Pollen Sterility—A Promising Approach to Gene Confinement and Breeding for Genetically Modified Bioenergy Crops
Abstract
:1. Introduction
2. Results and Discussion
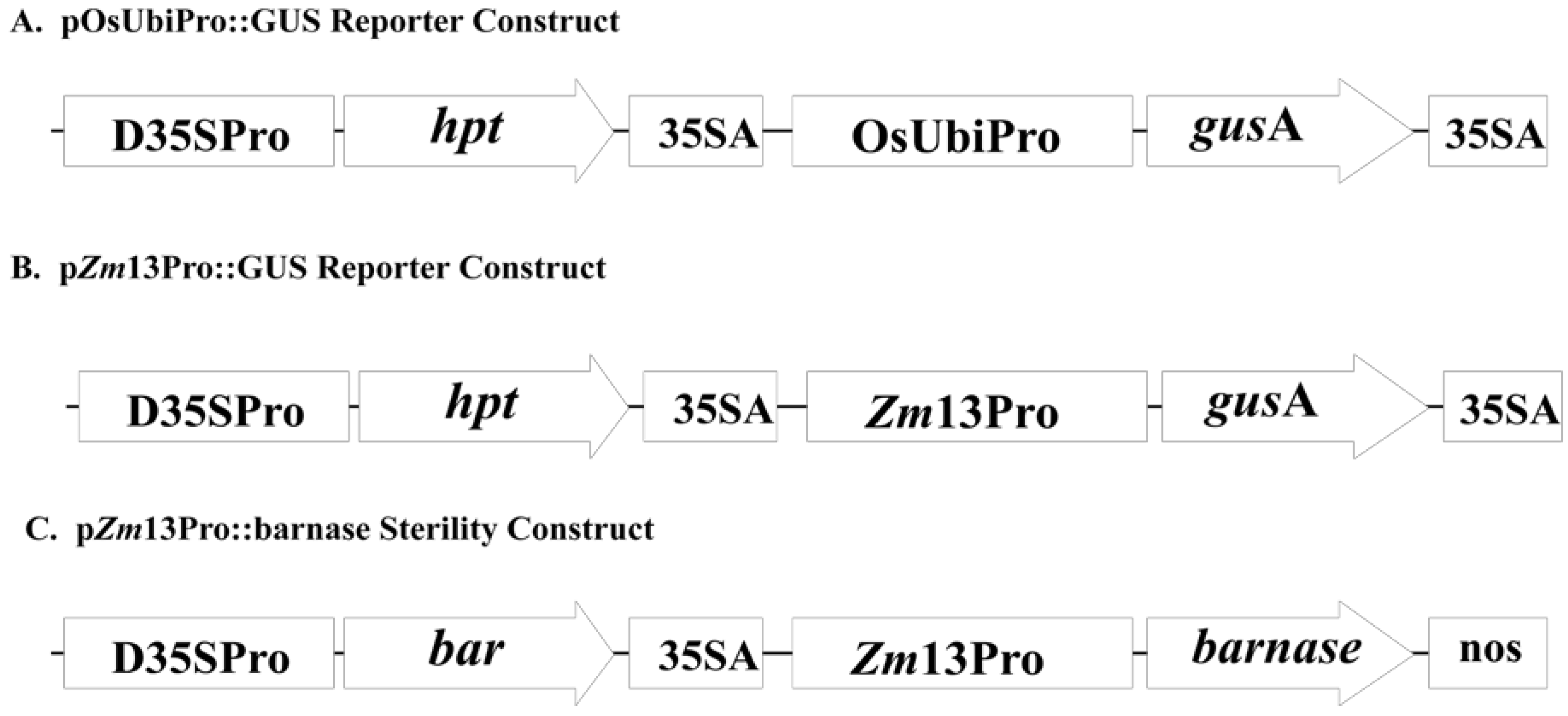
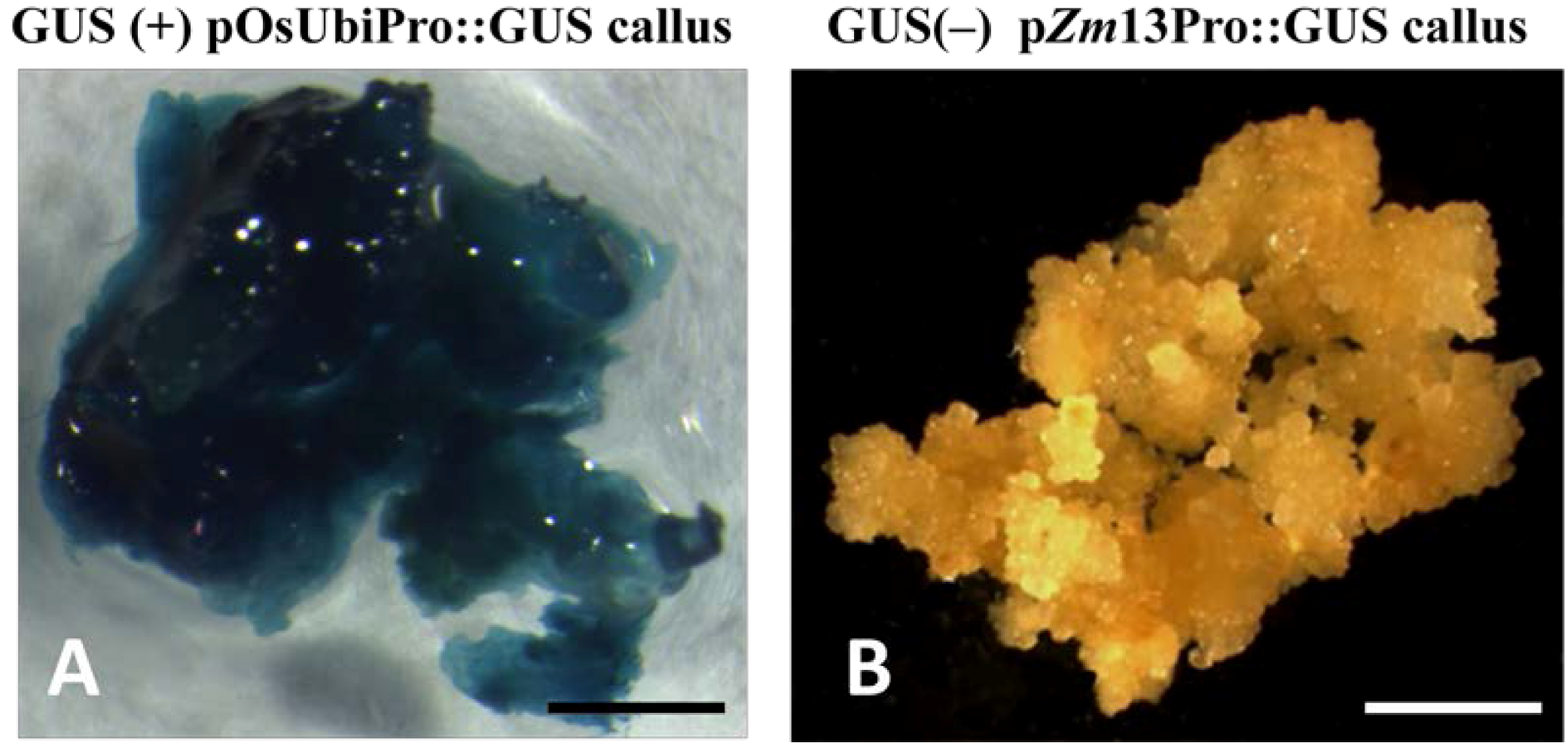
2.1. Co-Expression of hpt and gusA in Transgenic Rice Embryogenic Callus
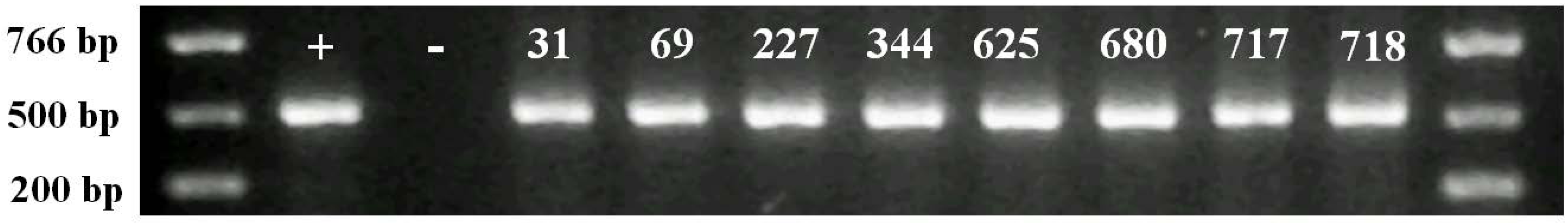
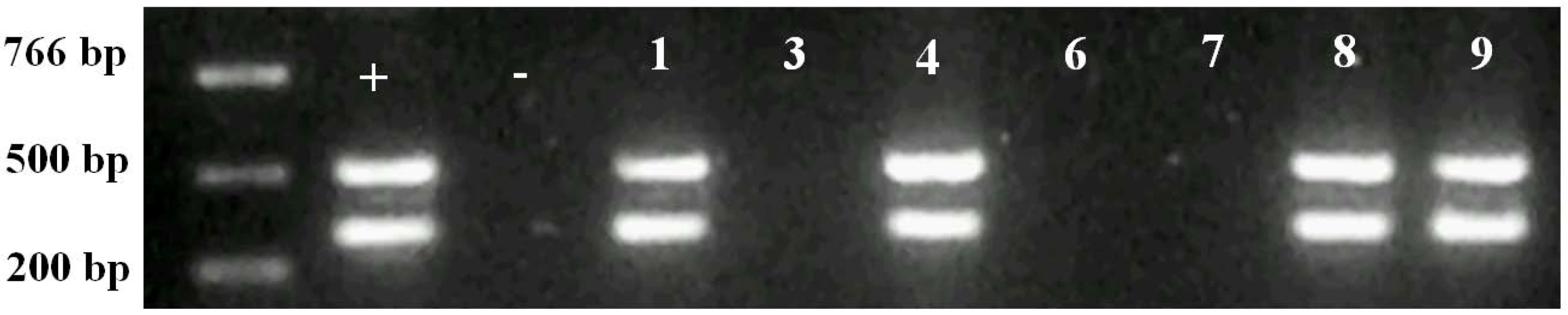
2.2. Molecular Analysis of Transgenic Plants

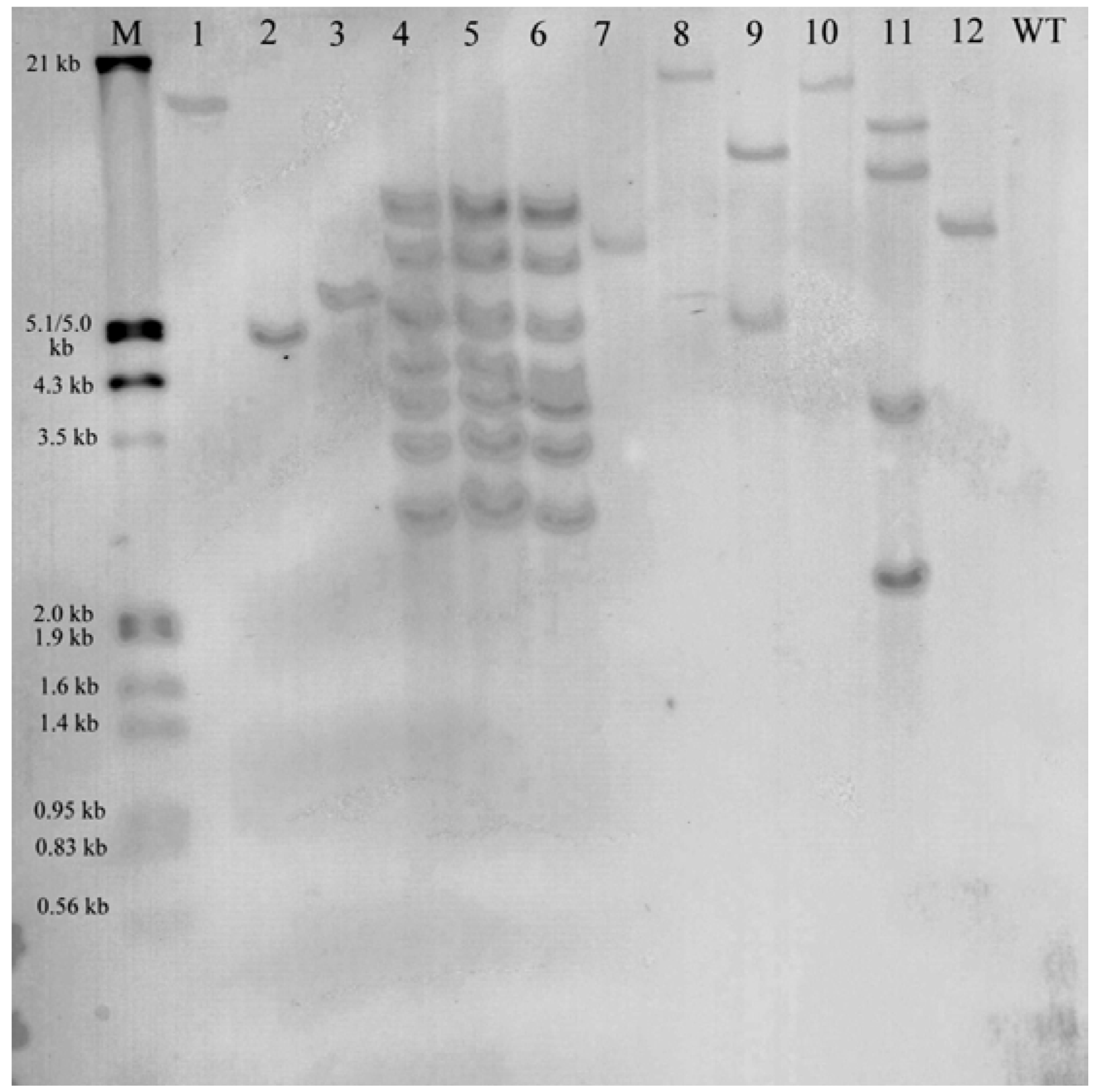
2.3. Histochemical Analysis of gusA Reporter Transgenic Plants
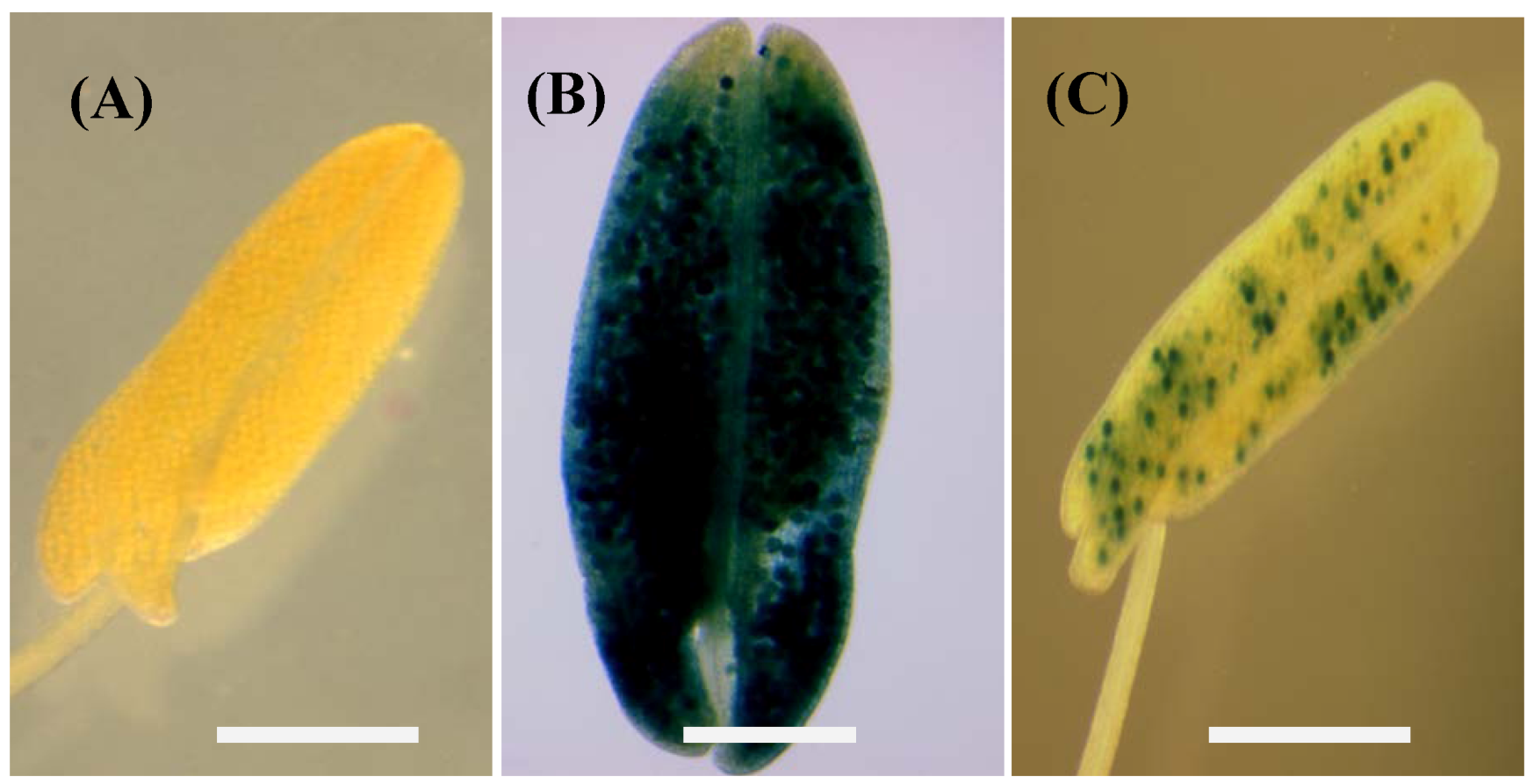
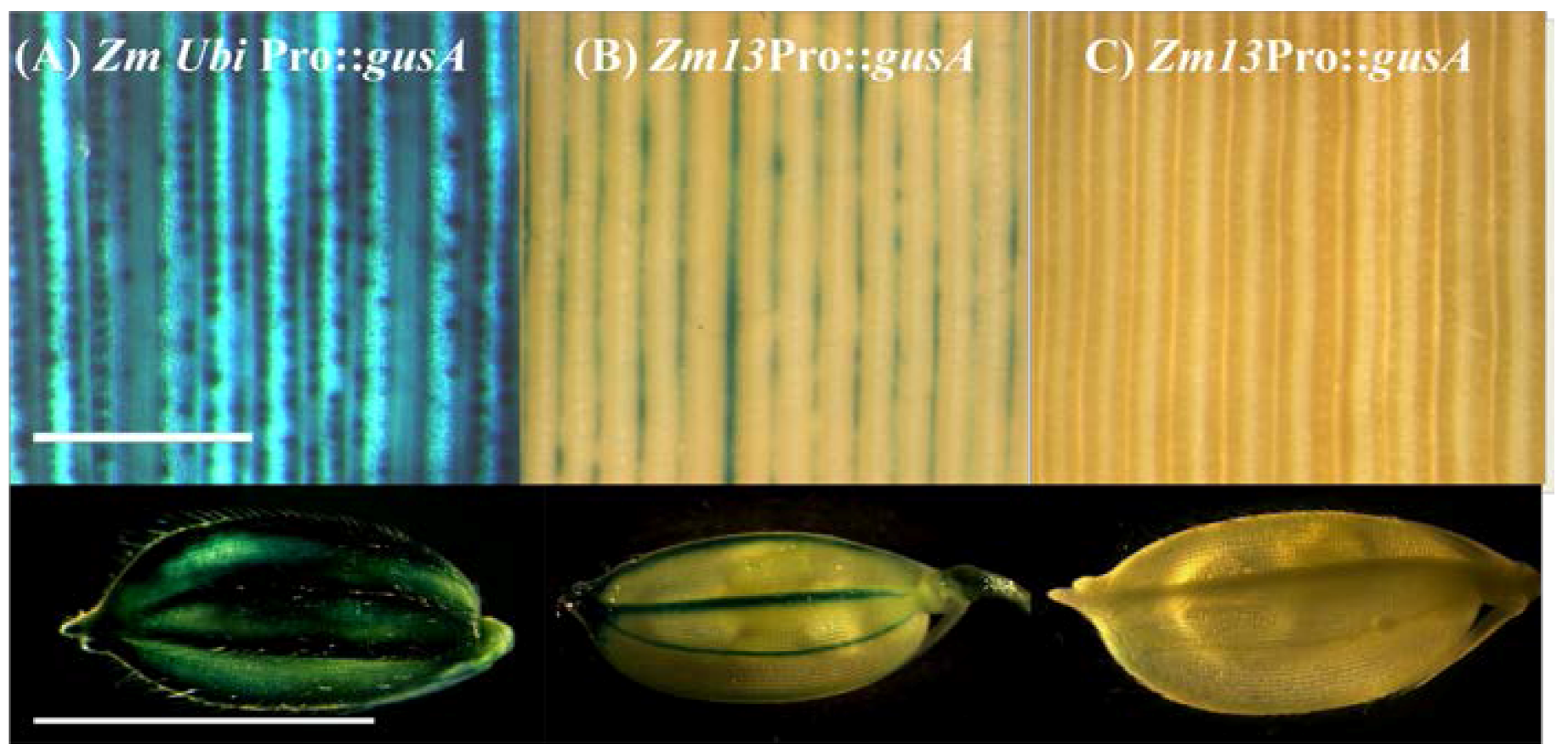


2.4. Expression Analysis of pZm13Pro::barnase Transgenic Plants

3. Experimental Section
3.1. Plant Material
3.2. Induction of Embryogenic Callus
3.3. Agrobacterium Strains and Plasmids
3.3.1. Agrobacterium-Mediated Transformation and Co-cultivation
3.4. Selection of Transformed Colonies and Regeneration of Transgenic Plants
3.5. Molecular Characterization of Transgenic Plants
3.6. Staining for GUS Activity
4. Conclusions
Acknowledgments
References
- Brutnell, T.; Frommer, W. Food for thought: Plant research remains grossly underfunded, despite the demand for increased crop production to support a growing population. The Scientist 2012, 23–24. [Google Scholar]
- Kausch, A.P.; Hague, J.; Oliver, M.; Li, Y.; Daniell, H.; Mascia, P.; Stewart, C.N. Genetic modification in dedicated bioenergy crops and strategies for gene confinement. In Plant Biotechnology for Sustainable Production of Energy and Co-products; Mascia, P.N., Scheffran, Jr., Widholm, J.M., Eds.; Springer Berlin Heidelberg: Berlin, Germany, 2010; Volume 66, pp. 299–315. [Google Scholar]
- Gressel, J. Transgenics are imperative for biofuel crops. Plant Sci. 2008, 174, 246–263. [Google Scholar] [CrossRef]
- Gruskin, D. Agbiotech 2.0. Nat. Biotechnol. 2012, 30, 211–214. [Google Scholar] [CrossRef]
- Lusser, M.; Parisi, C.; Plan, D.; Rodriguez-Cerezo, E. Deployment of new biotechnologies in plant breeding. Nat. Biotechnol. 2012, 30, 231–239. [Google Scholar]
- Stewart, C.N.J. Genetically Modified Planet: Environmental Impacts of Genetically Engineered Plants; Oxford University Press: New York, NY, USA, 2004; p. 240. [Google Scholar]
- Agnostic about agriculture. Nat. Biotechnol. 2012, 30, 197–197. [CrossRef]
- Wolfenbarger, L.L.; Phifer, P.R. The ecological risks and benefits of genetically engineered plants. Science 2000, 290, 2088–2093. [Google Scholar] [CrossRef]
- Kausch, A.P.; Hague, J.; Oliver, M.; Watrud, L.S.; Mallory-Smith, C.; Meier, V.; Stewart, C.N. Gene flow in genetically engineered perennial grasses: Lessons for modification of dedicated bioenergy crops. In Plant Biotechnology for Sustainable Production of Energy and Co-products; Mascia, P.N., Scheffran, Jr., Widholm, J.M., Eds.; Springer Berlin Heidelberg: Berlin, Germany, 2010; Volume 66, pp. 285–297. [Google Scholar]
- Moon, H.S.; Abercrombie, J.M.; Kausch, A.P.; Stewart, C.N. Sustainable use of biotechnology for bioenergy feedstocks. Environ. Manag. 2010, 46, 531–538. [Google Scholar] [CrossRef]
- Kausch, A.P.; Hague, J.; Oliver, M.; Li, Y.; Daniell, H.; Mascia, P.; Watrud, L.S.; Stewart, C.N. Transgenic perennial biofuel feedstocks and strategies for bioconfinement. Biofuels 2010, 1, 163–176. [Google Scholar] [CrossRef]
- Reichman, J.R.; Watrud, L.S.; Lee, E.H.; Burdick, C.A.; Bollman, M.A.; Storm, M.J.; King, G.A.; Mallory-Smith, C. Establishment of transgenic herbicide-resistant creeping bentgrass (Agrostis stolonifera L.) in nonagronomic habitats. Mol. Ecol. 2006, 15, 4243–4255. [Google Scholar] [CrossRef]
- Watrud, L.S.; Lee, E.H.; Fairbrother, A.; Burdick, C.; Reichman, J.R.; Bollman, M.; Storm, M.; King, G.; van de Water, P.K. Evidence for landscape-level, pollen-mediated gene flow from genetically modified creeping bentgrass with CP4 EPSPS as a marker. Proc. Nat. Acad. Sci. USA 2004, 101, 14533–14538. [Google Scholar]
- Wipff, J.K.; Fricker, C. Gene flow from transgenic creeping bentgrass (Agrostis stolonifera L.) in the Willamette valley, Oregon. Int. Turfgrass Soc. Res. J. 2001, 9, 224–242. [Google Scholar]
- Zapiola, M.L.; Campbell, C.K.; Butler, M.D.; Mallory-Smith, C.A. Escape and establishment of transgenic glyphosate-resistant creeping bentgrass agrostis stolonifera in oregon, USA: A 4-year study. J. Appl. Ecol. 2008, 45, 486–494. [Google Scholar] [CrossRef]
- Zapiola, M.L.; Mallory-Smith, C.A.; Thompson, J.H.; Rue, L.J.; Campbell, C.K.; Butler, M.D. Gene escape from glyphosate-resistent creeping bentgrass fields: Past, present, and future. In Proceedings of the Western Society of Weed Science, Portland, OR, USA, 13-15 March 2007; Western Society of Weed Science: Las Cruces, NM, USA; 60, p. 18.
- Luo, K.; Duan, H.; Zhao, D.; Zheng, X.; Deng, W.; Chen, Y.; Stewart, C.N., Jr.; McAvoy, R.; Jiang, X.; Wu, Y.; He, A.; Pei, Y.; Li, Y. “GM-gene-deletor”: Fused loxP-FRT recognition sequences dramatically improve the efficiency of FLP or CRE recombinase on transgene excision from pollen and seed of tobacco plants. Plant Biotechnol. J. 2007, 5, 263–274. [Google Scholar] [CrossRef]
- Oliver, M.J.; Quisenberry, J.E.; Trolinder, N.L.G.; Keim, D.L. Control of Plant Gene Expression. U.S. Patent 5925808, 20 July 1999. [Google Scholar]
- Oliver, M.J.; Quiseberry, J.E.; Trolinder, N.L.G.; Keim, D.L. Control of Plant Gene Expression. U.S. Patent 5723765, 3 March 1998. [Google Scholar]
- Oliver, M.J.; Quiseberry, J.E.; Trolinder, N.L.G.; Keim, D.L. Control of Plant Gene Expression. U.S. Patent 5977441, 2 November 1999. [Google Scholar]
- Djukanovic, V.; Lenderts, B.; Bidney, D.; Lyznik, L.A. A cre::FLP fusion protein recombines FRT or loxP sites in transgenic maize plants. Plant Biotechnol. J. 2008, 6, 770–781. [Google Scholar] [CrossRef]
- Ow, D.W. Gm maize from site-specific recombination technology, what next? Curr. Opin. Biotechnol. 2007, 18, 115–120. [Google Scholar] [CrossRef]
- Srivastava, V.; Ow, D.W. Rare instances of Cre-mediated deletion product maintained in transgenic wheat. Plant Mol. Biol. 2003, 52, 661–668. [Google Scholar] [CrossRef]
- Mallory-Smith, C.; Zapiola, M. Gene flow from glyphosate-resistant crops. Pest Manag. Sci. 2008, 64, 428–440. [Google Scholar] [CrossRef]
- Ellstrand, N.C.; Hoffman, C.A. Hybridization as an avenue of escape for engineered genes-Strategies for risk reduction. BioScience 1990, 40, 438–442. [Google Scholar] [CrossRef]
- Belanger, F.C.; Meagher, T.R.; Day, P.R.; Plumley, K.; Meyer, W.A. Interspecific hybridization between Agrostis stolonifera and related Agrostis species under field conditions. Crop Sci. 2003, 43, 240–246. [Google Scholar]
- Christoffer, P.M. Transgenic glyphosate resistant creeping bentgrass: Studies in pollen-mediated transgene flow. Master’s Thesis, Washington State University, 2003. [Google Scholar]
- Pineyro-Nelson, A.; Van Heerwaarden, J.; Perales, H.R.; Serratos-Hernandez, J.A.; Rangel, A.; Hufford, M.B.; Gepts, P.; Garay-Arroyo, A.; Rivera-Bustamante, R.; Alvarez-Buylla, E.R. Transgenes in mexican maize: Molecular evidence and methodological considerations for GMO detection in landrace populations. Mol. Ecol. 2009, 18, 750–761. [Google Scholar] [CrossRef]
- Snow, A. Unwanted transgenes re-discovered in oaxacan maize. Mol. Ecol. 2009, 18, 569–571. [Google Scholar] [CrossRef]
- Snow, A.A. Transgenic crops-Why gene flow matters. Nat. Biotechnol. 2002, 20, 542–542. [Google Scholar]
- Chandler, S.; Dunwell, J.M. Gene flow, risk assessment and the environmental release of transgenic plants. Crit. Rev. Plant Sci. 2008, 27, 25–49. [Google Scholar] [CrossRef]
- De Block, M.; Debrouwer, D.; and Moens, T. The development of nuclear male sterility system in wheat: Expression of the barnase gene under the control of tapetum specific promoters. Theor. Appl. Genet. 1997, 95, 125–131. [Google Scholar] [CrossRef]
- Jagannath, A.; Bandyopadhyay, P.; Arumugam, N.; Gupta, V.; Kumar, P.; Pental, D. The use of spacer DNA fragment insulates the tissue-specific expression of a cytotoxic gene (barnase) and allows high-frequency generation of transgenic male sterile lines in Brassica juncea L. Mol. Breed. 2001, 8, 11–23. [Google Scholar] [CrossRef]
- Mariani, C.; Beuckeleer, M.; Truettner, J.; Leemans, J.; Goldberg, R. Induction of male sterility in plants by a chimaeric ribonuclease gene. Nature 1990, 374, 737–738. [Google Scholar]
- Mariani, C.; Gossele, V.; Beuckeleer, M.D.; Block, M.D.; Goldberg, R.B.; Greef, W.D.; Leemans, J. A chimaeric ribonuclease-inhibitor gene restores fertility to male sterile plants. Nature 1992, 357, 384–387. [Google Scholar]
- Chase, C.D. Genetically engineered cytoplasmic male sterility. Trends Plant Sci 2006, 11, 7–9. [Google Scholar] [CrossRef]
- Kausch, A.P.; Hague, J.; Deresienski, A.; Tilelli, M.; Longo, C., Jr.; Nelson, K. Male sterility and hybrid plant systems for gene confinement. In Plant Gene Containment, 1st; Oliver, M.J., Li, Y., Eds.; Wiley-Blackwell: Ames, IA, USA, 2012; pp. 83–98. [Google Scholar]
- Tanaka, J. Transgenic male sterility permits efficient recurrent selection in autogamous crops. Crop Sci. 2010, 50, 1124–1127. [Google Scholar] [CrossRef]
- Khan, M.S. Plant biology: Engineered male sterility. Nature 2005, 436, 783–785. [Google Scholar] [CrossRef]
- Hamilton, D.A.; Bashe, D.M.; Stinson, J.R.; Mascarenhas, J.P. Characterization of a pollen-specific genomic clone in maize. Sex. Plant Reprod. 1989, 2, 208–212. [Google Scholar]
- Hanson, D.D.; Hamilton, D.A.; Travis, J.L.; Bashe, D.M.; Mascarenhas, J.P. Characterization of a pollen-specific cdna clone from zea mays and its expression. Plant Cell 1989, 1, 173–179. [Google Scholar]
- Twell, D.; Wing, R.; Yamaguchi, J.; McCormick, S. Isolation and expression of an anther-specific gene from tomato. Mol. Gen. Genet. 1989, 217, 240–245. [Google Scholar] [CrossRef]
- Ursin, V.M.; Yamaguchi, J.; McCormick, S. Gametophytic and sporophytic expression of anther-specific genes in developing tomato anthers. Plant Cell 1989, 1, 727–736. [Google Scholar]
- Bate, N.; Twell, D. Functional architecture of a late pollen promoter: Pollen-specific transcription is developmentally regulated by multiple stage-specific and co-dependent activator elements. Plant Mol. Biol. 1998, 37, 859–869. [Google Scholar] [CrossRef]
- Eyal, Y.; Curie, C.; McCormick, S. Pollen specificity elements reside in 30 bp of the proximal promoters of two pollen-expressed genes. Plant Cell 1995, 7, 373–384. [Google Scholar]
- Guerrero, F.D.; Crossland, L.; Smutzer, G.S.; Hamilton, D.A.; Mascarenhas, J.P. Promoter sequences from a maize pollen-specific gene direct tissue-specific transcription in tobacco. Mol. Gen. Genet. 1990, 224, 161–168. [Google Scholar] [CrossRef]
- Hamilton, D.A.; Schwarz, Y.H.; Rueda, J.; Mascarenhas, J.P. Comparison of transient and stable expression by a pollen-specific promoter: The transformation results do not always agree. Sex. Plant Reprod. 2000, 12, 292–295. [Google Scholar] [CrossRef]
- Hamilton, D.A.; Roy, M.; Rueda, J.; Sindhu, R.K.; Sanford, J.; Mascarenhas, J.P. Dissection of a pollen-specific promoter from maize by transient transformation assays. Plant Mol. Biol. 1992, 18, 211–218. [Google Scholar] [CrossRef]
- Hamilton, D.A.; Schwarz, Y.H.; Mascarenhas, J.P. A monocot pollen-specific promoter contains separable pollen-specific and quantitative elements. Plant Mol. Biol. 1998, 38, 663–669. [Google Scholar] [CrossRef]
- Twell, D.; Yamaguchi, J.; McCormick, S. Pollen-specific gene expression in transgenic plants: Coordinate regulation of two different tomato gene promoters during microsporogenesis. Development 1990, 109, 705–713. [Google Scholar]
- Twell, D.; Yamaguchi, J.; Wing, R.A.; Ushiba, J.; Mccormick, S. Promoter analysis of genes that are coordinately expressed during pollen development reveals pollen-specific enhancer sequences and shared regulatory elements. Genes & Dev. 1991, 5, 496–507. [Google Scholar]
- Wang, J.; Jiang, J.; Oard, J.H. Structure, expression and promoter activity of two polyubiquitin genes from rice (Oryza sativa L.). Plant Sci. 2000, 156, 201–211. [Google Scholar] [CrossRef]
- Chen, J.; Dellaporta, S. Urea-based plant DNA miniprep. In The Maize Handbook; Freeling, M., Walbot, V., Eds.; Springer-Verlag: New York, NY, USA, 1994; pp. 526–528. [Google Scholar]
- Weigel, D.; Ahn, J.H.; Blázquez, M.A.; Borevitz, J.O.; Christensen, S.K.; Fankhauser, C.; Ferrándiz, C.; Kardailsky, I.; Malancharuvil, E.J.; Neff, M.M.; et al. Activation tagging in arabidopsis. Plant Physiol. 2000, 122, 1003–1014. [Google Scholar] [CrossRef]
- Yoo, S.Y.; Bomblies, K.; Yoo, S.K.; Yang, J.W.; Choi, M.S.; Lee, J.S.; Weigel, D.; Ahn, J.H. The 35S promoter used in a selectable marker gene of a plant transformation vector affects the expression of the transgene. Planta 2005, 221, 523–530. [Google Scholar] [CrossRef]
- Zheng, X.; Deng, W.; Luo, K.; Duan, H.; Chen, Y.; McAvoy, R.; Song, S.; Pei, Y.; Li, Y. The cauliflower mosaic virus (CaMV) 35S promoter sequence alters the level and patterns of activity of adjacent tissue- and organ-specific gene promoters. Plant Cell Rep. 2007, 26, 1195–1203. [Google Scholar] [CrossRef]
- De Block, M.D.; Botterman, J.; Vandewiele, M.; Dockx, J.; Thoen, C.; Gossele, V.; Movva, N.R.; Thompson, C.; Montagu, M.V.; Leemans, J. Engineering herbicide resistance in plants by expression of a detoxifying enzyme. EMBO J. 1987, 6, 2513–2518. [Google Scholar]
- Chu, C.C.; Want, C.C.; Sun, C.S.; Hsu, C.; Yin, K.C.; Chu, C.Y.; Bi, F.Y. Establishment of an efficient medium for anther culture of rice, through comparative experiments on the nitrogen sources. Sci. Sin. 1975, 18, 659–668. [Google Scholar]
- Jefferson, R.A.; Kavanagh, T.A.; Bevan, M.W. Gus fusions-β-glucuronidase as a sensitive and versatile gene fusion marker in higher-plants. EMBO J. 1987, 6, 3901–3907. [Google Scholar]
- Murashige, T.; Skoog, F. A revised medium for rapid growth and bioassays with tobacco tissue cultures. Physiol. Plant 1962, 15, 473–497. [Google Scholar] [CrossRef]
- Sambrook, J.; Fritch, E.; Maniatis, T.; Russell, D. Molecular Cloning: A Laboratory Manual, 3rd ed; Cold Spring Harbor Laboratory Press: Long Island, NY, USA, 2001. [Google Scholar]
- Boch, J.; Scholze, H.; Schornack, S.; Landgraf, A.; Hahn, S.; Kay, S.; Lahaye, T.; Nickstadt, A.; Bonas, U. Breaking the code of DNA binding specificity of TAL-type III effectors. Science 2009, 326, 1509–1512. [Google Scholar]
- Bogdanove, A.J.; Voytas, D.F. Tal effectors: Customizable proteins for DNA targeting. Science 2011, 333, 1843–1846. [Google Scholar]
- Li, T.; Liu, B.; Spalding, M.H.; Weeks, D.P.; Yang, B. High-efficiency talen-based gene editing produces disease-resistant rice. Nat. Biotechnol. 2012, 30, 390–392. [Google Scholar] [CrossRef]
- Moscou, M.J.; Bogdanove, A.J. A simple cipher governs DNA recognition by TAL effectors. Science 2009, 326, 1501. [Google Scholar]
- Porteus, M.H. Plant biotechnology: Zinc fingers on target. Nature 2009, 459, 337–338. [Google Scholar] [CrossRef]
- Townsend, J.A.; Wright, D.A.; Winfrey, R.J.; Fu, F.; Maeder, M.L.; Joung, J.K.; Voytas, D.F. High-frequency modification of plant genes using engineered zinc-finger nucleases. Nature 2009, 459, 442–445. [Google Scholar]
- Shukla, V.K.; Doyon, Y.; Miller, J.C.; DeKelver, R.C.; Moehle, E.A.; Worden, S.E.; Mitchell, J.C.; Arnold, N.L.; Gopalan, S.; Meng, X.; et al. Precise genome modification in the crop species zea mays using zinc-finger nucleases. Nature 2009, 459, 437–441. [Google Scholar] [CrossRef]
- Cai, C.; Doyon, Y.; Ainley, W.; Miller, J.; DeKelver, R.; Moehle, E.; Rock, J.; Lee, Y.-L.; Garrison, R.; Schulenberg, L.; et al. Targeted transgene integration in plant cells using designed zinc finger nucleases. Plant Mol. Biol. 2009, 69, 699–709. [Google Scholar] [CrossRef]
- McVetty, P.B.E. Cytoplasmic male sterility. In Pollen Biotechnology and Crop Production and Improvement; Shivanna, K.R., Sawhney, V.K., Eds.; Cambridge University Press: Cambridge, UK, 1997; pp. 155–182. [Google Scholar]
- Schnable, P.S.; Wise, R.P. The molecular basis of cytoplasmic male sterility and fertility restoration. Trends Plant Sci. 1998, 3, 175–180. [Google Scholar] [CrossRef]
- Luo, H.; Lyznik, L.A.; Gidoni, D.; Hodges, T.K. Flp-mediated recombination for use in hybrid plant production. Plant J. 2000, 23, 423–430. [Google Scholar] [CrossRef]
- Perez-Prat, E.; van Lookeren Campagne, M.M. Hybrid seed production and the challenge of propagating male-sterile plants. Trends Plant Sci. 2002, 7, 199–203. [Google Scholar] [CrossRef]
- Dunwell, J.; Ford, C. Technologies for Biological Containment of GM and non-GM Crops. DEFRA Contract CPEC 47; The Department for Environment, Food and Rural Affairs (DEFRA): London, UK, 2005. Available online: http://www.gmo-safety.eu/pdf/biosafenet/Defra_2005.pdf accessed on 6 June 2012.
- Dale, P.J. Spread of engineered genes to wild relatives. Plant Physiol. 1992, 100, 13–15. [Google Scholar] [CrossRef]
- Eastham, K.; Sweet, J. Genetically Modified Organisms (GMOs): The Significance of Gene Flow through Pollen Transfer; European Environment Agency: Copenhagen, Denmark, 2002; pp. 1–74. Available online: http://www.eea.europa.eu/publications/environmental_issue_report _2002_28 (accessed on 6 June 2012).
- Ellstrand, N.C.; Prentice, H.C.; Hancock, J.F. Gene flow and introgression from domesticated plants into their wild relatives. Ann. Rev. Ecol. Syst. 1999, 30, 539–563. [Google Scholar] [CrossRef]
- Rogers, H.J.; Parkes, H.C. Transgenic plants and the environment. J. Exp. Bot. 1995, 46, 467–488. [Google Scholar] [CrossRef]
- Bae, T.W.; Vanjildorj, E.; Song, S.Y.; Nishiguchi, S.; Yang, S.S.; Song, I.J.; Chandrasekhar, T.; Kang, T.W.; Kim, J.I.; Koh, Y.J.; et al. Environmental risk assessment of genetically engineered herbicide-tolerant Zoysia japonica. J. Environ. Qual. 2008, 37, 207–218. [Google Scholar]
© 2012 by the authors; licensee MDPI, Basel, Switzerland. This article is an open-access article distributed under the terms and conditions of the Creative Commons Attribution license (http://creativecommons.org/licenses/by/3.0/).
Share and Cite
Hague, J.P.; Dellaporta, S.L.; Moreno, M.A.; Longo, C.; Nelson, K.; Kausch, A.P. Pollen Sterility—A Promising Approach to Gene Confinement and Breeding for Genetically Modified Bioenergy Crops. Agriculture 2012, 2, 295-315. https://doi.org/10.3390/agriculture2040295
Hague JP, Dellaporta SL, Moreno MA, Longo C, Nelson K, Kausch AP. Pollen Sterility—A Promising Approach to Gene Confinement and Breeding for Genetically Modified Bioenergy Crops. Agriculture. 2012; 2(4):295-315. https://doi.org/10.3390/agriculture2040295
Chicago/Turabian StyleHague, Joel P., Stephen L. Dellaporta, Maria A. Moreno, Chip Longo, Kimberly Nelson, and Albert P. Kausch. 2012. "Pollen Sterility—A Promising Approach to Gene Confinement and Breeding for Genetically Modified Bioenergy Crops" Agriculture 2, no. 4: 295-315. https://doi.org/10.3390/agriculture2040295



