Genome-Wide Identification and Characterization of Trihelix Gene Family in Asian and African Vigna Species
Abstract
:1. Introduction
2. Materials and Methods
2.1. Genome-Wide Identification of Trihelix Genes in Vigna Species
2.2. Phylogenetic Analysis
2.3. Identification of Basic Features, Conserved Motifs and Gene Structure
2.4. Chromosomal Location and Evolutionary Selection Pressure
2.5. Expression Analysis of Trihelix Genes in Cowpea
3. Results
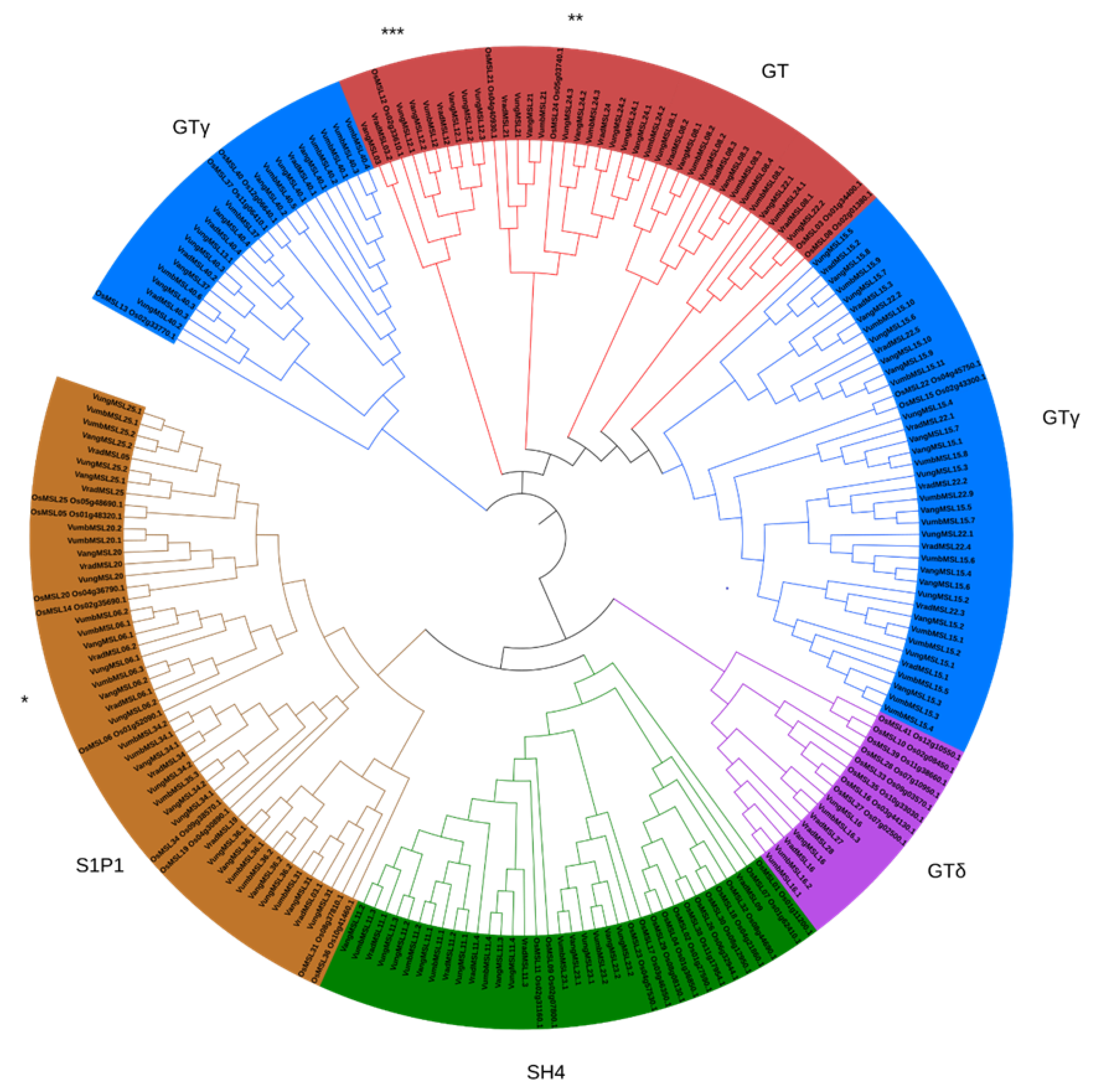
3.1. Genome-Wide Identification, Phylogenetic Analysis and Nomenclature
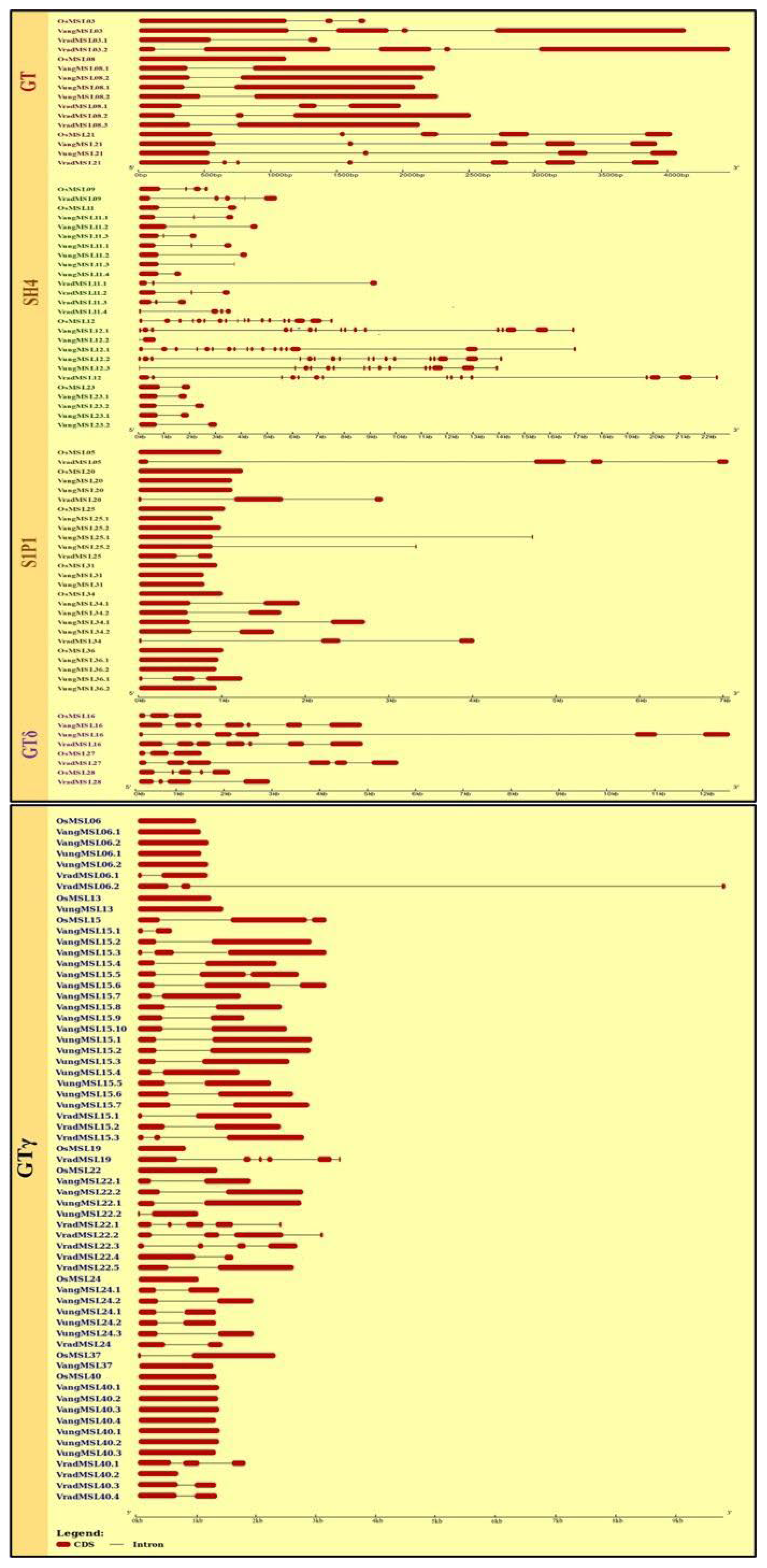
3.2. Identification of Basic Features, Conserved Motifs, Gene Structure and Selection Pressure
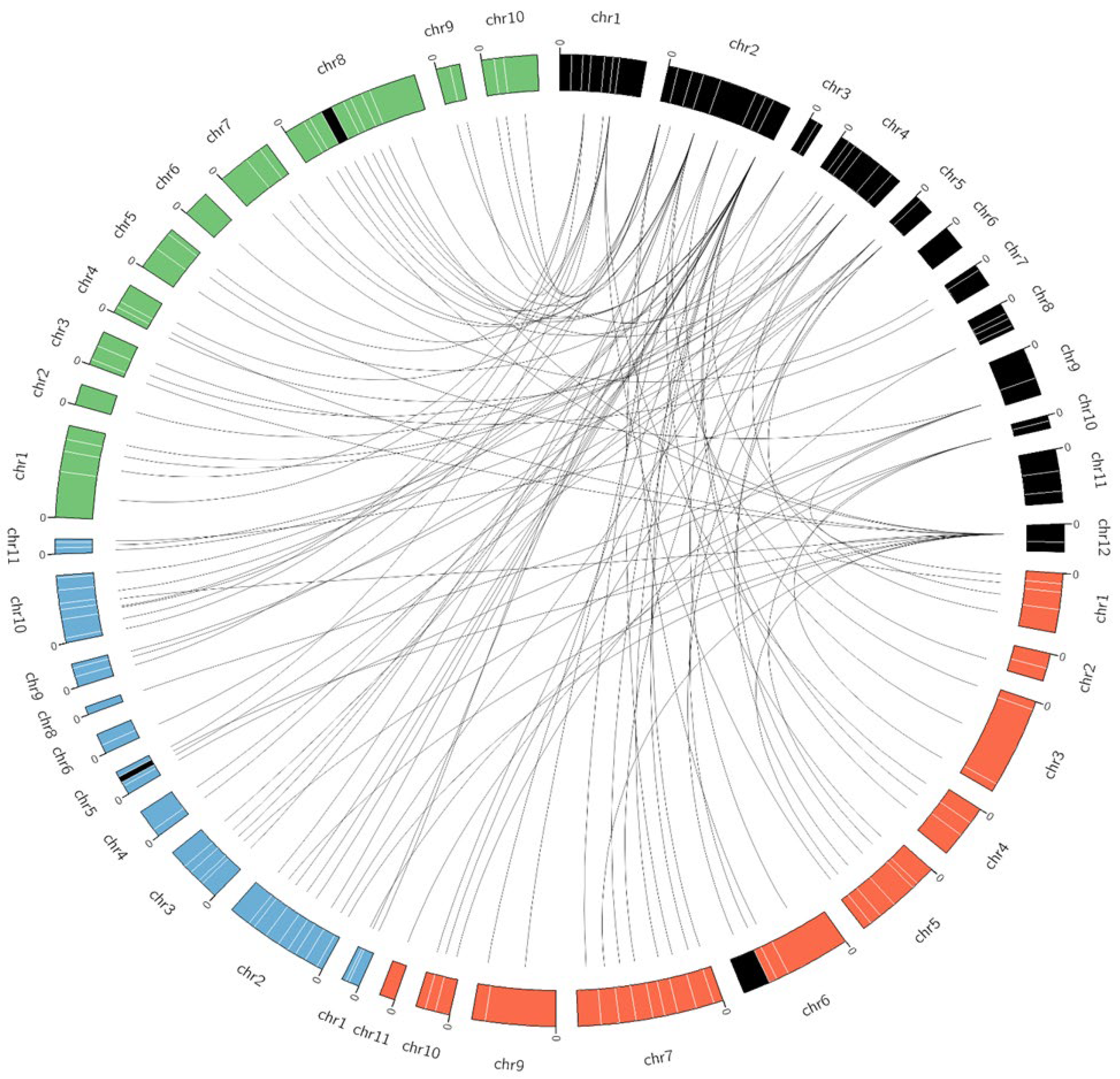
3.3. Chromosomal Distribution and Collinearity of Trihelix Genes among Vigna Species
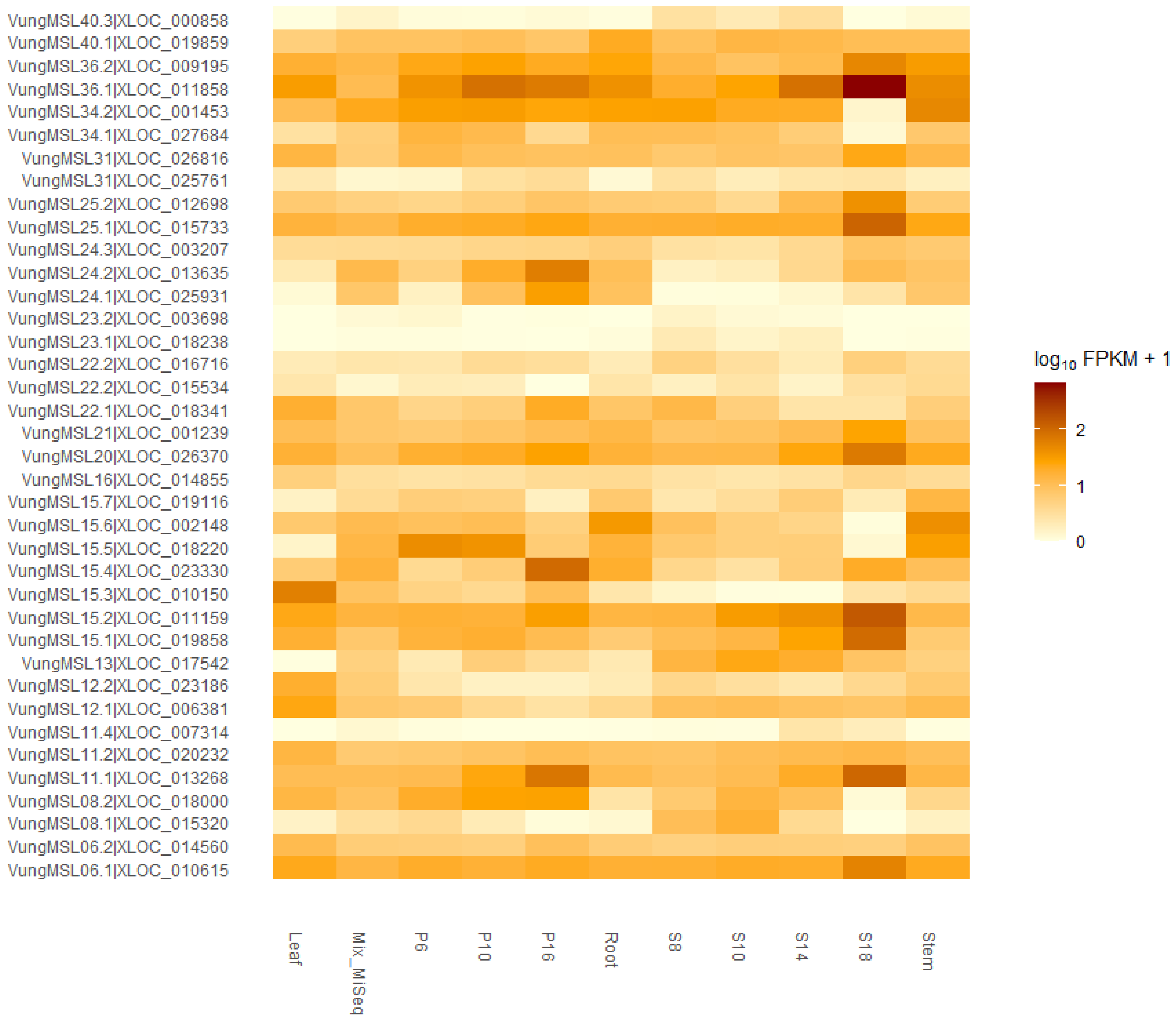
3.4. Expression Profile of Trihelix Genes in Different Tissue in Cowpea
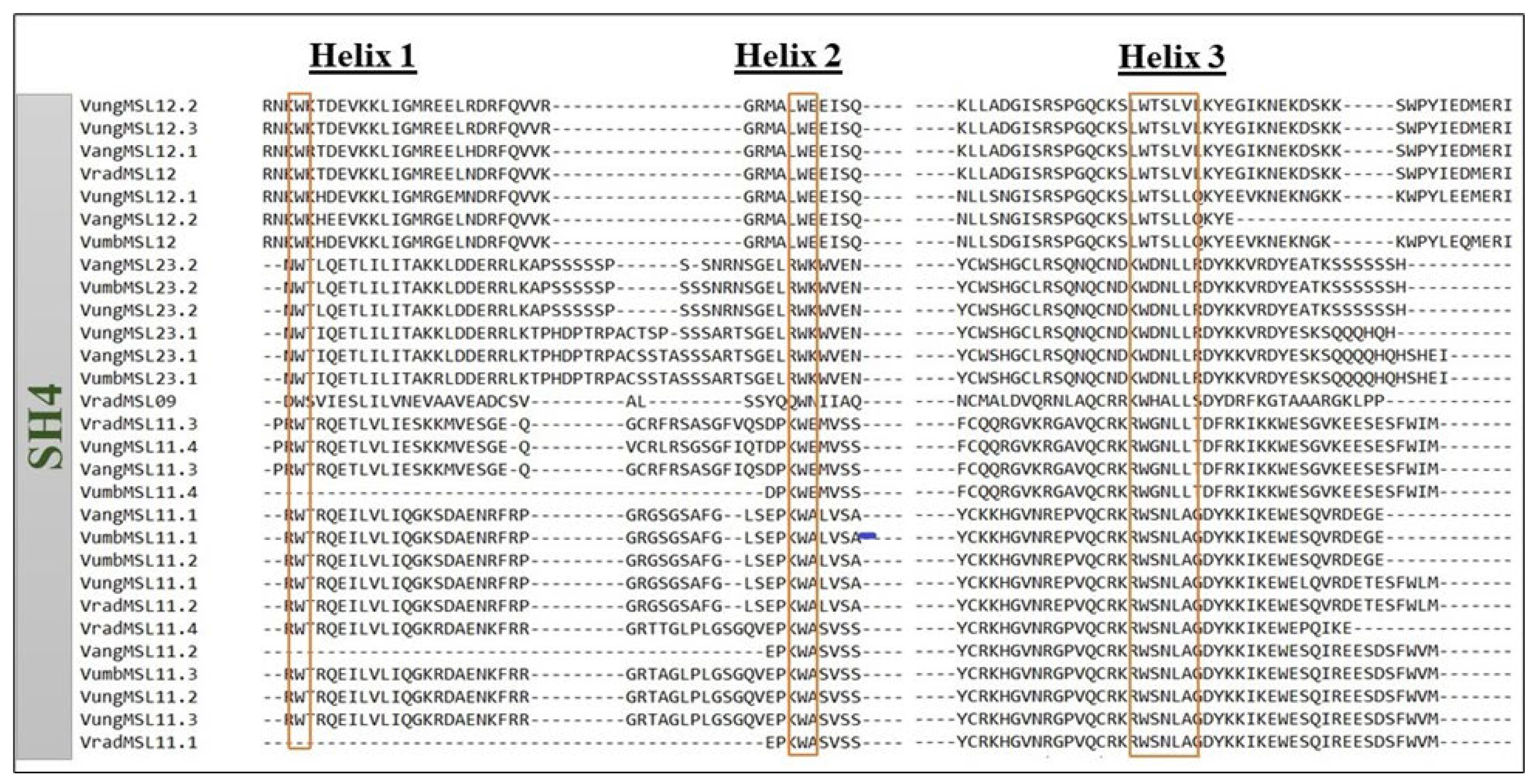
3.5. Analysis of Rice Shattering 1 (Sha1) Orthologs in Vigna Species
4. Discussion
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Data Availability Statement
Acknowledgments
Conflicts of Interest
Appendix A
References
- Ross-Ibarra, J.; Morrell, P.L.; Gaut, B.S. Plant domestication, a unique opportunity to identify the genetic basis of adaptation. Proc. Natl. Acad. Sci. USA 2007, 104 (Suppl. S1), 8641–8648. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Olsen, K.M.; Wendel, J.F. A bountiful harvest: Genomic insights into crop domestication phenotypes. Annu. Rev. Plant. Biol. 2013, 64, 47–70. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Dong, Y.; Wang, Y.Z. Seed shattering: From models to crops. Front. Plant. Sci. 2015, 6, 476. [Google Scholar] [CrossRef] [PubMed]
- Vittori, V.D.; Gioia, T.; Rodriguez, M.; Bellucci, E.; Bitocchi, E.; Nanni, L.; Attene, G.; Rau, D.; Papa, R. Convergent evolution of the seed shattering trait. Genes 2019, 10, 68. [Google Scholar] [CrossRef] [Green Version]
- Lin, Z.; Li, X.; Shannon, L.M.; Yeh, C.T.; Wang, M.L.; Bai, G.; Peng, Z.; Li, J.; Trick, H.N.; Clemente, T.E.; et al. Parallel domestication of the Shattering1 genes in cereals. Nat. Genet. 2012, 44, 720–724. [Google Scholar] [CrossRef] [Green Version]
- Li, C.; Zhou, A.; Sang, T. Genetic analysis of rice domestication syndrome with the wild annual species, Oryza nivara. New Phytol. 2006, 170, 185–193. [Google Scholar] [CrossRef]
- Li, C.; Zhou, A.; Sang, T. Rice domestication by reducing shattering. Science 2006, 311, 1936–1939. [Google Scholar] [CrossRef] [Green Version]
- Lin, Z.; Griffith, M.E.; Li, X.; Zhu, Z.; Tan, L.; Fu, Y.; Zhang, W.; Wang, X.; Xie, D.; Sun, C. Origin of seed shattering in rice (Oryza sativa L.). Planta 2007, 226, 11–20. [Google Scholar] [CrossRef]
- Zhou, D.X. Regulatory mechanism of plant gene transcription by GT-elements and GT-factors. Trends Plant. Sci. 1999, 4, 210–214. [Google Scholar] [CrossRef]
- Qin, Y.; Ma, X.; Yu, G.; Wang, Q.; Wang, L.; Kong, L.; Kim, W.; Wang, H.W. Evolutionary history of trihelix family and their functional diversification. DNA Res. 2014, 21, 499–510. [Google Scholar] [CrossRef]
- Nagano, Y. Several features of the GT-factor trihelix domain resemble those of the Myb DNA-binding domain. Plant Physiol. 2000, 124, 491–494. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Riechmann, J.L.; Heard, J.; Martin, G.; Reuber, L.; Jiang, C.; Keddie, J.; Adam, L.; Pineda, O.; Ratcliffe, O.J.; Samaha, R.R.; et al. Sequence-specific interactions of a pea nuclear factor with light-responsive elements upstream of the rbcS-3A gene. EMBO J. 1987, 6, 2543–2549. [Google Scholar]
- Kaplan-Levy, R.N.; Brewer, P.B.; Quon, T.; Smyth, D.R. The trihelix family of transcription factors-light, stress and development. Trends Plant. Sci. 2012, 17, 163–171. [Google Scholar] [CrossRef]
- Gilmartin, P.M.; Memelink, J.; Hiratsuka, K.; Kay, S.A.; Chua, N.H. Characterization of a gene encoding a DNA binding protein with specificity for a light-responsive element. Plant Cell. 1992, 4, 839–849. [Google Scholar] [PubMed] [Green Version]
- Kay, S.A.; Keith, B.; Shinozaki, K.; Chye, M.L.; Chua, N.H. The rice phytochrome gene: Structure, autoregulated expression, and binding of GT-1 to a conserved site in the 5’upstream region. Plant Cell. 1989, 1, 351–360. [Google Scholar] [PubMed]
- Dehesh, K.; Bruce, W.B.; Quail, P.H. A trans-acting factor that binds to a GT-motif in a phytochrome gene promoter. Science 1990, 250, 1397–1399. [Google Scholar] [CrossRef] [PubMed]
- Dehesh, K.; Hung, H.; Tepperman, J.M.; Quail, P.H. GT-2: A transcription factor with twin autonomous DNA-binding domains of closely related but different target sequence specificity. EMBO J. 1992, 11, 4131–4144. [Google Scholar] [CrossRef] [PubMed]
- Yu, C.; Cai, X.; Ye, Z.; Li, H. Genome-wide identification and expression profiling analysis of trihelix gene family in tomato. Biochem. Biophys. Res. Commun. 2015, 468, 653–659. [Google Scholar] [CrossRef]
- Yasmeen, E.; Riaz, M.; Sultan, S.; Azeem, F.; Abbas, A.; Riaz, K.; Ali, M.A. Genome-wide analysis of trihelix transcription factor gene family in Arabidopsis thaliana. Pak. J. Agric. Sci. 2016, 53. [Google Scholar]
- Song, A.; Wu, D.; Fan, Q.; Tian, C.; Chen, S.; Guan, Z.; Xin, J.; Zhao, K.; Chen, F. Transcriptome-wide identification and expression profiling analysis of chrysanthemum trihelix transcription factors. Int. J. Mol. Sci. 2016, 17, 198. [Google Scholar] [CrossRef] [Green Version]
- Li, J.; Zhang, M.; Sun, J.; Mao, X.; Wang, J.; Wang, J.; Liu, H.; Zheng, H.; Zhen, Z.; Zhao, H.; et al. Genome-Wide Characterization and Identification of Trihelix Transcription Factor and Expression Profiling in Response to Abiotic Stresses in Rice (Oryza sativa L.). Int. J. Mol. Sci. 2019, 20, 251. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Brewer, P.B.; Howles, P.A.; Dorian, K.; Griffith, M.E.; Ishida, T.; Kaplan-Levy, R.N.; Kilinc, A.; Smyth, D.R. PETAL LOSS, a trihelix transcription factor gene, regulates perianth architecture in the Arabidopsis flower. Development 2004, 131, 4035–4045. [Google Scholar] [CrossRef] [PubMed]
- Breuer, C.; Kawamura, A.; Ichikawa, T.; Tominaga-Wada, R.; Wada, T.; Kondou, Y.; Muto, S.; Matsui, M.; Sugimoto, K. The trihelix transcription factor GTL1 regulates ploidy-dependent cell growth in the Arabidopsis trichome. Plant Cell 2009, 21, 2307–2322. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Gao, M.J.; Lydiate, D.J.; Li, X.; Lui, H.; Gjetvaj, B.; Hegedus, D.D.; Rozwadowski, K. Repression of seed maturation genes by a trihelix transcriptional repressor in Arabidopsis seedlings. Plant Cell 2009, 21, 54–71. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Barr, M.S.; Willmann, M.R.; Jenik, P.D. Is there a role for trihelix transcription factors in embryo maturation? Plant Signal. Behav. 2012, 7, 205–209. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Pratap, A.; Malviya, N.; Tomar, R.; Gupta, D.S.; Kumar, J. Vigna. In Alien Gene Transfer in Crop Plants., 1st ed.; Pratap, A., Kumar, J., Eds.; Springer: New York, NY, USA, 2014; Volume 2, pp. 163–189. [Google Scholar]
- Kang, Y.J.; Satyawan, D.; Shim, S.; Lee, T.; Lee, J.; Hwang, W.J.; Lee, S.H. Draft genome sequence of adzuki bean, Vigna angularis. Sci. Rep. 2015, 5. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Kang, Y.J.; Kim, S.K.; Kim, M.Y.; Lestari, P.; Kim, K.H.; Ha, B.K.; Lee, S.H. Genome sequence of mung bean and insights into evolution within Vigna species. Nat. Commun. 2014, 5, 1–9. [Google Scholar] [CrossRef] [Green Version]
- Lonardi, S.; Muñoz-Amatriaín, M.; Liang, Q.; Shu, S.; Wanamaker, S.I.; Lo, S.; Close, T.J. The genome of cowpea (Vigna unguiculata [L.] Walp.). Plant J. 2019, 98, 767–782. [Google Scholar] [CrossRef] [Green Version]
- Kaul, T.; Eswaran, M.; Thangaraj, A.; Meyyazhagan, A.; Nehra, M.; Raman, N.M.; Balamurali, B. Rice Bean (Vigna umbellata) draft genome sequence: Unravelling the late flowering and unpalatability related genomic resources for efficient domestication of this underutilized crop. bioRxiv 2019, 816595. [Google Scholar] [CrossRef] [Green Version]
- Goodstein, D.M.; Shu, S.; Howson, R.; Neupane, R.; Hayes, R.D.; Fazo, J.; Mitros, T.; Dirks, W.; Hellsten, U.; Putnam, N.; et al. Phytozome: A comparative platform for green plant genomics. Nucleic Acids Res. 2011, 40, D1178–D1186. [Google Scholar] [CrossRef]
- Grabherr, M.G.; Haas, B.J.; Yassour, M.; Levin, J.Z.; Thompson, D.A.; Amit, I.; Adiconis, X.; Fan, L.; Raychowdhury, R.; Zeng, Q.; et al. Trinity: Reconstructing a full-length transcriptome without a genome from RNA-Seq data. Nat. Biotechnol. 2011, 29, 644–652. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Li, W.; Godzik, A. Cd-hit: A fast program for clustering and comparing large sets of protein or nucleotide sequences. J. Bioinform. 2006, 22, 1658–1659. [Google Scholar] [CrossRef] [PubMed]
- Cantarel, B.L.; Korf, I.; Robb, S.M.; Parra, G.; Ross, E.; Moore, B.; Yandell, M. MAKER: An easy-to-use annotation pipeline designed for emerging model organism genomes. Genome Res. 2008, 18, 188–196. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Stanke, M.; Tzvetkova, A.; Morgenstem, B. AUGUSTUS at EGASP: Using, EST, protein and genomic alignments for improved gene prediction in the human genome. Genome Biol. 2006, 7, S11. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Notredame, C.; Higgins, D.G.; Heringa, J. T-Coffee: A novel method for fast and accurate multiple sequence alignment. J. Mol. Biol. 2000, 302, 205–217. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Marchler-Bauer, A.; Derbyshire, M.K.; Gonzales, N.R.; Lu, S.; Chitsaz, F.; Geer, L.Y.; Geer, R.C.; He, J.; Gwadz, M.; Hurwitz, D.I.; et al. CDD: NCBI’s conserved domain database. Nucleic Acids Res. 2015, 43, D222–D226. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Finn, R.D.; Bateman, A.; Clements, J.; Coggill, P.; Eberhardt, R.Y. Pfam: The protein families database. Nucleic Acids Res. 2014, 42, D222–D230. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Kumar, S.; Nei, M.; Dudley, J.; Tamura, K. MEGA: A biologist-centric software for evolutionary analysis of DNA and protein sequences. Brief. Bioinform. 2008, 9, 299–306. [Google Scholar] [CrossRef] [Green Version]
- Gasteiger, E.; Gattiker, A.; Hoogland, C.; Ivanyi, I.; Appel, R.D.; Bairoch, A. ExPASy: The proteomics server for in-depth protein knowledge and analysis. Nucleic Acids Res. 2003, 31, 3784–3788. [Google Scholar] [CrossRef] [Green Version]
- Bailey, T.L.; Boden, M.; Buske, F.A.; Frith, M.; Grant, C.E.; Clementi, L. MEME SUITE: Tools for motif discovery and searching. Nucleic Acids Res. 2009, 37 (Suppl. S2), W202–W208. [Google Scholar] [CrossRef]
- Hu, B.; Jin, J.; Guo, A.Y.; Zhang, H.; Luo, J.; Gao, G. GSDS 2.0: An upgraded gene feature visualization server. Bioinformatics 2015, 31, 1296–1297. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Yu, C.S.; Cheng, C.W.; Su, W.C.; Chang, K.C.; Huang, S.W.; Hwang, J.K.; Lu, C.H. CELLO2GO: A web server for protein subCELlular LOcalization prediction with functional gene ontology annotation. PLoS ONE 2014, 9, e99368. [Google Scholar] [CrossRef] [PubMed]
- Voorrips, R.E. MapChart: Software for the graphical presentation of linkage maps and QTLs. J. Hered. 2002, 93, 77–78. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Krzywinski, M.; Schein, J.; Birol, I.; Connors, J.; Gascoyne, R. Circos: An information aesthetic for comparative genomics. Genome Res. 2009, 19, 1639–1645. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Suyama, M.; Torrents, D.; Bork, P. PAL2NAL: Robust conversion of protein sequence alignments into the corresponding codon alignments. Nucleic Acids Res. 2006, 34 (Suppl. 2), W609–W612. [Google Scholar] [CrossRef] [Green Version]
- Yao, S.; Jiang, C.; Huang, Z.; Torres-Jerez, I. The Vigna unguiculata Gene Expression Atlas (VuGEA) from de novo assembly and quantification of RNA-seq data provides insights into seed maturation mechanisms. Plant J. 2016, 88, 318–327. [Google Scholar] [CrossRef]
- Trapnell, C.; Roberts, A.; Goff, L.; Pertea, G. Differential gene and transcript expression analysis of RNA-seq experiments with TopHat and Cufflinks. Nat. Protoc. 2012, 7, 562–578. [Google Scholar] [CrossRef] [Green Version]
- Yang, J.; Zhang, Y. I-TASSER server: New development for protein structure and function predictions. Nucleic Acids Res. 2015, 43, W174–W181. [Google Scholar] [CrossRef] [Green Version]
- Abraham, M.J.; Murtola, T.; Schulz, R.; Páll, S.; Smith, J.C.; Hess, B.; Lindahl, E. GROMACS: High performance molecular simulations through multi-level parallelism from laptops to supercomputers. SoftwareX 2015, 1, 19–25. [Google Scholar] [CrossRef] [Green Version]
- Wang, W.; Wu, P.; Liu, T.; Ren, H.; Li, Y.; Hou, X. Genome-wide analysis and expression divergence of the trihelix family in Brassica rapa: Insight into the evolutionary patterns in plants. Sci. Rep. 2017, 7, 6463. [Google Scholar] [CrossRef] [Green Version]
- Ma, Z.; Liu, M.; Sun, W.; Huang, L.; Wu, Q.; Bu, T.; Chen, H. Genome-wide identification and expression analysis of the trihelix transcription factor family in tartary buckwheat (Fagopyrum tataricum). BMC Plant Biol. 2019, 19, 344. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Liu, W.; Zhang, Y.; Li, W.; Lin, Y.; Wang, C.; Xu, R.; Zhang, L. Genome-wide characterization and expression analysis of soybean trihelix gene family. PeerJ 2020, 8, 8753. [Google Scholar] [CrossRef] [PubMed]
- Xiao, J.; Hu, R.; Gu, T.; Han, J.; Qiu, D. Genome-wide identification and expression profiling of trihelix gene family under abiotic stresses in wheat. BMC Genom. 2019, 20, 287. [Google Scholar] [CrossRef] [PubMed]
- Hamel, L.P.; Nicole, M.C.; Sritubtim, S.; Morency, M.J.; Ellis, M.; Ehlting, J.; Beaudoin, N.; Barbazuk, B.; Klessig, D.; Lee, J.; et al. Ancient signals: Comparative genomics of plant MAPK and MAPKK gene families. Trends Plant Sci. 2006, 11, 192–198. [Google Scholar] [CrossRef] [PubMed]
- Hamel, L.P.; Sheen, J.; Seguin, A. Ancient signals: Comparative genomics of green plant CDPKs. Trends Plant Sci. 2014, 19, 79–89. [Google Scholar] [CrossRef] [Green Version]
- Wankhede, D.P.; Aravind, J.; Mishra, S.P. Identification of Genic SNPs from ESTs and Effect of Non-synonymous SNP on Proteins in Pigeonpea. Proc. Natl. Acad. Sci. USA 2019, 89, 595–603. [Google Scholar] [CrossRef]
- Fang, Y.; Xie, K.; Hou, X.; Hu, H.; Xiong, L. Systematic analysis of GT factor family of rice reveals a novel subfamily involved in stress responses. Mol. Genet. Genom. 2010, 283, 157–169. [Google Scholar] [CrossRef]
- Liu, X.; Zhang, H.; Ma, L.; Wang, Z.; Wang, K. Genome-Wide Identification and Expression Profiling Analysis of the Trihelix Gene Family Under Abiotic Stresses in Medicago truncatula. Genes 2020, 11, 1389. [Google Scholar] [CrossRef]
- Xin, K.; Pan, T.; Gao, S.; Yan, S.A. Transcription Factor Regulates Gene Expression in Chloroplasts. Int. J. Mol. Sci. 2021, 22, 6769. [Google Scholar] [CrossRef]
- Yadav, S.K.; Santosh Kumar, V.V.; Verma, R.K. Genome-wide identification and characterization of ABA receptor PYL gene family in rice. BMC Genom. 2020, 21, 676. [Google Scholar] [CrossRef]
- Rau, D.; Murgia, M.L.; Rodriguez, M.; Bitocchi, E.; Bellucci, E. Genomic dissection of pod shattering in common bean: Mutations at non-orthologous loci at the basis of convergent phenotypic evolution under domestication of leguminous species. Plant J. 2019, 97, 693–714. [Google Scholar] [CrossRef] [PubMed]
- Vittori, D.V.; Bitocchi, E.; Rodriguez, M.; Alseekh, S.; Bellucci, E. Pod indehiscence in common bean is associated to the fine regulation of PvMYB26 and a non-functional abscission layer. BioRxiv 2020, 021972. [Google Scholar] [CrossRef]
- Takahashi, Y.; Kongjaimun, A.; Muto, C.; Kobayashi, Y.; Kumagai, M. Same locus for non-shattering seed pod in two independently domesticated legumes, Vigna angularis and Vigna unguiculata. Front. Genet. 2020, 11, 748. [Google Scholar] [CrossRef] [PubMed]
- Isemura, T.; Kaga, A.; Konishi, S.; Ando, T.; Tomooka, N.; Han, O.K.; Vaughan, D.A. Genome dissection of traits related to domestication in azuki bean (Vigna angularis) and comparison with other warm-season legumes. Ann. Bot. 2007, 100, 1053–1071. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Vairam, N.; Lavanya, S.A.; Vanniarajan, C. Screening for pod shattering in mutant population of mung bean (Vigna radiata (L.) Wilczek). J. Appl. Nat. Sci. 2017, 9, 1787–1791. [Google Scholar] [CrossRef] [Green Version]
- Wankhede, D.P.; Kumari, M.; Richa, T.; Aravind, J.; Rajkumar, S. Genome wide identification and characterization of Calcium Dependent Protein Kinase gene family in Cajanus cajan. J. Environ. Biol. 2017, 38, 167. [Google Scholar] [CrossRef]
- Osterberg, J.T.; Xiang, W.; Olsen, L.I.; Edenbrandt, A.K.; Vedel, S.E. Accelerating the domestication of new crops: Feasibility and approaches. Trends Plant Sci. 2017, 22, 373–384. [Google Scholar] [CrossRef]
- Yu, H.; Li, J. Breeding future crops to feed the world through de novo domestication. Nat Commun. 2022, 13, 1171. [Google Scholar] [CrossRef]
- Zsogon, A.; Cermak, T.; Voytas, D.; Peres, L.E. Genome editing as a tool to achieve the crop ideotype and de novo domestication of wild relatives: Case study in tomato. Plant Sci. 2017, 256, 120–130. [Google Scholar] [CrossRef]
- Pisias, M.T.; Bakala, H.S.; McAlvay, A.C.; Mabry, M.E.; Birchler, J.A.; Yang, B.; Pires, J.C. Prospects of Feral Crop De Novo Re-Domestication. Plant Cell Physiol. 2022, 63, 1641–1653. [Google Scholar] [CrossRef]




| Subgroup | Assigned Name | Locus ID | Exon (No.) | Amino Acid (No.) | a MW (kDa) | b PI | GC% | c Loc. | Rice Orthologs |
|---|---|---|---|---|---|---|---|---|---|
| GT | VungMSL08.1 | Vigun06g111200 | 2 | 569 | 64.21 | 6.14 | 42.6 | N | OsMSL08 (Os02g01380.1) |
| VungMSL08.2 | Vigun07g154500 | 2 | 617 | 70.95 | 6.78 | 40.9 | N | ||
| VungMSL21 | Vigun01g244400 | 4 | 377 | 43.14 | 5.83 | 37.6 | N, C | OsMSL21 (Os04g40930.1) | |
| SH4 | VungMSL11.1 | Vigun05g020800 | 3 | 326 | 36.07 | 5.23 | 38 | N | OsMSL11 (Os02g31160.1) |
| VungMSL11.2 | Vigun07g298000 | 2 | 349 | 38.86 | 6.5 | 37.4 | N | ||
| VungMSL11.3 | Vigun07g298000 | 2 | 260 | 29.00 | 6.03 | 37.4 | N | ||
| VungMSL11.4 | Vigun03g052500 | 2 | 339 | 38.02 | 6.87 | 41.9 | N | ||
| VungMSL12.1 | Vigun03g314600 | 17 | 883 | 98.29 | 8.76 | 33.7 | Ch | OsMSL12 (Os02g33610.1) | |
| VungMSL12.2 | Vigun09g101800 | 17 | 723 | 79.93 | 8.64 | 35.5 | M | ||
| VungMSL12.3 | Vigun09g101800 | 15 | 723 | 79.93 | 8.64 | 35.5 | M | ||
| VungMSL23.1 | Vigun07g196800 | 2 | 346 | 39.75 | 9.03 | 43.6 | N | OsMSL23 (Os04g57530.1) | |
| VungMSL23.2 | Vigun02g002400 | 2 | 344 | 39.14 | 7.73 | 32.8 | N | ||
| S1P1 | VungMSL20 | Vigun10g160800 | 1 | 376 | 40.79 | 9.73 | 47.4 | N | OsMSL20 (Os04g36790.1) |
| VungMSL25.1 | Vigun06g191000 | 2 | 301 | 33.97 | 5.15 | 39 | N | OsMSL25 (Os05g48690.1) | |
| VungMSL25.2 | Vigun05g208200 | 2 | 300 | 33.84 | 9.76 | 35.7 | N | ||
| VungMSL31 | Vigun10g048800 | 1 | 262 | 30.16 | 6.68 | 40.7 | N | OsMSL31 (Os08g37810.1) | |
| VungMSL34.1 | Vigun11g011300 | 2 | 339 | 38.16 | 9.25 | 30.5 | N | OsMSL34 (Os09g38570.1) | |
| VungMSL34.2 | Vigun01g027600 | 2 | 349 | 38.86 | 8.9 | 38.2 | N | ||
| VungMSL36.1 | Vigun05g050600 | 3 | 245 | 27.32 | 10.47 | 61.3 | N, M | OsMSL36 (Os10g41460.1) | |
| VungMSL36.2 | Vigun03g429900 | 1 | 310 | 34.73 | 9.34 | 49.3 | N | ||
| GTδ | VungMSL16 | Vigun06g021100 | 5 | 496 | 57.10 | 7.99 | 35.6 | N, C | OsMSL16 (Os03g44130.1) |
| GTγ | VungMSL06.1 | Vigun04g017300 | 1 | 352 | 40.73 | 4.62 | 39.5 | N | OsMSL06 (Os01g52090.1) |
| VungMSL06.2 | Vigun05g286700 | 1 | 390 | 44.55 | 4.7 | 39.3 | N | ||
| VungMSL13 | Vigun07g065400 | 1 | 474 | 54.34 | 6.12 | 44 | N | OsMSL13 (Os02g33770.1) | |
| VungMSL15.1 | Vigun07g217300 | 2 | 655 | 72.32 | 5.94 | 44.7 | N | OsMSL15 (Os02g43300.1) | |
| VungMSL15.2 | Vigun04g131900 | 2 | 661 | 73.01 | 5.3 | 41.5 | N | ||
| VungMSL15.3 | Vigun04g131800 | 2 | 584 | 66.44 | 5.61 | 38.4 | N | ||
| VungMSL15.4 | Vigun09g125900 | 2 | 503 | 56.99 | 6.44 | 43.6 | N | ||
| VungMSL15.5 | Vigun07g193200 | 2 | 519 | 59.5 | 5.76 | 41.3 | N | ||
| VungMSL15.6 | Vigun01g170400 | 2 | 586 | 67.6 | 6.38 | 41.4 | N | ||
| VungMSL15.7 | Vigun07g067700 | 2 | 602 | 69.77 | 6.32 | 41.7 | N | ||
| VungMSL22.1 | Vigun07g217100 | 2 | 815 | 92.69 | 5.24 | 41.3 | N | OsMSL22 (Os04g45750.1) | |
| VungMSL22.2 | Vigun06g151000 | 2 | 268 | 31.34 | 7.63 | 35.5 | N | ||
| VungMSL24.1 | Vigun10g080500 | 2 | 274 | 32.528 | 6.65 | 38.5 | N | OsMSL24 (Os05g03740.1 | |
| VungMSL24.2 | Vigun05g100400 | 2 | 289 | 34.71 | 6.99 | 32.9 | N | ||
| VungMSL24.3 | Vigun02g104900 | 2 | 309 | 36.26 | 7.74 | 39 | N | ||
| VungMSL40.1 | Vigun07g217400 | 1 | 450 | 51.34 | 6.2 | 38.8 | N | OsMSL40 (Os12g06640.1) | |
| VungMSL40.2 | Vigun05g263000 | 1 | 447 | 50.73 | 5.78 | 39.2 | N | ||
| VungMSL40.3 | Vigun01g168100 | 1 | 429 | 49.48 | 6.22 | 44.7 | N |
| Sub-group | Assigned Name | Locus ID | Exon (No.) | Amino Acid (No.) | a MW (kDa) | b PI | GC% | c Loc. | Rice Orthologs |
|---|---|---|---|---|---|---|---|---|---|
| GT | VradMSL03.1 | Vradi09g07980.1 | 2 | 203 | 23.45 | 7.69 | 42.5 | N | OsMSL03 (Os01g34400.1) |
| VradMSL03.2 | Vradi01g07000.1 | 5 | 987 | 110.71 | 7.1 | 45 | N | ||
| VradMSL08.1 | Vradi10g07270.1 | 3 | 282 | 32.00 | 7.77 | 41.8 | N | OsMSL08 (Os02g01380.1) | |
| VradMSL08.2 | Vradi10g05520.1 | 3 | 556 | 62.56 | 8.18 | 43.9 | N | ||
| VradMSL08.3 | Vradi08g13180.1 | 2 | 590 | 67.75 | 7 | 42.9 | N | ||
| VradMSL21 | Vradi02g12120.1 | 7 | 396 | 45.15 | 5.9 | 49.37 | N, C | OsMSL21 (Os04g40930.1) | |
| SH4 | VradMSL09 | Vradi06g09390.1 | 5 | 451 | 51.29 | 5.2 | 37.8 | N | OsMSL09 (Os02g07800.1) |
| VradMSL11.1 | Vradi08g23420.1 | 3 | 231 | 26.23 | 9 | 35.4 | N | OsMSL11 (Os02g31160.1) | |
| VradMSL11.2 | Vradi04g10090.1 | 3 | 326 | 35.99 | 5.24 | 38.4 | N | ||
| VradMSL11.3 | Vradi07g29850.1 | 3 | 300 | 34.51 | 9.3 | 41.4 | N | ||
| VradMSL11.4 | Vradi07g01760.1 | 4 | 227 | 26.20 | 9.2 | 30.9 | N | ||
| VradMSL12 | Vradi0043s00060 | 15 | 873 | 96.84 | 8.5 | 35.6 | M | OsMSL12 (Os02g33610.1) | |
| S1P1 | VradMSL05 | Vradi10g10130.1 | 4 | 258 | 29.10 | 10.06 | 33.1 | N, M | OsMSL05 (Os01g48320.1) |
| VradMSL20 | Vradi09g02750.1 | 3 | 239 | 26.12 | 9.98 | 43.7 | N | OsMSL20 (Os04g36790.1) | |
| VradMSL25 | Vradi0023s00720.1 | 2 | 212 | 24.22 | 9.85 | 59.17 | N | OsMSL25 (Os05g48690.1) | |
| VradMSL34 | Vradi05g17220.1 | 3 | 149 | 16.78 | 9.86 | 33.2 | N | OsMSL34 (Os09g38570.1) | |
| GTδ | VradMSL16 | Vradi05g11080.1 | 7 | 877 | 101.45 | 5.4 | 39.48 | N | OsMSL16 (Os03g44130.1) |
| VradMSL27 | Vradi07g00070.1 | 6 | 762 | 87.92 | 6.25 | 40.37 | N, C | OsMSL27 (Os07g02500.1) | |
| VradMSL28 | Vradi03g06280.1 | 4 | 482 | 55.85 | 6.49 | 39.61 | N | OsMSL28 (Os07g10950.1) | |
| GTγ | VradMSL06.1 | Vradi05g20210.1 | 2 | 278 | 31.72 | 5.36 | 41.34 | N | OsMSL06 (Os01g52090.1) |
| VradMSL06.2 | Vradi01g04230.1 | 3 | 242 | 27.80 | 4.8 | 51.17 | N, C | ||
| VradMSL15.1 | Vradi08g18150.1 | 2 | 445 | 49.91 | 9.81 | 46.52 | N | OsMSL15 (Os02g43300.1) | |
| VradMSL15.2 | Vradi08g16250.1 | 2 | 518 | 59.58 | 5.78 | 45.54 | N | ||
| VradMSL15.3 | Vradi08g06550.1 | 3 | 497 | 58138 | 6.18 | 43.91 | N | ||
| VradMSL19 | Vradi08g05950.1 | 6 | 485 | 53.11 | 5.87 | 48.9 | N, Ch | OsMSL19 (Os04g30890.1) | |
| VradMSL22.1 | Vradi0160s00310.1 | 5 | 307 | 35.26 | 9.17 | 49.78 | N | OsMSL22 (Os04g45750.1) | |
| VradMSL22.2 | Vradi01g12370.1 | 4 | 446 | 51.30 | 9.3 | 44.82 | N | ||
| VradMSL22.3 | Vradi01g12360.1 | 4 | 277 | 32147 | 4.93 | 50.84 | N, C | ||
| VradMSL22.4 | Vradi08g18140.1 | 2 | 367 | 41.80 | 9.44 | 45.56 | N | ||
| VradMSL22.5 | Vradi03g04320.1 | 2 | 590 | 67.77 | 6.29 | 43.36 | N | ||
| VradMSL24 | Vradi04g00170.1 | 2 | 233 | 28.78 | 9.57 | 46.87 | N, M | OsMSL24 (Os05g03740.1) | |
| VradMSL40.1 | Vradi08g18160.1 | 3 | 351 | 40.07 | 5.49 | 42.05 | N | OsMSL40 (Os12g06640.1) | |
| VradMSL40.2 | Vradi03g04100.1 | 1 | 225 | 25.54 | 8.89 | 46.9 | N | ||
| VradMSL40.3 | Vradi04g01660.1 | 2 | 341 | 38.41 | 8.93 | 43.47 | N | ||
| VradMSL40.4 | Vradi08g05970.1 | 2 | 341 | 39.88 | 9.59 | 46.2 | N |
| Subgroup | Assigned Name | Locus ID | Exon (No.) | Amino Acid (No.) | a MW (kDa) | b PI | GC (%) | c Loc. | Rice Orthologs |
|---|---|---|---|---|---|---|---|---|---|
| GT | VangMSL03 | Vigan10g080900 | 4 | 1005 | 112.7 | 8 | 51.0 | N | OsMSL03 (Os01g34400.1) |
| VangMSL08.1 | Vigan09g161100 | 2 | 582 | 65.57 | 6.1 | 45.7 | N | OsMSL08 (Os02g01380.1) | |
| VangMSL08.2 | Vigan10g097100 | 2 | 588 | 67.28 | 6.5 | 46.5 | N | ||
| VangMSL21 | Vigan03g313100 | 5 | 392 | 44.86 | 5.9 | 49.1 | N, C | OsMSL21 (Os04g40930.1) | |
| SH4 | VangMSL11.1 | Vigan03g008100 | 3 | 322 | 35.66 | 5.4 | 54.4 | N | OsMSL11 (Os02g31160.1) |
| VangMSL11.2 | Vigan02g258700 | 2 | 357 | 39.58 | 6.1 | 39.3 | N | ||
| VangMSL11.3 | Vigan01g003800 | 3 | 359 | 40.52 | 5.6 | 48.3 | N | ||
| VangMSL12.1 | Vigan1112s000300 | 16 | 849 | 94.21 | 8.9 | 42.2 | M | OsMSL12 (Os02g33610.1) | |
| VangMSL12.2 | Vigan01g226300 | 2 | 167 | 18.56 | 5.3 | 38.8 | N | ||
| VangMSL23.1 | Vigan02g183900 | 2 | 346 | 39.30 | 9.2 | 52.8 | N | OsMSL23 (Os04g57530.1) | |
| VangMSL23.2 | Vigan06g146100 | 2 | 344 | 39.14 | 8.6 | 49.2 | N | ||
| S1P1 | VangMSL20 | Vigan11g006000 | 1 | 374 | 40.64 | 9.7 | 56.8 | N | OsMSL20 (Os04g36790.1) |
| VangMSL25.1 | Vigan05g139300 | 1 | 582 | 65.57 | 6.1 | 55.7 | N | OsMSL25 (Os05g48690.1) | |
| VangMSL25.2 | Vigan09g234100 | 1 | 330 | 37.53 | 5.1 | 55.7 | N | ||
| VangMSL31 | Vigan11g133500 | 1 | 261 | 29.99 | 6.3 | 57.6 | N | OsMSL31 (Os08g37810.1) | |
| VangMSL34.1 | Vigan05g004000 | 2 | 349 | 38.83 | 9.2 | 45.9 | N | OsMSL34 (Os09g38570.1) | |
| VangMSL34.2 | Vigan08g003100 | 2 | 325 | 36.90 | 8.9 | 47.6 | N | ||
| VangMSL36.1 | Vigan03g034300 | 1 | 317 | 35.20 | 9.5 | 61.6 | N | OsMSL36 (Os10g41460.1) | |
| VangMSL36.2 | Vigan01g314100 | 1 | 309 | 34.65 | 9.3 | 56.2 | N | ||
| GTδ | VangMSL16 | Vigan04g227300 | 7 | 829 | 96.05 | 5.3 | 39.7 | N | OsMSL16 (Os03g44130.1) |
| GTγ | VangMSL06.1 | Vigan10g017200 | 1 | 349 | 40.51 | 4.6 | 52.5 | N | OsMSL06 (Os01g52090.1) |
| VangMSL06.2 | Vigan05g214400 | 1 | 387 | 44.21 | 4.7 | 41.8 | N | ||
| VangMSL15.1 | Vigan10g174500 | 2 | 120 | 13.59 | 8.7 | 44.9 | N | OsMSL15 (Os02g43300.1) | |
| VangMSL15.2 | Vigan10g132200 | 2 | 657 | 72.52 | 5.3 | 50.7 | N | ||
| VangMSL15.3 | Vigan02g203500 | 3 | 682 | 75.06 | 6.0 | 50.4 | N | ||
| VangMSL15.4 | Vigan02g203300 | 2 | 489 | 54.30 | 9 | 45.9 | N | ||
| VangMSL15.5 | Vigan10g132400 | 3 | 625 | 71.03 | 5.5 | 45.2 | N | ||
| VangMSL15.6 | Vigan02g203400 | 3 | 605 | 68.21 | 5.6 | 46 | N | ||
| VangMSL15.7 | Vigan04g134500 | 2 | 514 | 57.98 | 6.6 | 49.6 | N | ||
| VangMSL15.8 | Vigan02g180100 | 2 | 516 | 59.46 | 5.7 | 45.3 | N | ||
| VangMSL15.9 | Vigan03g238700 | 2 | 325 | 36.68 | 8.4 | 48.5 | N | ||
| VangMSL15.10 | Vigan03g238600 | 2 | 557 | 63.51 | 6.3 | 46.8 | N | ||
| VangMSL22.1 | Vigan09g201300 | 2 | 329 | 37.71 | 6.3 | 47.2 | N | OsMSL22 (Os04g45750.1) | |
| VangMSL22.2 | Vigan02g080400 | 2 | 553 | 63.51 | 5.8 | 45.8 | N | ||
| VangMSL24.1 | Vigan11g113800 | 2 | 270 | 32.15 | 6.6 | 47.3 | N | OsMSL24 (Os05g03740.1) | |
| VangMSL24.2 | Vigan06g074900 | 2 | 309 | 36.33 | 8.3 | 49.3 | N | ||
| VangMSL37 | Vigan641s002000 | 1 | 410 | 47.62 | 6.4 | 46.5 | N | OsMSL37 (Os11g06410.1) | |
| VangMSL40.1 | Vigan02g203600 | 1 | 448 | 51.27 | 6.2 | 42.2 | N | OsMSL40 (Os12g06640.1) | |
| VangMSL40.2 | Vigan10g131700 | 1 | 442 | 50.57 | 6.2 | 42.5 | N | ||
| VangMSL40.3 | Vigan05g194200 | 1 | 448 | 51.02 | 5.9 | 42.6 | N | ||
| VangMSL40.4 | Vigan02g076800 | 1 | 430 | 49.57 | 6.1 | 46.9 | N |
| Subgroup | Assigned Name | Locus ID | Length | a MW | b PI | GC% | c Loc. | Rice Orthologs |
|---|---|---|---|---|---|---|---|---|
| GT | VumbMSL08.1 | Gene_28670 | 424 | 49.13 | 7.24 | 46.4 | N | OsMSL08 (Os02g01380.1) |
| VumbMSL08.2 | Gene_13448 | 582 | 65.56 | 6.18 | 45.7 | N | ||
| VumbMSL08.3 | Gene_13439 | 588 | 67.37 | 6.58 | 46.3 | N | ||
| VumbMSL08.4 | Gene_13440 | 459 | 52.32 | 7.77 | 45.6 | N | ||
| VumbMSL21 | Gene_22669 | 377 | 43.15 | 5.83 | 49.4 | N, C | OsMSL21 (Os04g40930.1) | |
| SH4 | VumbMSL11.1 | Gene_1713 | 210 | 23.08 | 5.40 | 60.5 | N | OsMSL11 (Os02g31160.1) |
| VumbMSL11.2 | Gene_1714 | 322 | 35.66 | 5.42 | 54.6 | N | ||
| VumbMSL11.3 | Gene_5885 | 363 | 40.07 | 6.10 | 50.4 | N | ||
| VumbMSL11.4 | Gene_30530 | 285 | 32.25 | 5.45 | 48 | N | ||
| VumbMSL12 | Gene_21287 | 867 | 96.37 | 8.76 | 41.8 | Ch | OsMSL12 (Os02g33610.1) | |
| VumbMSL23.1 | Gene_32537 | 340 | 38.61 | 9.03 | 53.2 | N | OsMSL23 (Os04g57530.1) | |
| VumbMSL23.2 | Gene_29588 | 340 | 38.77 | 8.85 | 49.5 | N | ||
| S1P1 | VumbMSL20.1 | Gene_23931 | 374 | 40.64 | 9.73 | 56.8 | N | OsMSL20 (Os04g36790.1) |
| VumbMSL20.2 | Gene_23932 | 374 | 40.64 | 9.73 | 56.8 | N | ||
| VumbMSL25.1 | Gene_14426 | 301 | 33.97 | 5.15 | 57.6 | N | OsMSL25 (Os05g48690.1) | |
| VumbMSL25.2 | Gene_14425 | 330 | 37.59 | 5.17 | 55.6 | N | ||
| VumbMSL31 | Gene_2041 | 261 | 30.14 | 6.62 | 57.5 | N | OsMSL31 (Os08g37810.1) | |
| VumbMSL34.1 | Gene_18487 | 206 | 22.51 | 9.16 | 49.5 | N | OsMSL34 (Os09g38570.1) | |
| VumbMSL34.2 | Gene_18488 | 349 | 38.83 | 9.15 | 46.4 | N | ||
| VumbMSL34.3 | Gene_18489 | 333 | 37.76 | 9.02 | 47.5 | N | ||
| VumbMSL36.1 | Gene_3480 | 317 | 35.20 | 9.55 | 61.6 | N | OsMSL36 (Os10g41460.1) | |
| VumbMSL36.2 | Gene_3479 | 309 | 34.68 | 9.34 | 56.6 | N | ||
| GTδ | VumbMSL16.1 | Gene_26106 | 487 | 56.85 | 6.63 | 40.4 | N | OsMSL16 (Os03g44130.1) |
| VumbMSL16.2 | Gene_26107 | 469 | 548.75 | 6.97 | 40.3 | N | ||
| VumbMSL16.3 | Gene_28699 | 505 | 58.67 | 9.26 | 39.2 | C | ||
| GTγ | VumbMSL06.1 | Gene_25832 | 349 | 40.50 | 4.69 | 52.4 | N | OsMSL06 (Os01g52090.1) |
| VumbMSL06.2 | Gene_25833 | 349 | 40.50 | 4.69 | 52.4 | N | ||
| VumbMSL06.3 | Gene_9047 | 387 | 44.21 | 4.68 | 41.7 | N | ||
| VumbMSL15.1 | Gene_16032 | 555 | 62.55 | 5.61 | 49.5 | N | OsMSL15 (Os02g43300.1) | |
| VumbMSL15.2 | Gene_16033 | 657 | 72.55 | 5.35 | 50.8 | N | ||
| VumbMSL15.3 | Gene_16028 | 549 | 61.88 | 6.25 | 48.6 | N | ||
| VumbMSL15.4 | Gene_16030 | 652 | 72.09 | 5.82 | 50.3 | N | ||
| VumbMSL15.5 | Gene_16029 | 130 | 15.13 | 4.43 | 52.7 | N, C | ||
| VumbMSL15.6 | Gene_16031 | 630 | 71.07 | 5.68 | 46.5 | N | ||
| VumbMSL15.7 | Gene_16026 | 601 | 68.34 | 5.69 | 45.5 | N | ||
| VumbMSL15.8 | Gene_24101 | 632 | 71.53 | 5.96 | 50 | N | ||
| VumbMSL15.9 | Gene_195 | 518 | 59.55 | 5.80 | 45.5 | N | ||
| VumbMSL15.10 | Gene_18868 | 607 | 70.29 | 6.33 | 46.5 | N | ||
| VumbMSL15.11 | Gene_18869 | 589 | 67.60 | 6.22 | 46.6 | N | ||
| VumbMSL22 | Gene_16025 | 123 | 14.53 | 4.26 | 42.9 | N | OsMSL22 (Os04g45750.1) | |
| VumbMSL24.1 | Gene_28671 | 256 | 29.88 | 7.66 | 42.4 | N | OsMSL24 (Os05g03740.1) | |
| VumbMSL24.2 | Gene_18886 | 269 | 31.96 | 6.39 | 47.4 | N | ||
| VumbMSL24.3 | Gene_10842 | 309 | 36.33 | 8.36 | 49.4 | N | ||
| VumbMSL37 | Gene_5137 | 149 | 17.21 | 9.55 | 45.9 | N | OsMSL37 (Os11g06410.1) | |
| VumbMSL40.1 | Gene_11435 | 448 | 51.27 | 6.20 | 42.3 | N | OsMSL40 (Os12g06640.1) | |
| VumbMSL40.2 | Gene_11436 | 448 | 51.27 | 6.20 | 42.3 | N | ||
| VumbMSL40.3 | Gene_11437 | 448 | 51.27 | 6.20 | 42.3 | N | ||
| VumbMSL40.4 | Gene_11438 | 448 | 51.27 | 6.20 | 42.3 | N | ||
| VumbMSL40.5 | Gene_11439 | 442 | 50.58 | 6.17 | 42.7 | N | ||
| VumbMSL40.6 | Gene_28728 | 448 | 50.99 | 5.94 | 42.5 | N |
| Candidate Protein | Minimum Potential Energy | Mutation Variants of Candidate Protein | Minimum Potential Energy | Effect of Mutation |
|---|---|---|---|---|
| VungMSL23.1 | −1,652,396.5 | VungMSL23.1_K70N | −1,878,625.2 | Stabilizing |
| VungMSL23.2 | −1,605,236.2 | VungMSL23.2_K39N | −1,676,144 | Stabilizing |
| VangMSL23.1 | −1,678,593.1 | VangMSL23.1_K69N | −1,734,201.9 | Stabilizing |
| VangMSL23.2 | −1,567,263.5 | VangMSL23.2_ K39N | −1,546,699.9 | Destabilizing |
| VumbMSL23.1 | −1,555,491.4 | VumbMSL23.1_K68N | −1,579,698.2 | Stabilizing |
| VumbMSL23.2 | −1,669,826.2 | VumbMSL23.2_K39N | −1,653,073.4 | Destabilizing |
| VradMSL12 | −3,030,210 | VradMSL12 _R794N | −3,738,055.5 | Stabilizing |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2022 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Kumari, S.; Wankhede, D.P.; Murmu, S.; Maurya, R.; Jaiswal, S.; Rai, A.; Archak, S. Genome-Wide Identification and Characterization of Trihelix Gene Family in Asian and African Vigna Species. Agriculture 2022, 12, 2172. https://doi.org/10.3390/agriculture12122172
Kumari S, Wankhede DP, Murmu S, Maurya R, Jaiswal S, Rai A, Archak S. Genome-Wide Identification and Characterization of Trihelix Gene Family in Asian and African Vigna Species. Agriculture. 2022; 12(12):2172. https://doi.org/10.3390/agriculture12122172
Chicago/Turabian StyleKumari, Shweta, Dhammaprakash Pandhari Wankhede, Sneha Murmu, Ranjeet Maurya, Sarika Jaiswal, Anil Rai, and Sunil Archak. 2022. "Genome-Wide Identification and Characterization of Trihelix Gene Family in Asian and African Vigna Species" Agriculture 12, no. 12: 2172. https://doi.org/10.3390/agriculture12122172






