Abundant Genetic Diversity of Yunling Cattle Based on Mitochondrial Genome
Abstract
:Simple Summary
Abstract
1. Introduction
2. Materials and Methods
2.1. Animal Sampling
2.2. Illumina Sequencing and Reconstruction of Mitochondrial Genomes
2.3. Data Analysis
3. Results
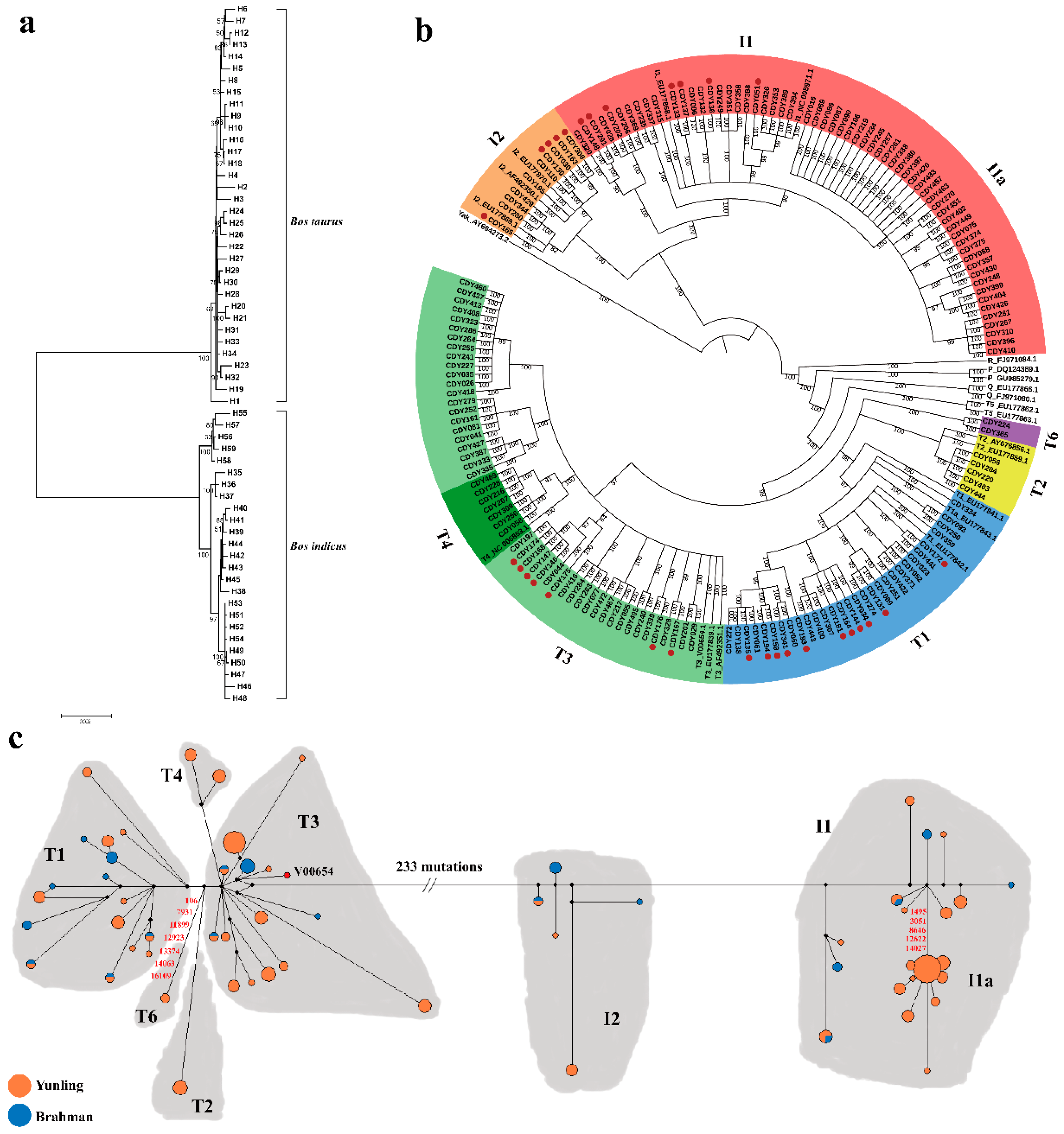
3.1. MtDNA Sequence Variation and Genetic Diversity
3.2. MtDNA Sequence Variation and Genetic Diversity
4. Discussion
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
References
- Qu, K.X.; Huang, B.Z.; Yang, G.R.; He, Z.X.; Zhang, Y.P.; Zan, L.S. Genetic diversity analysis of BMY cattle based on microsatellite DNA markers. Genetika 2012, 48, 522–528. [Google Scholar] [CrossRef] [PubMed]
- Huang, B.Z.; Wang, A.K.; Jin, X.D.; Zhang, J.C.; Qu, K.X.; He, Z.X.; Yang, G.R.; Yang, K.; Yang, S.P.; Fu, M.F.; et al. Crossbreeding in Yunling cattle. In Proceedings of the Ninth China Cattle Industry Development Conference, Hainan, China, 8–9 November 2014. [Google Scholar]
- Qu, K.X.; Zhang, J.C.; Yang, G.R.; He, Z.X.; Jin, X.D.; Wang, A.K.; Yuan, X.P.; Huang, B.Z.; Zan, L.S. Karyotypic analysis of BMY cattle and Brahman. J. Northwest A & F Univ. 2011, 39, 8–12. [Google Scholar]
- Xia, X.; Yao, Y.; Li, C.; Zhang, F.; Qu, K.; Chen, H.; Huang, B.; Lei, C. Genetic diversity of Chinese cattle revealed by Y-SNP and Y-STR markers. Anim. Genet. 2019, 50, 64–69. [Google Scholar] [CrossRef] [PubMed]
- Troy, C.S.; Machugh, D.E.; Bailey, J.F.; Magee, D.A.; Loftus, R.T.; Cunningham, P.; Chamberlain, A.T.; Sykes, B.C.; Bradley, D.G. Genetic evidence for Near-Eastern origins of European cattle. Nature 2001, 410, 1088–1091. [Google Scholar] [CrossRef]
- Mannen, H.; Kohno, M.; Nagata, Y.; Tsuji, S.; Bradley, D.G.; Yeo, J.S.; Nyamsamba, D.; Zagdsuren, Y.; Yokohama, M.; Nomura, K. Independent mitochondrial origin and historical genetic differentiation in North Eastern Asian cattle. Mol. Phylogenet. Evol. 2004, 32, 539–544. [Google Scholar] [CrossRef]
- Achilli, A.; Bonfiglio, S.; Olivieri, A.; Malusà, A.; Pala, M.; Hooshiar, K.B.; Perego, U.A.; Ajmonemarsan, P.; Liotta, L.; Semino, O. The Multifaceted Origin of Taurine Cattle Reflected by the Mitochondrial Genome. PLoS ONE 2009, 4, e5753. [Google Scholar] [CrossRef]
- Chen, S.; Lin, B.Z.; Baig, M.; Mitra, B.; Lopes, R.J.; Santos, A.M.; Magee, D.A.; Azevedo, M.; Tarroso, P.; Sasazaki, S. Zebu cattle are an exclusive legacy of the South Asia neolithic. Mol. Biol. Evol. 2010, 27, 1–6. [Google Scholar] [CrossRef]
- Briggs, H.M.; Briggs, D.M. Modern Breeds of Livestock, 4th ed.; Macmillan Publishing Co.: New York, NY, USA, 1980; pp. 479–487. [Google Scholar]
- Sambrook, J.; Russell, D.W. Molecular Cloning: A Laboratory Manual; Translated by Huang P. T.; Science Press: Beijing, China, 2002. [Google Scholar]
- Librado, P.; Rozas, J. DnaSP v5: A software for comprehensive analysis of DNA polymorphism data. Bioinformatics 2009, 25, 1451–1452. [Google Scholar] [CrossRef]
- Ronquist, F.; Huelsenbeck, J.P. MrBayes 3: Bayesian phylogenetic inference under mixed models. Bioinformatics 2003, 19, 1572–1574. [Google Scholar] [CrossRef] [Green Version]
- Tamura, K.; Peterson, D.; Peterson, N.; Stecher, G.; Nei, M.; Kumar, S. MEGA5: Molecular evolutionary genetics analysis using maximum likelihood, evolutionary distance, and maximum parsimony methods. Mol. Biol. Evol. 2011, 28, 2731–2739. [Google Scholar] [CrossRef]
- Bandelt, H.J.; Forster, P.; Röhl, A. Median-joining networks for inferring intraspecific phylogenies. Mol. Biol. Evol. 1999, 16, 37. [Google Scholar] [CrossRef] [PubMed]
- Achilli, A.; Olivieri, A.; Soares, P.; Lancioni, H.; Hooshiar Kashani, B.; Perego, U.A.; Nergadze, S.G.; Carossa, V.; Santagostino, M.; Capomaccio, S.; et al. Mitochondrial genomes from modern horses reveal the major haplogroups that underwent domestication. Proc. Natl. Acad. Sci. USA 2012, 109, 2449–2454. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Colli, L.; Lancioni, H.; Cardinali, I.; Olivieri, A.; Capodiferro, M.R.; Pellecchia, M.; Rzepus, M.; Zamani, W.; Naderi, S.; Gandini, F.; et al. Whole mitochondrial genomes unveil the impact of domestication on goat matrilineal variability. BMC Genom. 2015, 16, 1115. [Google Scholar] [CrossRef] [PubMed]
- Wang, S.; Chen, N.; Capodiferro, M.R.; Zhang, T.; Lancioni, H.; Zhang, H.; Miao, Y.; Chanthakhoun, V.; Wanapat, M.; Yindee, M.; et al. Whole mitogenomes reveal the history of swamp buffalo: Initially shaped by Glacial Periods and eventually modelled by domestication. Sci. Rep. 2017, 7, 4708. [Google Scholar] [CrossRef] [PubMed]
- Silvia, B.; Catarina, G.; Anna, D.G.; Alessandro, A.; Anna, O.; Licia, C.; Kassahun, T.; Saif Hassan, A.; Gama, L.T.; Federica, C. Origin and spread of Bos taurus: New clues from mitochondrial genomes belonging to haplogroup T1. PLoS ONE 2012, 7, e38601. [Google Scholar]
- Chen, N.; Cai, Y.; Chen, Q.; Li, R.; Wang, K.; Huang, Y.; Hu, S.; Huang, S.; Zhang, H.; Zheng, Z. Whole-genome resequencing reveals world-wide ancestry and adaptive introgression events of domesticated cattle in East Asia. Nat. Commun. 2018, 9, 2337. [Google Scholar] [CrossRef] [PubMed]
- Li, R.; Li, C.; Liu, H.; Zeng, B.; Xiao, H.; Chen, S. Mitochondrial diversity and phylogeographic structure of native cattle breeds from Yunnan, Southwestern China. Livest. Sci. 2018, 214, 129–134. [Google Scholar] [CrossRef]
- Xia, X.; Qu, K.; Zhang, G.; Jia, Y.; Ma, Z.; Zhao, X.; Huang, Y.; Chen, H.; Huang, B.; Lei, C. Comprehensive analysis of the mitochondrial DNA diversity in Chinese cattle. Anim. Genet. 2019, 50, 70–73. [Google Scholar] [CrossRef]
- Gou, X.; Wang, Y.; Yang, S.; Deng, W.; Mao, H. Genetic diversity and origin of Gayal and cattle in Yunnan revealed by mtDNA control region and SRY gene sequence variation. J. Anim. Breed. Genet. 2015, 127, 154–160. [Google Scholar] [CrossRef]
- Lei, C.Z.; Chen, H.; Zhang, H.C.; Cai, X.; Liu, R.Y.; Luo, L.Y.; Wang, C.F.; Zhang, W.; Ge, Q.L.; Zhang, R.F. Origin and phylogeographical structure of Chinese cattle. Anim. Genet. 2006, 37, 579–582. [Google Scholar] [CrossRef]
- Jia, S.G.; Zhou, Y.; Lei, C.Z.; Yao, R.; Zhang, Z.Y.; Fang, X.T.; Chen, H. A new insight into cattle’s maternal origin in six Asian countries. J. Genet. Genom. 2010, 37, 173–180. [Google Scholar] [CrossRef]
- Kim, J.H.; Lee, S.S.; Kim, S.C.; Choi, S.B.; Kim, S.H.; Chang, W.L.; Jung, K.S.; Kim, E.S.; Choi, Y.S.; Kim, S.B. Haplogroup classification of Korean cattle breeds based on sequence variations of mtDNA control region. Asian-Australas. J. Anim. Sci. 2016, 29, 624–630. [Google Scholar] [CrossRef] [PubMed]
- Ginja, C.; Penedo, M.C.; Melucci, L.; Quiroz, J.; Martínez López, O.R.; Revidatti, M.A.; Martínez-Martínez, A.; Delgado, J.V.; Gama, L.T. Origins and genetic diversity of New World Creole cattle: Inferences from mitochondrial and Y chromosome polymorphisms. Anim. Genet. 2010, 41, 128–141. [Google Scholar] [CrossRef] [PubMed]
| Breed | Sample Size | Haplogroup | S | H | k | Hd ± SD | Pi ± SD | ||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| T1 | T2 | T3 | T4 | T6 | I1 | I2 | |||||||
| Yunling | 129 | 19 | 5 | 37 | 7 | 2 | 54 | 5 | 405 | 47 | 128.074 | 0.964 ± 0.007 | 0.00784 ± 0.00013 |
| Brahman | 31 | 11 | 0 | 8 | 0 | 0 | 7 | 5 | 312 | 20 | 123.394 | 0.959 ± 0.020 | 0.00756 ± 0.00067 |
| Total | 160 | 30 | 5 | 45 | 7 | 2 | 61 | 10 | 429 | 59 | 127.055 | 0.974 ± 0.005 | 0.00778 ± 0.00014 |
© 2019 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Xia, X.; Qu, K.; Li, F.; Jia, P.; Chen, Q.; Chen, N.; Zhang, J.; Chen, H.; Huang, B.; Lei, C. Abundant Genetic Diversity of Yunling Cattle Based on Mitochondrial Genome. Animals 2019, 9, 641. https://doi.org/10.3390/ani9090641
Xia X, Qu K, Li F, Jia P, Chen Q, Chen N, Zhang J, Chen H, Huang B, Lei C. Abundant Genetic Diversity of Yunling Cattle Based on Mitochondrial Genome. Animals. 2019; 9(9):641. https://doi.org/10.3390/ani9090641
Chicago/Turabian StyleXia, Xiaoting, Kaixing Qu, Fangyu Li, Peng Jia, Qiuming Chen, Ningbo Chen, Jicai Zhang, Hong Chen, Bizhi Huang, and Chuzhao Lei. 2019. "Abundant Genetic Diversity of Yunling Cattle Based on Mitochondrial Genome" Animals 9, no. 9: 641. https://doi.org/10.3390/ani9090641





