Multi-Spectroscopic Analysis of Seed Quality and 13C-Stable-Iotopologue Monitoring in Initial Growth Metabolism of Jatropha curcas L.
Abstract
:1. Introduction
2. Results and Discussion
2.1. Metabolic-Analysis-Based Quality Control Methods for Jatropha Seed
| 1R12 | 2R09 | 2R11 | 2R12 | 2F12 | 3R12 | 3F12 | |
|---|---|---|---|---|---|---|---|
| number of germinated seeds [-] | 60 | 138 | 6 | 0 | 63 | 2 | 48 |
| number of seeds [-] | 80 | 208 | 13 | 30 | 79 | 39 | 79 |
| germination rate [%] | 75.0 | 66.3 | 46.2 | 0.0 | 79.7 | 5.1 | 60.8 |
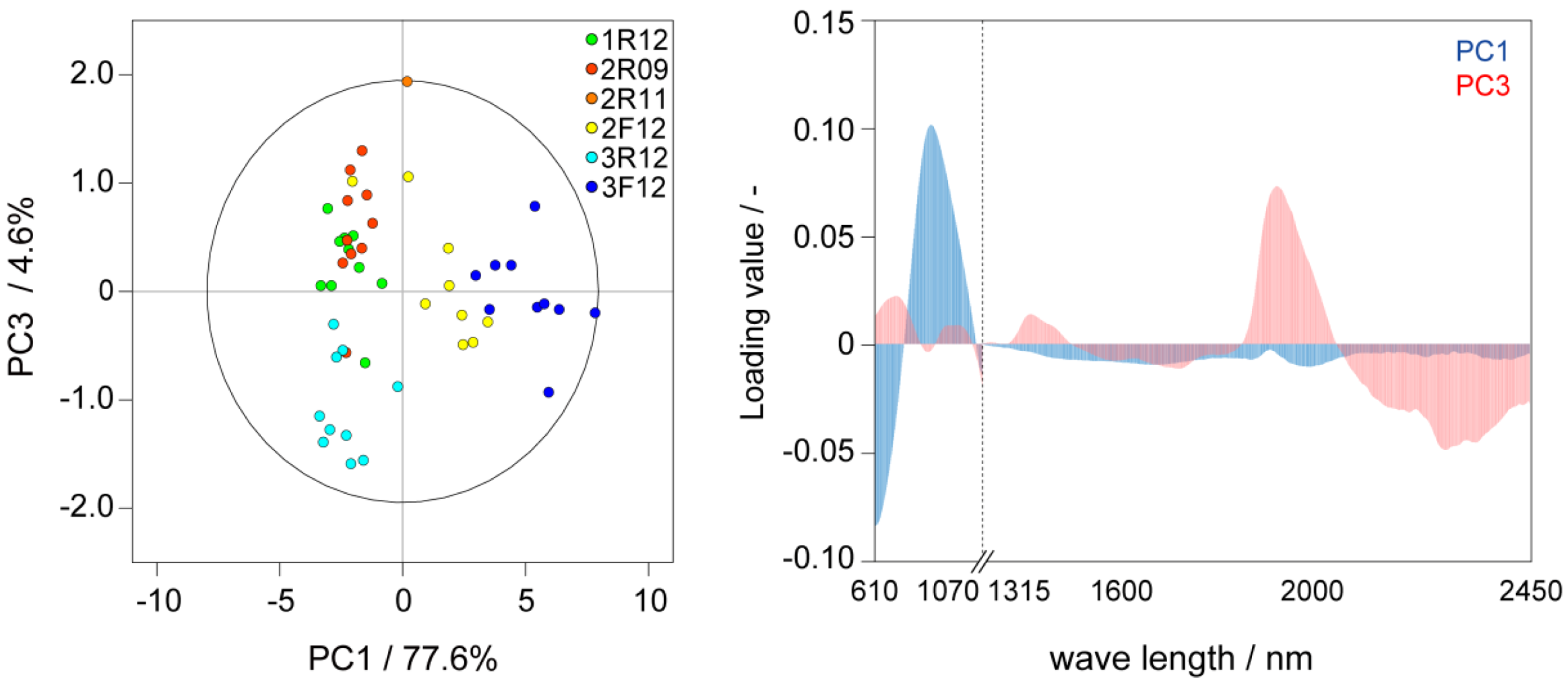
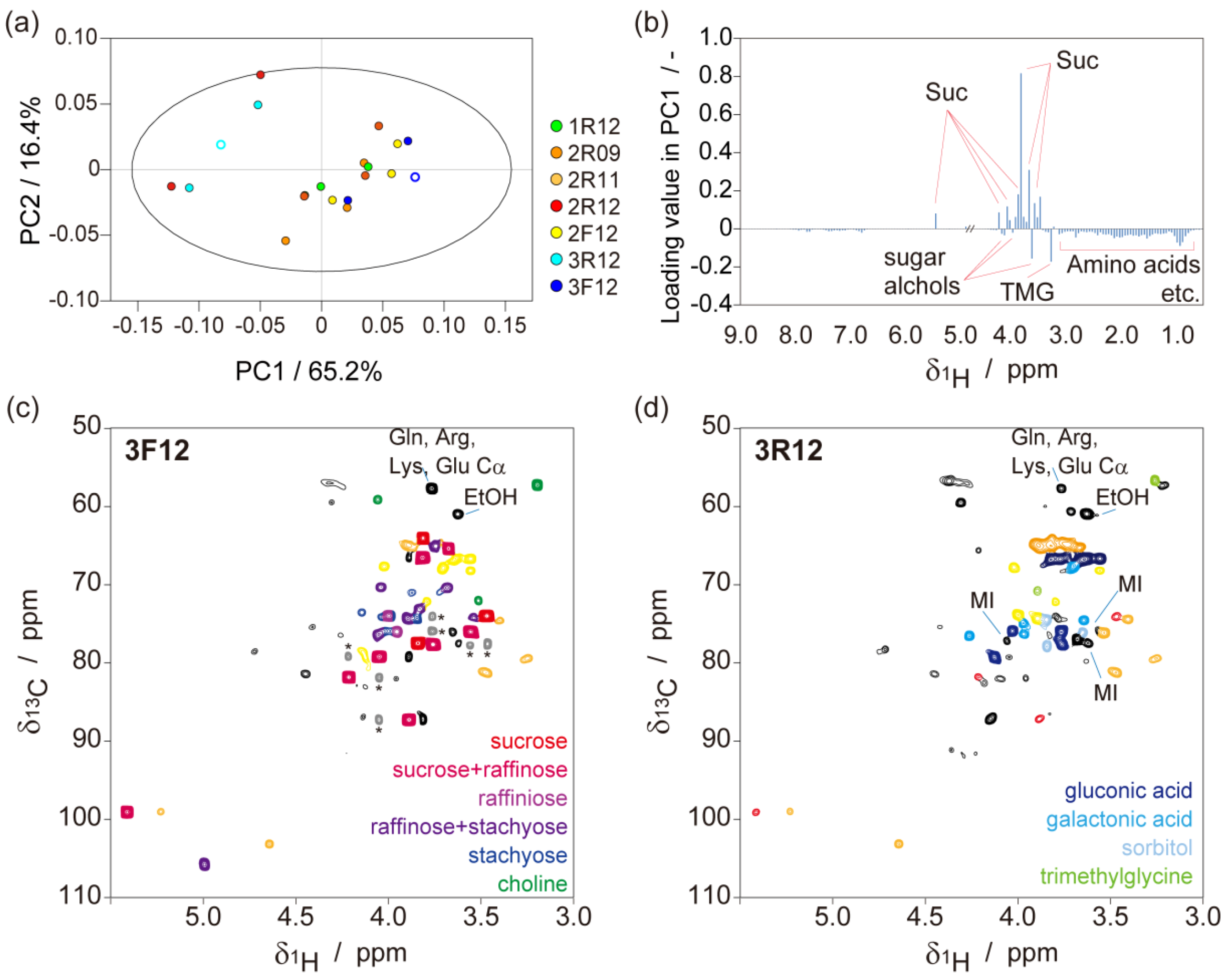
2.2. NMR-Based Metabolic Analysis in Primary Growth of J. curcas.
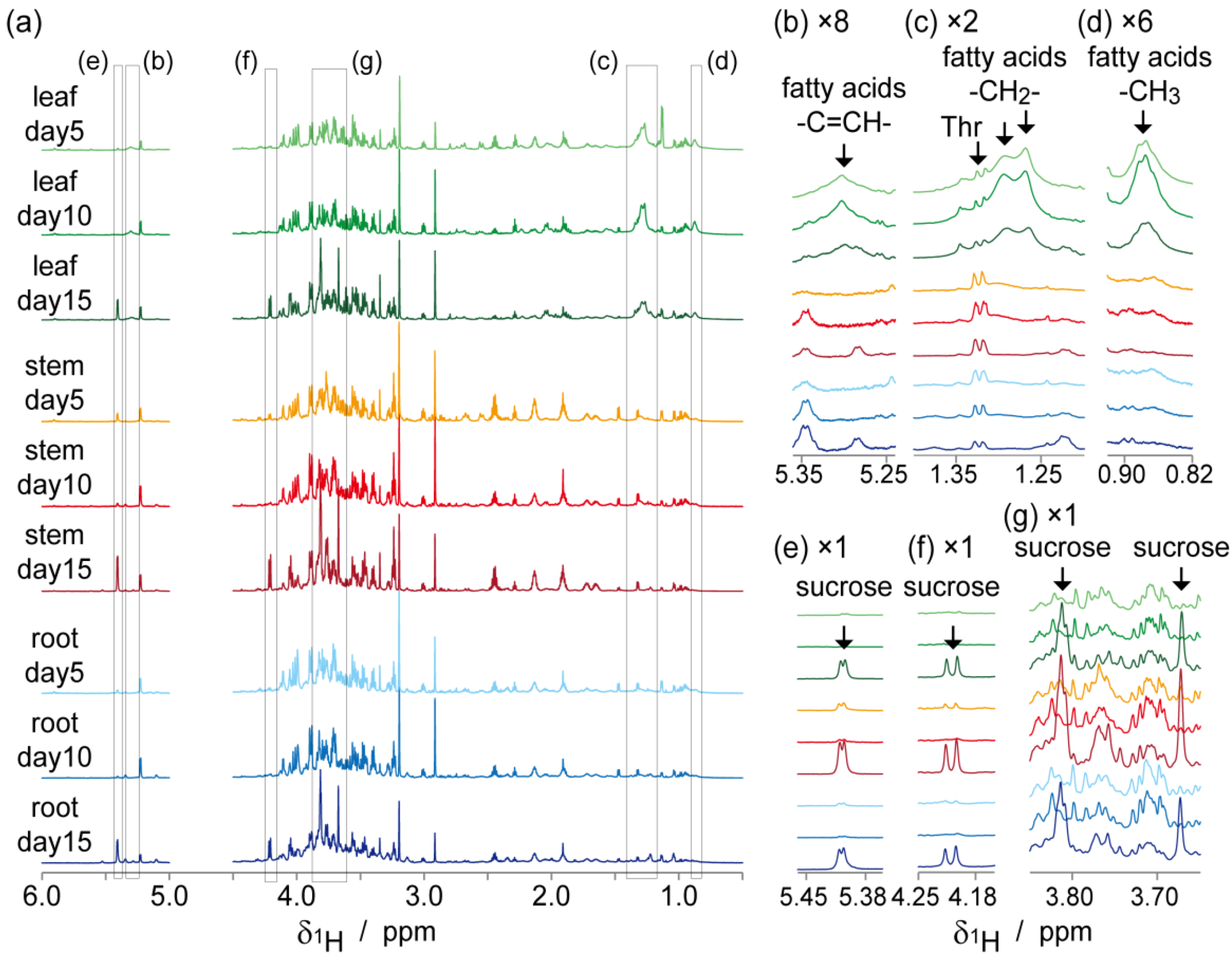
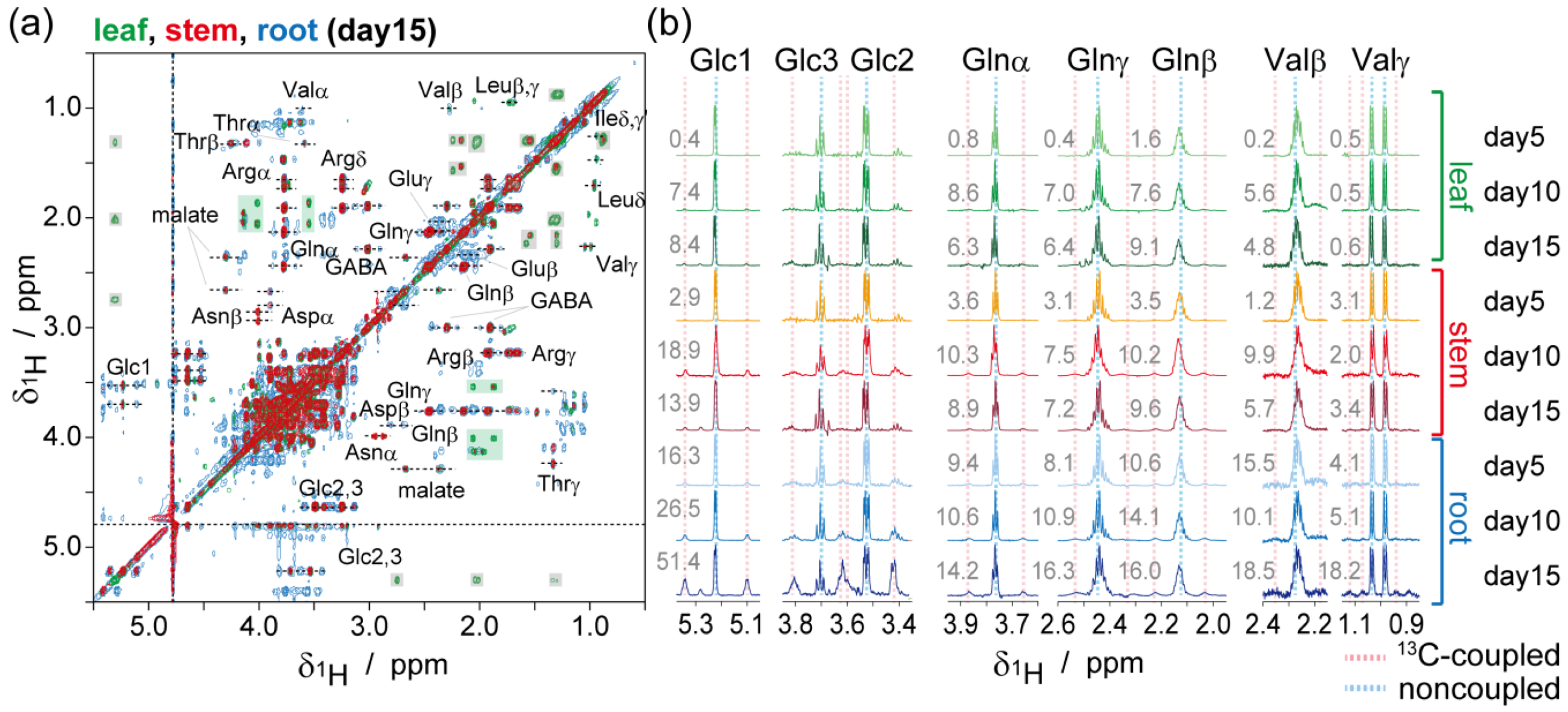
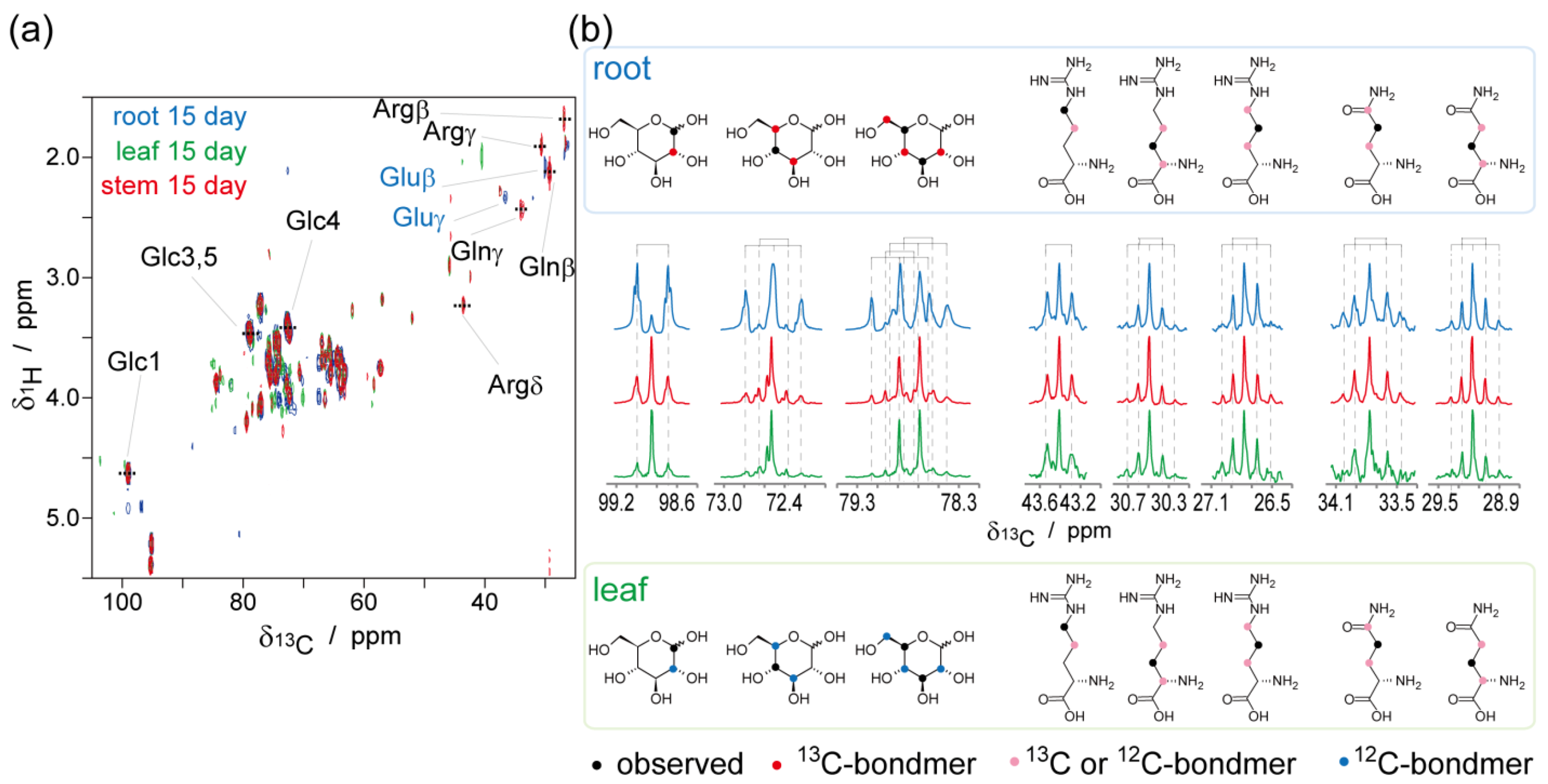
3. Experimental Section
3.1. Chemicals and Plant Materials
3.2. Spectroscopic Analysis
3.3. Multivariable Analysis of NIR and NMR Spectra
4. Conclusions

Supplementary Files
Supplementary File 1Acknowledgments
Author Contributions
Conflicts of Interest
References
- Fairless, D. Biofuel: The little shrub that could—maybe. Nature 2007, 449, 652–655. [Google Scholar] [CrossRef]
- Achten, W.M.J.; Verchot, L.; Franken, Y.J.; Mathijs, E.; Singh, V.P.; Aerts, R.; Muys, B. Jatropha bio-diesel production and use. Biomass Bioenergy 2008, 32, 1063–1084. [Google Scholar] [CrossRef] [Green Version]
- Martinez-Herrera, J.; Siddhuraju, P.; Francis, G.; Davila-Ortiz, G.; Becker, K. Chemical composition, toxic/antimetabolic constituents, and effects of different treatments on their levels, in four provenances of Jatropha curcas L. From mexico. Food Chem. 2006, 96, 80–89. [Google Scholar] [CrossRef]
- Maghuly, F.; Laimer, M. Jatropha curcas, a biofuel crop: Functional genomics for understanding metabolic pathways and genetic improvement. Biotechnol. J. 2013, 8, 1172–1182. [Google Scholar] [CrossRef] [PubMed]
- Sato, S.; Hirakawa, H.; Isobe, S.; Fukai, E.; Watanabe, A.; Kato, M.; Kawashima, K.; Minami, C.; Muraki, A.; Nakazaki, N.; et al. Sequence analysis of the genome of an oil-bearing tree, Jatropha curcas L. DNA Res. 2011, 18, 65–76. [Google Scholar] [CrossRef] [PubMed]
- Hirakawa, H.; Tsuchimoto, S.; Sakai, H.; Nakayama, S.; Fujishiro, T.; Kishida, Y.; Kohara, M.; Watanabe, A.; Yamada, M.; Aizu, T.; et al. Upgraded genomic information of Jatropha curcas L. Plant Biotechnol. 2012, 29, 123–130. [Google Scholar] [CrossRef]
- Finch-Savage, W.E.; Leubner-Metzger, G. Seed dormancy and the control of germination. New Phytol. 2006, 171, 501–523. [Google Scholar] [CrossRef] [PubMed]
- Moncaleano-Escandon, J.; Silva, B.C.F.; Silva, S.R.S.; Granja, J.A.A.; Alves, M.C.J.L.; Pompelli, M.F. Germination responses of Jatropha curcas L. Seeds to storage and aging. Ind. Crops Prod. 2013, 44, 684–690. [Google Scholar] [CrossRef]
- Costa, G.G.L.; Cardoso, K.C.; Del Bem, L.E.V.; Lima, A.C.; Cunha, M.A.S.; de Campos-Leite, L.; Vicentini, R.; Papes, F.; Moreira, R.C.; Yunes, J.A.; et al. Transcriptome analysis of the oil-rich seed of the bioenergy crop Jatropha curcas L. BMC Genomics 2010, 11. [Google Scholar] [CrossRef] [PubMed]
- Natarajan, P.; Parani, M. De novo assembly and transcriptome analysis of five major tissues of Jatropha curcas L. Using gs flx titanium platform of 454 pyrosequencing. BMC Genomics 2011, 12. [Google Scholar] [CrossRef] [PubMed]
- Yang, M.F.; Liu, Y.J.; Liu, Y.; Chen, H.; Chen, F.; Shen, S.H. Proteomic analysis of oil mobilization in seed germination and postgermination development of Jatropha curcas. J. Proteome Res. 2009, 8, 1441–1451. [Google Scholar] [CrossRef] [PubMed]
- Liu, H.; Liu, Y.J.; Yang, M.F.; Shen, S.H. A comparative analysis of embryo and endosperm proteome from seeds of Jatropha curcas. J. Integr. Plant Biol. 2009, 51, 850–857. [Google Scholar] [CrossRef] [PubMed]
- Liu, H.; Yang, Z.L.; Yang, M.F.; Shen, S.H. The differential proteome of endosperm and embryo from mature seed of Jatropha curcas. Plant Sci. 2011, 181, 660–666. [Google Scholar] [CrossRef] [PubMed]
- Bingol, K.; Bruschweiler, R. Multidimensional approaches to NMR-based metabolomics. Anal. Chem. 2014, 86, 47–57. [Google Scholar] [CrossRef] [PubMed]
- Bingol, K.; Zhang, F.; Bruschweiler-Li, L.; Bruschweiler, R. Carbon backbone topology of the metabolome of a cell. J. Am. Chem. Soc. 2012, 134, 9006–9011. [Google Scholar] [CrossRef] [PubMed]
- Chikayama, E.; Sekiyama, Y.; Okamoto, Y.; Nakanishi, Y.; Tsuboi, Y.; Akiyama, K.; Saito, K.; Shinozaki, K.; Kikuchi, J. Statistical indices for simultaneous large-scale metabolite detections for a single NMR spectrum. Anal. Chem. 2010, 82, 1653–1658. [Google Scholar] [CrossRef] [PubMed]
- Chikayama, E.; Suto, M.; Nishihara, T.; Shinozaki, K.; Kikuchi, J. Systematic NMR analysis of stable isotope labeled metabolite mixtures in plant and animal systems: Coarse grained views of metabolic pathways. PLoS One 2008, 3, e3805. [Google Scholar] [CrossRef] [PubMed]
- Sekiyama, Y.; Chikayama, E.; Kikuchi, J. Profiling polar and semipolar plant metabolites throughout extraction processes using a combined solution-state and high-resolution magic angle spinning NMR approach. Anal. Chem. 2010, 82, 1643–1652. [Google Scholar] [CrossRef] [PubMed]
- Sekiyama, Y.; Chikayama, E.; Kikuchi, J. Evaluation of a semipolar solvent system as a step toward heteronuclear multidimensional NMR-based metabolomics for 13C-labeled bacteria, plants, and animals. Anal. Chem. 2011, 83, 719–726. [Google Scholar] [CrossRef] [PubMed]
- An, Y.J.; Xu, W.J.; Jin, X.; Wen, H.; Kim, H.; Lee, J.; Park, S. Metabotyping of the C. elegans sir-2.1 mutant using in vivo labeling and 13C-heteronuclear multidimensional NMR metabolomics. ACS Chem. Biol. 2012, 7, 2012–2018. [Google Scholar] [CrossRef] [PubMed]
- Ratcliffe, R.G.; Shachar-Hill, Y. Revealing metabolic phenotypes in plants: Inputs from NMR analysis. Biol. Rev. Camb. Philos. Soc. 2005, 80, 27–43. [Google Scholar] [CrossRef] [PubMed]
- Allen, D.K.; Shachar-Hill, Y.; Ohlrogge, J.B. Compartment-specific labeling information in 13C metabolic flux analysis of plants. Phytochemistry 2007, 68, 2197–2210. [Google Scholar] [CrossRef] [PubMed]
- Flores, S.; Gosset, G.; Flores, N.; de Graaf, A.A.; Bolivar, F. Analysis of carbon metabolism in escherichia coli strains with an inactive phosphotransferase system by 13C labeling and NMR spectroscopy. Metab. Eng. 2002, 4, 124–137. [Google Scholar] [CrossRef] [PubMed]
- Masakapalli, S.K.; Ratcliffe, R.G.; Williams, T.C. Quantification of 13C enrichments and isotopomer abundances for metabolic flux analysis using 1D NMR spectroscopy. Methods Mol. Biol. 2014, 1090, 73–86. [Google Scholar] [PubMed]
- Thrippleton, M.J.; Keeler, J. Elimination of zero-quantum interference in two-dimensional NMR spectra. Angew. Chem.-Int. Ed. 2003, 42, 3938–3941. [Google Scholar] [CrossRef]
- Massou, S.; Nicolas, C.; Letisse, F.; Portais, J.C. NMR-based fluxomics: Quantitative 2D NMR methods for isotopomers analysis. Phytochemistry 2007, 68, 2330–2340. [Google Scholar] [CrossRef] [PubMed]
- Massou, S.; Nicolas, C.; Letisse, F.; Portais, J.C. Application of 2D-TOCSY NMR to the measurement of specific 13C-enrichments in complex mixtures of 13C-labeled metabolites. Metab. Eng. 2007, 9, 252–257. [Google Scholar] [CrossRef] [PubMed]
- Sekiyama, Y.; Kikuchi, J. Towards dynamic metabolic network measurements by multi-dimensional NMR-based fluxomics. Phytochemistry 2007, 68, 2320–2329. [Google Scholar] [CrossRef]
- Tian, C.; Chikayama, E.; Tsuboi, Y.; Kuromori, T.; Shinozaki, K.; Kikuchi, J.; Hirayama, T. Top-down phenomics of arabidopsis thaliana: Metabolic profiling by one- and two-dimensional nuclear magnetic resonance spectroscopy and transcriptome analysis of albino mutants. J. Biol. Chem. 2007, 282, 18532–18541. [Google Scholar] [CrossRef] [PubMed]
- Castillo, E.M.; Delumen, B.O.; Reyes, P.S.; Delumen, H.Z. Raffinose synthase and galactinol synthase in developing seeds and leaves of legumes. J. Agric. Food Chem. 1990, 38, 351–355. [Google Scholar] [CrossRef]
- ElSayed, A.I.; Rafudeen, M.S.; Golldack, D. Physiological aspects of raffinose family oligosaccharides in plants: Protection against abiotic stress. Plant Biol. 2014, 16, 1–8. [Google Scholar] [CrossRef] [PubMed]
- Saravitz, D.M.; Pharr, D.M.; Carter, T.E. Galactinol synthase activity and soluble sugars in developing seeds of 4 soybean genotypes. Plant Physiol. 1987, 83, 185–189. [Google Scholar] [CrossRef] [PubMed]
- Taji, T.; Ohsumi, C.; Iuchi, S.; Seki, M.; Kasuga, M.; Kobayashi, M.; Yamaguchi-Shinozaki, K.; Shinozaki, K. Important roles of drought- and cold-inducible genes for galactinol synthase in stress tolerance in arabidopsis thaliana. Plant J. 2002, 29, 417–426. [Google Scholar] [CrossRef] [PubMed]
- Dierking, E.C.; Bilyeu, K.D. Raffinose and stachyose metabolism are not required for efficient soybean seed germination. J. Plant Physiol. 2009, 166, 1329–1335. [Google Scholar] [CrossRef] [PubMed]
- Fukao, Y.; Hayashi, M.; Hara-Nishimura, I.; Nishimura, M. Novel glyoxysomal protein kinase, gpk1, identified by proteomic analysis of glyoxysomes in etiolated cotyledons of arabidopsis thaliana. Plant Cell Physiol. 2003, 44, 1002–1012. [Google Scholar] [CrossRef] [PubMed]
- Kikuchi, J.; Hirayama, T. Practical aspects of uniform stable isotope labeling of higher plants for heteronuclear NMR-based metabolomics. Methods Mol. Biol. 2007, 358, 273–286. [Google Scholar] [PubMed]
- Kikuchi, J.; Shinozaki, K.; Hirayama, T. Stable isotope labeling of arabidopsis thaliana for an NMR-based metabolomics approach. Plant Cell Physiol. 2004, 45, 1099–1104. [Google Scholar] [CrossRef]
- Eisenreich, W.; Menhard, B.; Hylands, P.J.; Zenk, M.H.; Bacher, A. Studies on the biosynthesis of taxol: The taxane carbon skeleton is not of mevalonoid origin. Proc. Natl. Acad. Sci. USA 1996, 93, 6431–6436. [Google Scholar] [PubMed]
- Goudar, C.; Biener, R.; Boisart, C.; Heidemann, R.; Piret, J.; de Graaf, A.; Konstantinov, K. Metabolic flux analysis of CHO cells in perfusion culture by metabolite balancing and 2D [13C, 1H] COSY NMR spectroscopy. Metab. Eng. 2010, 12, 138–149. [Google Scholar] [CrossRef] [PubMed]
- Szyperski, T.; Bailey, J.E.; Wuthrich, K. Detecting and dissecting metabolic fluxes using biosynthetic fractional 13C labeling and two-dimensional NMR spectroscopy. Trends Biotechnol. 1996, 14, 453–459. [Google Scholar] [CrossRef]
- Bingol, K.; Zhang, F.; Bruschweiler-Li, L.; Bruschweiler, R. Quantitative analysis of metabolic mixtures by two-dimensional 13C constant-time tocsy NMR spectroscopy. Anal. Chem. 2013, 85, 6414–6420. [Google Scholar] [CrossRef] [PubMed]
- Keun, H.C.; Beckonert, O.; Griffin, J.L.; Richter, C.; Moskau, D.; Lindon, J.C.; Nicholson, J.K. Cryogenic probe 13C NMR spectroscopy of urine for metabonomic studies. Anal. Chem. 2002, 74, 4588–4593. [Google Scholar] [CrossRef] [PubMed]
- Lea, P.J.; Ireland, R.J. Nitrogen metabolism in higher plants. In Plant Amino Acids: Biochemistry and Biotechnology; Marcel Dekker Inc: New York, NY, USA, 1999; pp. 1–47. [Google Scholar]
- Betti, M.; Garcia-Calderon, M.; Perez-Delgado, C.M.; Credali, A.; Estivill, G.; Galvan, F.; Vega, J.M.; Marquez, A.J. Glutamine synthetase in legumes: Recent advances in enzyme structure and functional genomics. Int. J. Mol. Sci. 2012, 13, 7994–8024. [Google Scholar] [CrossRef] [PubMed]
- Bernard, S.M.; Habash, D.Z. The importance of cytosolic glutamine synthetase in nitrogen assimilation and recycling. New Phytol. 2009, 182, 608–620. [Google Scholar] [CrossRef] [PubMed]
- Sauter, J.J.; Vancleve, B. Seasonal-variation of amino-acids in the xylem sap of populus x canadensis and its relation to protein body mobilization. Trees-Struct. Funct. 1992, 7, 26–32. [Google Scholar] [CrossRef]
- Dilworth, M.F.; Dure, L. Developmental biochemistry of cotton seed embryogenesis and germination 10. Nitrogen flow from arginine to asparagine in germination. Plant Physiol. 1978, 61, 698–702. [Google Scholar] [CrossRef] [PubMed]
- Escher, P.; Eiblmeier, M.; Hetzger, I.; Rennenberg, H. Spatial and seasonal variation in amino compounds in the xylem sap of a mistletoe (viscum album) and its hosts (populus spp. And abies alba). Tree Physiol. 2004, 24, 639–650. [Google Scholar] [CrossRef] [PubMed]
- Vanetten, C.H.; Kwolek, W.F.; Peters, J.E.; Barclay, A.S. Plant seeds as protein sources for food or feed. Evaluation based on amino acid composition of 379 species. J. Agric. Food Chem. 1967, 15, 1077. [Google Scholar] [CrossRef]
- Takase, T.; Ishikawa, H.; Murakami, H.; Kikuchi, J.; Sato-Nara, K.; Suzuki, H. The circadian clock modulates water dynamics and aquaporin expression in arabidopsis roots. Plant Cell Physiol. 2011, 52, 373–383. [Google Scholar] [CrossRef] [PubMed]
- Melkus, G.; Rolletschek, H.; Fuchs, J.; Radchuk, V.; Grafahrend-Belau, E.; Sreenivasulu, N.; Rutten, T.; Weier, D.; Heinzel, N.; Schreiber, F.; et al. Dynamic 13C/1H NMR imaging uncovers sugar allocation in the living seed. Plant Biotechnol. J. 2011, 9, 1022–1037. [Google Scholar] [CrossRef] [PubMed]
- Rolletschek, H.; Melkus, G.; Grafahrend-Belau, E.; Fuchs, J.; Heinzel, N.; Schreiber, F.; Jakob, P.M.; Borisjuk, L. Combined noninvasive imaging and modeling approaches reveal metabolic compartmentation in the barley endosperm. Plant Cell 2011, 23, 3041–3054. [Google Scholar] [CrossRef] [PubMed]
- Borisjuk, L.; Rolletschek, H.; Neuberger, T. Surveying the plant’s world by magnetic resonance imaging. Plant J. 2012, 70, 129–146. [Google Scholar] [CrossRef] [PubMed]
- Liu, M.L.; Mao, X.A.; Ye, C.H.; Huang, H.; Nicholson, J.K.; Lindon, J.C. Improved watergate pulse sequences for solvent suppression in NMR spectroscopy. J. Magn. Reson. 1998, 132, 125–129. [Google Scholar] [CrossRef]
- The R Project for Statistical Computing. Available online: http://www.r-project.org/ (accessed on 11 November 2014).
© 2014 by the authors; licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Komatsu, T.; Ohishi, R.; Shino, A.; Akashi, K.; Kikuchi, J. Multi-Spectroscopic Analysis of Seed Quality and 13C-Stable-Iotopologue Monitoring in Initial Growth Metabolism of Jatropha curcas L. Metabolites 2014, 4, 1018-1033. https://doi.org/10.3390/metabo4041018
Komatsu T, Ohishi R, Shino A, Akashi K, Kikuchi J. Multi-Spectroscopic Analysis of Seed Quality and 13C-Stable-Iotopologue Monitoring in Initial Growth Metabolism of Jatropha curcas L. Metabolites. 2014; 4(4):1018-1033. https://doi.org/10.3390/metabo4041018
Chicago/Turabian StyleKomatsu, Takanori, Risa Ohishi, Amiu Shino, Kinya Akashi, and Jun Kikuchi. 2014. "Multi-Spectroscopic Analysis of Seed Quality and 13C-Stable-Iotopologue Monitoring in Initial Growth Metabolism of Jatropha curcas L." Metabolites 4, no. 4: 1018-1033. https://doi.org/10.3390/metabo4041018





