Composition, Diversity, and Stability of Microbial Assemblages in Seasonal Lake Ice, Miquelon Lake, Central Alberta
Abstract
:1. Introduction
2. Methods
2.1. Study Site
2.2. Sample Collection and Processing
2.3. Cell Enumeration
2.4. Chlorophyll a Measurements
2.5. Nucleic Acid Extraction
2.6. Denaturing Gradient Gel Electrophoresis (DGGE)
| Primer set | Target | Sequence (5'-3') | Reference | |
|---|---|---|---|---|
| General Primers | 341F | Bact 16S rRNA | CCTACGGGAGGCAGCAG | [32] |
| 518R | Bact 16S rRNA | ATTACCGCGGCTGCTGG | [32] | |
| Euk1A | Euk 18S rRNA | CTGGTTGATCCTGCCAG | [28] | |
| Euk516R | Euk 18S rRNA | ACCAGACTTGCCCTCC | [28] | |
| A21F external | Arc 16S rRNA | TTCCGGTTGATCCYGCCGGA | [33] | |
| 344F internal | Arc 16S rRNA | ACGGGGCGCAGCAGGCGCGA | [30] | |
| 519R internal | Arc 16S rRNA | GGTDTTACCGCGGCKGCTG | [20] | |
| DGGE Primers | 341F * | Bact 16S rRNA | CCTACGGGAGGCAGCAG | [32] |
| 518R | Bact 16S rRNA | ATTACCGCGGCTGCTGG | [32] | |
| Euk1A | Euk 18S rRNA | CTGGTTGATCCTGCCAG | [28] | |
| Euk516R * | Euk 18S rRNA | ACCAGACTTGCCCTCC | [28] |
2.7. qPCR Analysis
2.8. Clone Library Construction
2.9. Restriction Fragment Length Polymorphism (RFLP)
2.10. Phylogenetic Analysis
2.11. Nucleotide Sequence Accession Numbers
3. Results
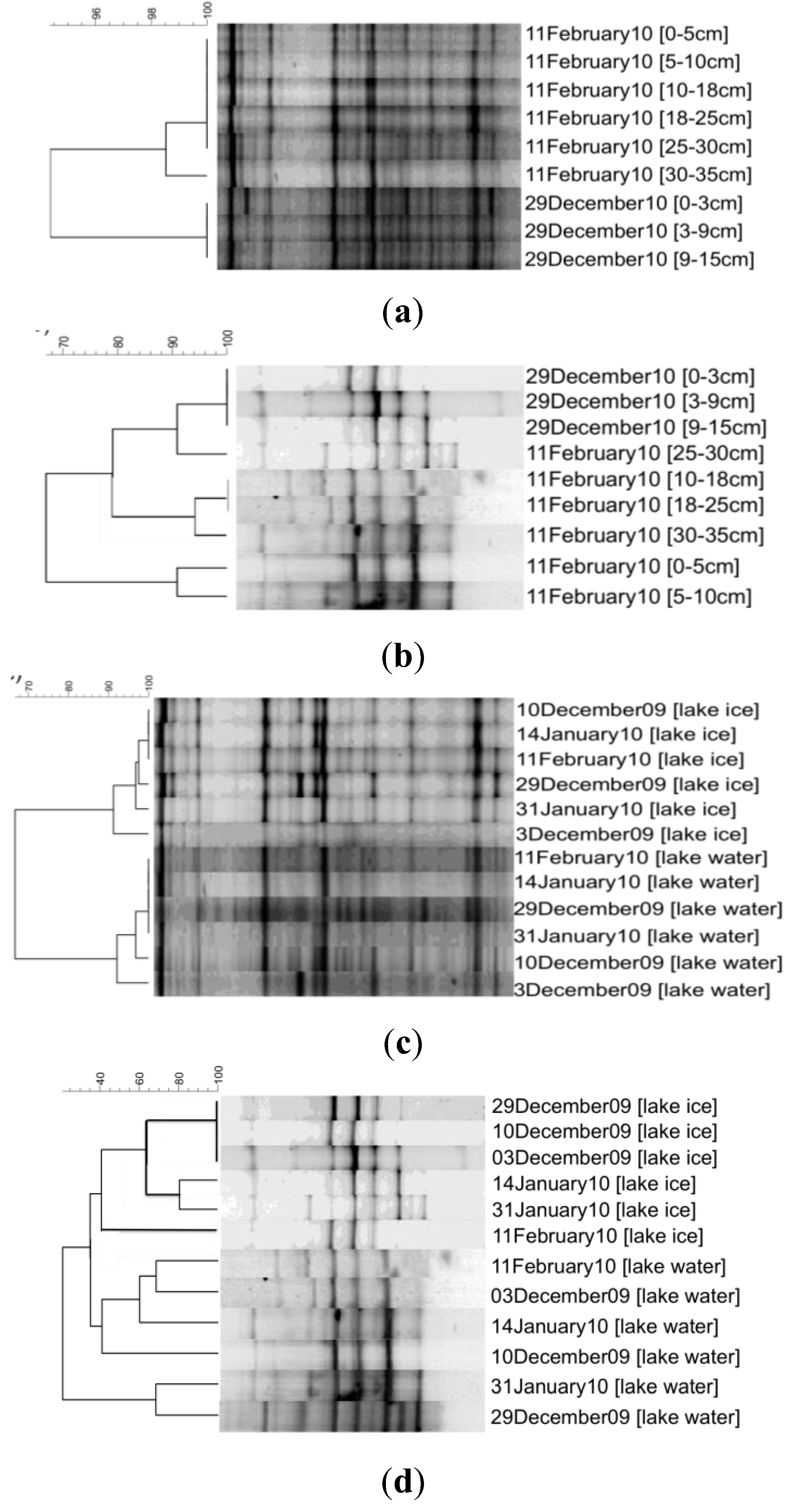
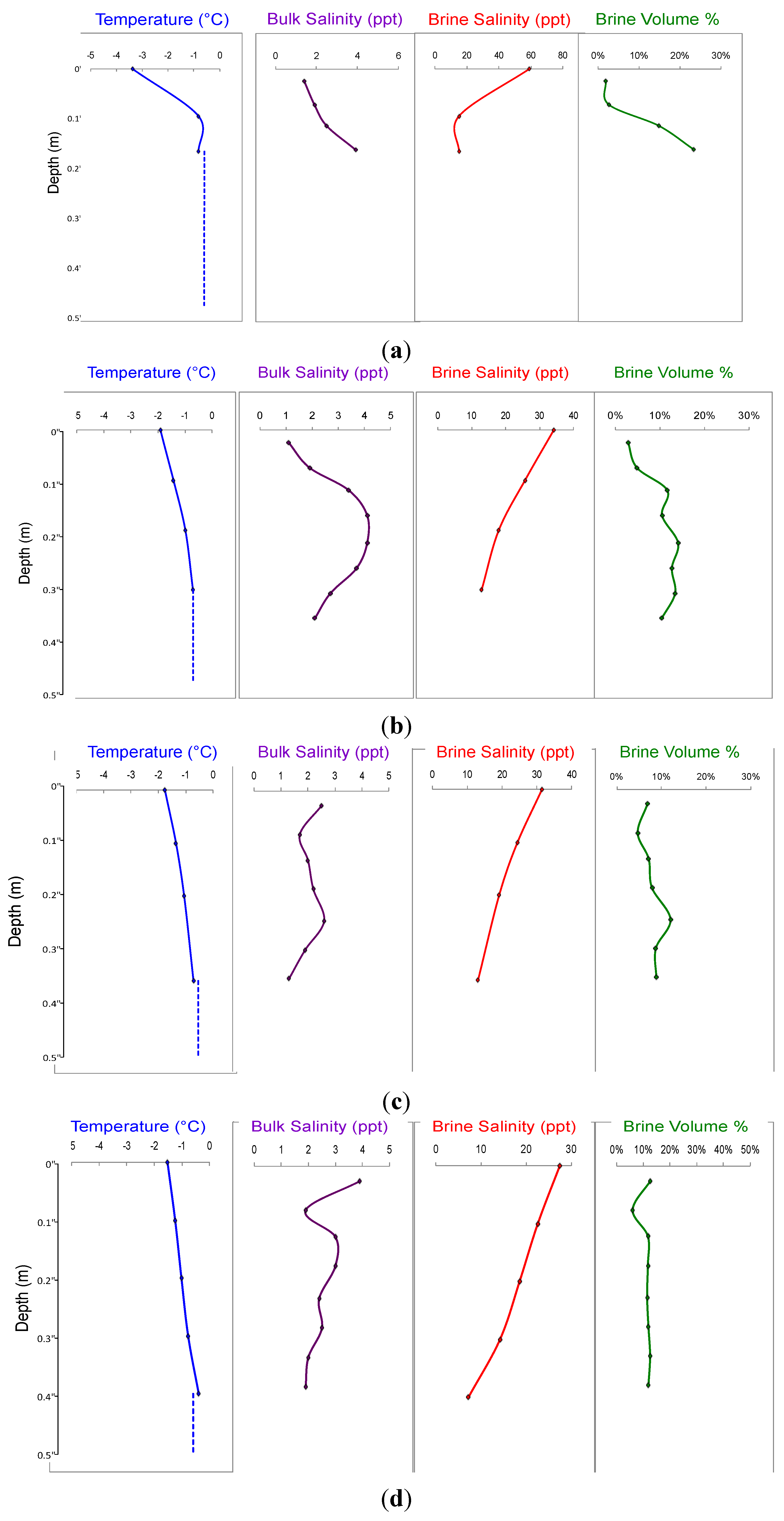
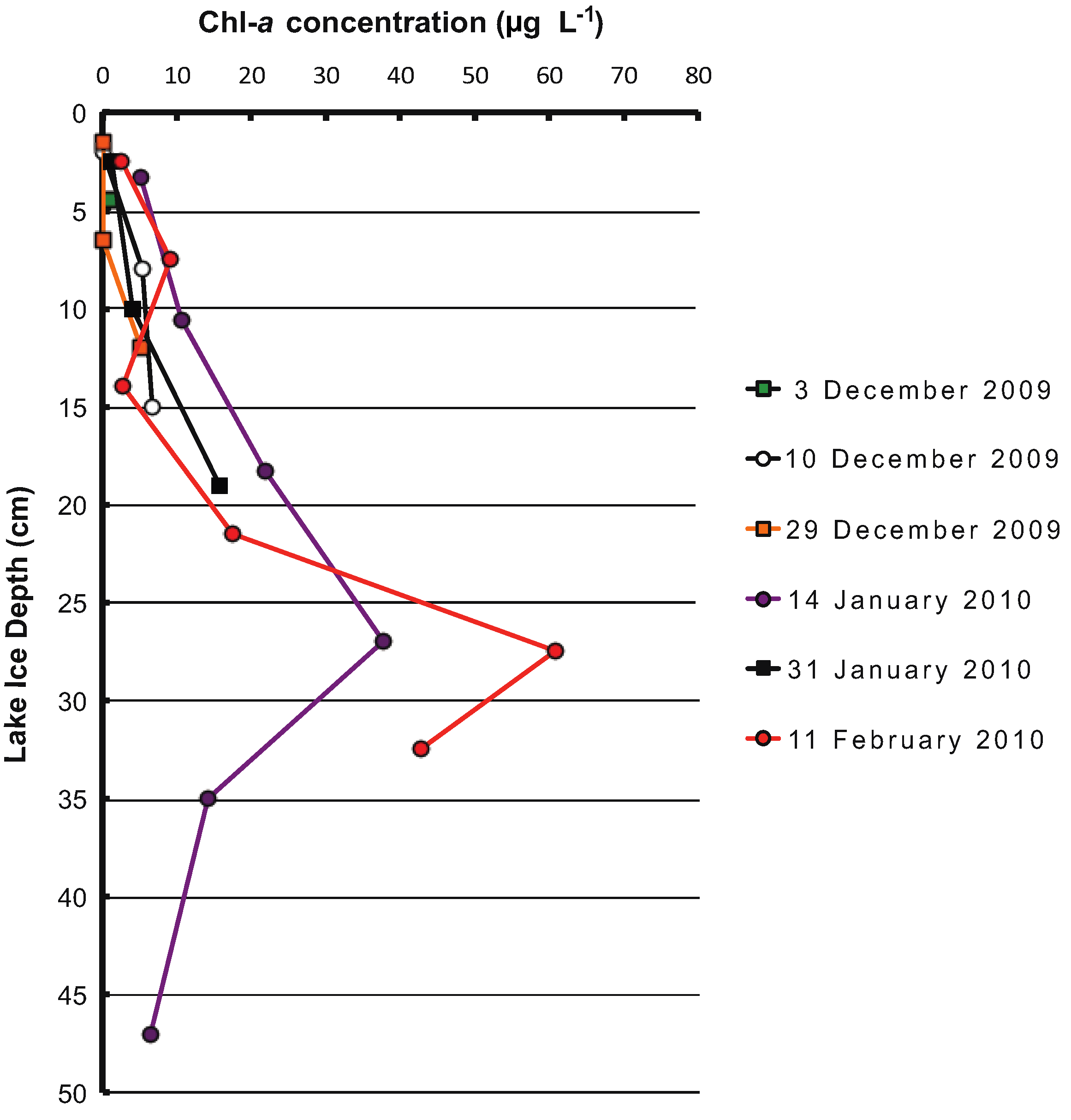


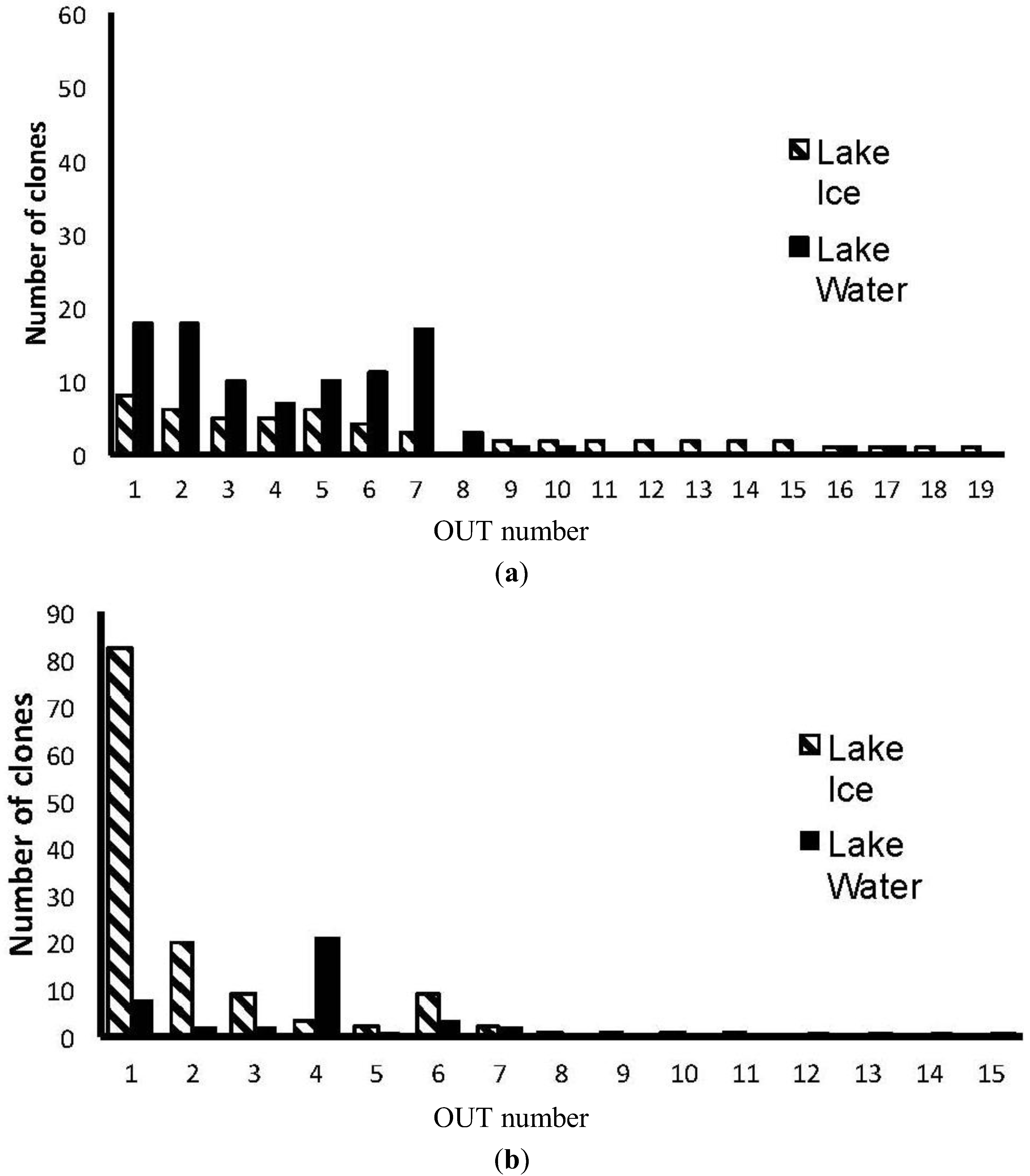
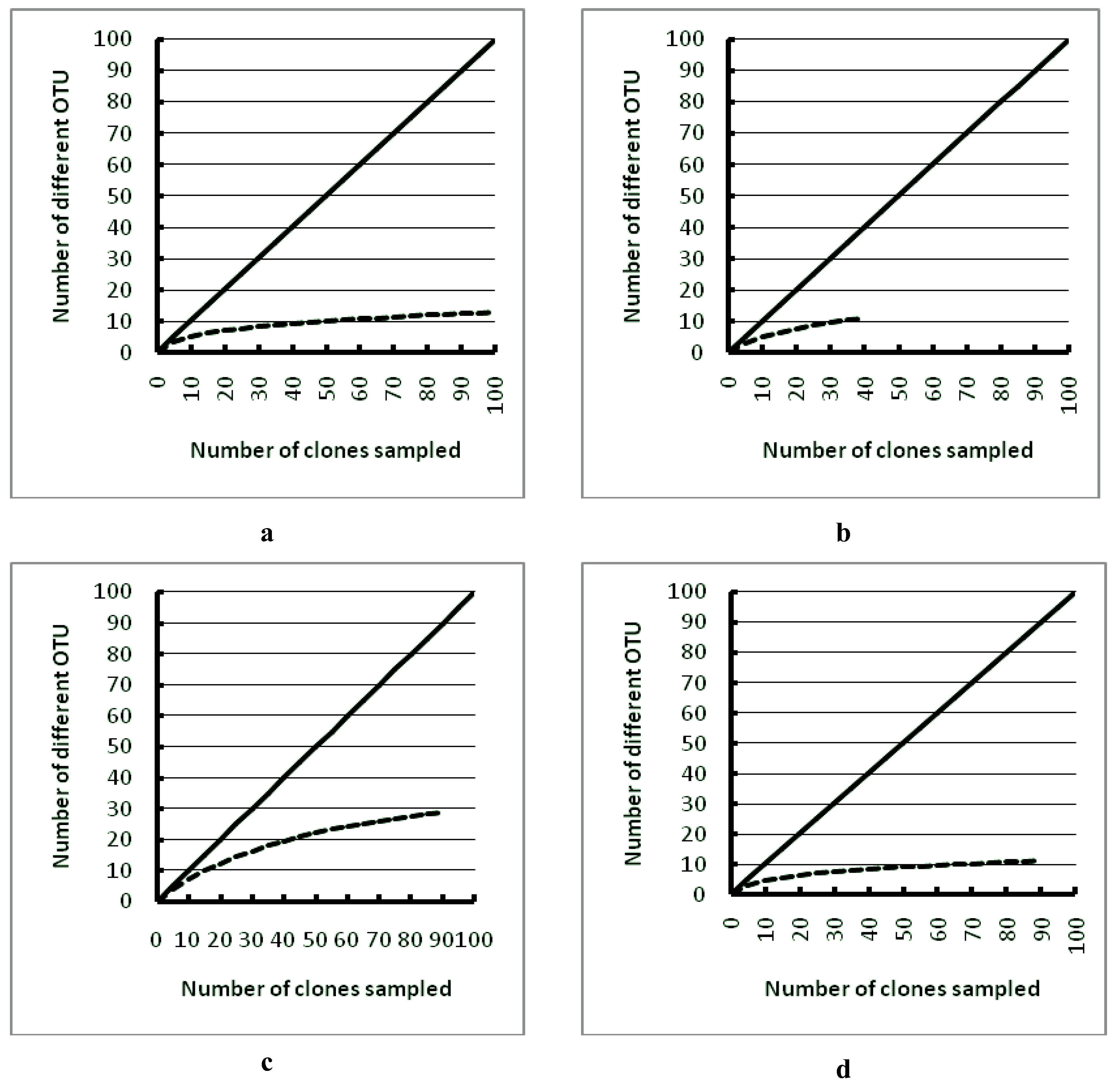
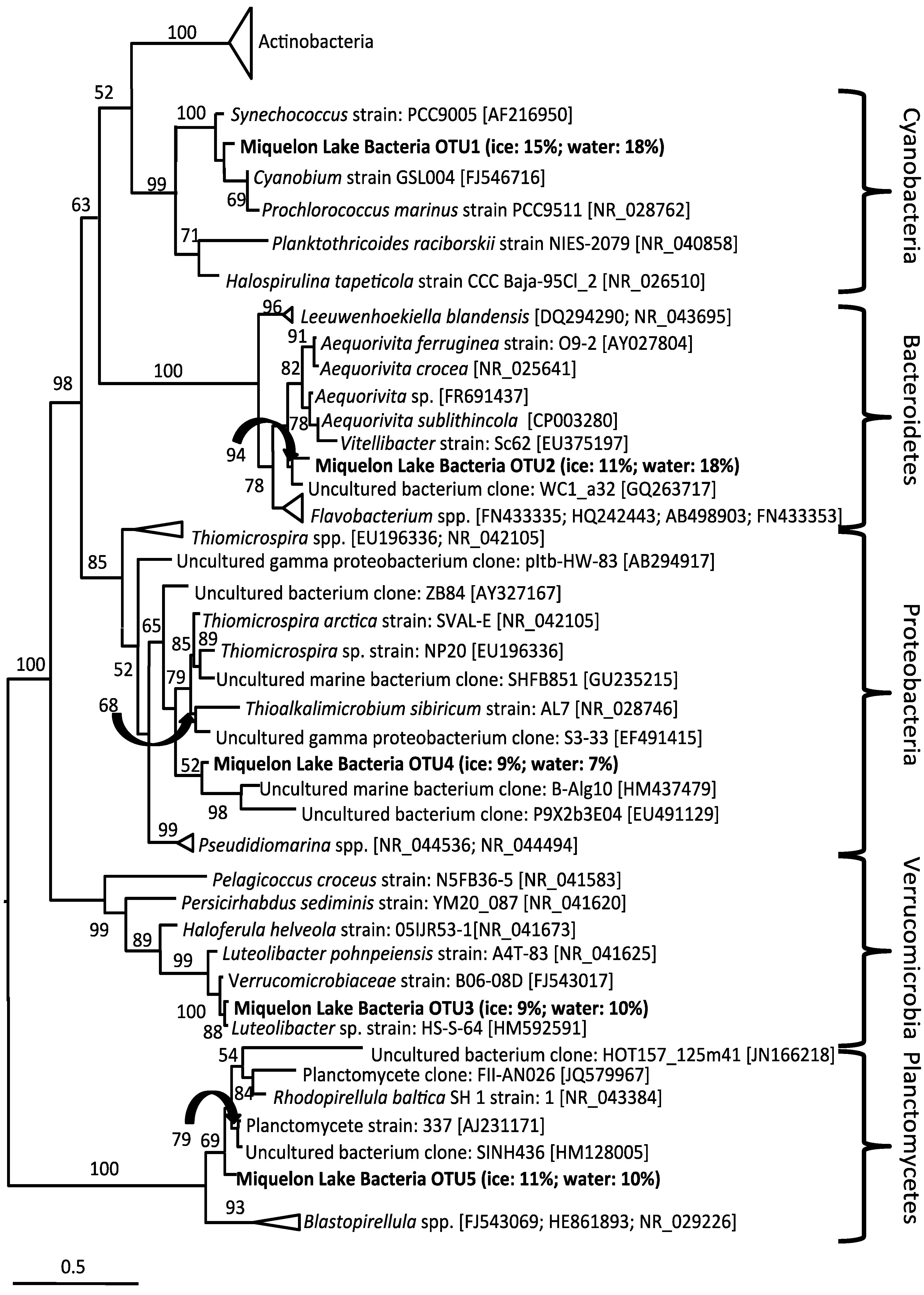
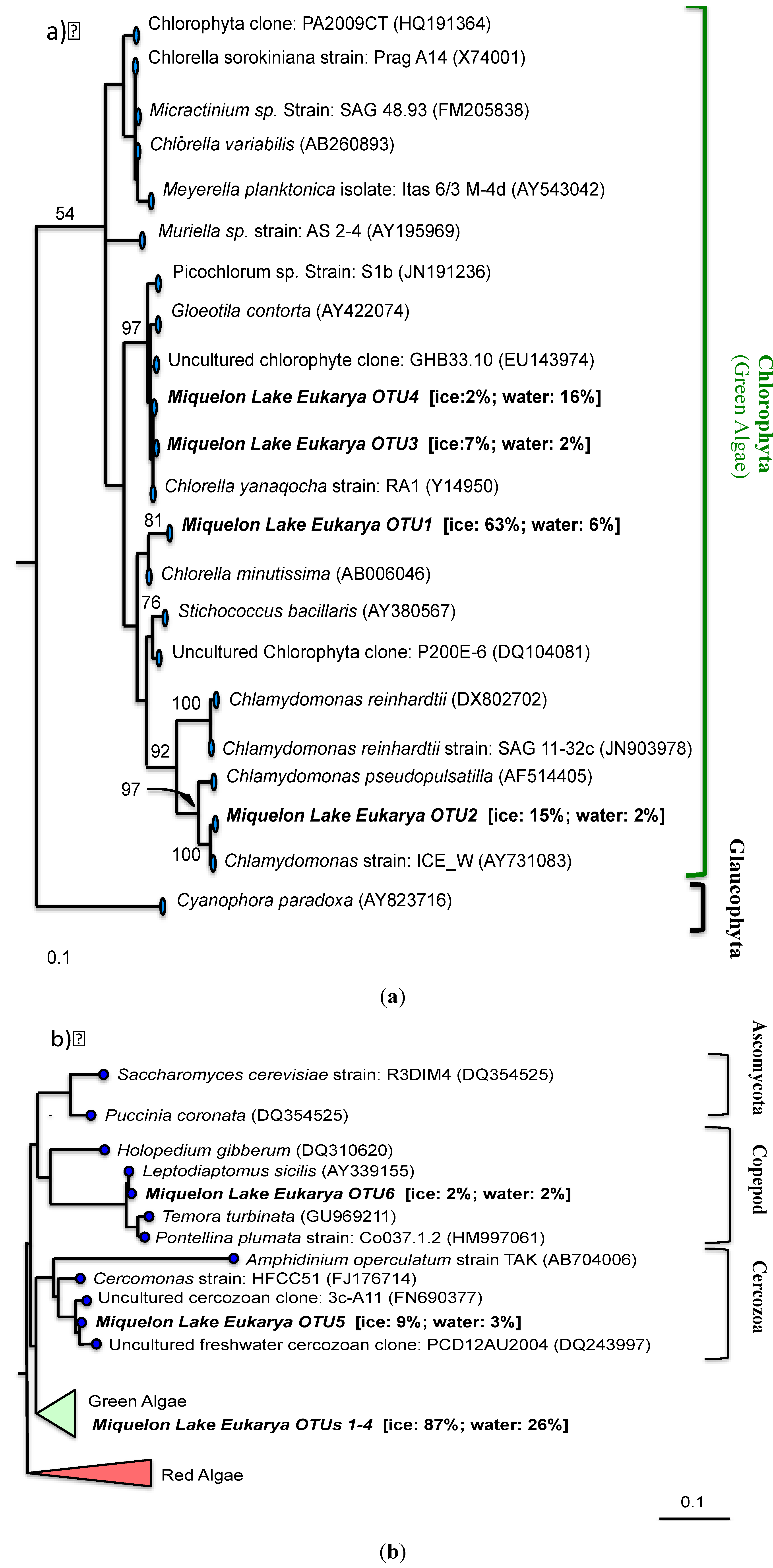
4. Discussion
5. Conclusions
Acknowledgments
References
- Mock, T.; Thomas, D.N. Recent advances in sea-ice microbiology. Environ. Microbiol. 2005, 7, 605–619. [Google Scholar] [CrossRef]
- Dieser, M.; Nocker, A.; Priscu, J.C.; Foreman, C.M. Viable microbes in ice: Application of molecular assays to McMurdo Dry Valley lake ice communities. Antarct. Sci. 2010, 22, 470–476. [Google Scholar] [CrossRef]
- Foreman, C.M.; Dieser, M.; Greenwood, M.; Cory, R.M.; Laybourn-Parry, J.; Lisle, J.T.; Jaros, C.; Miller, P.L.; Chin, Y.P.; Mcknight, D.M. When a habitat freezes solid: Microorganisms over-winter within the ice column of a coastal Antarctic lake. FEMS Microbiol. Ecol. 2011, 76, 401–412. [Google Scholar]
- Priscu, J.C.; Fritsen, C.H.; Adams, E.E.; Giovannoni, S.J.; Paerl, H.W.; McKay, C.P.; Doran, P.T.; Gordon, D.A.; Lanoil, B.D.; Pinckney, J.L. Perennial antarctic lake ice: An oasis for life in a polar desert. Science 1998, 280, 2095–2098. [Google Scholar] [CrossRef]
- Bowers, J.A.; Cooper, W.E.; Hall, D.J. Midwater and epibenthic behaviors of Mysis relicta Loven: Observations from the Johnson-Sea-Link II submersible in Lake Superior and from a remotely operated vehicle in northern Lake Michigan. J. Plankton Res. 1990, 12, 1279–1286. [Google Scholar] [CrossRef]
- Vanderploeg, H.A.; Bolsenga, S.J.; Fahnenstiel, G.L.; Liebig, J.R.; Gardner, W.S. Plankton ecology in an ice-covered bay of Lake Michigan: Utilization of a winter phytoplankton bloom by reproducing copepods. Hydrobiologia 1992, 243/244, 175–183. [Google Scholar] [CrossRef]
- Lizotte, M.P. The contributions of sea ice algae to Antarctic marine primary production. Am. Zool. 2001, 41, 57–73. [Google Scholar] [CrossRef]
- Kottmeier, S.T.; Sullivan, C.W. Sea ice microbial communities (SIMCO)—9. Effects of temperature and salinity on rates of metabolism and growth of autotrophs and heterotrophs. Polar Biol. 1988, 8, 293–304. [Google Scholar] [CrossRef]
- McGrath Grossi, S.; Kottmeier, S.T.; Sullivan, C.W. Sea ice microbial communities. III. Seasonal abundance of microalgae and associated bacteria, McMurdo Sound, Antarctica. Microb. Ecol. 1984, 10, 231–242. [Google Scholar] [CrossRef]
- Garrison, D.L.; Buck, K.R.; Silver, M.W. Microheterotrophs in the ice-edge zone. Antarct. J. US 1984, 19, 109–111. [Google Scholar]
- Kottmeier, S.T.; Sullivan, C.W. Bacterial biomass and production in pack ice of Antarctic marginal ice edge zones. Deep Sea Res. Oceanogr. Res. Pap. 1990, 37, 1311–1330. [Google Scholar] [CrossRef]
- Last, W.M. Geolimnology of salt lakes. Geosci. J. 2002, 6, 347–369. [Google Scholar] [CrossRef]
- Bowman, J.S.; Sachs, J.P. Chemical and physical properties of some saline lakes in Alberta and Saskatchewan. Saline Syst. 2008, 4, 1–17. [Google Scholar] [CrossRef]
- Swanson, H.; Zurawell, R. Miquelon Lake water quality monitoring report. Provincial Park Lakes Monitoring Program. Monitoring and Evaluation Branch, Environmental Assurance Division, Alberta Environment: Edmonton, Canada, February 2006. Available online: http://environment.gov.ab.ca/info/library/7730.pdf (accessed on 12 December 2011). [Google Scholar]
- Bierhuizen, J.F.H.; Prepas, E.E. Relationships between nutrients, dominant ions, and phytoplankton standing crop in prairie saline lakes. Can. J. Fish. Aquat. Sci. 1985, 42, 1588. [Google Scholar] [CrossRef]
- Evans, J.C.; Prepas, E.E. Potential effects of climate change on ion chemistry and phytoplankton communities in prairie saline lakes. Limnol. Oceanogr. 1996, 41, 1063. [Google Scholar] [CrossRef]
- Hammer, U.T. The Saline Lakes of Saskatchewan I. Background and Rationale for Saline Lakes Research. Int. Rev. Gesamten Hydrobiol. Hydrog. 1978, 63, 173. [Google Scholar] [CrossRef]
- Haynes, R.C.; Hammer, U.T. The saline lakes of Saskatchewan. IV Primary production of phytoplankton in selected saline ecosystems. Int. Rev. Gesamten Hydrobiol. 1978, 63, 337. [Google Scholar] [CrossRef]
- Grasby, S.E.; Londry, K.L. Supporting a Mid-Continent Marine Ecosystem: An Analogue for Martian Springs? Astrobiology 2007, 7, 662. [Google Scholar] [CrossRef]
- Sørensen, K.B.; Teske, A. Stratified communities of active archaea in deep marine subsurface sediments. Appl. Environ. Microbiol. 2006, 72, 4596–4603. [Google Scholar] [CrossRef]
- Sorokin, D.Y.; Tourova, T.P.; Lysenko, A.M.; Muyzer, G. Diversity of culturable halophilic sulfur-oxidizing bacteria in hypersaline habitats. Microbiology 2006, 152, 3013–3023. [Google Scholar] [CrossRef]
- Richter-Menge, J.A.; Perovich, D.K.; Elder, B.C.; Claffey, K.; Rigor, I.; Ortmeyer, M. Ice mass-balance buoys: A tool for measuring and attributing changes in the thickness of the Arctic sea-ice cover. Ann. Glaciol. 2006, 44, 205–210. [Google Scholar] [CrossRef]
- Cox, G.F.N.; Weeks, W.F. Equations for determining the gas and brine volumes in sea ice samples. J. Glaciol. 1983, 29, 306–316. [Google Scholar]
- Porter, K.G.; Feig, Y.S. The use of DAPI for identifying and counting aquatic microflora. Limnol. Oceanogr. 1980, 25, 943–948. [Google Scholar] [CrossRef]
- Bergmann, M.; Peters, R.H. A simple reflectance method for the measurement of particulate pigment in lake water and its application to phosphorus-chlorophyll-seston relationships. Can. J. Fish. Aquat. Sci. 1980, 37, 111–114. [Google Scholar] [CrossRef]
- Kulp, T.R.; Han, S.; Saltikov, C.W.; Lanoil, B.D.; Zargar, K.; Oremland, R.S. Effects of imposed salinity gradients on dissimilatory arsenate reduction, sulfate reduction, and other microbial processes in sediments from California soda lakes. Appl. Environ. Microbiol. 2008, 74, 3618–3618. [Google Scholar] [CrossRef]
- Myers, R.; Fischer, S.G.; Lerman, L.S.; Maniatis, T. Nearly all single base substitutions in DNA fragments joined to a GC-clamp can be detected by denaturing gradient gel electrophoresis. Nucleic Acids Res. 1985, 139, 3131–3145. [Google Scholar]
- Díez, B.; Pedrós-Alió, C.; Marsh, T.L.; Massana, R. Application of denaturing gradient gel electrophoresis (DGGE) to study the diversity of marine picoeukaryotic assemblages and comparison of DGGE with other molecular techniques. Appl. Environ. Microbiol. 2001, 67, 2942–2951. [Google Scholar] [CrossRef]
- Bano, N.; Ruffin, S.; Ransom, B.; Hollibaugh, J.T. Phylogenetic Composition of Arctic Ocean Archaeal Assemblages and Comparison with Antarctic Assemblages. Appl. Environ. Microbiol. 2004, 70, 781–789. [Google Scholar] [CrossRef]
- Labrenz, M.; Sintes, E.; Toetzke, F.; Zumsteg, A.; Herndl, G.J.; Seidler, M.; Jürgens, K. Relevance of a crenarchaeotal subcluster related to Candidatus Nitrosopumilus maritimus to ammonia oxidation in the suboxic zone of the central Baltic Sea. ISME J. 2010, 4, 1496–1508. [Google Scholar] [CrossRef]
- Polz, M.F.; Cavanaugh, C.M. Bias in template-to-product ratios in multitemplate PCR. Appl. Environ. Microbiol. 1998, 64, 3724–3730. [Google Scholar]
- Muyzer, G.; de Waal, E.C.; Uitterlinden, A.G. Profiling of complex microbial populations by denaturing gradient gel electrophoresis analysis of polymerase chain reaction-amplified genes coding for 16S rRNA. Appl. Environ. Microbiol. 1993, 59, 695–700. [Google Scholar]
- Delong, E.F. Archaea in coastal marine environments. PNAS 1992, 89, 5685–5689. [Google Scholar] [CrossRef]
- Skidmore, M.; Anderson, S.P.; Sharp, M.; Foght, J.; Lanoil, B.D. Comparison of microbial community compositions of two subglacial environments reveals a possible role for microbes in chemical weathering processes. Appl. Environ. Microbiol. 2005, 71, 6986–6997. [Google Scholar]
- Good, I.J. The population frequencies of species and the estimation of population parameters. Biometrika 1953, 40, 237. [Google Scholar]
- Lanoil, B.D.; Sassen, R.; La Duc, M.T.; Sweet, S.T.; Nealson, K.H. Bacteria and Archaea Physically Associated with Gulf of Mexico Gas Hydrates. Appl. Environ. Microbiol. 2001, 67, 5143–5153. [Google Scholar] [CrossRef]
- Decipher. Available online: http://decipher.cee.wisc.edu/ (accessed on 12 September 2012).
- Drummond, A.; Ashton, B.; Buxton, S.; Cheung, M.; Cooper, A.; Duran, C.; Field, M.; Heled, J.; Kearse, M.; Markowitz, S.; et al. Geneious Pro: Geneious v5.4.5; Biomatters. Available online: http://www.geneious.com/ (accessed on 12 December 2011).
- Junge, K.; Eicken, H.; Deming, J.W. Bacterial Activity at −2 to −20 °C in Arctic Wintertime Sea Ice. Appl. Environ. Microbiol. 2004, 70, 550–557. [Google Scholar] [CrossRef]
- Murray, A.E.; Wu, K.Y.; Moyer, C.L.; Karl, D.M.; Delong, E.F. Evidence for circumpolar distribution of planktonic Archaea in the Southern Ocean. Aquat. Microb. Ecol. 1999, 18, 263–273. [Google Scholar] [CrossRef]
© 2013 by the authors; licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution license (http://creativecommons.org/licenses/by/3.0/).
Share and Cite
Bramucci, A.; Han, S.; Beckers, J.; Haas, C.; Lanoil, B. Composition, Diversity, and Stability of Microbial Assemblages in Seasonal Lake Ice, Miquelon Lake, Central Alberta. Biology 2013, 2, 514-532. https://doi.org/10.3390/biology2020514
Bramucci A, Han S, Beckers J, Haas C, Lanoil B. Composition, Diversity, and Stability of Microbial Assemblages in Seasonal Lake Ice, Miquelon Lake, Central Alberta. Biology. 2013; 2(2):514-532. https://doi.org/10.3390/biology2020514
Chicago/Turabian StyleBramucci, Anna, Sukkyun Han, Justin Beckers, Christian Haas, and Brian Lanoil. 2013. "Composition, Diversity, and Stability of Microbial Assemblages in Seasonal Lake Ice, Miquelon Lake, Central Alberta" Biology 2, no. 2: 514-532. https://doi.org/10.3390/biology2020514




