Recruitment of Transcription Complexes to Enhancers and the Role of Enhancer Transcription
Abstract
:1. Introduction
2. Enhancer and Locus Control Region (LCR)
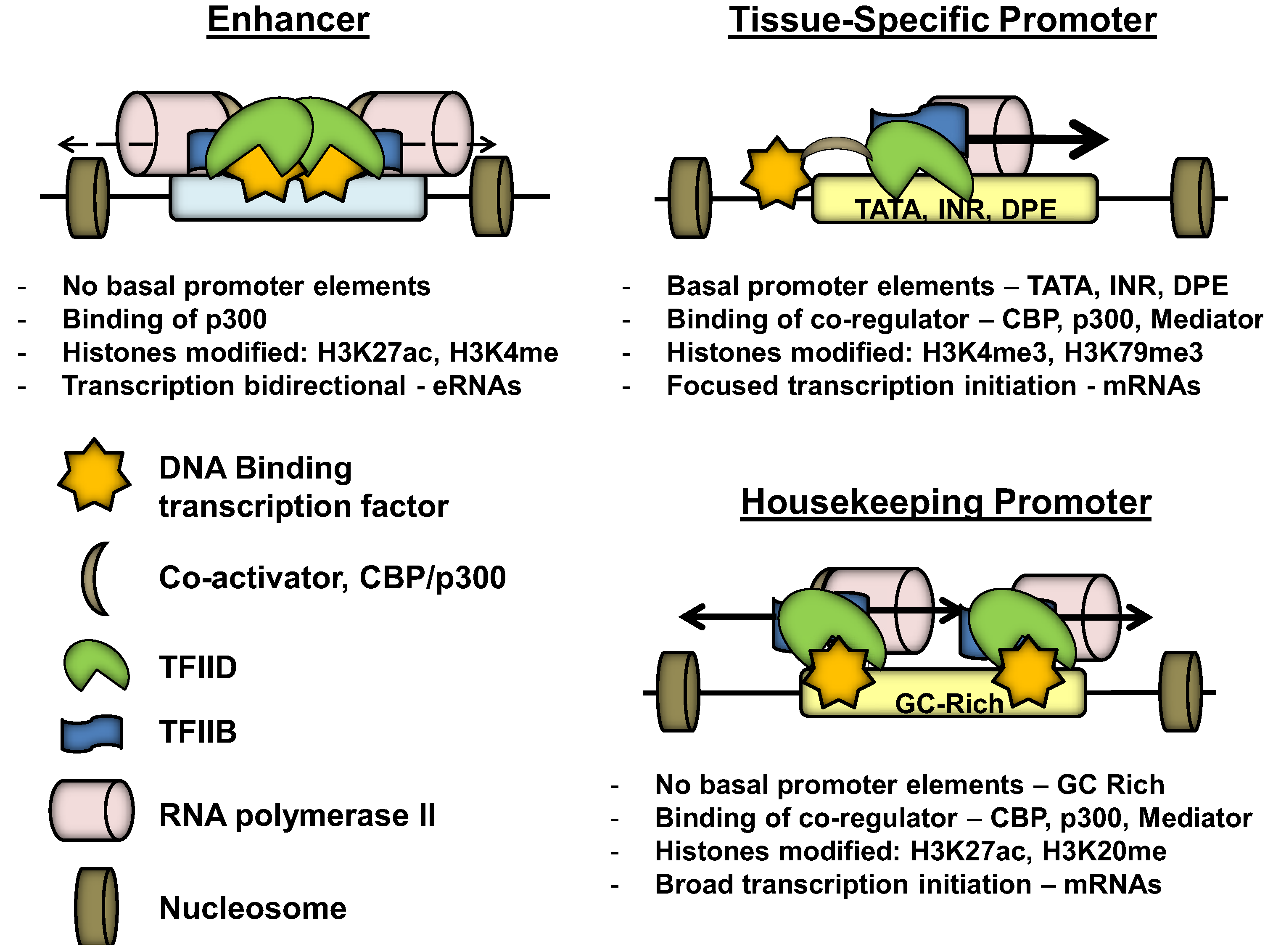
3. Promoter
4. Nuclear Transcription Domains
5. Enhancer Function

6. Transcription Initiation at Enhancers
7. Possible Functions of eRNAs
8. Poised Enhancers During Cellular Differentiation
9. The Role of Enhancer-Mediated Transcription in Chromatin Modification
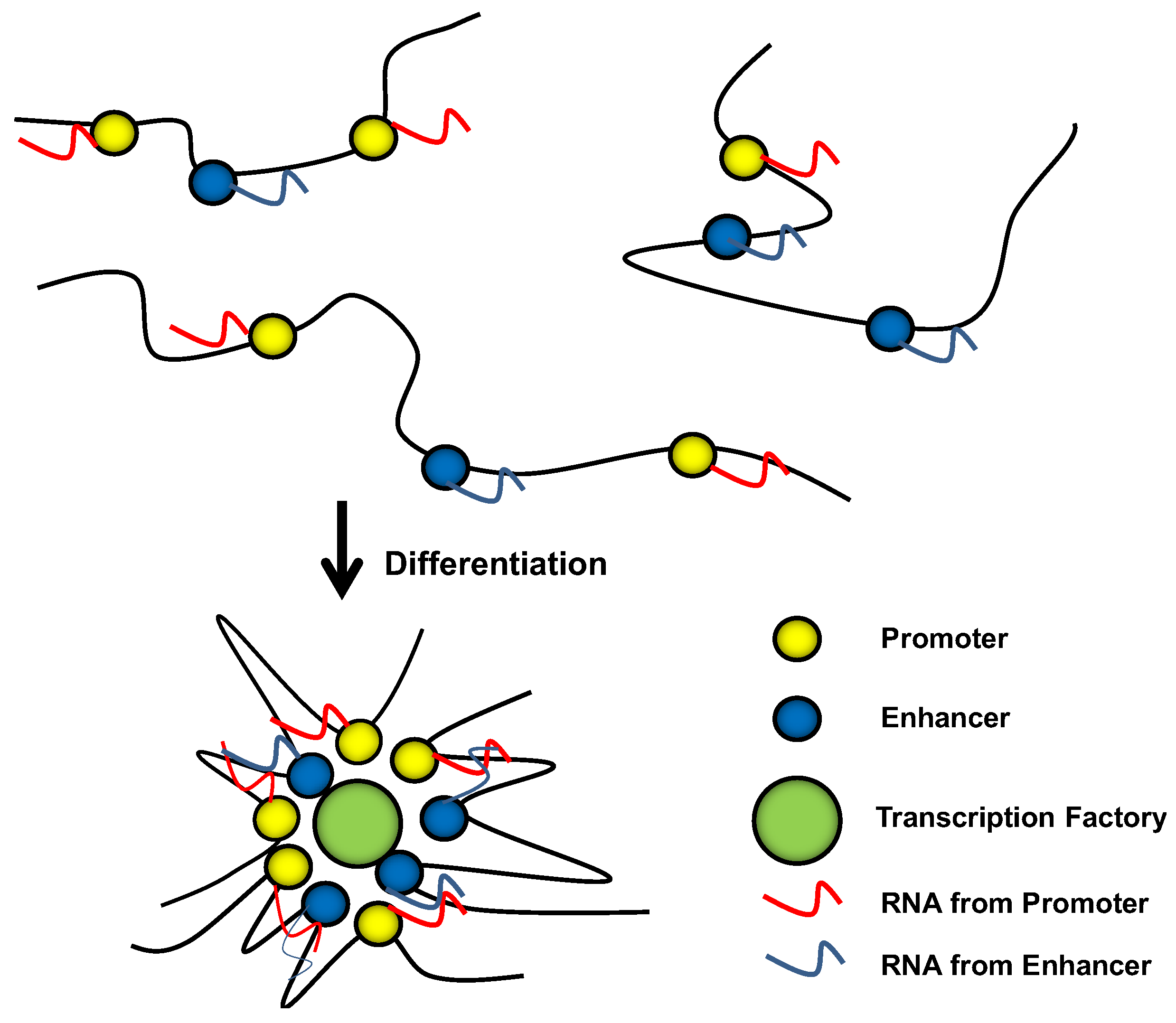
10. Conclusions
Acknowledgements
References
- Orphanides, G.; Reinberg, D. A unified theory of gene expression. Cell 2002, 108, 439–451. [Google Scholar] [CrossRef]
- Bernstein, B.E.; Birney, E.; Dunham, I.; Green, E.D.; Gunter, C.; Snyder, M. An integrated encyclopedia of DNA elements in the human genome. Nature 2012, 489, 57–74. [Google Scholar]
- Juven-Gershon, T.; Kadonaga, J.T. Regulation of gene expression via the core promoter and the basal transcriptional machinery. Dev. Biol. 2010, 339, 225–229. [Google Scholar] [CrossRef]
- Ghirlando, R.; Giles, K.; Gowher, H.; Xiao, T.; Xu, Z.; Yao, H.; Felsenfeld, G. Chomatin domains, insulators, and the regulation of gene expression. Biochim. Biophys. Acta 2012, 1819, 644–651. [Google Scholar] [CrossRef]
- Bulger, M.; Groudine, M. Functional and mechanistic diversity of distal transcription enhancers. Cell 2011, 144, 327–339. [Google Scholar] [CrossRef]
- Shen, Y.; Yue, F.; McCleary, D.F.; Ye, Z.; Edsall, L.; Kuan, S.; Wagner, U.; Dixon, J.; Lee, L.; Lobanenkov, V.V.; Ren, B. A map of the cis-regulatory sequences in the mouse genome. Nature 2012, 488, 116–120. [Google Scholar]
- Banerji, J.; Rusconi, S.; Schaffner, W. Expression of a beta-globin gene is enhanced by remote SV40 sequences. Cell 1981, 27, 299–308. [Google Scholar] [CrossRef]
- Müller, M.M.; Gerster, T.; Schaffner, W. Enhancer sequences and the regulation of gene transcription. Eur. J. Biochem. 1988, 176, 485–495. [Google Scholar] [CrossRef]
- Wu, C. The 5’ends of Drosophila heat shock genes in chromatin are hypersensitive to DNase I. Nature 1980, 286, 854–860. [Google Scholar] [CrossRef]
- Collis, P.; Antoniou, M.; Grosveld, F. Definition of the minimal requirements within the human beta-globin gene and the dominant control region for high level expression. EMBO J. 1990, 9, 233–240. [Google Scholar]
- Ellis, J.; Tan-Un, K.C.; Harper, A.; Michalovich, D.; Yannoutsos, N.; Philipsen, S.; Grosveld, F. A dominant chromatin-opening activity in 5'hypersensitive site 3 of the human beta-globin locus control region. EMBO J. 1996, 15, 562–568. [Google Scholar]
- Valen, E.; Sandelin, A. Genomic and chromatin signals underlying transcription start-site selection. Trends Genet. 2011, 27, 475–485. [Google Scholar] [CrossRef]
- Li, Q.; Peterson, K.R.; Fang, X. Stamatoyannopoulos G. Locus control regions. Blood 2002, 100, 3077–3086. [Google Scholar] [CrossRef]
- Bulger, M.; Groudine, M. Looping versus linking: toward a model for long-distance gene activation. Genes Dev. 1999, 13, 2465–2477. [Google Scholar] [CrossRef]
- Engel, J.D.; Tanimoto, K. Looping, linking, and chromatin activity: new insights into beta-globin locus regulation. Cell 2000, 100, 499–502. [Google Scholar] [CrossRef]
- Fleetwood, M.R.; Ho, Y.; Cooke, N.E.; Liebhaber, S.A. DNase I hypersensitive site II of the human growth hormone locus control region mediates an essential and distinct long-range enhancer function. J. Biol. Chem. 2012, 287, 25454–25465. [Google Scholar]
- Tanimoto, K.; Liu, Q.; Bungert, J.; Engel, J.D. Effects of altered gene order or orientation of the locus control region on human beta-globin gene expression in mice. Nature 1999, 398, 344–348. [Google Scholar]
- Heintzman, N.D.; Hon, G.C.; Hawkins, R.D.; Kheradpour, P.; Stark, A.; Harp, L.F.; Ye, Z.; Lee, L.K.; Stuart, R.K.; Ching, C.W.; et al. Histone modifications at human enhancers reflect global cell type-specific gene expression. Nature 2009, 459, 108–112. [Google Scholar]
- Ernst, J.; Kheradpour, P.; Mikkelsen, T.S.; Shoresh, N.; Ward, L.D.; Epstein, C.B.; Zhang, X.; Wang, L.; Issner, R.; Coyne, M.; Ku, M.; et al. Mapping and analysis of chromatin state dynamics in nine human cell types. Nature 2011, 473, 43–49. [Google Scholar]
- Zentner, G.E.; Scacheri, P.C. The chromatin fingerprint of gene enhancer elements. J. Biol. Chem. 2012, 287, 30888–30896. [Google Scholar] [CrossRef]
- Xi, H.; Shulha, H.P.; Lin, J.M.; Vales, T.R.; Fu, Y.; Bodine, D.; McKay, R.D.; Chenoweth, J.G.; Tesar, P.J.; Furey, T.S.; et al. Identification and characterization of cell type-specific and ubiquitous chromatin regulatory structures in the human genome. PLoS Genet. 2007, 3. [Google Scholar] [CrossRef]
- Koch, F.; Fenouil, R.; Gut, M.; Cauchy, P.; Albert, T.K.; Zacarias-Cabeza, J.; Spicuglia, S.; de la Chapelle, A.L.; Heidemann, M.; Hintermair, C.; et al. Transcription initiation platforms and GTF recruitment at tissue-specific enhancers and promoters. Nat. Struct. Mol. Biol. 2011, 18, 956–963. [Google Scholar]
- Creyghton, M.P.; Cheng, A.W.; Welstead, G.G.; Kooistra, T.; Carey, B.W.; Steine, E.J.; Hanna, J.; Lodato, M.A.; Frampton, G.M.; Sharp, P.A.; et al. Histone H3K27ac separates active from poised enhancers and predicts developmental state. Proc. Natl. Acad. Sci. USA 2010, 107, 21931–21936. [Google Scholar]
- Zentner, G.E.; Tesar, P.J.; Scacheri, P.C. Epigenetic signatures distinguish multiple classes of enhancers with distinct cellular functions. Genome Res. 2011, 21, 1273–1283. [Google Scholar] [CrossRef]
- Zhou, Z.; Li, X.; Deng, C.; Ney, P.A.; Huang, S.; Bungert, J. USF and NF-E2 cooperate to regulate the recruitment and activity of RNA polymerase II in the beta-globin gene locus. J. Biol. Chem. 2010, 285, 15894–15905. [Google Scholar]
- Wang, J.; Zhuang, J.L.; Iyer, S.; Lin, X.Y.; Whitfield, T.W.; Greven, M.C.; Pierce, B.G.; Dong, X.J.; Kundaje, A.; Cheng, Y.; et al. Sequence features and chromatin structure around the genomic regions bound by 119 human transcription factors. Genome Res. 2012, 22, 1798–1812. [Google Scholar]
- Neph, S.; Vierstra, J.; Stergachis, A.B.; Reynolds, A.P.; Haugen, E.; Vernot, B.; Thurman, R.E.; John, S.; Sandstrom, R.; Johnson, A.K.; et al. An expansive human regulatory lexicon encoded in transcription factor footprints. Nature 2012, 489, 83–90. [Google Scholar]
- Johnson, K.D.; Grass, J.A.; Park, C.; Im, H.; Choi, K.; Bresnick, E.H. Highly restricted localization of RNA polymerase II within a locus control region of a tissue-specific chromatin domain. Mol. Cell Biol. 2003, 23, 6484–6493. [Google Scholar] [CrossRef]
- Vieira, K.F.; Levings, P.P.; Hill, M.A.; Crusselle, V.J.; Kang, S.H.; Engel, J.D.; Bungert, J. Recruitment of transcription complexes to the beta-globin gene locus control region in vivo and in vitro. J. Biol. Chem. 2004, 279, 50350–50357. [Google Scholar]
- Kim, T.K.; Hemberg, M.; Gray, J.M.; Costa, A.M.; Bear, D.M.; Wu, J.; Harmin, D.A.; Laptewicz, M.; Barbara-Haley, K.; Kuersten, S.; et al. Widespread transcription at neuronal activity-regulated enhancers. Nature 2010, 465, 182–187. [Google Scholar]
- de Santa, F.; Barozzi, I.; Mietton, F.; Ghisletti, S.; Polletti, S.; Tusi, B.K.; Muller, H.; Ragoussis, J.; Wei, C.L.; Natoli, G. A large fraction of extragenic RNA pol II transcription sites overlap enhancers. PLoS Biol. 2010, 8. [Google Scholar] [CrossRef]
- Djebali, S.; Davis, C.A.; Merkel, A.; Dobin, A.; Lassmann, T.; Mortazavi, A.; Tanzer, A.; Lagarde, J.; Lin, W.; Schlesinger, F.; et al. Landscape of transcription in human cells. Nature 2012, 489, 101–108. [Google Scholar]
- Jimenez, G.; Griffiths, S.D.; Ford, A.M.; Greaves, M.F.; Enver, T. Activation of the beta-globin locus control region precedes commitment to the erythroid lineage. Proc. Natl. Acad. Sci. USA 1992, 89, 10618–10622. [Google Scholar]
- Buratowski, S.; Hahn, S.; Guarente, L.; Sharp, P.A. Five intermediate complexes in transcription initiation by RNA polymerase II. Cell 1989, 56, 549–561. [Google Scholar] [CrossRef]
- Roeder, R.G. The role of general transcription factors in transcription by RNA polymerase II. Trends Biochem. Sci. 1996, 21, 327–335. [Google Scholar]
- Kim, T.H.; Barrera, L.O.; Zheng, M.; Qu, C.; Singer, M.A.; Richmond, T.A.; Wu, Y.; Green, R.D.; Ren, B. A high-resolution map of active promoters in the human genome. Nature 2005, 436, 876–880. [Google Scholar]
- Gershenzon, N.I.; Ioshikhes, I.P. Synergy of human Pol II core promoter elements revealed by statistical sequence analysis. Bioinformatics 2005, 21, 1295–1300. [Google Scholar] [CrossRef]
- Karlic, R.; Chung, H.R.; Lassere, J.; Vlahovicek, K.; Vingron, M. Histone modification levels are predictive for gene expression. Proc. Natl. Acad. Sci. USA 2010, 107, 2926–2931. [Google Scholar]
- Iborra, F.J.; Pombo, A.; McManus, J.; Jackson, D.A.; Cook, P.R. The topology of transcription by immobilized polymerases. Exp. Cell Res. 1996, 229, 167–173. [Google Scholar] [CrossRef]
- Deng, B.; Melnik, S.; Cook, P.R. Transcription factories, chromatin loops, and the dysregulation of gene expression in malignancy. Semin. Cancer Biol. 2012, in press. [Google Scholar]
- Edelman, L.B.; Fraser, P. Transcription factories: Genetic programming in three dimensions. Curr. Opin. Genet. Dev. 2012, 22, 110–114. [Google Scholar] [CrossRef]
- Schoenfelder, S.; Sexton, T.; Chakalova, L.; Cope, N.F.; Horton, A.; Andrews, S.; Kurukuti, S.; Mitchell, J.A.; Umlauf, D.; Dimitrova, D.S.; et al. Preferential association between co-regulated genes reveal a transcriptional interactome in erythroid cells. Nat. Genet. 2010, 42, 53–61. [Google Scholar]
- Lanctot, C.; Cheutin, T.; Cremer, M.; Cavalli, G.; Cremer, T. Dynamic genome architecture in the nuclear space: regulation of gene expression in three dimensions. Nat. Rev. Genet. 2007, 8, 104–115. [Google Scholar] [CrossRef]
- Niedojadlo, J.; Perret-Vivancos, C.; Kalland, K.H.; Cmarko, D.; Cremer, T.; van Driel, R.; Fakan, S. Transcribed DNA is preferentially located in the perichromatin region of mammalian nuclei. Exp. Cell Res. 2011, 317, 433–444. [Google Scholar] [CrossRef]
- Fromm, G.; Cadiz-Rivera, B.; de Vries, C.; Getman, M.; McGrath, K.E.; Kingsley, P.D.; Fields, J.; Fiering, S.; Bulger, M. An embryonic stage-specific enhancer within the murine beta-globin locus mediates domain-wide histone hyperacetylation. Blood 2011, 117, 5207–5214. [Google Scholar] [CrossRef]
- Vernimmen, D.; Lynch, M.D.; DeGobbi, M.; Garrick, D.; Sharpe, J.A.; Sloane-Stanley, J.A.; Smith, A.J.H.; Higgs, D.R. Polycomb eviction as a new distant enhancer function. Genes Dev. 2011, 25, 1583–1588. [Google Scholar]
- Morey, L.; Helin, K. Polycomb group protein-mediated repression of transcription. Trends Biochem. Sci. 2010, 35, 323–332. [Google Scholar] [CrossRef]
- Taberlay, P.C.; Kelly, T.K.; Liu, C.C.; You, J.S.; de Carvalho, D.D.; Miranda, T.B.; Zhou, X.J.; Liang, G.N.; Jones, P.A. Polycomb-repressed genes have permissive enhancers that initiate reprogramming. Cell 2011, 147, 1283–1294. [Google Scholar] [CrossRef]
- Rodriguez-Jato, S.; Nicholls, R.D.; Driscoll, D.J.; Yang, T.P. Characterization of cis- and trans-acting elements in the imprinted human SNURF-SNRPN locus. Nucl. Acids. Res. 2005, 33, 4740–4753. [Google Scholar]
- Zhao, H.; Friedman, R.D.; Fournier, R.E.K. The locus control region activates serpin gene expression through recruitment of liver-specific transcription factors and RNA polymerase II. Mol. Cell Biol. 2007, 27, 5286–5295. [Google Scholar] [CrossRef]
- Levings, P.P.; Bungert, J. The human beta-globin locus control region. Eur. J. Biochem. 2002, 269, 1589–1599. [Google Scholar] [CrossRef]
- Ragoczy, T.; Bender, M.A.; Telling, A.; Byron, R.; Groudine, M. The locus control region is required for association of the murine beta-globin locus with engaged transcription factories during erythroid maturation. Genes Dev. 2006, 20, 1447–1457. [Google Scholar] [CrossRef]
- Francastel, C.; Walters, M.C.; Groudine, M.; Martin, D.I. A functional enhancer suppresses silencing of a transgene and prevents its localization close to centromeric heterochromatin. Cell 1999, 99, 259–269. [Google Scholar] [CrossRef]
- Sawado, T.; Halow, J.; Bender, M.A.; Groudine, M. The beta-globin locus control region (LCR) functions primarily by enhancing the transition from transcription initiation to elongation. Genes Dev. 2003, 17, 1009–1018. [Google Scholar] [CrossRef]
- Song, S.H.; Kim, A.; Ragoczy, T.; Bender, M.A.; Groudine, M.; Dean, A. Multiple functions of Ldb1 required for beta-globin activation during erythroid differentiation. Blood 2010, 116, 2356–2364. [Google Scholar] [CrossRef]
- Nechaev, S.; Adelman, K. Pol II waiting in the starting gates: Regulating the transition from transcription initiation into productive elongation. Biochim. Biophys. Acta. 2011, 1809, 34–45. [Google Scholar] [CrossRef]
- Marks, H.; Kalkan, T.; Denissov, S.; Jones, K.; Hofemeister, H.; Nichols, J.; Stewart, A.F.; Smith, A.; Stunnenberg, H.G. The transcriptional and epigenomic foundations of ground state pluripotency. Cell 2012, 149, 590–604. [Google Scholar] [CrossRef]
- Li, G.; Ruan, X.; Auerbach, R.K.; Sandhu, K.S.; Zheng, M.; Wang, P.; Poh, H.M.; Goh, Y.; Lim, J.; Zhang, J.; et al. Extensive promoter-centered chromatin interactions provide a topological basis for transcription regulation. Cell 2012, 148, 84–98. [Google Scholar] [CrossRef]
- Ong, C.T.; Corces, V.G. Enhancer function: new insights into the regulation of tissue-specific gene expression. Nat. Rev. Genet. 2011, 12, 283–293. [Google Scholar]
- Tuan, D.; Kong, S.; Hu, K. Transcription of the hypersensitive site HS2 enhancer in erythroid cells. Proc. Natl. Acad. Sci. USA 1992, 89, 11219–11223. [Google Scholar] [CrossRef]
- Zhu, X.; Ling, J.; Zhang, L.; Pi, W.; Wu, M.; Tuan, D. A facilitated tracking and transcription mechanism of long-range enhancer function. Nucleic Acids Res. 2007, 35, 5532–5544. [Google Scholar] [CrossRef]
- Leach, K.M.; Nightingale, K.; Igarashi, K.; Levings, P.P.; Engel, J.D.; Becker, P.P.; Bungert, J. Reconstitution of human beta-globin locus control region hypersensitive sites in the absence of chromatin assembly. Mol. Cell Biol. 2001, 21, 2629–2640. [Google Scholar] [CrossRef]
- Routledge, S.J.; Proudfoot, N.J. Definition of transcriptional promoters in the human beta globin locus control region. J. Mol. Biol. 2002, 323, 601–611. [Google Scholar] [CrossRef]
- Schmitt, S.; Prestel, M.; Paro, R. Intergenic transcription through a polycomb group response element counteracts silencing. Genes Dev. 2005, 19, 697–708. [Google Scholar] [CrossRef]
- Walia, H.; Chen, H.Y.; Sun, J.M.; Holth, L.T.; Davie, J.R. Histone acetylation is required to maintain the unfolded nucleosome structure associated with transcribing DNA. J. Biol. Chem. 1998, 273, 14516–14522. [Google Scholar]
- Hatzis, P.; Talianidis, I. Dynamics of enhancer-promoter communication during differentiation-induced gene activation. Mol. Cell 2002, 10, 1467–1477. [Google Scholar] [CrossRef]
- Rogers, R.J.; Chesrown, S.E.; Kuo, S.; Monnier, J.M.; Nick, H.S. Cytokine-inducible enhancer with promoter activity in both the rat and human manganese-superoxide dismutase genes. Biochem. J. 2000, 347, 233–242. [Google Scholar] [CrossRef]
- Kowalczyk, M.S.; Hughes, J.R.; Garrick, D.; Lynch, M.D.; Sharpe, J.A.; Sloane-Stanley, J.A.; McGowan, S.J.; de Gobbi, M.; Hosseini, M.; Vernimmen, D.; et al. Intragenic enhancers act as alternative promoters. Cell 2012, 45, 447–458. [Google Scholar]
- Wang, D.; Garcia-Bassets, I.; Benner, C.; Li, W.; Sue, X.; Zhou, Y.; Qiu, J.; Lie, W.; Kaikkonen, M.U.; Ohgi, K.A.; et al. Reprogramming transcription by distinct classes of enhancers functionally defined by eRNA. Nature 2011, 474, 390–394. [Google Scholar]
- Sanyal, A.; Lajoie, B.R.; Jain, G.; Dekker, J. The long-range interaction landscape of gene promoters. Nature 2012, 489, 109–113. [Google Scholar]
- Khalil, A.; Guttman, M.; Huarte, M.; Garber, M.; Raj, A.; Rivea-Morales, D.; Thomas, K.; Presser, A.; Bernstein, B.E.; van Oudenaarden, A.; et al. Many human large intergenic noncoding RNAs associate with chromatin-modifying complexes and affect gene expression. Proc. Natl. Acad. Sci. USA 2009, 106, 11667–11672. [Google Scholar]
- Orum, U.A.; Derrien, T.; Beringer, M.; Gumireddy, K.; Gardini, A.; Bussotti, G.; Lai, F.; Zytnicki, M.; Notredame, C.; Huang, Q.; et al. Long noncoding RNAs with enhancer-like function in human cells. Cell 2010, 143, 46–58. [Google Scholar] [CrossRef]
- Flynn, R.A.; Chang, H.Y. Active chromatin and noncoding RNAs: An intimate relationship. Curr. Opin. Genet. Dev. 2012, 22, 172–178. [Google Scholar] [CrossRef]
- Caudron-Herger, M.; Rippe, K. Nuclear architecture by RNA. Curr. Opin. Genet. Dev. 2012, 22, 179–187. [Google Scholar] [CrossRef]
- Shevtsov, S.P.; Dundr, M. Nucleation of nuclear bodies by RNA. Nat. Cell Biol. 2011, 13, 167–173. [Google Scholar] [CrossRef]
- Fnu, S.; Williamson, E.A.; de Haro, L.P.; Brenneman, M.; Wray, J.; Shaheen, M.; Radhakrishnan, K.; Lee, S.H.; Nickoloff, J.A.; Hromas, R. Methylation of histone H3 lysine 36 enhances DNA repair by non-homologous end-joining. Proc. Natl. Acad. Sci. USA 2011, 108, 540–545. [Google Scholar]
- Rada-Iglesias, A.; Bajpai, R.; Swigut, T.; Brugmann, S.A.; Flynn, R.A.; Wysocka, J. A unique chromatin signature uncovers early developmental enhancers in humans. Nature 2011, 470, 279–283. [Google Scholar]
- Magnani, L.; Eeckhoute, J.; Lupien, M. Pioneer factors: directing transcriptional regulators within the chromatin environment. Trends Genet. 2011, 27, 465–474. [Google Scholar] [CrossRef]
- Zhu, Y.; van Essen, D.; Saccani, S. Cell type-specific control of enhancer activity by H3K9 trimethylation. Mol. Cell 2012, 46, 408–423. [Google Scholar] [CrossRef]
- Chong, M.M.; Simpson, N.; Ciofani, M.; Chen, G.; Collins, A.; Littman, D.R. Epigenetic propagation of CD4 expression is established by the Cd4 proximal enhancer in helper T-cells. Genes Dev. 2010, 24, 659–669. [Google Scholar] [CrossRef]
- DuBose, A.J.; Smith, E.Y.; Johnstone, K.A.; Resnick, J.L. Temporal and developmental requirements for the Prader-Willi imprinting center. Proc. Natl. Acad. Sci. USA 2012, 109, 3446–3450. [Google Scholar]
- Klein, S.; Gerster, T.; Picard, D.; Radbruch, A.; Schaffner, W. Evidence for transient requirement of the IgH enhancer. Nucl. Acids Res. 1985, 13, 8901–8912. [Google Scholar] [CrossRef]
- Yoo, E.J.; Cooke, N.E.; Liebhaber, S.A. An RNA-independent linkage of noncoding transcription to long-range enhancer function. Mol. Cell Biol. 2012, 32, 2020–2029. [Google Scholar] [CrossRef]
- Sequeira-Mendes, J.; Gomez, M. On the opportunistic nature of transcription and replication initiation of the metazoan genome. Bioessays. 2011, 34, 119–125. [Google Scholar] [CrossRef] [Green Version]
- Thanos, D.; Maniatis, T. Virus induction of human IFN beta gene expression requires the assembly of an enhanceosome. Cell 1995, 83, 1091–1100. [Google Scholar] [CrossRef]
© 2012 by the authors; licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution license (http://creativecommons.org/licenses/by/3.0/).
Share and Cite
Stees, J.S.; Varn, F.; Huang, S.; Strouboulis, J.; Bungert, J. Recruitment of Transcription Complexes to Enhancers and the Role of Enhancer Transcription. Biology 2012, 1, 778-793. https://doi.org/10.3390/biology1030778
Stees JS, Varn F, Huang S, Strouboulis J, Bungert J. Recruitment of Transcription Complexes to Enhancers and the Role of Enhancer Transcription. Biology. 2012; 1(3):778-793. https://doi.org/10.3390/biology1030778
Chicago/Turabian StyleStees, Jared S., Fred Varn, Suming Huang, John Strouboulis, and Jörg Bungert. 2012. "Recruitment of Transcription Complexes to Enhancers and the Role of Enhancer Transcription" Biology 1, no. 3: 778-793. https://doi.org/10.3390/biology1030778



