Characterizing Microbial Diversity and the Potential for Metabolic Function at −15 °C in the Basal Ice of Taylor Glacier, Antarctica
Abstract
:1. Introduction
2. Methods
2.1. Site Information and Field Sampling
2.2. Ice Decontamination and Sampling
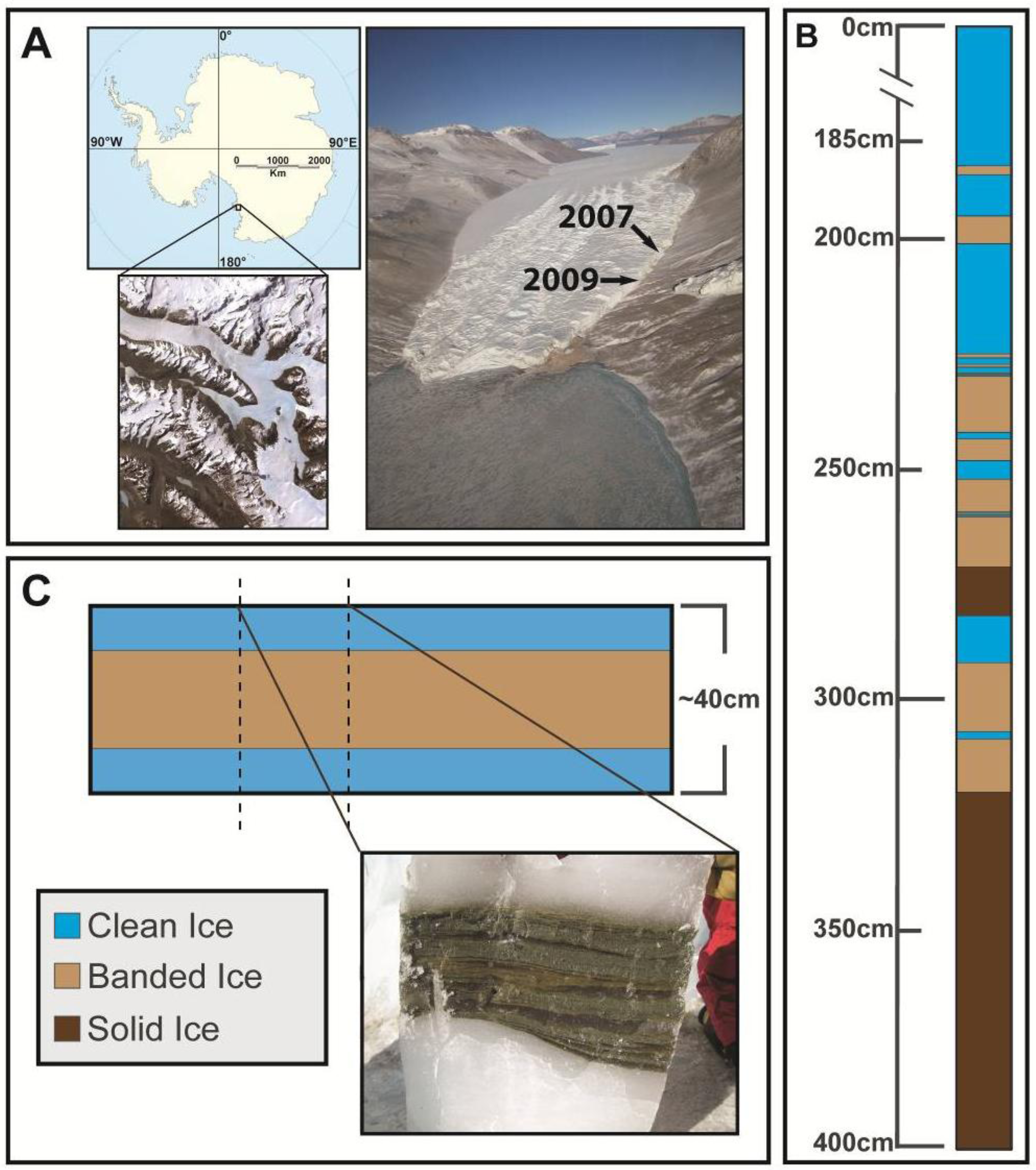
2.3. Microbial Cell Density
2.4. Enrichment and Isolate Culturing
2.5. Molecular Analysis of Bacterial 16S rRNA Genes

| Isolate | Closest Relative | % Identity | Sequence Length (bp) | Isolation Media | Isolation temp | Optimal temp (±5 °C) | Halotolerance (% NaCl) | Description |
|---|---|---|---|---|---|---|---|---|
| TG14 | Paenisporosarcina antarctica N-05 | 99 | 1,379 | R2A | 10 °C | 15 °C | ~7.9% (w/v) | bright yellow, circular, flat |
| TG24 | Paenisporosarcina antarctica N-05 | 99 | 1,363 | marine | 10 °C | 15 °C | ~9.9% (w/v) | tan, circular, convex |
| TG27 | Paenisporosarcina antarctica N-05 | 99 | 1,366 | M9 glucose | 4 °C | 15 °C | ~7.9% (w/v) | off-white, shiny, convex |
| TG29 | Paenisporosarcina antarctica N-05 | 99 | 1,364 | M9 glucose | 4 °C | 22 °C | N.D. | yellow, shiny, convex |
| TG30 | Paenisporosarcina antarctica N-05 | 99 | 1,369 | M9 pyruvate | 4 °C | 22 °C | N.D. | white, shiny, convex |
| TG32 | Paenisporosarcina antarctica N-05 | 99 | 1,364 | 10% R2A | 4 °C | 22 °C | N.D. | off-white, circular, shiny |
| TG25 | Paenisporosarcina antarctica N-05 | 99 | 1,363 | marine | 10 °C | 15 °C | N.D. | cream-yellow, shiny, convex |
| TG34 | Paenisporosarcina antarctica N-05 | 100 | 1,368 | 1% R2A | 4 °C | 22 °C | N.D. | off-white, circular |
| TG21 | Paenisporosarcina macmurdoensis CMS21w | 99 | 1,367 | marine | 22 °C | 22 °C | ~7.9% (w/v) | bright yellow, rough |
| TG3 | Paenisporosarcina macmurdoensis CMS21w | 99 | 1,375 | R2A | 22 °C | 22 °C | ~5.9% (w/v) | off-white center, mucoid |
| TG6 | Paenisporosarcina macmurdoensis CMS21w | 99 | 1,382 | 1% R2A | 22 °C | 22 °C | ~5.9% (w/v) | white-yellow, mucoid |
| TG7 | Paenisporosarcina macmurdoensis CMS21w | 99 | 1,387 | 1% R2A | 22 °C | 22 °C | N.D. | off-white, mucoid |
| TG18 | Paenisporosarcina macmurdoensis CMS21w | 99 | 1,349 | R2A | 22 °C | 22 °C | N.D. | off-white, mucoid |
| TG11 | Paenisporosarcina macmurdoensis CMS21w | 99 | 1,373 | R2A | 22 °C | 15 °C | N.D. | cream yellow, circular |
| TG2 | Paenisporosarcina macmurdoensis CMS21w | 99 | 1,382 | R2A | 22 °C | 22 °C | N.D. | cream yellow, mucoid |
| TG15 | Paenisporosarcina macmurdoensis CMS21w | 99 | 1,376 | R2A | 10 °C | 22 °C | N.D. | yellow, mucoid |
| TG17 | Paenisporosarcina macmurdoensis CMS21w | 99 | 1,377 | R2A | 10 °C | 22 °C | N.D. | white-yellow, convex, mucoid |
| TG26 | Paenisporosarcina macmurdoensis CMS21w | 99 | 1,369 | marine | 10 °C | 22 °C | N.D. | off-white, mucoid |
| TG19 | Paenisporosarcina indica | 99 | 1,369 | marine | 22 °C | 22 °C | ~7.9% (w/v) | dark brown, convex |
| TG9 | Paenisporosarcina indica | 99 | 1,372 | R2A | 22 °C | 22 °C | ~7.9% (w/v) | dull yellow, translucent, flat, mucoid |
| TG20 | Paenisporosarcina indica | 99 | 1,386 | marine | 22 °C | 22 °C | N.D. | cream-yellow, convex |
| TG39 | Paenisporosarcina indica | 99 | 1,370 | marine | 4 °C | 22 °C | N.D. | white, shiny, convex |
| TG10 | Paenisporosarcina quisquiliarum SK 55 | 99 | 1,383 | R2A | 22 °C | 22 °C | ~5.9% (w/v) | off-white, shiny, convex |
| TG8 | Bacillus humi LMG18435 | 97 | 1,367 | 1% R2A | 22 °C | 22 °C | ~5.9% (w/v) | tan with brown center, circular |
| TG4 | Paraliobacillus quinghaiensis YIMC158 | 99 | 1,418 | R2A | 22 °C | 15 °C | ~11.9% (w/v) | yellow, convex, rough |
2.6. DNA and Protein Synthesis of Isolated Bacteria at −15 °C
3. Results
3.1. Cell Concentration and Viability within Horizons of the Basal Ice
3.2. Enrichment Culturing from the Basal Ice
3.3. Phylogenetic Analysis of Clone Sequences and Isolates
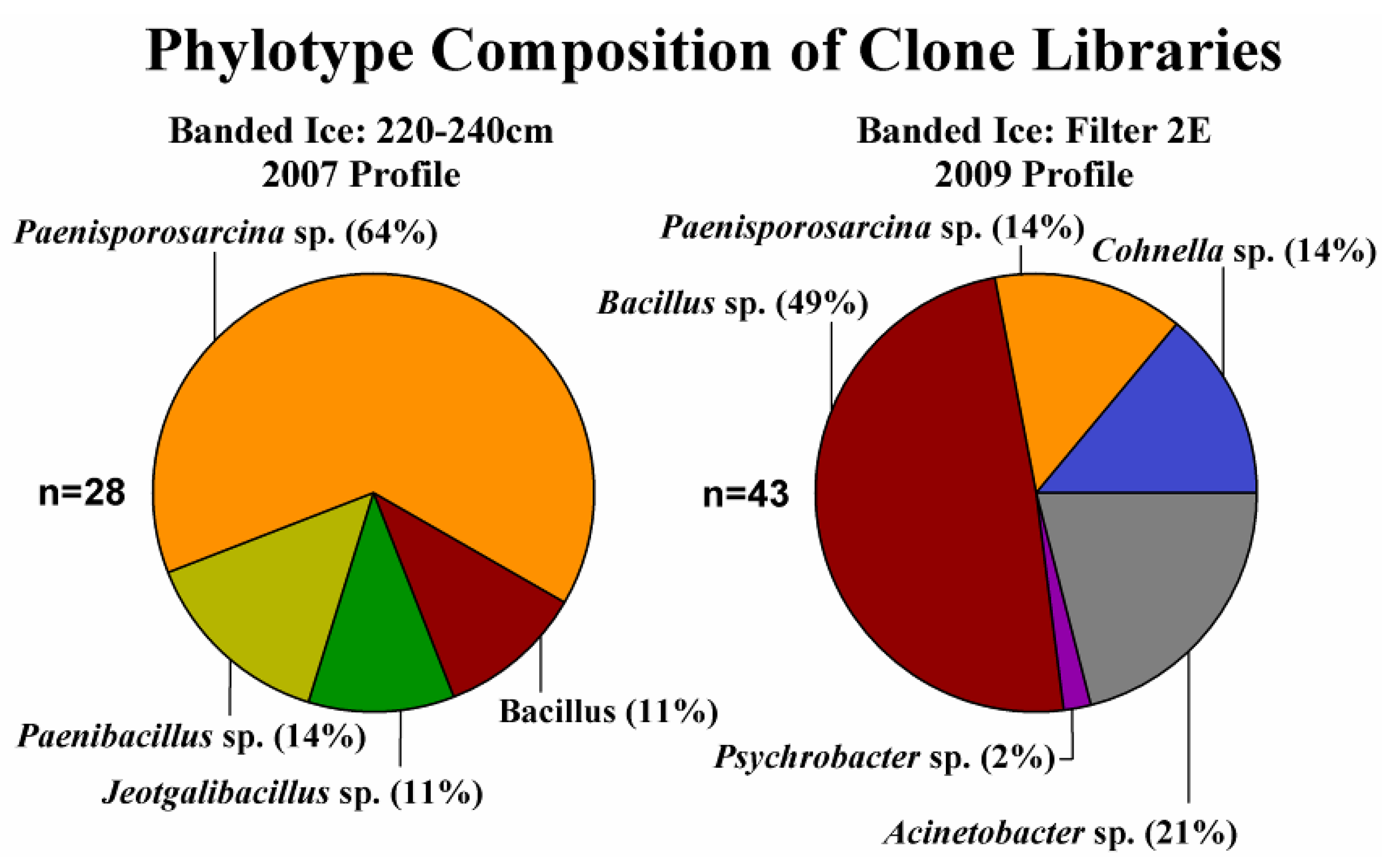
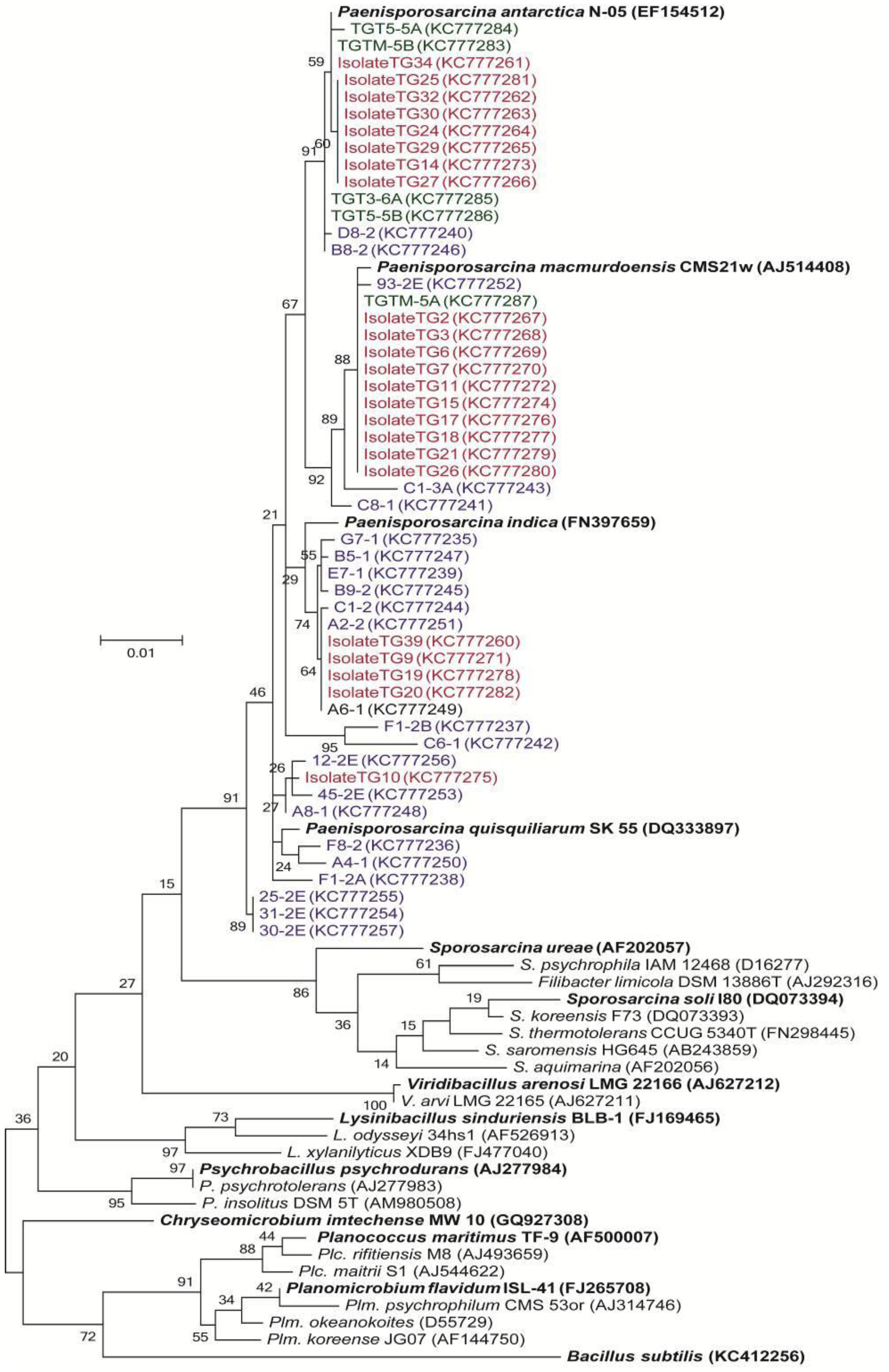
3.4. Incorporation of Macromolecular Precursors in Melt water at −15 °C
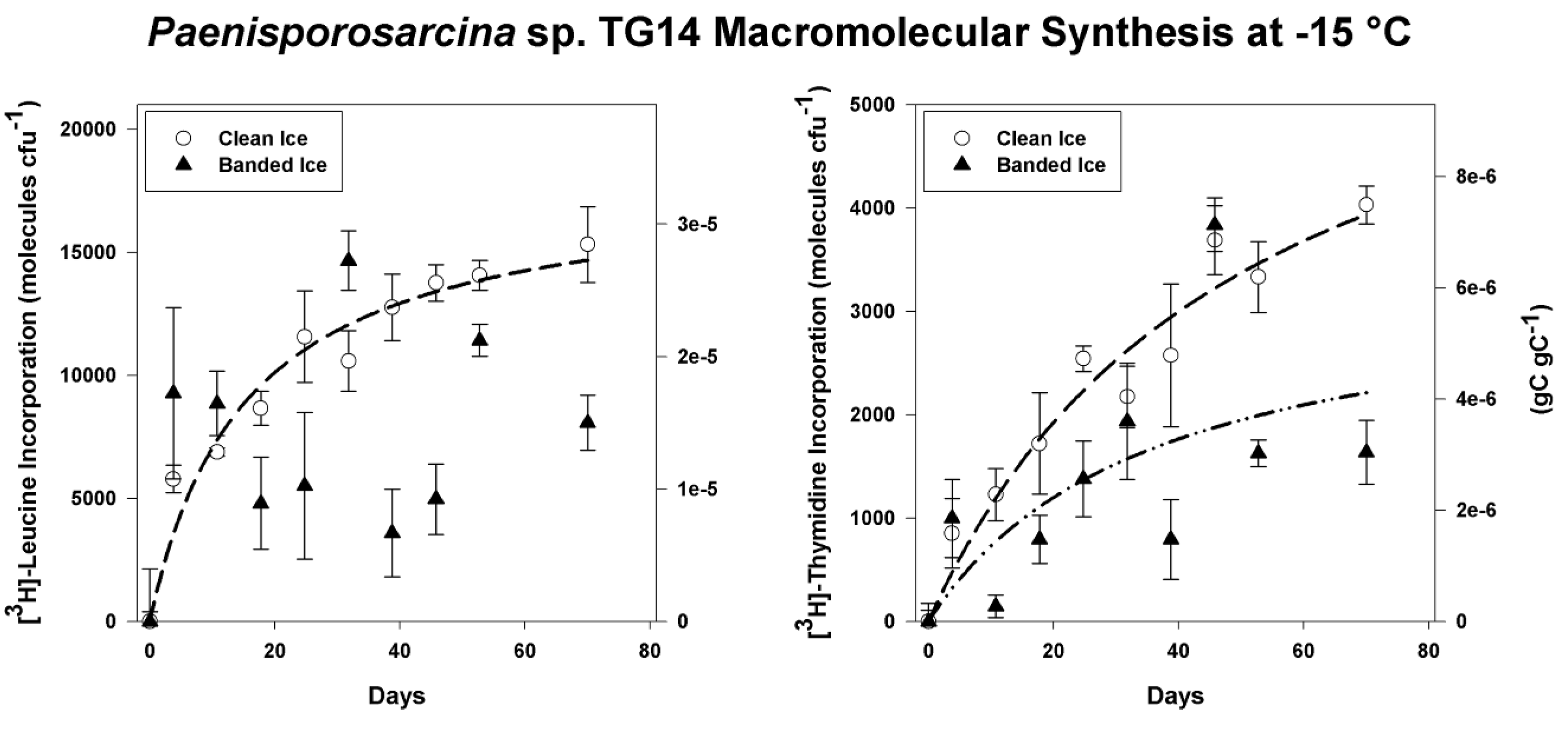
4. Discussion
Acknowledgements
Conflict of Interest
References
- Price, P.B. A habitat for psychrophiles in deep Antarctic ice. Proc. Natl. Acad. Sci. USA 2000, 97, 1247–1251. [Google Scholar] [CrossRef]
- Doyle, S.M.; Dieser, M.; Broemsen, E.; Christner, B.C. General characteristics of cold-adapted microorganisms. In Polar Microbiology: Life In a Deep Freeze; Whyte, L., Miller, R.V., Eds.; ASM Press: Washington, DC, USA, 2012; pp. 103–125. [Google Scholar]
- Miteva, V.I.; Sheridan, P.P.; Brenchley, J.E. Phylogenetic and physiological diversity of microorganisms isolated from a deep Greenland glacier ice core. Appl. Environ. Microbiol. 2004, 70, 202–213. [Google Scholar] [CrossRef]
- Xiang, S.-R.; Shang, T.-C.; Chen, Y.; Yao, T.-D. Deposition and postdeposition mechanisms as possible drivers of microbial population variability in glacier ice. FEMS Microbiol. Ecol. 2009, 70, 165–176. [Google Scholar] [CrossRef]
- Amato, P.; Doyle, S.M.; Battista, J.R.; Christner, B.C. Implications of subzero metabolic activity on long-term microbial survival in terrestrial and extraterrestrial permafrost. Astrobiology 2010, 10, 789–798. [Google Scholar] [CrossRef]
- Abyzov, S.S. Microorganisms in the Antarctic ice. Antarct. Microbiol. 1993, 1, 265–296. [Google Scholar]
- Abyzov, S.S.; Lipenkov, V.Y.; Bobin, N.E.; Koudryashov, B.B. The microflora of the central Antarctic glacier and the control methods of the sterile isolation of the ice core for microbiological analysis. Akad. Nauk SSSR Izv. Ser. Biol. 1982, 4, 537–548. [Google Scholar]
- Antony, R.; Krishnan, K.P.; Laluraj, C.M.; Thamban, M.; Dhakephalkar, P.K.; Engineer, A.S.; Shivaji, S. Diversity and physiology of culturable bacteria associated with a coastal Antarctic ice core. Microbiol. Res. 2012, 167, 372–380. [Google Scholar] [CrossRef]
- Christner, B.C.; Mosley-Thompson, E.; Thompson, L.G.; Reeve, J.N. Bacterial recovery from ancient ice. Environ. Microbiol. 2003, 5, 433–436. [Google Scholar] [CrossRef]
- Christner, B.C.; Mosley-Thompson, E.; Thompson, L.G.; Zagorodnov, V.; Sandman, K.; Reeve, J.N. Recovery and identification of viable bacteria immured in glacial ice. Icarus 2000, 144, 479–485. [Google Scholar] [CrossRef]
- Miteva, V.I.; Brenchley, J.E. Detection and isolation of ultrasmall microorganisms from a 120,000-year-old Greenland glacier ice core. Appl. Environ. Microbiol. 2005, 71, 7806–7818. [Google Scholar] [CrossRef]
- Skidmore, M.L.; Foght, J.M.; Sharp, M.J. Microbial life beneath a high Arctic glacier. Appl. Environ. Microbiol. 2000, 66, 3214–3220. [Google Scholar] [CrossRef]
- Ahn, J.; Wahlen, M.; Deck, B.L.; Brook, J.; Mayewski, P.A.; Taylor, K.C.; White, J.W.C. A record of atmospheric CO2 during the last 40,000 years from the siple dome, Antarctica ice core. J. Geophys. Res. 2004, 109. [Google Scholar] [CrossRef]
- Flückiger, J.; Blunier, T.; Stauffer, B.; Chappellaz, J.; Spahni, R.; Kawamura, K.; Schwander, J.; Stocker, T.F.; Dahl-Jensen, D. N2O and CH4 variations during the last glacial epoch: Insight into global processes. Glob. Biogeochem. Cy. 2004, 18. [Google Scholar] [CrossRef]
- Souchez, R.; Janssens, L.; Lemmens, M.; Stauffer, B. Very low oxygen concentration in basal ice from summit, central Greenland. Geophys. Res. Lett. 1995, 22, 2001–2004. [Google Scholar] [CrossRef]
- Sowers, T. N2O record spanning the penultimate deglaciation from the vostok ice core. J. Geophys. Res. 2001, 106, 31903–31914. [Google Scholar] [CrossRef]
- Campen, R.K.; Sowers, T.; Alley, R.B. Evidence of microbial consortia metabolizing within a low-latitude mountain glacier. Geology 2003, 31, 231–234. [Google Scholar] [CrossRef]
- Souchez, R.; Jouzel, J.; Landais, A.; Chappellaz, J.; Lorrain, R.; Tison, J.-L. Gas isotopes in ice reveal a vegetated central Greenland during ice sheet invasion. Geophys. Res. Lett. 2006, 33. [Google Scholar] [CrossRef]
- Rohde, R.A.; Price, P.B.; Bay, R.C.; Bramall, N.E. In situ microbial metabolism as a cause of gas anomalies in ice. Proc. Natl. Acad. Sci. USA 2008, 105, 8667–8672. [Google Scholar] [CrossRef]
- Tung, H.C.; Bramall, N.E.; Price, P.B. Microbial origin of excess methane in glacial ice and implications for life on Mars. Proc. Natl. Acad. Sci. USA 2005, 102, 18292–18296. [Google Scholar] [CrossRef]
- Bakermans, C.; Skidmore, M. Microbial respiration in ice at subzero temperatures (−4 °C to −33 °C). Environ. Microbiol. Rep. 2011, 3, 774–782. [Google Scholar] [CrossRef]
- Panikov, N.; Flanagan, P.; Oechel, W.; Mastepanov, M.; Christensen, T. Microbial activity in soils frozen to below −39 °C. Soil Biol. Biochem. 2006, 38, 785–794. [Google Scholar] [CrossRef]
- Christner, B.C. Incorporation of DNA and protein precursors into macromolecules by bacteria at −15 °C. Appl. Environ. Microbiol. 2002, 68, 6435–6438. [Google Scholar] [CrossRef]
- Junge, K.; Eicken, H.; Swanson, B.D.; Deming, J.W. Bacterial incorporation of leucine into protein down to −20 °C with evidence for potential activity in sub-eutectic saline ice formations. Cryobiology 2006, 52, 417–429. [Google Scholar] [CrossRef]
- Knight, P.G. The basal ice layer of glaciers and ice sheets. Quat. Sci. Rev. 1997, 16, 975–993. [Google Scholar] [CrossRef]
- Skidmore, M.; Anderson, S.P.; Sharp, M.; Foght, J.; Lanoil, B.D. Comparison of microbial community compositions of two subglacial environments reveals a possible role for microbes in chemical weathering processes. Appl. Environ. Microbiol. 2005, 71, 6986–6997. [Google Scholar] [CrossRef]
- Sharp, M.; Parkes, J.; Cragg, B.; Fairchild, I.J.; Lamb, H.; Tranter, M. Widespread bacterial populations at glacier beds and their relationship to rock weathering and carbon cycling. Geology 1999, 27, 107–110. [Google Scholar] [CrossRef]
- Tung, H.C.; Price, P.B.; Bramall, N.E.; Vrdoljak, G. Microorganisms metabolizing on clay grains in 3-km-deep Greenland basal ice. Astrobiology 2006, 6, 69–86. [Google Scholar] [CrossRef]
- Hubbard, B.; Cook, S.; Coulson, H. Basal Ice facies: A review and unifying approach. Quat. Sci. Rev. 2009, 28, 1956–1969. [Google Scholar] [CrossRef]
- Montross, S.N. Biogeochemistry of Basal Ice from Taylor Glacier, Antarctica. Ph.D. Dissertation, Montana State University, Bozeman, MT, USA, 2012. [Google Scholar]
- Montross, S.M.; Skidmore, M.; Christner, B.; Samyn, D.; Tison, J.L.; Lorrain, R.; Doyle, S.; Fitzsimons, S. Debris-rich basal ice as a microbial habitat, Taylor Glacier, Antarctica. Geomicrobiol. J. 2013. [Google Scholar] [CrossRef]
- Christner, B.C.; Mikucki, J.A.; Foreman, C.M.; Denson, J.; Priscu, J.C. Glacial ice cores: A model system for developing extraterrestrial decontamination protocols. Icarus 2005, 174, 572–584. [Google Scholar] [CrossRef]
- Trevors, J.T.; Cook, S. A Comparison of plating media and diluents for enumeration of aerobic bacteria in a loam soil. J. Microbiol. Meth. 1992, 14, 271–275. [Google Scholar] [CrossRef]
- Samyn, D.; Svensson, A.; Fitzsimons, S. Dynamic implications of discontinuous recrystallization in cold basal ice: Taylor Glacier, Antarctica. J. Geophys. Res. 2008, 113. [Google Scholar] [CrossRef]
- Robe, P.; Nalin, R.; Capellano, C.; Vogel, T.M.; Simonet, P. Extraction of DNA from soil. Eur. J. Soil. Biol. 2003, 39, 183–190. [Google Scholar] [CrossRef]
- Lane, D.J. 16S/23S rRNA sequencing. In Nucleic Acid Techniques in Bacterial Systematics; Stackebrandt, E., Goodfellow, M., Eds.; Wiley: New York, NY, USA, 1991; pp. 115–175. [Google Scholar]
- Pruesse, E.; Peplies, J.; Glöckner, F.O. SINA: Accurate high-throughput multiple sequence alignment of ribosomal RNA genes. Bioinformatics 2012, 28, 1823–1829. [Google Scholar] [CrossRef]
- Pruesse, E.; Quast, C.; Knittel, K.; Fuchs, B.M.; Ludwig, W.; Peplies, J.; Glöckner, F.O. SILVA: A comprehensive online resource for quality checked and aligned ribosomal RNA sequence data compatible with ARB. Nucleic Acids Res. 2007, 35, 7188–7196. [Google Scholar]
- Schloss, P.D.; Westcott, S.L.; Ryabin, T.; Hall, J.R.; Hartmann, M.; Hollister, E.B.; Lesniewski, R.A.; Oakley, B.B.; Parks, D.H.; Robinson, C.J.; et al. Introducing mothur: Open-source, platform-independent, community-supported software for describing and comparing microbial communities. Appl. Environ. Microbiol. 2009, 75, 7537–7541. [Google Scholar] [CrossRef]
- Edgar, R.C.; Haas, B.J.; Clemente, J.C.; Quince, C.; Knight, R. UCHIME improves sensitivity and speed of chimera detection. Bioinformatics 2011, 27, 2194–2200. [Google Scholar] [CrossRef]
- Tamura, K.; Peterson, D.; Peterson, N.; Stecher, G.; Nei, M.; Kumar, S. MEGA5: Molecular evolutionary genetics analysis using maximum likelihood, evolutionary distance, and maximum parsimony methods. Mol. Biol. Evol. 2011, 28, 2731–2739. [Google Scholar] [CrossRef]
- Kim, O.S.; Cho, Y.J.; Lee, K.; Yoon, S.H.; Kim, M.; Na, H.; Park, S.C.; Jeon, Y.S.; Lee, J.H.; Yi, H.; et al. Introducing EzTaxon-E: A prokaryotic 16S rRNA gene sequence database with phylotypes that represent uncultured species. Int. J. Syst. Evol. Microbiol. 2012, 62, 716–721. [Google Scholar] [CrossRef]
- Wang, Q.; Garrity, G.M.; Tiedje, J.M.; Cole, J.R. Naive bayesian classifier for rapid assignment of rRNA sequences into the new bacterial taxonomy. Appl. Environ. Microbiol. 2007, 73, 5261–5267. [Google Scholar] [CrossRef]
- Felsenstein, J. Confidence limits on phylogenies: An approach using the bootstrap. Evolution 1985, 783–791. [Google Scholar]
- Bell, R.E.; Ferraccioli, F.; Creyts, T.T.; Braaten, D.; Corr, H.; Das, I.; Damaske, D.; Frearson, N.; Jordan, T.; Rose, K.; et al. Widespread persistent thickening of the east Antarctic ice sheet by freezing from the base. Science 2011, 331, 1592–1595. [Google Scholar] [CrossRef]
- Wadham, J.; Arndt, S.; Tulaczyk, S.; Stibal, M.; Tranter, M.; Telling, J.; Lis, G.; Lawson, E.; Ridgwell, A.; Dubnick, A. Potential methane reservoirs beneath Antarctica. Nature 2012, 488, 633–637. [Google Scholar] [CrossRef]
- Byrne, S.; Dundas, C.M.; Kennedy, M.R.; Mellon, M.T.; McEwen, A.S.; Cull, S.C.; Daubar, I.J.; Shean, D.E.; Seelos, K.D.; Murchie, S.L. Distribution of mid-latitude ground ice on Mars from new impact craters. Science 2009, 325, 1674–1676. [Google Scholar] [CrossRef]
- Samyn, D.; Fitzsimons, S.J.; Lorrain, R.D. Strain-induced phase changes within cold basal ice from Taylor Glacier, Antarctica, indicated by textural and gas analyses. J. Glaciol. 2005, 51, 611–619. [Google Scholar] [CrossRef]
- Souchez, R.; Lemmens, M.; Chappellaz, J. Flow-induced mixing in the GRIP basal ice deduced from the CO2 and CH4 records. Geophys. Res. Lett. 1995, 22, 41–44. [Google Scholar] [CrossRef]
- Steven, B.; Briggs, G.; McKay, C.P.; Pollard, W.H.; Greer, C.W.; Whyte, L.G. Characterization of the microbial diversity in a permafrost sample from the Canadian High Arctic using culture-dependent and culture-independent methods. FEMS Microbiol. Ecol. 2007, 59, 513–523. [Google Scholar]
- Brinkmeyer, R.; Knittel, K.; Jürgens, J.; Weyland, H.; Amann, R.; Helmke, E. Diversity and structure of bacterial communities in Arctic versus Antarctic pack ice. Appl. Environ. Microbiol. 2003, 69, 6610–6619. [Google Scholar] [CrossRef]
- Cheng, S.M.; Foght, J.M. Cultivation-independent and -dependent characterization of bacteria resident beneath John Evans Glacier. FEMS Microbiol. Ecol. 2007, 59, 318–330. [Google Scholar] [CrossRef]
- Yde, J.C.; Finster, K.W.; Raiswell, R.; Steffensen, J.P.; Heinemeier, J.; Olsen, J.; Gunnlaugsson, H.P.; Nielsen, O.B. Basal ice microbiology at the margin of the Greenland ice sheet. Ann. Glaciol. 2010, 51, 71–79. [Google Scholar]
- Lacelle, D.; Radtke, K.; Clark, I.D.; Fisher, D.; Lauriol, B.; Utting, N.; Whyte, L.G. Geomicrobiology and occluded O2-CO2-Ar gas analyses provide evidence of microbial respiration in ancient terrestrial ground ice. Earth Planet. Sci. Lett. 2011, 306, 46–54. [Google Scholar] [CrossRef]
- Steven, B.; Pollard, W.H.; Greer, C.W.; Whyte, L.G. Microbial diversity and activity through a permafrost/ground ice core profile from the Canadian High Arctic. Environ. Microbiol. 2008, 10, 3388–3403. [Google Scholar] [CrossRef]
- Christner, B.C.; Skidmore, M.L. Montana State University: Bozeman, MT, USA, 2005; Unpublished work.
- Perreault, N.N.; Greer, C.W.; Andersen, D.T.; Tille, S.; Lacrampe-Couloume, G.; Lollar, B.S.; Whyte, L.G. Heterotrophic and autotrophic microbial populations in cold perennial springs of the high Arctic. Appl. Environ. Microbiol. 2008, 74, 6898–6907. [Google Scholar] [CrossRef]
- Reddy, G.; Manasa, B.P.; Singh, S.K.; Shivaji, S. Paenisporosarcina indica sp. nov., a psychrophilic bacterium from pindari glacier of the Himalayan Mountain Ranges and reclassification of sporosarcina Antarctica Yu et al., 2008 as Paenisporosarcina antarctica comb. nov. and emended description of the genus Paenisporosarcina. Int. J. Syst. Evol. Microbiol. 2013, in press. [Google Scholar]
- Reddy, G.; Matsumoto, G.; Shivaji, S. Sporosarcina macmurdoensis sp. nov., from a cyanobacterial mat sample from a pond in the McMurdo Dry Valleys, Antarctica. Int. J. Syst. Evol. Microbiol. 2003, 53, 1363–1367. [Google Scholar] [CrossRef]
- Price, P.B.; Sowers, T. Temperature dependence of metabolic rates for microbial growth, maintenance, and survival. Proc. Natl. Acad. Sci. USA 2004, 101, 4631–4636. [Google Scholar] [CrossRef]
- Barletta, R.E.; Priscu, J.C.; Mader, H.M.; Jones, W.L.; Roe, C.H. Chemical analysis of ice vein microenvironments: II. analysis of glacial samples from Greenland and Antarctica. J. Glaciol. 2012, 58, 1109–1118. [Google Scholar]
- Priscu, J.C.; Hand, K.P. Microbial habitability of icy worlds. Microbe 2012, 7, 167–172. [Google Scholar]
© 2013 by the authors; licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution license (http://creativecommons.org/licenses/by/3.0/).
Share and Cite
Doyle, S.M.; Montross, S.N.; Skidmore, M.L.; Christner, B.C. Characterizing Microbial Diversity and the Potential for Metabolic Function at −15 °C in the Basal Ice of Taylor Glacier, Antarctica. Biology 2013, 2, 1034-1053. https://doi.org/10.3390/biology2031034
Doyle SM, Montross SN, Skidmore ML, Christner BC. Characterizing Microbial Diversity and the Potential for Metabolic Function at −15 °C in the Basal Ice of Taylor Glacier, Antarctica. Biology. 2013; 2(3):1034-1053. https://doi.org/10.3390/biology2031034
Chicago/Turabian StyleDoyle, Shawn M., Scott N. Montross, Mark L. Skidmore, and Brent C. Christner. 2013. "Characterizing Microbial Diversity and the Potential for Metabolic Function at −15 °C in the Basal Ice of Taylor Glacier, Antarctica" Biology 2, no. 3: 1034-1053. https://doi.org/10.3390/biology2031034
APA StyleDoyle, S. M., Montross, S. N., Skidmore, M. L., & Christner, B. C. (2013). Characterizing Microbial Diversity and the Potential for Metabolic Function at −15 °C in the Basal Ice of Taylor Glacier, Antarctica. Biology, 2(3), 1034-1053. https://doi.org/10.3390/biology2031034




