Proteins Directly Interacting with Mammalian 20S Proteasomal Subunits and Ubiquitin-Independent Proteasomal Degradation
Abstract
:1. Introduction
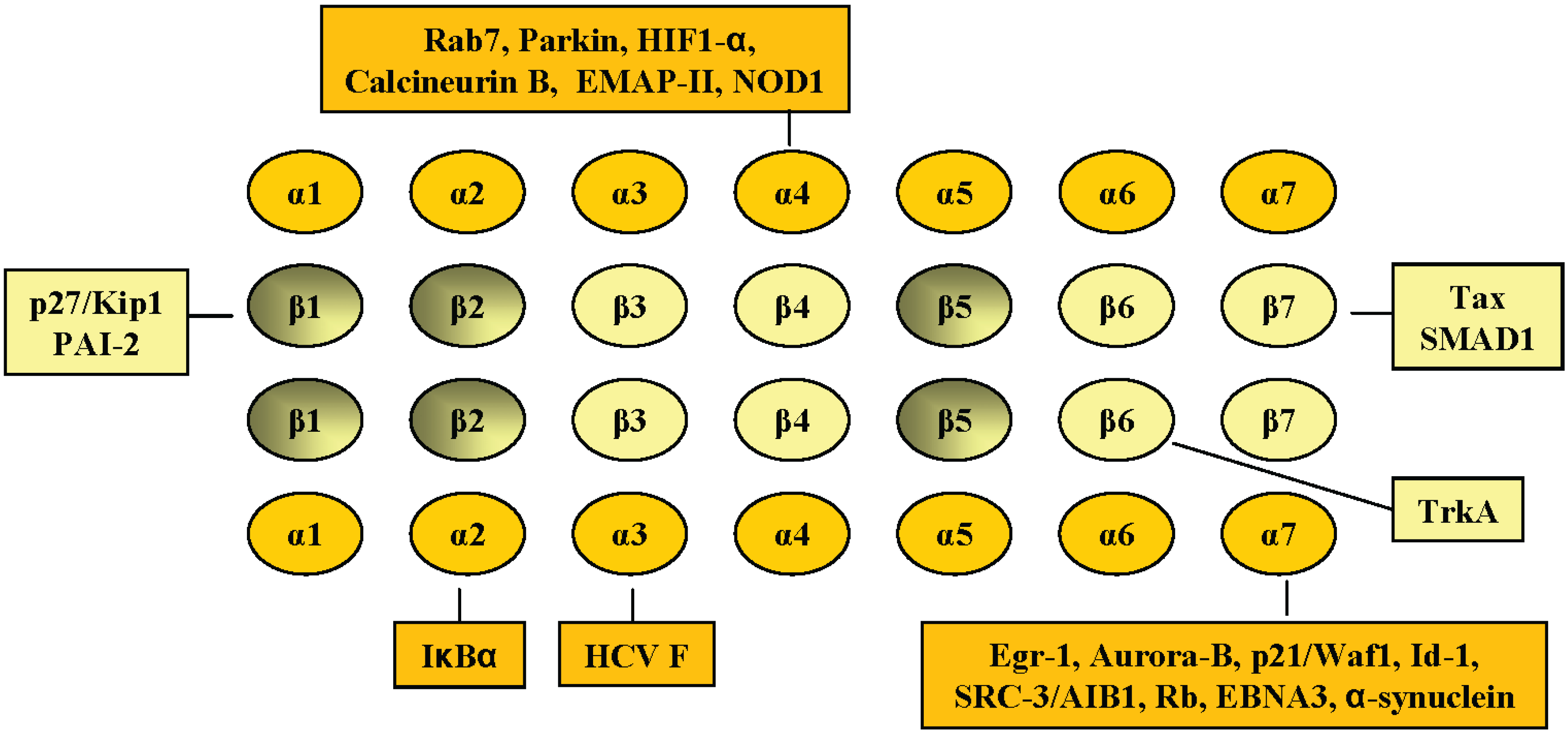
2. Interaction of Cellular Proteins with Specific Proteasomal α and β Subunits of the 20S Proteasome Complex
2.1. PSMA2, C3, α2
2.2. PSMA4, C9, α3
2.3. PSMA7, XAPC7, α4
2.4. PSMA3, C8, α7 Subunit
2.5. PSMB6, Y, β1
2.6. PSMB1, C5, β6
2.7. PSMB4, N3, β7
3. Ubiquitin Independent Proteasomal Degradation
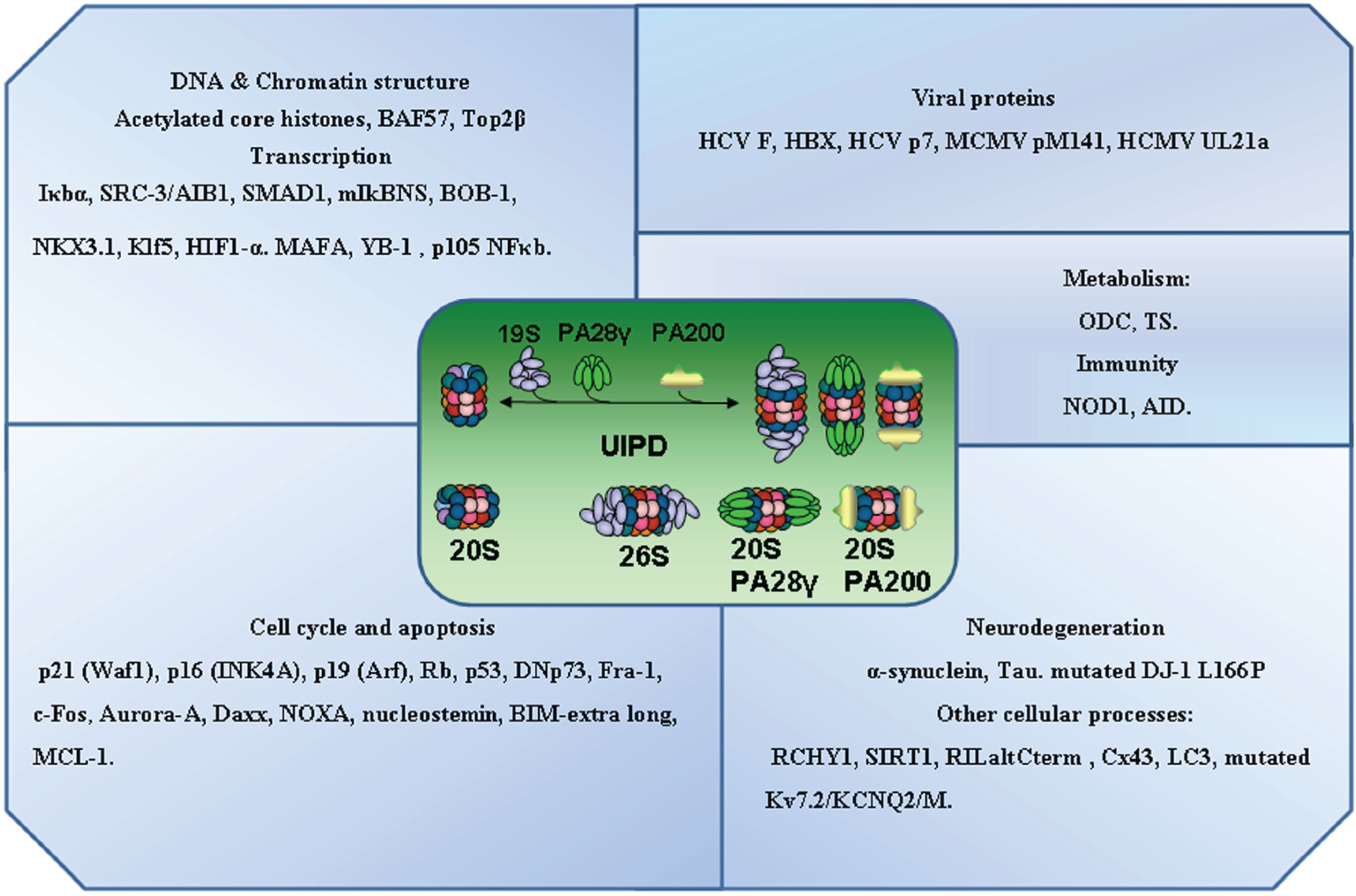
4. A Critical Assessment of Specific Protein Interactions of Proteasomal Subunits and UIPD
5. Conclusions
Acknowledgments
Conflicts of Interest
References
- Tomko, R.J., Jr.; Hochstrasser, M. Molecular architecture and assembly of the eukaryotic proteasome. Annu. Rev. Biochem. 2013, 82, 415–445. [Google Scholar] [CrossRef] [PubMed]
- Jariel-Encontre, I.; Bossis, G.; Piechaczyk, M. Ubiquitin-independent degradation of proteins by the proteasome. Biochim. Biophys. Acta 2008, 1786, 153–177. [Google Scholar] [PubMed]
- Alvarez-Castelao, B.; Castano, J.G. Mechanism of direct degradation of IkappaBalpha by 20S proteasome. FEBS Lett. 2005, 579, 4797–4802. [Google Scholar] [CrossRef] [PubMed]
- Zhang, W.; Wei, Q. Calcineurin stimulates the expression of inflammatory factors in RAW 264.7 cells by interacting with proteasome subunit alpha type 6. Biochem. Biophys. Res. Commun. 2011, 407, 668–673. [Google Scholar] [CrossRef] [PubMed]
- Yuksek, K.; Chen, W.L.; Chien, D.; Ou, J.H. Ubiquitin-independent degradation of hepatitis C virus F protein. J. Virol. 2009, 83, 612–621. [Google Scholar] [CrossRef] [PubMed]
- Stohwasser, R.; Holzhutter, H.G.; Lehmann, U.; Henklein, P.; Kloetzel, P.M. Hepatitis B virus HBx peptide 116–138 and proteasome activator PA28 compete for binding to the proteasome alpha4/MC6 subunit. Biol. Chem. 2003, 384, 39–49. [Google Scholar] [CrossRef] [PubMed]
- Dong, J.; Chen, W.; Welford, A.; Wandinger-Ness, A. The proteasome alpha-subunit XAPC7 interacts specifically with Rab7 and late endosomes. J. Biol. Chem. 2004, 279, 21334–21342. [Google Scholar] [CrossRef] [PubMed]
- Dachsel, J.C.; Lucking, C.B.; Deeg, S.; Schultz, E.; Lalowski, M.; Casademunt, E.; Corti, O.; Hampe, C.; Patenge, N.; Vaupel, K.; et al. Parkin interacts with the proteasome subunit alpha4. FEBS Lett. 2005, 579, 3913–3919. [Google Scholar] [CrossRef]
- Cho, S.; Choi, Y.J.; Kim, J.M.; Jeong, S.T.; Kim, J.H.; Kim, S.H.; Ryu, S.E. Binding and regulation of HIF-1alpha by a subunit of the proteasome complex, PSMA7. FEBS Lett. 2001, 498, 62–66. [Google Scholar] [CrossRef] [PubMed]
- Li, N.; Zhang, Z.; Zhang, W.; Wei, Q. Calcineurin B subunit interacts with proteasome subunit alpha type 7 and represses hypoxia-inducible factor-1alpha activity via the proteasome pathway. Biochem. Biophys. Res. Commun. 2011, 405, 468–472. [Google Scholar] [CrossRef] [PubMed]
- Tandle, A.T.; Calvani, M.; Uranchimeg, B.; Zahavi, D.; Melillo, G.; Libutti, S.K. Endothelial monocyte activating polypeptide-II modulates endothelial cell responses by degrading hypoxia-inducible factor-1alpha through interaction with PSMA7, a component of the proteasome. Exp. Cell Res. 2009, 315, 1850–1859. [Google Scholar] [CrossRef] [PubMed]
- Yang, L.; Tang, Z.; Zhang, H.; Kou, W.; Lu, Z.; Li, X.; Li, Q.; Miao, Z. PSMA7 directly interacts with NOD1 and regulates its function. Cell. Physiol. Biochem. 2013, 31, 952–959. [Google Scholar] [CrossRef]
- Gerards, W.L.; Enzlin, J.; Haner, M.; Hendriks, I.L.; Aebi, U.; Bloemendal, H.; Boelens, W. The human alpha-type proteasomal subunit HsC8 forms a double ringlike structure, but does not assemble into proteasome-like particles with the beta-type subunits HsDelta or HsBPROS26. J. Biol. Chem. 1997, 272, 10080–10086. [Google Scholar] [CrossRef]
- Boelens, W.C.; Croes, Y.; de Jong, W.W. Interaction between alphaB-crystallin and the human 20S proteasomal subunit C8/alpha7. Biochim. Biophys. Acta 2001, 1544, 311–319. [Google Scholar] [CrossRef] [PubMed]
- Kania, M.A.; DeMartino, G.N.; Baumeister, W.; Goldberg, A.L. The proteasome subunit, C2, contains an important site for binding of the PA28 (11S) activator. Eur. J. Biochem. 1996, 236, 510–516. [Google Scholar] [CrossRef] [PubMed]
- Bae, M.H.; Jeong, C.H.; Kim, S.H.; Bae, M.K.; Jeong, J.W.; Ahn, M.Y.; Bae, S.K.; Kim, N.D.; Kim, C.W.; Kim, K.R.; et al. Regulation of Egr-1 by association with the proteasome component C8. Biochim. Biophys. Acta 2002, 1592, 163–167. [Google Scholar]
- Shu, F.; Guo, S.; Dang, Y.; Qi, M.; Zhou, G.; Guo, Z.; Zhang, Y.; Wu, C.; Zhao, S.; Yu, L. Human aurora-B binds to a proteasome alpha-subunit HC8 and undergoes degradation in a proteasome-dependent manner. Mol. Cell. Biochem. 2003, 254, 157–162. [Google Scholar] [CrossRef]
- Touitou, R.; Richardson, J.; Bose, S.; Nakanishi, M.; Rivett, J.; Allday, M.J. A degradation signal located in the C-terminus of p21WAF1/CIP1 is a binding site for the C8 alpha-subunit of the 20S proteasome. EMBO J. 2001, 20, 2367–2375. [Google Scholar] [CrossRef] [PubMed]
- Zhang, Z.; Wang, H.; Li, M.; Agrawal, S.; Chen, X.; Zhang, R. MDM2 is a negative regulator of p21WAF1/CIP1, independent of p53. J. Biol. Chem. 2004, 279, 16000–16006. [Google Scholar] [CrossRef] [PubMed]
- Wang, B.; Liu, K.; Lin, H.Y.; Bellam, N.; Ling, S.; Lin, W.C. 14-3-3Tau regulates ubiquitin-independent proteasomal degradation of p21, a novel mechanism of p21 downregulation in breast cancer. Mol. Cell. Biol. 2010, 30, 1508–1527. [Google Scholar] [CrossRef] [PubMed]
- Chen, X.; Barton, L.F.; Chi, Y.; Clurman, B.E.; Roberts, J.M. Ubiquitin-independent degradation of cell-cycle inhibitors by the REGγ proteasome. Mol. Cell 2007, 26, 843–852. [Google Scholar] [CrossRef] [PubMed]
- Li, X.; Amazit, L.; Long, W.; Lonard, D.M.; Monaco, J.J.; O’Malley, B.W. Ubiquitin- and ATP-independent proteolytic turnover of p21 by the REGγ-proteasome pathway. Mol. Cell 2007, 26, 831–842. [Google Scholar] [CrossRef] [PubMed]
- Ling, M.T.; Chiu, Y.T.; Lee, T.K.; Leung, S.C.; Fung, M.K.; Wang, X.; Wong, K.F.; Wong, Y.C. Id-1 induces proteasome-dependent degradation of the HBX protein. J. Mol. Biol. 2008, 382, 34–43. [Google Scholar] [CrossRef]
- Yi, P.; Feng, Q.; Amazit, L.; Lonard, D.M.; Tsai, S.Y.; Tsai, M.J.; O’Malley, B.W. Atypical protein kinase C regulates dual pathways for degradation of the oncogenic coactivator SRC-3/AIB1. Mol. Cell 2008, 29, 465–476. [Google Scholar] [CrossRef]
- Li, X.; Lonard, D.M.; Jung, S.Y.; Malovannaya, A.; Feng, Q.; Qin, J.; Tsai, S.Y.; Tsai, M.J.; O’Malley, B.W. The SRC-3/AIB1 coactivator is degraded in a ubiquitin- and ATP-independent manner by the REGγ proteasome. Cell 2006, 124, 381–392. [Google Scholar] [CrossRef]
- Sdek, P.; Ying, H.; Chang, D.L.; Qiu, W.; Zheng, H.; Touitou, R.; Allday, M.J.; Xiao, Z.X. MDM2 promotes proteasome-dependent ubiquitin-independent degradation of retinoblastoma protein. Mol. Cell 2005, 20, 699–708. [Google Scholar] [CrossRef]
- Ying, H.; Xiao, Z.X. Targeting retinoblastoma protein for degradation by proteasomes. Cell Cycle 2006, 5, 506–508. [Google Scholar] [CrossRef] [PubMed]
- Touitou, R.; O'Nions, J.; Heaney, J.; Allday, M.J. Epstein-Barr virus EBNA3 proteins bind to the C8/alpha7 subunit of the 20S proteasome and are degraded by 20S proteasomes in vitro, but are very stable in latently infected B cells. J. Gen. Virol. 2005, 86, 1269–1277. [Google Scholar] [CrossRef] [PubMed]
- Fedorova, O.A.; Moiseeva, T.N.; Nikiforov, A.A.; Tsimokha, A.S.; Livinskaya, V.A.; Hodson, M.; Bottrill, A.; Evteeva, I.N.; Ermolayeva, J.B.; Kuznetzova, I.M.; et al. Proteomic analysis of the 20S proteasome (PSMA3)-interacting proteins reveals a functional link between the proteasome and mRNA metabolism. Biochem. Biophys. Res. Commun. 2011, 416, 258–265. [Google Scholar] [CrossRef]
- Alvarez-Castelao, B.; Goethals, M.; Vandekerckhove, J.; Castano, J.G. Mechanism of cleavage of alpha-synuclein by the 20S proteasome and modulation of its degradation by the RedOx state of the N-terminal methionines. Biochim. Biophys. Acta 2014, 1843, 352–365. [Google Scholar] [CrossRef] [PubMed]
- Fan, J.; Zhang, Y.Q.; Li, P.; Hou, M.; Tan, L.; Wang, X.; Zhu, Y.S. Interaction of plasminogen activator inhibitor-2 and proteasome subunit, beta type 1. Acta Biochim. Biophys. Sin. 2004, 36, 42–46. [Google Scholar] [CrossRef]
- Tambyrajah, W.S.; Bowler, L.D.; Medina-Palazon, C.; Sinclair, A.J. Cell cycle-dependent caspase-like activity that cleaves p27KIP1 is the β1 subunit of the 20S proteasome. Arch. Biochem. Biophys. 2007, 466, 186–193. [Google Scholar] [CrossRef] [PubMed]
- Yuan, F.; Ma, Y.; You, P.; Lin, W.; Lu, H.; Yu, Y.; Wang, X.; Jiang, J.; Yang, P.; Ma, Q.; et al. A novel role of proteasomal beta1 subunit in tumorigenesis. Biosci. Rep. 2013, 33, 555–565. [Google Scholar] [CrossRef]
- MacDonald, J.I.; Verdi, J.M.; Meakin, S.O. Activity-dependent interaction of the intracellular domain of rat Trka with intermediate filament proteins, the β-6 proteasomal subunit, Ras-GRF1, and the p162 subunit of eIF3. J. Mol. Neurosci. 1999, 13, 141–158. [Google Scholar] [CrossRef] [PubMed]
- Beraud, C.; Greene, W.C. Interaction of HTLV-I Tax with the human proteasome: Implications for NF-κB induction. J. Acquir. Immune Defic. Syndr. Hum. Retrovirol. 1996, 13, S76–S84. [Google Scholar] [CrossRef] [PubMed]
- Gruendler, C.; Lin, Y.; Farley, J.; Wang, T. Proteasomal degradation of Smad1 induced by bone morphogenetic proteins. J. Biol. Chem. 2001, 276, 46533–46543. [Google Scholar] [CrossRef] [PubMed]
- Erales, J.; Coffino, P. Ubiquitin-independent proteasomal degradation. Biochim. Biophys. Acta 2014, 1843, 216–221. [Google Scholar] [CrossRef] [PubMed]
- Adler, J.; Reuven, N.; Kahana, C.; Shaul, Y. c-Fos proteasomal degradation is activated by a default mechanism, and its regulation by NAD(P)H:quinone oxidoreductase 1 determines c-Fos serum response kinetics. Mol. Cell Biol. 2010, 30, 3767–3778. [Google Scholar] [CrossRef] [PubMed]
- Pakay, J.L.; Diesch, J.; Gilan, O.; Yip, Y.Y.; Sayan, E.; Kolch, W.; Mariadason, J.M.; Hannan, R.D.; Tulchinsky, E.; Dhillon, A.S. A 19S proteasomal subunit cooperates with an ERK MAPK-regulated degron to regulate accumulation of Fra-1 in tumour cells. Oncogene 2012, 31, 1817–1824. [Google Scholar] [CrossRef] [PubMed]
- Keppler, B.R.; Archer, T.K. Ubiquitin-dependent and ubiquitin-independent control of subunit stoichiometry in the SWI/SNF complex. J. Biol. Chem. 2010, 285, 35665–35674. [Google Scholar] [CrossRef] [PubMed]
- Ban, Y.; Ho, C.W.; Lin, R.K.; Lyu, Y.L.; Liu, L.F. Activation of a novel ubiquitin-independent proteasome pathway when RNA polymerase II encounters a protein roadblock. Mol. Cell Biol. 2013, 33, 4008–4016. [Google Scholar] [CrossRef] [PubMed]
- Chen, C.; Zhou, Z.; Guo, P.; Dong, J.T. Proteasomal degradation of the KLF5 transcription factor through a ubiquitin-independent pathway. FEBS Lett. 2007, 581, 1124–1130. [Google Scholar] [CrossRef] [PubMed]
- Dulloo, I.; Gopalan, G.; Melino, G.; Sabapathy, K. The antiapoptotic DeltaNp73 is degraded in a c-Jun-dependent manner upon genotoxic stress through the antizyme-mediated pathway. Proc. Natl. Acad. Sci. USA 2010, 107, 4902–4907. [Google Scholar] [CrossRef] [PubMed]
- Lindner, J.M.; Wong, C.S.; Moller, A.; Nielsen, P.J. A C-terminal acidic domain regulates degradation of the transcriptional coactivator Bob1. Mol. Cell Biol. 2013, 33, 4628–4640. [Google Scholar] [CrossRef] [PubMed]
- Park, K.C.; Jeong, J.; Kim, K.I. Regulation of mIkappaBNS stability through PEST-mediated degradation by proteasome. Biochem. Biophys. Res. Commun. 2014, 443, 1291–1295. [Google Scholar] [CrossRef] [PubMed]
- Lim, S.K.; Gopalan, G. Antizyme1 mediates AURKAIP1-dependent degradation of Aurora-A. Oncogene 2007, 26, 6593–6603. [Google Scholar] [CrossRef] [PubMed]
- Hwang, J.; Kalejta, R.F. Proteasome-dependent, ubiquitin-independent degradation of Daxx by the viral pp71 protein in human cytomegalovirus-infected cells. Virology 2007, 367, 334–338. [Google Scholar] [CrossRef] [PubMed]
- Stewart, D.P.; Koss, B.; Bathina, M.; Perciavalle, R.M.; Bisanz, K.; Opferman, J.T. Ubiquitin-independent degradation of antiapoptotic MCL-1. Mol. Cell Biol. 2010, 30, 3099–3110. [Google Scholar] [CrossRef] [PubMed]
- Wiggins, C.M.; Tsvetkov, P.; Johnson, M.; Joyce, C.L.; Lamb, C.A.; Bryant, N.J.; Komander, D.; Shaul, Y.; Cook, S.J. BIM(EL), an intrinsically disordered protein, is degraded by 20S proteasomes in the absence of poly-ubiquitylation. J. Cell Sci. 2011, 124, 969–977. [Google Scholar] [CrossRef] [PubMed]
- Craxton, A.; Butterworth, M.; Harper, N.; Fairall, L.; Schwabe, J.; Ciechanover, A.; Cohen, G.M. NOXA, a sensor of proteasome integrity, is degraded by 26S proteasomes by an ubiquitin-independent pathway that is blocked by MCL-1. Cell Death Differ. 2012, 19, 1424–1434. [Google Scholar] [CrossRef] [PubMed]
- Rao, V.; Guan, B.; Mutton, L.N.; Bieberich, C.J. Proline-mediated proteasomal degradation of the prostate-specific tumor suppressor NKX3.1. J. Biol. Chem. 2012, 287, 36331–36340. [Google Scholar] [CrossRef] [PubMed]
- Lo, D.; Dai, M.S.; Sun, X.X.; Zeng, S.X.; Lu, H. Ubiquitin- and MDM2 E3 ligase-independent proteasomal turnover of nucleostemin in response to GTP depletion. J. Biol. Chem. 2012, 287, 10013–10020. [Google Scholar] [CrossRef] [PubMed]
- Kim, J.H.; Sohn, S.Y.; Benedict Yen, T.S.; Ahn, B.Y. Ubiquitin-dependent and -independent proteasomal degradation of hepatitis B virus X protein. Biochem. Biophys. Res. Commun. 2008, 366, 1036–1042. [Google Scholar] [CrossRef] [PubMed]
- Fehr, A.R.; Yu, D. Human cytomegalovirus gene UL21a encodes a short-lived cytoplasmic protein and facilitates virus replication in fibroblasts. J. Virol. 2010, 84, 291–302. [Google Scholar] [CrossRef] [PubMed]
- Bolin, L.L.; Hanson, L.K.; Slater, J.S.; Kerry, J.A.; Campbell, A.E. Murine cytomegalovirus US22 protein pM140 protects its binding partner, pM141, from proteasome-dependent but ubiquitin-independent degradation. J. Virol. 2010, 84, 2164–2168. [Google Scholar] [CrossRef] [PubMed]
- Haqshenas, G. The p7 protein of hepatitis C virus is degraded via the proteasome-dependent pathway. Virus Res. 2013, 176, 211–215. [Google Scholar] [CrossRef] [PubMed]
- Gao, Z.; Gammoh, N.; Wong, P.M.; Erdjument-Bromage, H.; Tempst, P.; Jiang, X. Processing of autophagic protein LC3 by the 20S proteasome. Autophagy 2010, 6, 126–137. [Google Scholar] [CrossRef] [PubMed]
- Su, V.; Nakagawa, R.; Koval, M.; Lau, A.F. Ubiquitin-independent proteasomal degradation of endoplasmic reticulum-localized connexin43 mediated by CIP75. J. Biol. Chem. 2010, 285, 40979–40990. [Google Scholar] [CrossRef] [PubMed]
- Guryanova, O.A.; Drazba, J.A.; Frolova, E.I.; Chumakov, P.M. Actin cytoskeleton remodeling by the alternatively spliced isoform of PDLIM4/RIL protein. J. Biol. Chem. 2011, 286, 26849–26859. [Google Scholar] [CrossRef] [PubMed]
- Su, J.; Cao, X.; Wang, K. A novel degradation signal derived from distal C-terminal frameshift mutations of KCNQ2 protein which cause neonatal epilepsy. J. Biol. Chem. 2011, 286, 42949–42958. [Google Scholar] [CrossRef] [PubMed]
- Alvarez-Castelao, B.; Munoz, C.; Sanchez, I.; Goethals, M.; Vandekerckhove, J.; Castano, J.G. Reduced protein stability of human DJ-1/PARK7 L166P, linked to autosomal recessive Parkinson disease, is due to direct endoproteolytic cleavage by the proteasome. Biochim. Biophys. Acta 2012, 1823, 524–533. [Google Scholar] [CrossRef] [PubMed]
- Bergiers, I.; Bridoux, L.; Nguyen, N.; Twizere, J.C.; Rezsohazy, R. The homeodomain transcription factor Hoxa2 interacts with and promotes the proteasomal degradation of the E3 ubiquitin protein ligase RCHY1. PLoS One 2013, 8, e80387. [Google Scholar] [CrossRef] [PubMed]
- Uchimura, Y.; Barton, L.F.; Rada, C.; Neuberger, M.S. REG-gamma associates with and modulates the abundance of nuclear activation-induced deaminase. J. Exp. Med. 2011, 208, 2385–2391. [Google Scholar] [CrossRef] [PubMed]
- Kanai, K.; Aramata, S.; Katakami, S.; Yasuda, K.; Kataoka, K. Proteasome activator PA28γ stimulates degradation of GSK3-phosphorylated insulin transcription activator MAFA. J. Mol. Endocrinol. 2011, 47, 119–127. [Google Scholar] [CrossRef] [PubMed]
- Zhang, Z.; Zhang, R. Proteasome activator PA28γ regulates p53 by enhancing its MDM2-mediated degradation. EMBO J. 2008, 27, 852–864. [Google Scholar] [CrossRef] [PubMed]
- Moriishi, K.; Okabayashi, T.; Nakai, K.; Moriya, K.; Koike, K.; Murata, S.; Chiba, T.; Tanaka, K.; Suzuki, R.; Suzuki, T.; et al. Proteasome activator PA28γ-dependent nuclear retention and degradation of hepatitis C virus core protein. J. Virol. 2003, 77, 10237–10249. [Google Scholar] [CrossRef] [PubMed]
- Suzuki, R.; Moriishi, K.; Fukuda, K.; Shirakura, M.; Ishii, K.; Shoji, I.; Wakita, T.; Miyamura, T.; Matsuura, Y.; Suzuki, T. Proteasomal turnover of hepatitis C virus core protein is regulated by two distinct mechanisms: A ubiquitin-dependent mechanism and a ubiquitin-independent but PA28γ-dependent mechanism. J. Virol. 2009, 83, 2389–2392. [Google Scholar] [CrossRef] [PubMed]
- Dange, T.; Smith, D.; Noy, T.; Rommel, P.C.; Jurzitza, L.; Cordero, R.J.; Legendre, A.; Finley, D.; Goldberg, A.L.; Schmidt, M. Blm10 protein promotes proteasomal substrate turnover by an active gating mechanism. J. Biol. Chem. 2011, 286, 42830–42839. [Google Scholar] [CrossRef] [PubMed]
- David, D.C.; Layfield, R.; Serpell, L.; Narain, Y.; Goedert, M.; Spillantini, M.G. Proteasomal degradation of tau protein. J. Neurochem. 2002, 83, 176–185. [Google Scholar] [CrossRef] [PubMed]
- Qian, M.X.; Pang, Y.; Liu, C.H.; Haratake, K.; Du, B.Y.; Ji, D.Y.; Wang, G.F.; Zhu, Q.Q.; Song, W.; Yu, Y.; et al. Acetylation-mediated proteasomal degradation of core histones during DNA repair and spermatogenesis. Cell 2013, 153, 1012–1024. [Google Scholar] [CrossRef] [PubMed]
- Gille, C.; Goede, A.; Schloetelburg, C.; Preissner, R.; Kloetzel, P.M.; Gobel, U.B.; Frommel, C. A comprehensive view on proteasomal sequences: Implications for the evolution of the proteasome. J. Mol. Biol. 2003, 326, 1437–1448. [Google Scholar] [CrossRef] [PubMed]
- Kumatori, A.; Tanaka, K.; Inamura, N.; Sone, S.; Ogura, T.; Matsumoto, T.; Tachikawa, T.; Shin, S.; Ichihara, A. Abnormally high expression of proteasomes in human leukemic cells. Proc. Natl. Acad. Sci. USA 1990, 87, 7071–7075. [Google Scholar] [CrossRef] [PubMed]
- Nandi, D.; Woodward, E.; Ginsburg, D.B.; Monaco, J.J. Intermediates in the formation of mouse 20S proteasomes: Implications for the assembly of precursor beta subunits. EMBO J. 1997, 16, 5363–5375. [Google Scholar] [CrossRef] [PubMed]
- Cuervo, A.M.; Palmer, A.; Rivett, A.J.; Knecht, E. Degradation of proteasomes by lysosomes in rat liver. Eur. J. Biochem. 1995, 227, 792–800. [Google Scholar] [CrossRef] [PubMed]
- Borris, T.J.; Gebhart, D.L. An unusual case of an erupting microsupernumerary tooth. Gen. Dent. 1991, 39, 286–287. [Google Scholar] [PubMed]
- Hwang, C.S.; Shemorry, A.; Varshavsky, A. N-terminal acetylation of cellular proteins creates specific degradation signals. Science 2010, 327, 973–977. [Google Scholar] [CrossRef] [PubMed]
- Kim, H.K.; Kim, R.R.; Oh, J.H.; Cho, H.; Varshavsky, A.; Hwang, C.S. The N-terminal methionine of cellular proteins as a degradation signal. Cell 2014, 156, 158–169. [Google Scholar] [CrossRef] [PubMed]
- Melo, S.P.; Barbour, K.W.; Berger, F.G. Cooperation between an intrinsically disordered region and a helical segment is required for ubiquitin-independent degradation by the proteasome. J. Biol. Chem. 2011, 286, 36559–36567. [Google Scholar] [CrossRef] [PubMed]
- Singh Gautam, A.K.; Balakrishnan, S.; Venkatraman, P. Direct ubiquitin independent recognition and degradation of a folded protein by the eukaryotic proteasomes-origin of intrinsic degradation signals. PLoS One 2012, 7, e34864. [Google Scholar] [CrossRef] [PubMed]
- Alfassy, O.S.; Cohen, I.; Reiss, Y.; Tirosh, B.; Ravid, T. Placing a disrupted degradation motif at the C terminus of proteasome substrates attenuates degradation without impairing ubiquitylation. J. Biol. Chem. 2013, 288, 12645–12653. [Google Scholar] [CrossRef] [PubMed]
- Dyson, H.J.; Wright, P.E. Intrinsically unstructured proteins and their functions. Nat. Rev. Mol. Cell Biol. 2005, 6, 197–208. [Google Scholar] [CrossRef] [PubMed]
- Suskiewicz, M.J.; Sussman, J.L.; Silman, I.; Shaul, Y. Context-dependent resistance to proteolysis of intrinsically disordered proteins. Protein Sci. 2011, 20, 1285–1297. [Google Scholar] [CrossRef] [PubMed]
- Groll, M.; Ditzel, L.; Lowe, J.; Stock, D.; Bochtler, M.; Bartunik, H.D.; Huber, R. Structure of 20S proteasome from yeast at 2.4 Å resolution. Nature 1997, 386, 463–471. [Google Scholar] [CrossRef] [PubMed]
© 2014 by the authors; licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution license ( http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Sánchez-Lanzas, R.; Castaño, J.G. Proteins Directly Interacting with Mammalian 20S Proteasomal Subunits and Ubiquitin-Independent Proteasomal Degradation. Biomolecules 2014, 4, 1140-1154. https://doi.org/10.3390/biom4041140
Sánchez-Lanzas R, Castaño JG. Proteins Directly Interacting with Mammalian 20S Proteasomal Subunits and Ubiquitin-Independent Proteasomal Degradation. Biomolecules. 2014; 4(4):1140-1154. https://doi.org/10.3390/biom4041140
Chicago/Turabian StyleSánchez-Lanzas, Raúl, and José G. Castaño. 2014. "Proteins Directly Interacting with Mammalian 20S Proteasomal Subunits and Ubiquitin-Independent Proteasomal Degradation" Biomolecules 4, no. 4: 1140-1154. https://doi.org/10.3390/biom4041140




