [Google Scholar] [CrossRef] [PubMed]
- Sawyer, E.B.; Claessen, D.; Haas, M.; Hurgobin, B.; Gras, S.L. The assembly of individual chaplin peptides from Streptomyces coelicolor into functional amyloid fibrils. PLoS ONE 2011, 6, e18839. [Google Scholar] [CrossRef] [PubMed]
- Di Berardo, C.; Capstick, D.S.; Bibb, M.J.; Findlay, K.C.; Buttner, M.J.; Elliot, M.A. Function and redundancy of the chaplin cell surface proteins in aerial hypha formation, rodlet assembly, and viability in Streptomyces coelicolor. J. Bacteriol. 2008, 190, 5879–5889. [Google Scholar] [CrossRef] [PubMed]
- Claessen, D.; Stokroos, I.; Deelstra, H.J.; Penninga, N.A.; Bormann, C.; Salas, J.A.; Dijkhuizen, L.; Wösten, H.A.B. The formation of the rodlet layer of streptomycetes is the result of the interplay between rodlins and chaplins. Mol. Microbiol. 2004, 53, 433–443. [Google Scholar] [CrossRef] [PubMed]
- Capstick, D.S.; Jomaa, A.; Hanke, C.; Ortega, J.; Elliot, M.A. Dual amyloid domains promote differential functioning of the chaplin proteins during Streptomyces aerial morphogenesis. Proc. Natl. Acad. Sci. USA 2011, 108, 9821–9826. [Google Scholar] [CrossRef] [PubMed]
- Capstick, D.S.; Willey, J.M.; Buttner, M.J.; Elliot, M.A. SapB and the chaplins: Connections between morphogenetic proteins in Streptomyces coelicolor. Mol. Microbiol. 2007, 64, 602–613. [Google Scholar] [CrossRef] [PubMed]
- Willey, J.M.; Santamaria, R.; Guijarro, J.; Geistlich, M.; Losick, R. Extracellular complementation of a developmental mutation implicates a small sporulation protein in aerial mycelium formation by S. coelicolor. Cell 1991, 65, 641–650. [Google Scholar] [CrossRef]
- Kodani, S.; Lodato, M.A.; Durrant, M.C.; Picart, F.; Willey, J.M. SapT, a lanthionine-containing peptide involved in aerial hyphae formation in the streptomycetes. Mol. Microbiol. 2005, 58, 1368–1380. [Google Scholar] [CrossRef] [PubMed]
- Willey, J.M.; Willems, A.; Kodani, S.; Nodwell, J.R. Morphogenetic surfactants and their role in the formation of aerial hyphae in Streptomyces coelicolor. Mol. Microbiol. 2006, 59, 731–742. [Google Scholar] [CrossRef] [PubMed]
- Tillotson, R.D.; Wösten, H.A.B.; Richter, M.; Willey, J.M. A surface active protein involved in aerial hyphae formation in the filamentous fungus Schizophillum commune restores the capacity of a bald mutant of the filamentous bacterium Streptomyces coelicolor to erect aerial structures. Mol. Microbiol. 1998, 30, 595–602. [Google Scholar] [CrossRef] [PubMed]
- De Jong, W.; Vijgenboom, E.; Dijkhuizen, L.; Wösten, H.A.B.; Claessen, D. SapB and the rodlins are required for development of Streptomyces coelicolor in high osmolarity media. FEMS Microbiol. Lett. 2012, 329, 154–159. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Petrus, M.L.C.; Claessen, D. Pivotal roles for Streptomyces cell surface polymers in morphological differentiation, attachment and mycelial architecture. Antonie van Leeuwenhoek 2014, 106, 127–139. [Google Scholar] [CrossRef] [PubMed]
- Yang, W.; Willemse, J.; Sawyer, E.B.; Lou, F.; Gong, W.; Zhang, H.; Gras, S.L.; Claessen, D.; Perrett, S. The propensity of the bacterial rodlin protein RdlB to form amyloid fibrils determines its function in Streptomyces coelicolor. Sci. Rep. 2017, 7, 42867. [Google Scholar] [CrossRef] [PubMed]
- Smucker, R.A.; Pfister, R.M. Characteristics of Streptomyces coelicolor A3(2) aerial spore rodlet mosaic. Can. J. Microbiol. 1978, 24, 397–408. [Google Scholar] [CrossRef] [PubMed]
- Claessen, D.; Wösten, H.A.B.; van Keulen, G.; Faber, O.G.; Alves, A.M.C.R.; Meijer, W.G.; Dijkhuizen, L. Two novel homologous proteins of Streptomyces coelicolor and Streptomyces lividans are involved in the formation of the rodlet layer and mediate attachment to a hydrophobic surface. Mol. Microbiol. 2002, 44, 1483–1492. [Google Scholar] [CrossRef] [PubMed]
- Wildermuth, H.; Wehrli, E.; Horne, R.W. The surface structure of spores and aerial mycelium in Streptomyces coelicolor. J. Ultrastruct. Res. 1971, 35, 168–180. [Google Scholar] [CrossRef]
- Romero, D.; Aguilar, C.; Losick, R.; Kolter, R. Amyloid fibers provide structural integrity to Bacillus subtilis biofilms. Proc. Natl. Acad. Sci. USA 2010, 107, 2230–2234. [Google Scholar] [CrossRef] [PubMed]
- Epstein, E.A.; Reizian, M.A.; Chapman, M.R. Spatial clustering of the curlin secretion lipoprotein requires curli fiber assembly. J. Bacteriol. 2009, 191, 608–615. [Google Scholar] [CrossRef] [PubMed]
- Schwartz, K.; Syed, A.K.; Stephenson, R.E.; Rickard, A.H.; Boles, B.R. Functional amyloids composed of phenol soluble modulins stabilize Staphylococcus aureus biofilms. PLoS Pathog. 2012, 8, e1002744. [Google Scholar] [CrossRef] [PubMed]
- Romero, D.; Sanabria-Valentín, E.; Vlamakis, H.; Kolter, R. Biofilm inhibitors that target amyloid proteins. Chem. Biol. 2013, 20, 102–110. [Google Scholar] [CrossRef] [PubMed]
- Romero, D.; Kolter, R. Functional amyloids in bacteria. Int. Microbiol. 2014, 17, 65–73. [Google Scholar] [PubMed]
- Romero, D.; Vlamakis, H.; Losick, R.; Kolter, R. Functional analysis of the accessory protein TapA in Bacillus subtilis amyloid fiber assembly. J. Bacteriol. 2014, 196, 1505–1513. [Google Scholar] [CrossRef] [PubMed]
- Kearns, D.B.; Chu, F.; Branda, S.S.; Kolter, R.; Losick, R. A master regulator for biofilm formation by Bacillus subtilis. Mol. Microbiol. 2005, 55, 739–749. [Google Scholar] [CrossRef] [PubMed]
- Romero, D.; Vlamakis, H.; Losick, R.; Kolter, R. An accessory protein required for anchoring and assembly of amyloid fibres in B. subtilis biofilms. Mol. Microbiol. 2011, 80, 1155–1168. [Google Scholar] [CrossRef] [PubMed]
- Mhatre, E.; Sundaram, A.; Hölscher, T.; Mühlstädt, M.; Bossert, J.; Kovács, A.T. Presence of calcium lowers the expansion of Bacillus subtilis colony biofilms. Microorganisms 2017, 5, 7. [Google Scholar] [CrossRef] [PubMed]
- Kesel, S.; Grumbein, S.; Gümperlein, I.; Tallawi, M.; Marel, A.K.; Lieleg, O.; Opitz, M. Direct comparison of physical properties of Bacillus subtilis NCIB 3610 and B-1 biofilms. Appl. Environ. Microbiol. 2016, 82, 2424–2432. [Google Scholar] [CrossRef] [PubMed]
- Kobayashi, K.; Iwano, M. BslA(YuaB) forms a hydrophobic layer on the surface of Bacillus subtilis biofilms. Mol. Microbiol. 2012, 85, 51–66. [Google Scholar] [CrossRef] [PubMed]
- Arnaouteli, S.; Ferreira, A.S.; Schor, M.; Morris, R.J.; Bromley, K.M.; Jo, J.K.; Cortez, K.L.; Sukhodub, T.; Prescott, A.R.; Dietrich, L.E.P.; et al. Bifunctionality of a biofilm matrix protein controlled by redox state. bioRxiv 2017. [Google Scholar] [CrossRef] [PubMed]
- Beauregard, P.B.; Chai, Y.; Vlamakis, H.; Losick, R.; Kolter, R. Bacillus subtilis biofilm induction by plant polysaccharides. Proc. Natl. Acad. Sci. USA 2013, 110, E1621–E1630. [Google Scholar] [CrossRef] [PubMed]
- Van Gestel, J.; Vlamakis, H.; Kolter, R. From cell differentiation to cell collectives: Bacillus subtilis uses division of labor to migrate. PLoS Biol. 2015, 13, e1002141. [Google Scholar] [CrossRef] [PubMed]
- Tsompanidou, E.; Denham, E.L.; Becher, D.; de Jong, A.; Buist, G.; van Oosten, M.; Manson, W.L.; Back, J.W.; van Dijl, J.M.; Dreisbach, A. Distinct roles of phenol-soluble modulins in spreading of Staphylococcus aureus on wet surfaces. Appl. Environ. Microbiol. 2013, 79, 886–895. [Google Scholar] [CrossRef] [PubMed]
- Reynolds, N.P.; Charnley, M.; Mezzenga, R.; Hartley, P.G. Engineered lysozyme amyloid fibril networks support cellular growth and spreading. Biomacromolecules 2014, 15, 599–608. [Google Scholar] [CrossRef] [PubMed]
- Kagan, B.L.; Jang, H.; Capone, R.; Teran Arce, F.; Ramachandran, S.; Lal, R.; Nussinov, R. Antimicrobial properties of amyloid peptides. Mol. Pharm. 2012, 9, 708–717. [Google Scholar] [CrossRef] [PubMed]
- Schwartz, K.; Boles, B.R. Microbial amyloids—Functions and interactions within the host. Curr. Opin. Microbiol. 2013, 16, 93–99. [Google Scholar] [CrossRef] [PubMed]
- Stover, A.G.; Driks, A. Secretion, localization, and antibacterial activity of TasA, a Bacillus subtilis spore-associated protein. J. Bacteriol. 1999, 181, 1664–1672. [Google Scholar] [PubMed]
- Khezri, M.; Jouzani, G.S.; Ahmadzadeh, M. Fusarium culmorum affects expression of biofilm formation key genes in Bacillus subtilis. Braz. J. Microbiol. 2016, 47, 47–54. [Google Scholar] [CrossRef] [PubMed]
- Lyons, N.A.; Kraigher, B.; Stefanic, P.; Mandic-Mulec, I.; Kolter, R. A combinatorial kin discrimination system in Bacillus subtilis. Curr. Biol. 2016, 26, 733–742. [Google Scholar] [CrossRef] [PubMed]
- Štefanić, P.; Kraigher, B.; Lyons, N.A.; Kolter, R.; Mandić-Mulec, I. Kin discrimination between sympatric Bacillus subtilis isolates. Proc. Natl. Acad. Sci. USA 2015, 112, 14042–14047. [Google Scholar] [CrossRef] [PubMed]
- Serrano, M.; Zilhao, R.; Ricca, E.; Ozin, A.J.; Moran, C.P., Jr.; Henriques, A.O. A Bacillus subtilis secreted protein with a role in endospore coat assembly and function. J. Bacteriol. 1999, 181, 3632–3643. [Google Scholar] [PubMed]
- Plomp, M.; Carroll, A.M.; Setlow, P.; Malkin, A.J. Architecture and assembly of the Bacillus subtilis spore coat. PLoS ONE 2014, 9, e108560. [Google Scholar] [CrossRef] [PubMed]
- Caro-Astorga, J.; Pérez-García, A.; de Vicente, A.; Romero, D. A genomic region involved in the formation of adhesin fibers in Bacillus cereus biofilms. Front. Microbiol. 2014, 5, 745. [Google Scholar] [CrossRef] [PubMed]
- Di Luccia, B.; Riccio, A.; Vanacore, A.; Baccigalupi, L.; Molinaro, A.; Ricca, E. Matrix production, pigment synthesis, and sporulation in a marine isolated strain of Bacillus pumilus. Mar. Drugs 2015, 13, 6472–6488. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Taylor, J.D.; Zhou, Y.; Salgado, P.S.; Patwardhan, A.; McGuffie, M.; Pape, T.; Grabe, G.; Ashman, E.; Constable, S.C.; Simpson, P.J.; et al. Atomic resolution insights into curli fiber biogenesis. Structure 2011, 19, 1307–1316. [Google Scholar] [CrossRef] [PubMed]
- Dueholm, M.S.; Sondergaard, M.T.; Nilsson, M.; Christiansen, G.; Stensballe, A.; Overgaard, M.T.; Givskov, M.; Tolker-Nielsen, T.; Otzen, D.E.; Nielsen, P.H. Expression of Fap amyloids in Pseudomonas aeruginosa, P. fluorescens, and P. putida results in aggregation and increased biofilm formation. Microbiologyopen 2013, 2, 365–382. [Google Scholar] [CrossRef] [PubMed]
- Bousset, L.; Thomson, N.H.; Radford, S.E.; Melki, R. The yeast prion Ure2p retains its native alpha-helical conformation upon assembly into protein fibrils in vitro. EMBO J. 2002, 21, 2903–2911. [Google Scholar] [CrossRef] [PubMed]
- Fei, L.; Perrett, S. New insights into the molecular mechanism of amyloid formation from cysteine scanning. Prion 2010, 4, 9–12. [Google Scholar] [CrossRef] [PubMed]
- Choi, M.S.; Kim, W.; Lee, C.; Oh, C.S. Harpins, multifunctional proteins secreted by gram-negative plant-pathogenic bacteria. Mol. Plant Microbe Interact. 2013, 26, 1115–1122. [Google Scholar] [CrossRef] [PubMed]
- Nielsen, P.H.; Dueholm, M.S.; Thomsen, T.R.; Nielsen, J.L.; Otzen, D. Functional bacterial amyloids in biofilms. In Biofilm Highlights; Flemming, H.-C., Wingender, J., Szewzyk, U., Eds.; Springer: Berlin/Heidelberg, Germany, 2011; pp. 41–62. [Google Scholar]
- Taglialegna, A.; Lasa, I.; Valle, J. Amyloid structures as biofilm matrix scaffolds. J. Bacteriol. 2016, 198, 2579–2588. [Google Scholar] [CrossRef] [PubMed]
- Zhou, Y.; Blanco, L.P.; Smith, D.R.; Chapman, M.R. Bacterial amyloids. Methods Mol. Biol. 2012, 849, 303–320. [Google Scholar] [PubMed]
- Arranz, R.; Mercado, G.; Martín-Benito, J.; Giraldo, R.; Monasterio, O.; Lagos, R.; Valpuesta, J.M. Structural characterization of microcin E492 amyloid formation: Identification of the precursors. J. Struct. Biol. 2012, 178, 54–60. [Google Scholar] [CrossRef] [PubMed]
- Dueholm, M.S.; Petersen, S.V.; Sonderkaer, M.; Larsen, P.; Christiansen, G.; Hein, K.L.; Enghild, J.J.; Nielsen, J.L.; Nielsen, K.L.; Nielsen, P.H.; et al. Functional amyloid in Pseudomonas. Mol. Microbiol. 2010, 77, 1009–1020. [Google Scholar] [CrossRef] [PubMed]
- Dutta, A.; Bhattacharyya, S.; Kundu, A.; Dutta, D.; Das, A.K. Macroscopic amyloid fiber formation by staphylococcal biofilm associated SuhB protein. Biophys. Chem. 2016, 217, 32–41. [Google Scholar] [CrossRef] [PubMed]
- Shewmaker, F.; McGlinchey, R.P.; Thurber, K.R.; McPhie, P.; Dyda, F.; Tycko, R.; Wickner, R.B. The functional curli amyloid is not based on in-register parallel beta-sheet structure. J. Biol. Chem. 2009, 284, 25065–25076. [Google Scholar] [CrossRef] [PubMed]
- Chai, L.; Romero, D.; Kayatekin, C.; Akabayov, B.; Vlamakis, H.; Losick, R.; Kolter, R. Isolation, characterization, and aggregation of a structured bacterial matrix precursor. J. Biol. Chem. 2013, 288, 17559–17568. [Google Scholar] [CrossRef] [PubMed]
- Tayeb-Fligelman, E.; Tabachnikov, O.; Moshe, A.; Goldshmidt-Tran, O.; Sawaya, M.R.; Coquelle, N.; Colletier, J.P.; Landau, M. The cytotoxic Staphylococcus aureus PSMα3 reveals a cross-α amyloid-like fibril. Science 2017, 355, 831–833. [Google Scholar] [CrossRef] [PubMed]
- Branda, S.S.; Chu, F.; Kearns, D.B.; Losick, R.; Kolter, R. A major protein component of the Bacillus subtilis biofilm matrix. Mol. Microbiol. 2006, 59, 1229–1238. [Google Scholar] [CrossRef] [PubMed]
- Martin, M.; Dragoš, A.; Hölscher, T.; Maróti, G.; Bálint, B.; Westermann, M.; Kovács, Á.T. De novo evolved interference competition promotes the spread of biofilm defectors. Nat. Commun. 2017, 8, 15127. [Google Scholar]

© 2017 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Dragoš, A.; Kovács, Á.T.; Claessen, D. The Role of Functional Amyloids in Multicellular Growth and Development of Gram-Positive Bacteria. Biomolecules 2017, 7, 60. https://doi.org/10.3390/biom7030060
Dragoš A, Kovács ÁT, Claessen D. The Role of Functional Amyloids in Multicellular Growth and Development of Gram-Positive Bacteria. Biomolecules. 2017; 7(3):60. https://doi.org/10.3390/biom7030060
Chicago/Turabian StyleDragoš, Anna, Ákos T. Kovács, and Dennis Claessen. 2017. "The Role of Functional Amyloids in Multicellular Growth and Development of Gram-Positive Bacteria" Biomolecules 7, no. 3: 60. https://doi.org/10.3390/biom7030060




