Rapid Development of Microsatellite Markers for Plantago ovata Forsk.: Using Next Generation Sequencing and Their Cross-Species Transferability
Abstract
:1. Introduction
2. Material and Methods
2.1. Germplasm Collection and DNA Extraction
| Serial No. | Species | Genotypes | Characteristics/Features |
|---|---|---|---|
| 1 | Plantago ovata | GI-3 | High yielding, susceptible to downy mildew, aphids, seed shattering |
| GI-2 | Susceptible to downy mildew, aphids, seed shattering | ||
| RI-89 | - | ||
| DPO13 | Early maturing mutant | ||
| DPO14 | Early maturing mutant with desirable traits, such as high seed yield, early and uniform seed maturation and high harvest index. | ||
| EC124345 | Resistant to downy mildew; line from Pakistan | ||
| 2 | Plantago arneria | - | Resistant to downy mildew, aphids |
| 3 | Plantago coronopus | - | Resistant to downy mildew, aphids, seed shattering |
| 4 | Plantago psyllium | - | - |
| 5 | Plantago indica | - | Resistant to downy mildew, aphids, seed shattering |
| 6 | Plantago serraria | - | - |
| 7 | Plantago lanceolata | - | Resistant to downy mildew, aphids, frost, drought |
2.2. Ion Torrent PGM™ Library Preparation, Enrichment of SSR and Sequencing
2.3. Searching for SSR-Containing Sequences and Primer Design
2.4. Validation and Cross-Species Amplification of SSR Markers
2.5. Characteristics of Microsatellites, SSR Marker Scoring and Data Analysis
3. Results and Discussion
3.1. Sequencing by Ion Torrent PGM™
| Sequencing | Reads | 930,940 |
| Total bases (bp) | 176,641,378 | |
| Average read length (bp) | 190 | |
| Highest read length (bp) | 324 | |
| Assembly | No. of contigs | 3,447 |
| Assembled sequence | 257,993 | |
| Unassembled sequences | 672,947 | |
| Contig N50 (bases) | 1,346 | |
| Average lengths contigs | 659 |
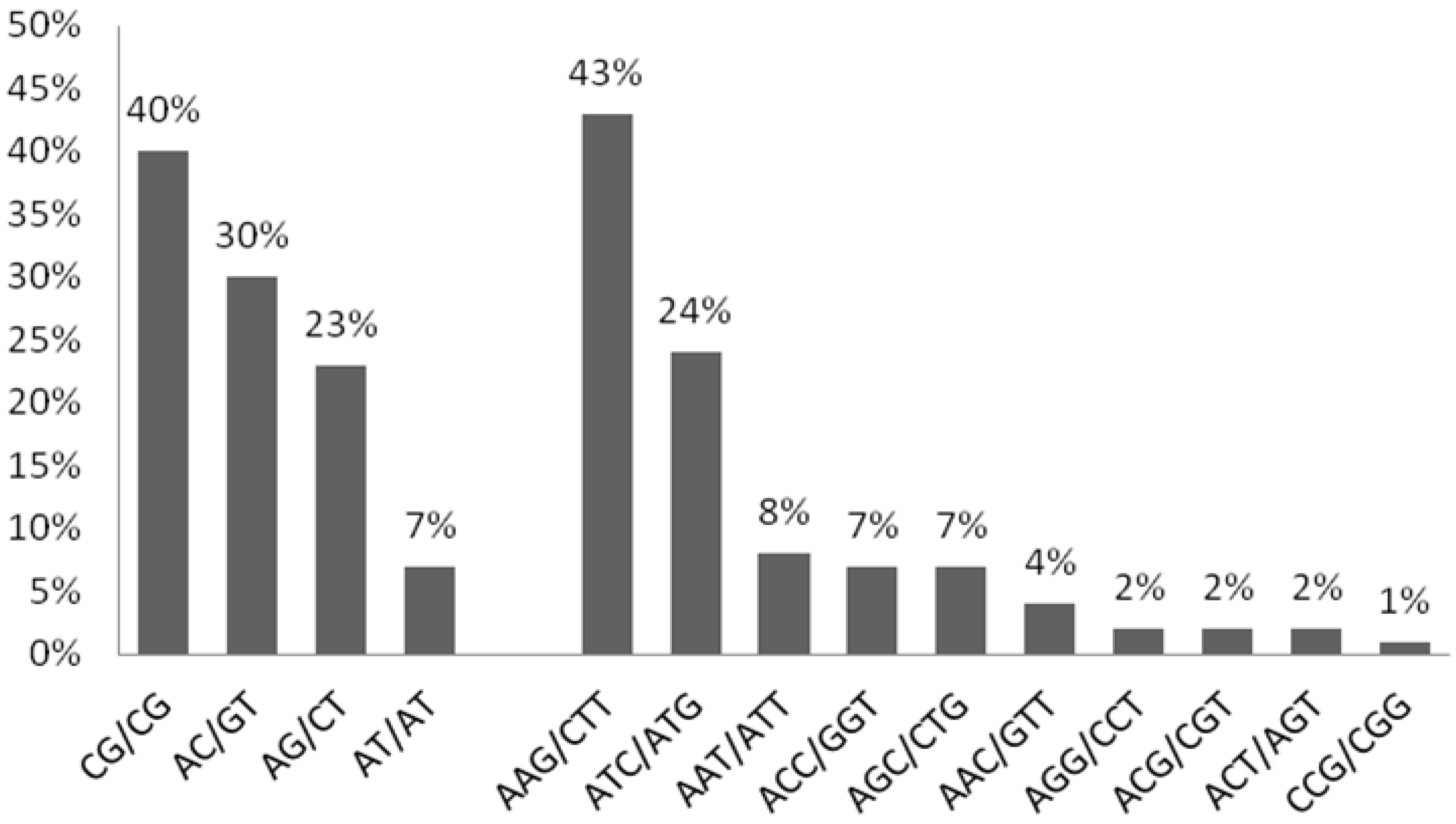
3.2. Summary for SSR-Containing Sequences
| Repeat | Number of Loci Identified | Percentage (%) | Frequency (%) | |
|---|---|---|---|---|
| Class I | Dinucleotide | 25 | 10.96 | 10.04 |
| ≥20 nts | Trinucleotide | 203 | 89.03 | 81.52 |
| Class II | Dinucleotide | 4 | 19.04 | 1.60 |
| ≥10 but <20 nts | Trinucleotide | 17 | 80.95 | 6.82 |
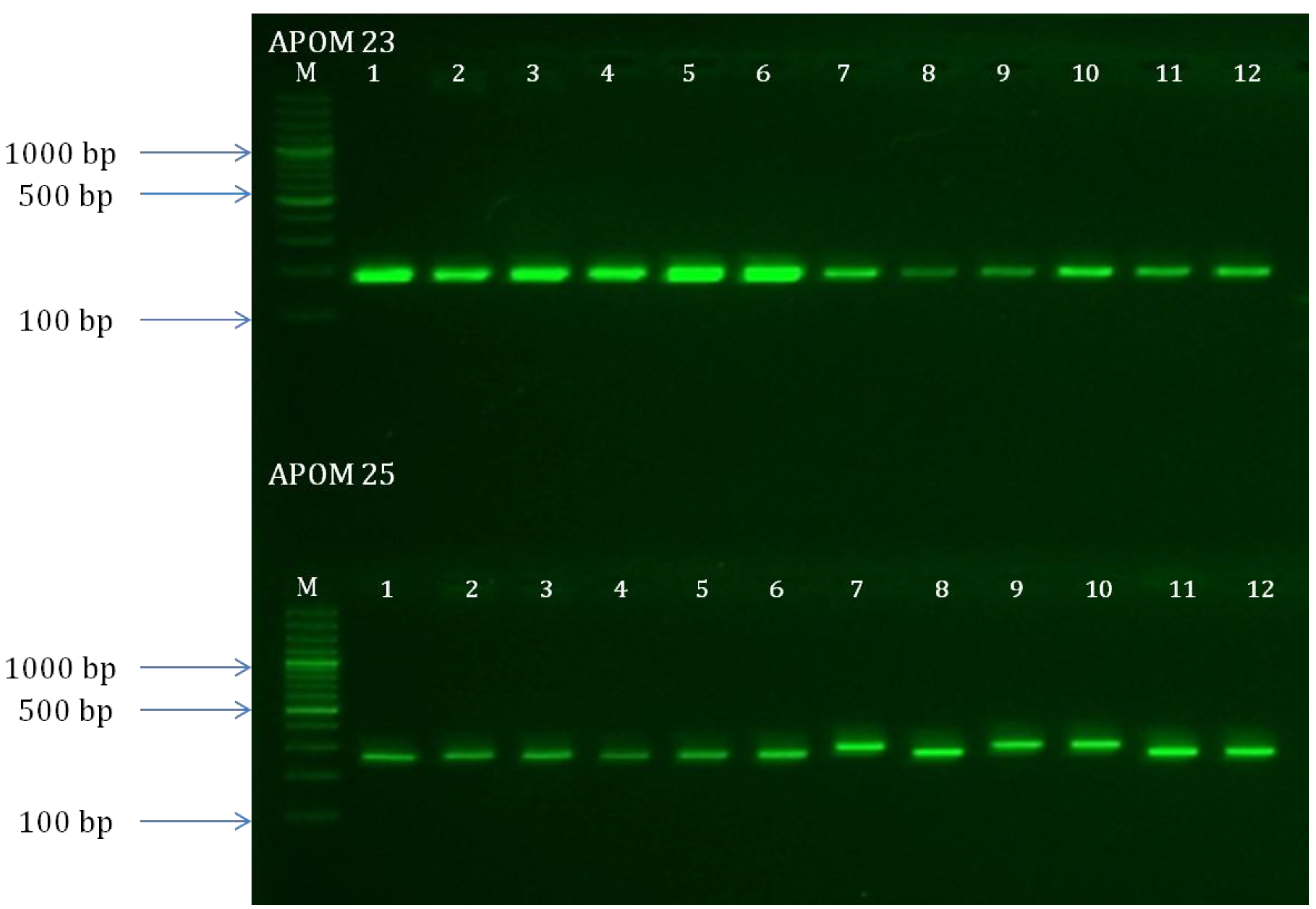
3.3. SSR Amplification and Cross-Species Amplification
| Sr. No | Primer | Primer Sequence (F/R) | Tm * | Repeat Motif | Amplified Fragment Size | No. of Observed Alleles | PIC ** |
|---|---|---|---|---|---|---|---|
| 1 | APOM 1 | TGGCACTTGGGCAAATCTACTTGG | 60 | GC6 | 235 | 6 | 0.75 |
| 2 | APOM3 | GTTTACCTTGCTCAAGTGCTTGCT | 52 | TG10 | 301 | 9 | 0.00 |
| 3 | APOM4 | TGTCACACACACACACACACACAC | 52 | CA10 | 269 | 12 | 0.00 |
| 4 | APOM 5 | ATGGAAGGAGGGTGGTGGAAGTTT | 52 | AG7 | 377 | 6 | 0.75 |
| 5 | APOM 6 | AATTGAAGACTGTGCACTTGGGCG | 60 | TC7 | 181 | 6 | 0.75 |
| 6 | APOM 7 | AGGAAGGAGTCCCGAAGAGTAAGA | 55 | GC7 | 431 | 6 | 0.75 |
| 7 | APOM 8 | ATCTAACTTATCCCTTGGAC | 52 | GC7 | 140 | 12 | 0.00 |
| 8 | APOM 9 | TTATATATCCGTCGATTTTC | 52 | CA10 | 197 | 12 | 0.00 |
| 9 | APOM 10 | TTATTGATTTCGGATCTTTA | 60 | AG7 | 192 | 12 | 0.00 |
| 10 | APOM 11 | ATCTAACTTATCCCTTGGAC | 55 | AG6 | 199 | 6 | 0.75 |
| 11 | APOM 12 | ATTAACTGAGGAAGAGGAAG | 55 | GAA5 | 212 | 12 | 0.00 |
| 12 | APOM 13 | CTATTTGTCCTTCTTTGCTA | 60 | AGC7 | 245 | 12 | 0.00 |
| 13 | APOM 14 | AAATTCTAAGGCTTGTACCT | 60 | TTC4 | 249 | 9 | 0.43 |
| 14 | APOM 15 | ATGGAGTAGGAAGTGAGAGT | 60 | GAT4 | 183 | 12 | 0.00 |
| 15 | APOM 16 | CATAGAGTATGCTGGAAAAT | 60 | TCC4 | 190 | 6 | 0.75 |
| 16 | APOM 17 | TGTAAGGTATTGAGATTGCT | 55 | GTG5 | 156 | 12 | 0.00 |
| 17 | APOM 19 | TGCAATGAGTTTTATTCTCT | 60 | CTC4 | 196 | 6 | 0.75 |
| 18 | APOM 20 | ATT AAA TTC GTA GTT TGA CAC ATC | 62 | CTT4 | 715 | 7 | 0.66 |
| 19 | APOM 21 | AATGTACAAGAAGTTTTCCA | 58 | TCT4 | 281 | 7 | 0.66 |
| 20 | APOM 23 | AGCTAGTATCAATGCACAAT | 47 | CTT4 | 198 | 12 | 0.00 |
| 21 | APOM 24 | TTGCTGAAACTACTCAATCT | 47 | CTG4 | 126 | 6 | 0.75 |
| 22 | APOM 25 | TCCGGAGTATAATAAGTCAA | 47 | TAA5 | 278 | 12 | 0.00 |
| 23 | APOM 27 | ACATTTTCCAACATTTAAGA | 47 | TCA5 | 191 | 11 | 0.16 |
| 24 | APOM 28 | TCAACCGATGGACAGT | 47 | GCT4 | 279 | 6 | 0.00 |
| 25 | APOM 29 | CATAGAGTATGCTGGAAAAT | 47 | TTCT4 | 199 | 12 | 0.75 |
| 26 | APOM 30 | TCAACCGATGGACAGT | 47 | GCT4 | 279 | 6 | 0.75 |
| 27 | APOM 31 | CGAGTTATCTTTACTCGATG | 47 | TGA4 | 192 | 12 | 0.00 |
| 28 | APOM 34 | GCGACTCCTCCCTATCTTCTTCAT | 57 | AC8 | 273 | 12 | 0.00 |
| 29 | APOM 36 | AATAAGGGAGGTGCACAGACCGAT | 57 | TA6 | 241 | 12 | 0.00 |
| 30 | APOM 37 | TGGTTTAGGCTTTCCTGCCTTTGC | 57 | TG7 | 307 | 9 | 0.43 |
| Primer | P. arneria | P. coronopus | P. psyllium | P. indica | P. serraria | P. lanceolata |
|---|---|---|---|---|---|---|
| APOM 3 | − | + | − | − | + | + |
| APOM 4 | + | + | + | + | + | + |
| APOM 8 | + | + | + | + | + | + |
| APOM 9 | + | + | + | + | + | + |
| APOM 10 | + | + | + | + | + | + |
| APOM 12 | + | + | + | + | + | + |
| APOM 13 | + | + | + | + | + | + |
| APOM 14 | − | + | − | − | + | + |
| APOM 15 | + | + | + | + | + | + |
| APOM 17 | + | + | + | + | + | + |
| APOM 20 | + | − | − | − | − | − |
| APOM 21 | + | − | − | − | − | − |
| APOM 23 | + | + | + | + | + | + |
| APOM 25-1 | − | + | − | − | + | + |
| APOM 25-2 | + | − | + | + | − | − |
| APOM 27 | − | + | + | + | + | + |
| APOM 29 | + | + | + | + | + | + |
| APOM 31 | + | + | + | + | + | + |
| APOM 34 | + | + | + | + | + | + |
| APOM 36 | + | + | + | + | + | + |
| APOM 37 | + | − | + | − | − | + |
| Total | 17 | 16 | 15 | 15 | 16 | 17 |
| Transferability (%) Percentage | 85 | 80 | 75 | 75 | 80 | 85 |
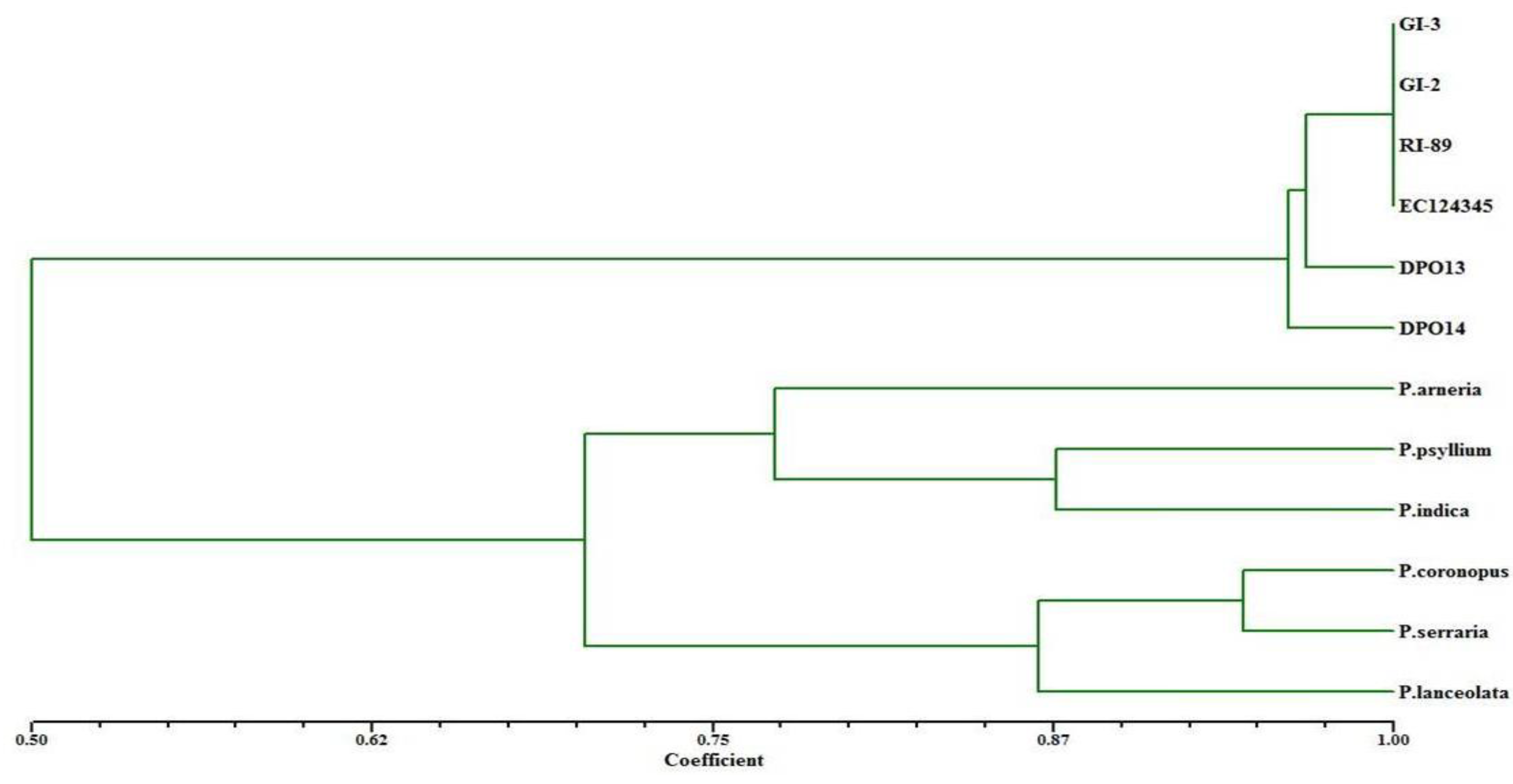
3.4. SSR Marker Scoring and Data Analysis
Acknowledgments
Author Contributions
Conflicts of Interest
References
- Dalal, K.C.; Sriram, S. Psyllium. In Advances in Horticulture. 11. Medicinal and Aromatic Plants; Chadha, K.L., Gupta, R., Eds.; Malhotra Publishing House: New Delhi, Indian, 1995; Volume 11, pp. 575–604. [Google Scholar]
- Chandler, C. Improvement of Plantago for mucilage production and growth in United States. Contr. Boyce Thomoson. Inst. 1954, 17, 495–505. [Google Scholar]
- Zargari, A. Medicinal Plants, 4th ed.; University of Tehran: Tehran, Iran, 1990; Volume 4, pp. 18–19. [Google Scholar]
- Ansari, S.H.; Ali, M. Chemical, pharmacological and clinical evaluation of Plantago ovata Forsk. Hamdard Med. 1996, 39, 63–85. [Google Scholar]
- Patel, A.; Saravanan, R. Screening of Plantago species for physiological parameters in relation to seed yield. Electron. J. Plant Breed. 2010, 1, 1454–1460. [Google Scholar]
- Dhar, M.K.; Kaul, S.; Friebe, B.; Gill, B.S. Chromosome identification in Plantago ovata Forsk. through C-banding and FISH. Curr. Sci. 2002, 83, 150–152. [Google Scholar]
- Punia, M.S.; Sharma, G.D.; Verma, P.K. Genetics and breeding of Plantago ovata Forsk. A review. Inter. J. Trop. Agri. 1985, 3, 255–264. [Google Scholar]
- Lal, R.K.; Sharma, J.R.; Misra, H.O. Induced variability and varietal selection in Isabgol (Plantago ovata). J. Med. Arom. Plant Sci. 1999, 21, 34–37. [Google Scholar]
- Mittal, S.P.; Bhagat, N.R.; Maheshwari, M.L. Improvement of Plantago ovata Forsk. through tetraploidy and mutation breeding. Indian J. Agric. Sci. 1975, 45, 426–429. [Google Scholar]
- Bhagat, N.R.; Hardas, M.W. Studies on induce and natural variation in Plantago ovata Forsk. Indian Drugs 1980, 17, 376–380. [Google Scholar]
- Bhan, A.; Dhar, M.K.; Langer, A. Genetic diversity among Plantagos XX. A tetraploid Plantago lagopus L. Cytologia 1990, 55, 511–517. [Google Scholar] [CrossRef]
- Dalal, K.C.; Sriram, S. Psyllium. In Advances in Horticulture medicinal and Aromatic Plants; Chaddha, K.L., Gupta, R., Eds.; Malhotra Publishing House: New Delhi, India, 1995; Volume 2, pp. 575–604. [Google Scholar]
- Dhar, M.K.; Kaul, S.; Sareen, S.; Koul, A.K. Plantago ovata: Genetic diversity, cultivation, utilization and chemistry. Plant Gen. Resour. 2005, 3, 252–263. [Google Scholar] [CrossRef]
- Samantaray, S.; Dhagat, U.M.; Maiti, S. Evaluation of genetic relationships in Plantago species using Random Amplified Polymorphic DNA (RAPD) markers. Plant Biotechnol. 2010, 27, 297–303. [Google Scholar] [CrossRef]
- Vala, A.G.; Fougat, R.S.; Jadeja, G.C. Genetic diversity of Plantago ovata Forsk. through RAPD markers. Elec. J. Plant Breed. 2011, 2, 592–596. [Google Scholar]
- Rohila, A.K.; Kumar, M.; Sindhu, A.; Boora, K.S. Genetic diversity analysis of the medicinal herb Plantago ovata (Forsk.). Afr. J. Biotechnol. 2012, 11, 15835–15842. [Google Scholar]
- Wolff, K.; Morgan, R.M. PCR markers distinguish Plantago major subspecies. Theor. Appl. Genet. 1998, 96, 282–286. [Google Scholar] [CrossRef]
- Wolff, K.; Morgan, R.M.; Davison, A.W. Patterns of molecular genetic variation in Plantago major and P. intermedia in relation to ozone resistance. New Phytologist. 2000, 145, 501–509. [Google Scholar] [CrossRef]
- Kaswan, V.; Joshi, A.; Maloo, S.R. Assessment of genetic diversity in Isabgol (Plantago ovata Forsk.) using random amplified polymorphic DNA (RAPD) and inter-simple sequence repeat (ISSR) markers for developing crop improvement strategies. Afr. J. Biotechnol 2013, 12, 3622–3635. [Google Scholar]
- Koorevaar, G.N.; Ivanovic, S.; van Damme, J.M.M.; Koelewijn, H.P.; van’T Westende, W.P.C.; Smulders, M.J.M.; Vosman, B. Dinucleotide repeat microsatellite markers for buck’s-horn Plantain (Plantago coronopus). Mol. Ecol. Note. 2002, 2, 524–526. [Google Scholar] [CrossRef]
- Hale, M.L.; Wolff, K. Polymorphic microsatellite loci in Plantago lanceolata. Mol. Ecol. Notes. 2003, 3, 134–135. [Google Scholar] [CrossRef]
- Squirrell, J.; Wolff, K. Isolation of polymorphic microsatellite loci in Plantago major and P. intermedia. Mol. Ecol. Notes. 2001, 1, 179–181. [Google Scholar] [CrossRef]
- Parida, S.K.; Kalia, S.K.; Kaul, S.; Dalal, V.; Hemaprabha, G.; Selvi, A.; Pandit, A.; Singh, A.; Gaikwad, K.; Sharma, T.R.; et al. Informative genomic microsatellite markers for efficient genotyping applications in sugarcane. Theor. Appl. Genet. 2009, 118, 327–338. [Google Scholar] [CrossRef]
- Zane, L.; Bargelloni, L.; Patarnello, T. Strategies for microsatellites isolation, a review. Mol. Ecol. 2002, 44, 1–16. [Google Scholar]
- Schadt, E.E.; Turner, S.; Kasarskis, A. A window into third-generation sequencing. Hum. Mol. Gen. 2010, 19, R227–R240. [Google Scholar] [CrossRef]
- Doyle, J.J.; Doyle, J.L. Isolation of Plant DNA from Fresh Tissue. Focus 1990, 12, 13–15. [Google Scholar]
- Gnirke, A.; Melnikov, A.; Maguire, J.; Rogov, P.; LeProust, E.M.; Brockman, W.; Fennell, T.; Giannoukos, G.; Fisher, S.; Russ, C.; et al. Solution hybrid selection with ultra-long oligonucleotides for massively parallel targeted sequencing. Nat. Biotechnol 2009, 27, 182–189. [Google Scholar] [CrossRef]
- Thiel, T.; Michalek, W.; Varshney, R.K.; Graner, A. Exploiting, EST databases for the development and characterization of gene-derived SSR-markers in barley (Hordeum vulgare L.). Theor. Appl. Genet 2003, 106, 411–422. [Google Scholar]
- Perl-based Microsatellite Search Module. Available online: http://pgrc.ipk-gatersleben.de/Misa (accessed on 1 December 2012).
- Senthilvel, S.; Jayashree, B.; Mahalakshmi, V.; Kumar, P.S.; Nakka, S.; Nepolean, T.; Hash, C.T. Development and mapping of simple sequence repeat markers for pearl millet from data mining of expressed sequence tags. BMC Plant Biol. 2008, 8, 119. [Google Scholar] [CrossRef]
- Rozen, S.; Skaletsky, H.J. Primer3 on the WWW for general users and for biologist programmers. In Methods in Molecular Biology; Misener, S., Krawetz, S.A., Eds.; Humana Press Inc.: Totowa, NJ, USA, 2000; pp. 365–386. [Google Scholar]
- Integrated DNA Technologies. Available online: http://www.idtdna.com/pages/scitools (accessed on 15 December 2012).
- Rohlf, F.J. NTSYS-PC Numerical Taxonomy and Multivariate Analysis System; Version 2.2.; State University of New York: Stony brook, NY, USA, 1994. [Google Scholar]
- Anderson, J.A.; Churchill, G.A.; Autrique, J.E.; Tanksley, S.D.; Sorrells, M.E. Optimizing parental selection for genetic linkage maps. Genome 1993, 36, 181–186. [Google Scholar]
- Rafalski, J.A.; Vogel, J.M.; Morgante, M.; Powell, W.; Andre, C.; Tingey, S.V. Generating and using DNA markers in plants. In Non-Mammalian Genomic Analysis: A Practical Guide; Birren, B., Lai, E., Eds.; Academic: San Diego, CA, USA, 1996; pp. 75–135. [Google Scholar]
- Santana, Q.; Coetzee, M.; Steenkamp, E.; Mlonyeni, O.; Hammond, G.; Wingfield, M.; Wingfield, B. Microsatellite discovery by deep sequencing of enriched genomic libraries. Biotechniques 2009, 46, 217–223. [Google Scholar] [CrossRef]
- Gardner, M.G.; Fitch, A.J.; Bertozzi, T.; Lowe, A.J. Rise of the machines recommendations for ecologists when using next generation sequencing for microsatellite development. Mol. Ecol. Resour. 2011, 11, 1093–1101. [Google Scholar] [CrossRef]
- Elliott, C.P.; Enriqht, N.J.; Allcock, R.J.; Gardner, M.G.; Meglécz, E.; Anthony, J.; Krauss, S.L. Microsatellite markers from the Ion Torrent: A multi-species contrast to 454 shotgun sequencing. Mol. Ecol. Resour. 2014, 14, 554–568. [Google Scholar] [CrossRef]
- Ellegren, H. Microsatellites: Simple sequences with complex evolution. Nat. Rev. Genet. 2004, 6, 435–445. [Google Scholar] [CrossRef]
- Temnykh, S.; DeClerck, G.; Lukashova, A.; Lipovich, L.; Cartinhour, S.; McCouch, S. Computational and experimental analysis of microsatellites in rice (Oryza sativa L.): Frequency, length variation, transposon associations, and genetic marker potential. Genome Res. 2001, 11, 1441–1452. [Google Scholar] [CrossRef]
- Pupko, T.; Graur, D. Evolution of microsatellites in the yeast Saccharomyces cerevisiae: Role of length and number of repeated units. J. Mol. Evol. 1999, 48, 313–316. [Google Scholar] [CrossRef]
- Zhao, Z.; Guo, C.; Sutharzan, S.; Li, P.; Echt, C.S.; Zhang, J.; Liang, C. Genome-wide analysis of tandem repeats in plants and green algae. G3(Bethesda) 2014, 4, 67–78. [Google Scholar]
- Liu, S.R.; Li, W.Y.; Long, D.; Hu, C.G.; Zhang, J.Z. Development and characterization of genomic and expressed SSRs in Citrus by genome-wide analysis. PLoS One 2013, 8, e75149. [Google Scholar]
- Cavagnaro, P.F.; Senalik, D.A.; Yang, L.; Simon, P.W.; Harkins, T.T.; Kodira, C.D.; Huang, S.; Weng, Y. Genome-wide characterization of simple sequence repeats in cucumber (Cucumis sativus L.). BMC Genomics 2010, 11, 569. [Google Scholar] [CrossRef]
- Sonah, H.; Deshmukh, R.K.; Sharma, A.; Singh, V.P.; Gupta, D.K.; Gacche, R.N.; Rana, J.C.; Singh, N.K.; Sharma, T.R. Genome-wide distribution and organization of microsatellites in plants, an insight into marker development in Brachypodium. PLoS One 2011, 6, e21298. [Google Scholar]
- Lin, H.; Chiang, C.Y.; Chang, S.B.; Kuoh, C.S. Development of Simple Sequence Repeats (SRR) markers in Setaria italica (Poaceae) and cross-amplification in related species. Int. J. Mol. Sci. 2011, 12, 7835–7845. [Google Scholar] [CrossRef]
- Dhar, M.K.; Fuchs, J.; Houben, A. Distribution of eu- and heterochromatin in Plantago ovata. Cytogenet. Gen. Res. 2009, 125, 235–240. [Google Scholar] [CrossRef]
- Yonemaru, J.; Ando, T.; Mizubayashi, T.; Kasuga, S.; Matsumoto, T.; Yano, M. Development of genome-wide simple sequence repeat markers using whole-genome shotgun sequences of sorghum (Sorghum bicolor (L.) Moench). DNA Res. 2009, 16, 187–193. [Google Scholar] [CrossRef]
- Katti, M.V.; Ranjekar, P.K.; Gupta, V.S. Differential distribution of simple sequence repeats in eukaryotic genome sequences. Mol. Biol. Evol. 2001, 18, 1161–1167. [Google Scholar] [CrossRef]
- Kalia, R.K.; Rai, M.K.; Kalia, S.; Singh, R.; Dhawan, A.K. Microsatellite markers, an overview of the recent progress in plants. Euphytica 2011, 177, 309–334. [Google Scholar] [CrossRef]
- Mun, J.; Kim, D.J.; Choi, H.K.; Gish, J.; Debellé, F.; Mudge, J.; Denny, R.; Endré, G.; Saurat, O.; Dudez, A.M.; et al. Distribution of microsatellites in the genome of Medicago truncatula: A resource of genetic markers that integrate genetic and physical maps. Genetics 2006, 172, 2541–2555. [Google Scholar]
- Victoria, F.C.; Maia, L.; Oliveira, A.C. In silico comparative analysis of SSR markers in plants. BMC Plant Biol. 2011, 11, 15. [Google Scholar] [CrossRef]
- Singh, N.; Lal, R.K.; Shasany, A.K. Phenotypic and RAPD diversity among 80 germplasm accessions of the medicinal plant Isabgol (Plantago ovata, Plantaginaceae). Genet. Mol. Res. 2009, 8, 1273–1284. [Google Scholar] [CrossRef]
- Liebhard, R.; Gianfranceschi, L.; Koller, B.; Ryder, C.D.; Tarchini, R.; van de Weg, E.; Gessler, C. Development and characterization of 140 new microsatellites in apple (Malus X domestica Borkh.). Mol. Breed. 2002, 10, 217–241. [Google Scholar] [CrossRef]
- Yamamoto, T.; Srivastava, A.; Rai, K.M.; Ranjan, A.; Singh, S.K.; Nisar, T.; Srivastava, M.; Bag, S.K.; Mantri, S.; Asif, M.H.; et al. Identification of quince varieties using SSR markers developed from pear and apple. Breed. Sci. 2004, 54, 239–244. [Google Scholar] [CrossRef]
- Dayanandan, S.; Bawa, K.; Kesseli, R. Conservation of microsatellites among tropical trees (Leguminosae). Amer. J. Bot. 1997, 84, 1658–1663. [Google Scholar] [CrossRef]
- Jena, S.N.; Srivastava, A.; Rai, K.M.; Ranjan, A.; Singh, S.K.; Nisar, T.; Srivastava, M.; Bag, S.K.; Mantri, S.; Asif, M.H.; et al. Development and characterization of genomic and expressed SSRs for levant cotton (Gossypium herbaceum L.). Theor. Appl. Genet. 2012, 124, 565–576. [Google Scholar]
- Singh, R.K. Development, cross-species/genera transferability of novel EST-SSR markers and their utility in revealing population structure and genetic diversity in sugarcane. Gene 2013, 524, 309–329. [Google Scholar] [CrossRef]
- Kotwal, S.; Dhar, M.K.; Kour, B.; Raj, K.; Kaul, S. Molecular markers unravel intraspecific and interspecific genetic variability in Plantago ovata and some of its wild allies. J. Genet. 2013, 92, 293–298. [Google Scholar] [CrossRef]
© 2014 by the authors; licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution license (http://creativecommons.org/licenses/by/3.0/).
Share and Cite
Fougat, R.S.; Joshi, C.; Kulkarni, K.; Kumar, S.; Patel, A.; Sakure, A.; Mistry, J. Rapid Development of Microsatellite Markers for Plantago ovata Forsk.: Using Next Generation Sequencing and Their Cross-Species Transferability. Agriculture 2014, 4, 199-216. https://doi.org/10.3390/agriculture4020199
Fougat RS, Joshi C, Kulkarni K, Kumar S, Patel A, Sakure A, Mistry J. Rapid Development of Microsatellite Markers for Plantago ovata Forsk.: Using Next Generation Sequencing and Their Cross-Species Transferability. Agriculture. 2014; 4(2):199-216. https://doi.org/10.3390/agriculture4020199
Chicago/Turabian StyleFougat, Ranbir Singh, Chaitanya Joshi, Kalyani Kulkarni, Sushil Kumar, Anand Patel, Amar Sakure, and Jigar Mistry. 2014. "Rapid Development of Microsatellite Markers for Plantago ovata Forsk.: Using Next Generation Sequencing and Their Cross-Species Transferability" Agriculture 4, no. 2: 199-216. https://doi.org/10.3390/agriculture4020199
APA StyleFougat, R. S., Joshi, C., Kulkarni, K., Kumar, S., Patel, A., Sakure, A., & Mistry, J. (2014). Rapid Development of Microsatellite Markers for Plantago ovata Forsk.: Using Next Generation Sequencing and Their Cross-Species Transferability. Agriculture, 4(2), 199-216. https://doi.org/10.3390/agriculture4020199






